Quantification of Circular RNAs Using Digital Droplet PCR
Summary
This protocol describes the detailed method of digital droplet PCR (dd-PCR) for precise quantification of circular RNA (circRNA) levels in cells using divergent primers.
Abstract
Digital droplet polymerase chain reaction (dd-PCR) is one of the most sensitive quantification methods; it fractionates the reaction into nearly 20,000 water-in-oil droplets, and the PCR occurs in the individual droplets. The dd-PCR has several advantages over conventional real-time qPCR, including increased accuracy in detecting low-abundance targets, omitting reference genes for quantification, eliminating technical replicates for samples, and showing high resilience to inhibitors in the samples. Recently, dd-PCR has become one of the most popular methods for accurately quantifying target DNA or RNA for gene expression analysis and diagnostics. Circular RNAs (circRNAs) are a large family of recently discovered covalently closed RNA molecules lacking 5′ and 3′ ends. They have been shown to regulate gene expression by acting as sponges for RNA-binding proteins and microRNAs. Furthermore, circRNAs are secreted into body fluids, and their resistance to exonucleases makes them serve as biomarkers for disease diagnosis. This article aims to show how to perform divergent primer design, RNA extraction, cDNA synthesis, and dd-PCR analysis to accurately quantify specific circular RNA (circRNA) levels in cells. In conclusion, we demonstrate the precise quantification of circRNAs using dd-PCR.
Introduction
Recent advancements in RNA sequencing technologies and novel computational algorithms have discovered a new member of the growing family of non-coding RNAs, called circular RNA (circRNA)1. As the name suggests, circRNAs are a family of single-stranded RNA molecules with no free ends. They are formed by non-canonical head-to-tail splicing called back-splicing, where the upstream splice acceptor site joins covalently with the downstream splice donor site to form a stable RNA circle1,2. This process could be mediated by several factors, including inverted Alu repetitive elements present in the upstream and downstream of circularized exons, or can be mediated by some splicing factors or RNA binding proteins (RBPs)2,3. Circular RNAs generated exclusively from the exonic or intronic sequence are classified as exonic circRNA and circular intronic RNAs or ci-RNAs, whereas some exonic circRNAs retain the intron and are called exon-intron circRNAs (EIcircRNAs)3,4. The functions of circRNAs are multifaceted, including sponging miRNA and/or RBP, regulating transcription, and regulating cellular function by translating into peptides3,5,6,7. Several reports have highlighted the significance of circRNAs in various diseases and physiological processes8. Furthermore, tissue-specific expression patterns and resistance to exonuclease digestion make it a functional biomarker for disease diagnosis and it can also be used as a suitable therapeutic target8. Considering its importance in regulating health and diseases, the accurate quantification of circRNA expression is the need of the hour.
Several biochemical methods have been developed to quantify circRNAs in biological samples9. One of the most convenient and widely accepted methods for circRNA quantification is reverse transcription followed by quantitative polymerase chain reaction (RT-qPCR) using divergent primer pairs10. However, the majority of circRNAs are in low abundance compared to linear mRNAs, which makes it challenging to quantify them11. To overcome this issue, we sought to use digital droplet PCR (dd-PCR) to quantify the number of circRNAs in a given sample accurately. The dd-PCR is an advanced PCR technology that follows the microfluidics principle; it generates multiple aqueous droplets in oil, and the PCR occurs in each droplet as an individual reaction12. The reaction occurs in individual droplets and is analyzed using a droplet reader, which gives the number of positive or negative droplets for the gene of interest12. It is the most sensitive technique to accurately quantify a gene of interest, even if there is only a single copy in a given sample. Decreased sensitivity toward inhibitors, better precision, and omitting the reference gene for quantification make it more advantageous than conventional qPCR13,14,15. It has been widely used as a research and diagnostic tool for the absolute quantification of a gene of interest16,17. Here, we describe the detailed dd-PCR protocol for circRNA quantification in differentiating mouse C2C12 myotubes and proliferating mouse C2C12 myoblasts using divergent primers.
Protocol
RNA is sensitive to RNases; therefore, all reagents, instruments, and workspaces should be RNase-free and handled with care.
1. Divergent primer design for circRNA (see Figure 1)
- Retrieve the mature sequence from circRNA annotation data using BEDTools or from the UCSC genome browser by joining the exon/intron sequence present between the backsplice junction coordinates18,19.
- Prepare a PCR template sequence of 200 nucleotides in length by joining the last 100 nucleotides of the total circRNA sequence length to the first 100 nucleotides of the circRNA sequence10.
NOTE: If the circRNA length is less than 200 nucleotides, then divide it into two equal halves and join the nucleotides from the last half to the beginning of the first half. - Use the above template sequence for primer design by using the Primer3 web tool and setting a PCR product length ranging from 120-180 nucleotides in length20.
- Use the default settings of Primer3 for other parameters such as Tm and primer length. The primer sequences for circBnc2 are listed in Table 1.
- Order the divergent primer sequences for synthesis from any oligo synthesizing company.
2. RNA isolation
NOTE: Isolate total RNA from the mouse C2C12 cells using any commercially available kits or in-house RNA isolation method. The in-house RNA isolation method used here has been described previously21. The magnetic silica beads are prepared in the lab using the previously published protocol22. These magnetic beads can also be procured from various vendors.
- Take one 10 cm dish containing about 5 x 106 proliferating C2C12 myoblast cells and one 4-day differentiated C2C12 myotube for RNA isolation.
- Wash the cells with 10 mL of 1x PBS three times and discard the PBS using a pipette.
- Scrape the cells in 1 mL of 1x PBS using a cell scraper, centrifuge them at 4 ˚C for 5 min at 750 x g, and discard the PBS using a pipette.
- Add 1 mL of RNA isolation reagent (RIR) and lyse the cells by vigorous pipetting21.
- Add 200 µL (1/5 of the volume of RIR) of chloroform and vortex for 15 s. Centrifuge the tube at 4 ˚C for 10 min at 12,000 x g.
- Take 400 µL of the upper aqueous layer, load it on a silica column, and centrifuge for 1 min at 12,000 x g at room temperature. Take the flow-through to a new tube and add 600 µL of 100% ethanol.
- Add 20 µL of magnetic silica beads and place the tube on a thermomixer set at 25 ˚C and 1,200 rpm for 5 min.
NOTE: The bead volume can be increased or decreased depending on the initial cell numbers taken for lysis. Magnetic silica beads can be procured from commercial vendors. - Place the tube on a magnetic stand for 30 s or until the solution becomes clear. Discard the supernatant carefully using a pipette.
- Resuspend the magnetic silica beads in a 500 µL of wash buffer containing 90% ethanol. Place the tube on the magnetic stand for 30 s. Rotate the tube twice on the magnetic stand to wash the beads. Let the beads settle toward the magnet and discard the buffer using a pipette.
- Repeat step 2.9 twice. After the wash, briefly spin the tube and place it back on the magnetic stand for 30 s. Discard the remaining wash buffer. Air-dry the tube at 50 ˚C for 3 min on a thermomixer, keeping the lid open.
- Add 20 µL of nuclease-free water and resuspend the beads.
NOTE: The RNA elution volume can be decreased down to 10 µL to scale up the RNA concentration. - Place the tubes for 2-3 min at room temperature. Place the tube back on the magnetic stand and let it sit for 30 s. Collect the dissolved RNA in a new tube for quantity and quality assessment using a spectrophotometer.
NOTE: RNA can be stored at −20 ˚C or −80 ˚C for long-term storage. To get an optimum result, it is recommended to proceed with the cDNA synthesis step the next day.
3. cDNA synthesis
- Measure the RNA concentration using a spectrophotometer and take 1 µg of RNA for cDNA synthesis.
- Mix 1 µg of total RNA with 1 µL of dNTP mix (10 mM each of dATP, dTTP, dGTP, and dCTP), 4 µL of 5x reverse transcriptase (RT) buffer, 2 µL of 10x RT random primer, 0.25 µL of reverse transcriptase enzyme (200 U/µL), and 0.5 µL of RNase inhibitor (40 U/µL), and make up the volume up to 20 µL using nuclease-free water.
NOTE: The reverse transcriptase enzyme volume can be altered depending on the initial RNA concentration taken for cDNA synthesis. - Tap the tube to mix the reaction and spin briefly. Place the tube at 25 ˚C for 10 min followed by at 50 ˚C for 1 h.
- To inactivate the enzyme, place the tube at 85 ˚C for 5 min.
- Ice chill the tube for 2 min and add 480 µL of nuclease-free water to the tube to make the cDNA concentration to 2 ng/µL.
NOTE: cDNA can be stored at -20 ˚C or used immediately for dd-PCR analysis.
4. Digital droplet PCR (dd-PCR) workflow
NOTE: The workflow of dd-PCR involves multiple steps, starting from sample preparation, followed by droplet generation, PCR amplification, droplet counting, and data analysis. Each step is crucial for accurate data generation as dd-PCR involves the absolute quantification of products and does not need any standard curve. Hence, each of the different steps of dd-PCR has been described below.
- Preparation of the dd-PCR reaction.
- Set up the PCR reaction of 22 µL in 0.2 mL PCR tubes or strips along with a negative control tube and NTC (non-template control) tubes.
NOTE: Each different divergent primer pair for the detection of different circRNAs must have an NTC reaction along with test samples. - Add 11 µL of dd-PCR master-mix (e.g., EvaGreen Supermix) and 11 µL of nuclease-free water to set up the negative control reaction tube.
NOTE: Thaw the dd-PCR master-mix at room temperature followed by a brief vortex to make a homogeneous mixture. Use the same nuclease-free water to set up the negative control reaction tube and NTC tubes that were used for diluting the cDNA. - Add 11 µL of dd-PCR master-mix, 5.5 µL of circRNA specific forward and reverse primer mix of 1 µM concentration, and 5.5 µL of nuclease-free water to set up the NTC reaction tubes.
NOTE: Dilute the forward and reverse circular RNA-specific divergent primers of 100 µM stock and mix to prepare a 1 µM working concentration of forward and reverse circRNA primer mix; thus, the final primer concentration is 250 nM in the 22 µL reaction. - Add 11 µL of dd-PCR master-mix, 5.5 µL of circRNA specific forward and reverse primer mix of 1 µM concentration, and 5.5 µL of cDNA (2 ng/µL) to set up the test reaction tubes.
- Vortex thoroughly followed by briefly centrifuging all the reaction tubes to get a homogeneous reaction mixture at the bottom of the tubes.
- Set up the PCR reaction of 22 µL in 0.2 mL PCR tubes or strips along with a negative control tube and NTC (non-template control) tubes.
- Droplet generation.
- Transfer the 20 µL of PCR mixture from 0.2 mL PCR tubes to the sample wells of the droplet generator cartridge.
- Add 70 µL of droplet generation oil to the oil wells of the droplet generator cartridge carefully using a multi-channel pipette.
NOTE: Incubate the droplet generation oil at room temperature for 15 min before use. - Cover the droplet generator cartridge with the rubber gasket and place it in the droplet generator machine to get the sample-oil droplet mix, generated in the droplet wells of the cartridge.
- PCR amplification.
- Transfer 40 µL of the sample-oil droplet mix from the droplet wells of the droplet generator cartridge to a 96-well PCR plate using a multi-channel pipette.
NOTE: Carefully transfer the sample-oil droplet mix while making an angle to the wells through the tips of the multi-channel pipette to avoid breaking the droplets while transferring them to the dd-PCR 96-well plate. - Seal the 96-well PCR plate with the aluminum foil sealer, place it in the plate sealer machine block preheated at 80 ˚C, and click seal for sealing the PCR plate.
- Now, place the plate in the dd-PCR thermal cycler and set the program as mentioned in Table 2 with the lid temperature set at 105 ˚C and a ramp rate of 2 ˚C/s.
- Transfer 40 µL of the sample-oil droplet mix from the droplet wells of the droplet generator cartridge to a 96-well PCR plate using a multi-channel pipette.
- Droplet counting.
- Once PCR is over, place the plate on the dd-PCR droplet counter machine with the plate holder in the correct position for counting the droplets.
- Open the droplet analysis software and set up the run by giving input for the sample information.
- Click on the setup option. Then, click on the template option to open a new template.
- Define each well by putting the details of the experiment, such as sample name (cDNA used or negative control or NTC), target name (circRNA primer name) and type (unknown), and experiment type as ABS (absolute quantification) and Supermix (dd-PCR EvaGreen Supermix) used, etc.
- Finally, save the template and start the run by clicking on the run option and select counting by row or column-wise. Allow the machine to complete the droplet counting, which takes about 1 min/well.
- Click the Analyze button to analyze the data after the completion of the droplet reading.
- Click the 1D Amplitude button to see the positive and negative droplets
- To segregate the positive droplets from the negative droplets, put a common threshold line for all samples with the same target.
NOTE: The total droplets in each sample should be above 10,000 to be considered for absolute quantification. The NTC should not show positive droplet counts. The presence of positive droplets in the NTC indicates contamination of the reagents or amplification of the primer dimers. It is difficult to find out whether the positive droplets have primer dimers or non-specific products in NTC. It is advised to check the primers and avoid any contamination of the reagents as a general practice for any PCR including dd-PCR. - Click the Export button to export the circRNA counts data as a .csv file. The exported data show the number of circRNAs present in each sample.
- Manually calculate the number of circRNAs in each sample per nanogram of RNA by considering the total amount of cDNA used in each reaction.
Representative Results
The absolute number of circRNAs in each sample can be derived from the exported dd-PCR data. Real-time quantitative PCR analysis suggested differential expression of circBnc2 in the differentiated C2C12 myotubes (data not shown). Here, we wanted to check the absolute copy number of circBnc2 in proliferating C2C12 myoblasts and myotubes. Since the expression of circBnc2 is compared in two conditions, it is really important to process all samples for RNA isolation, cDNA synthesis, and PCR simultaneously using the same reagents and procedure. To perform the absolute quantification of circRNAs using the dd-PCR reaction, an equal amount of initial total RNA should be used for cDNA synthesis to calculate the exact number of target RNA molecules per nanogram of total RNA. For example, 1 µg of total RNA was used for cDNA synthesis and diluted to 500 µL using nuclease-free water to get a final concentration of 2 ng/µL before performing the dd-PCR assay (Figure 1).
As shown in Figure 2A, cDNA from 10 ng of RNA from proliferating C2C12 myoblasts cells (MB) and a 4-day differentiated myotube (MT) was used to check the difference in expression of circBnc2 in these two conditions. The samples were processed to generate the droplet and perform PCR, followed by counting the positive and negative droplets using the droplet analysis software as per the manufacturer's instructions (Figure 2B). Since primers can amplify non-specific products and form primer dimers, NTC should be used for all primer sets. Ideally, NTC should have no positive droplet counts.
As shown in Figure 3A, all samples except MT1 showed more than 12,000 droplet counts. Since the total droplet count was low in MT1, this sample was not considered for final data analysis. The low total droplet count could be due to an error in droplet generation, rupture of the droplets during transfer to the PCR plate, or pipetting issues. Interestingly, there was a clear difference in the expression pattern of circBnc2 in the 4-day differentiated myotube condition compared to the myoblast cells. The analysis of circBnc2 abundance in terms of copy number indicated that the three replicates of C2C12 myoblasts had 76.8, 67, and 46 copies/ng of RNA, while the two myotube samples had 558 and 610 copies/ng of RNA (Figure 3B). Since one myotube sample did not work well, it is better to have a greater number of biological replicates to study the statistical differences in their expression patterns in the two conditions.
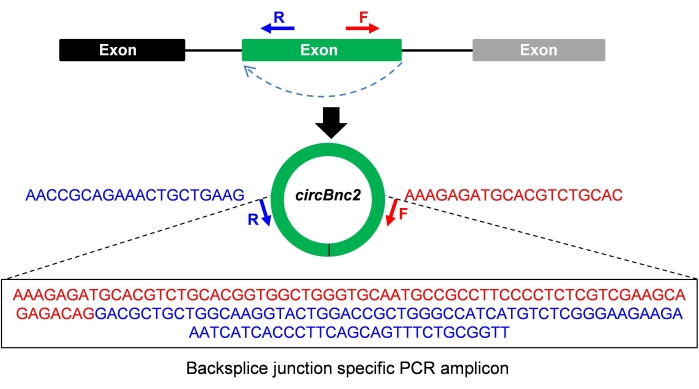
Figure 1: Schematic representation of the divergent primer design for the PCR amplification of circBnc2. Please click here to view a larger version of this figure.
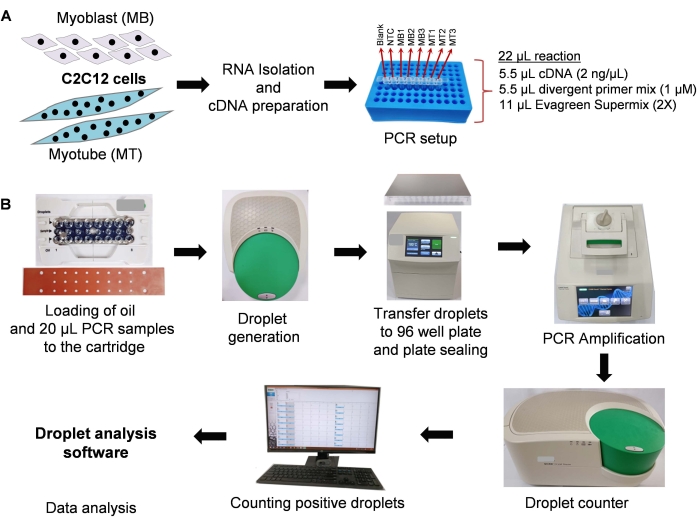
Figure 2: The dd-PCR workflow for circRNA quantification. The complete dd-PCR workflow includes many steps: (A) RNA isolation and cDNA preparation from C2C12 myoblast and 4-day differentiated myotube cells, PCR mixture preparation, (B) droplet generation, PCR amplification, droplet counting, and data analysis. Please click here to view a larger version of this figure.
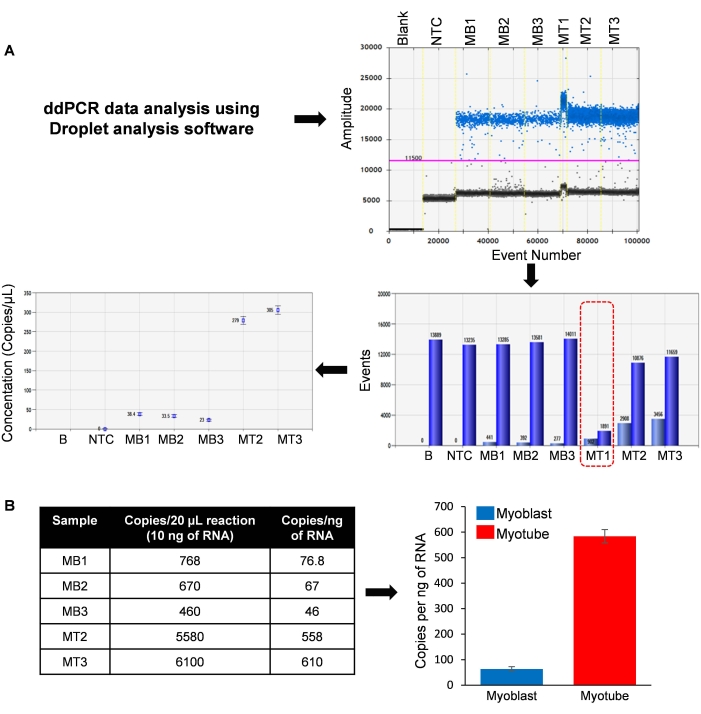
Figure 3: Data analysis using QuantaSoft software. (A) Data analysis includes the identification of positive and negative droplets using the quantitation software. (B) Calculating the number of circRNAs/ng of RNA in C2C12 myoblasts (MB) and myotubes (MT). The data in the panel B bar graph represent the mean ± STDEV of two to three biological replicates. Please click here to view a larger version of this figure.
| Primer Name | Sequence |
| circBnc2 Forward primer | AAAGAGATGCACGTCTGCAC |
| circBnc2 Reverse primer | AACCGCAGAAACTGCTGAAG |
Table 1: Divergent primer sequences used for the amplification of circBnc2.
| Step | Temperature (˚C) | Time | No of cycles | Ramp rate |
| Enzyme activation | 95 | 10 min | 1 | ~ 2 ˚C/s |
| Denaturation | 94 | 30 s | 40 | ~ 2 ˚C/s |
| Annealing/Extension | 60 | 1 min | 40 | ~ 2 ˚C/s |
| Signal stabilisation | 4 | 5 min | 1 | ~ 2 ˚C/s |
| Signal stabilisation | 90 | 5 min | 1 | ~ 2 ˚C/s |
Table 2: Conditions for PCR amplification of circBnc2 on a thermal cycler.
Discussion
CircRNA research has grown in the last decade with the discovery of high-throughput sequencing technologies. As a result, it has been considered a potential molecule for future RNA therapeutics. In addition, it is known to act as a biomarker in several diseases, including cancer and cardiovascular diseases4,8. However, the identification of circRNA is tricky because of its low abundance and it having only one specific backsplice junction sequence that differentiates it from the parental mRNA9. RT-PCR using divergent primers has been the gold standard technique to validate circRNA since its discovery9,10. Although several molecular methods have been developed for circRNA quantification, it is challenging to measure circRNA expressions accurately because of their low abundance. Since dd-PCR can amplify very low-abundant RNA/DNA molecules from a given sample14, it is one of the best methods for the accurate quantification of circRNAs.
The dd-PCR is an end-point assay that gives absolute quantification of the target gene15. Unlike conventional qPCR, it is time-consuming, tedious, and expensive, making it less accepted compared to real-time PCR even a decade after its discovery15. However, it offers several advantages over the widely used qPCR to study gene expression changes13,23. First, dd-PCR can provide the exact number of copies of DNA present in a given sample. Second, a change in housekeeping gene expression alters the calculation of the target gene population in a given sample. This can be avoided in dd-PCR, which does not depend on standard curves or housekeeping genes for quantifying target DNA15. Third, as the PCR reaction occurs in small compartments, it provides higher flexibility toward the presence of non-specific inhibitors or background DNA, making it a more accurate method for quantifying target genes23,24.
We used the dd-PCR technology to quantify circBnc2 expression in proliferating C2C12 myoblasts and differentiated myotubes using divergent primers. However, specific circRNA amplification with divergent primer pairs without primer-dimers must be checked in regular PCR before performing dd-PCR. Also, the dd-PCR protocol must be followed carefully for the accurate measurement of circRNA expression. The workflow presented here can be easily adapted for the quantification of any circRNA of interest. Recent studies have highlighted the diagnostic values of circRNAs present in tissue and body fluids for the detection of diseases8. Together, this method will be an essential tool in the research and diagnostics industry and will accelerate circRNA research.
Declarações
The authors have nothing to disclose.
Acknowledgements
This work was supported by intramural funding from the Institute of Life Sciences, the DBT research grant (BT/PR27622/MED/30/1961/2018), and the Wellcome Trust/DBT India Alliance Fellowship [IA/I/18/2/504017] awarded to Amaresh C. Panda. We thank other lab members for proofreading the article.
Materials
| 1.5 ml microcentifuge tube | Tarson | 500010 | |
| 0.2 ml tube strips with cap | Tarson | 610020, 510073 | |
| Filter Tips | Tarson | 528104 | |
| DNase/RNase-Free Distilled Water | Thermo Fisher Scientific | 10977023 | |
| Phosphate-buffered saline (PBS) | Sigma | P4417 | |
| Cell scrapper | HiMedia | TCP223 | |
| Chloroform | SRL | 96764 | |
| DNA diluent | HiMedia | MB228 | |
| Random primers | Thermo Fisher Scientific | 48190011 | |
| dNTP set | Thermo Fisher Scientific | R0181 | |
| Murine RNase inhibitor | NEB | M0314S | |
| Maxima reverse transcriptase | Thermo Fisher Scientific | EP0743 | |
| QX200 dd-PCR Evagreen Supermix | Bio-Rad | 1864033 | |
| Droplet generation oil for Evagreen | Bio-Rad | 1864006 | |
| PCR Plate Heat Seal, foil, pierceable | Bio-Rad | 1814040 | |
| DG8 Cartridges and Gaskets | Bio-Rad | 1864007 | |
| DG8 Cartridge holder | Bio-Rad | 1863051 | |
| QX200 Droplet Generator | Bio-Rad | 1864002 | |
| ddPCR 96-Well Plates | Bio-Rad | 12001925 | |
| PX1 PCR Plate Sealer | Bio-Rad | 1814000 | |
| C1000 Touch Thermal Cycler with 96–Deep Well Reaction Module | Bio-Rad | 1851197 | |
| QX200 Droplet Reader | Bio-Rad | 1864003 | |
| Quantasoft Software | Bio-Rad | 1864011 | |
| Silica column | Umbrella Life Science | 38220090 | |
| UCSC Genome Browser | https://genome.ucsc.edu/ | ||
| Primer 3 | https://primer3.ut.ee/ |
Referências
- Salzman, J., Gawad, C., Wang, P. L., Lacayo, N., Brown, P. O. Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. PLoS One. 7 (2), 30733 (2012).
- Chen, L. L., Yang, L. Regulation of circRNA biogenesis. RNA Biology. 12 (4), 381-388 (2015).
- Kristensen, L. S., et al. The biogenesis, biology and characterization of circular RNAs. Nature Reviews Genetics. 20 (11), 675-691 (2019).
- Chen, X., Zhou, M., Yant, L., Huang, C. Circular RNA in disease: Basic properties and biomedical relevance. Wiley Interdisciplinary Reviews. RNA. , (2022).
- Sinha, T., Panigrahi, C., Das, D., Chandra Panda, A. Circular RNA translation, a path to hidden proteome. Wiley Interdisciplinary Reviews. RNA. 13 (1), 1685 (2022).
- Panda, A. C., Xiao, J. Circular RNAs Act as miRNA Sponges. Circular RNAs: Biogenesis and Functions. , 67-79 (2018).
- Das, A., Sinha, T., Shyamal, S., Panda, A. C. Emerging role of circular RNA-protein interactions. Non-Coding RNA. 7 (3), 48 (2021).
- Verduci, L., Tarcitano, E., Strano, S., Yarden, Y., Blandino, G. CircRNAs: Role in human diseases and potential use as biomarkers. Cell Death & Disease. 12 (5), 468 (2021).
- Pandey, P. R., et al. Methods for analysis of circular RNAs. Wiley Interdisciplinary Reviews: RNA. 11 (1), 1566 (2020).
- Das, A., Das, D., Panda, A. C. Validation of circular RNAs by PCR. PCR Primer Design. Methods in Molecular Biology. 2392, 103-114 (2022).
- Jeck, W. R., et al. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA. 19 (2), 141-157 (2013).
- Pinheiro, L. B., et al. Evaluation of a droplet digital polymerase chain reaction format for DNA copy number quantification. Analytical Chemistry. 84 (2), 1003-1011 (2012).
- Hayden, R. T., et al. Comparison of droplet digital PCR to real-time PCR for quantitative detection of cytomegalovirus. Journal of Clinical Microbiology. 51 (2), 540-546 (2013).
- Taylor, S. C., Laperriere, G., Germain, H. Droplet digital PCR versus qPCR for gene expression analysis with low abundant targets: From variable nonsense to publication quality data. Scientific Reports. 7, 2409 (2017).
- Hindson, B. J., et al. High-throughput droplet digital PCR system for absolute quantitation of DNA copy number. Analytical Chemistry. 83 (22), 8604-8610 (2011).
- Ishak, A., AlRawashdeh, M. M., Esagian, S. M., Nikas, I. P. Diagnostic, prognostic, and therapeutic value of droplet digital PCR (ddPCR) in COVID-19 patients: A systematic review. Journal of Clinical Medicine. 10 (23), 5712 (2021).
- Li, H., et al. Application of droplet digital PCR to detect the pathogens of infectious diseases. Bioscience Reports. 38 (6), (2018).
- Lee, B. T., et al. The UCSC genome browser database: 2022 update. Nucleic Acids Research. 50, 1115-1122 (2022).
- Quinlan, A. R., Hall, I. M. BEDTools: A flexible suite of utilities for comparing genomic features. Bioinformatics. 26 (6), 841-842 (2010).
- Untergasser, A., et al. Primer3–new capabilities and interfaces. Nucleic Acids Research. 40 (15), 115 (2012).
- Das, A., Das, D., Das, A., Panda, A. C. A quick and cost-effective method for DNA-free total RNA isolation using magnetic silica beads. bioRxiv. , (2020).
- Oberacker, P., et al. Simple Synthesis of Functionalized Paramagnetic Beads for Nucleic Acid Purification and Manipulation. Bio-protocol. 9 (20), 3394 (2019).
- Rački, N., Dreo, T., Gutierrez-Aguirre, I., Blejec, A., Ravnikar, M. Reverse transcriptase droplet digital PCR shows high resilience to PCR inhibitors from plant, soil and water samples. Plant Methods. 10 (1), 42 (2014).
- Taylor, S. C., Carbonneau, J., Shelton, D. N., Boivin, G. Optimization of droplet digital PCR from RNA and DNA extracts with direct comparison to RT-qPCR: Clinical implications for quantification of Oseltamivir-resistant subpopulations. Journal of Virological Methods. 224, 58-66 (2015).

