A Swin Transformer-Based Model for Thyroid Nodule Detection in Ultrasound Images
Summary
Here, a new model for thyroid nodule detection in ultrasound images is proposed, which uses Swin Transformer as the backbone to perform long-range context modeling. Experiments prove that it performs well in terms of sensitivity and accuracy.
Abstract
In recent years, the incidence of thyroid cancer has been increasing. Thyroid nodule detection is critical for both the detection and treatment of thyroid cancer. Convolutional neural networks (CNNs) have achieved good results in thyroid ultrasound image analysis tasks. However, due to the limited valid receptive field of convolutional layers, CNNs fail to capture long-range contextual dependencies, which are important for identifying thyroid nodules in ultrasound images. Transformer networks are effective in capturing long-range contextual information. Inspired by this, we propose a novel thyroid nodule detection method that combines the Swin Transformer backbone and Faster R-CNN. Specifically, an ultrasound image is first projected into a 1D sequence of embeddings, which are then fed into a hierarchical Swin Transformer.
The Swin Transformer backbone extracts features at five different scales by utilizing shifted windows for the computation of self-attention. Subsequently, a feature pyramid network (FPN) is used to fuse the features from different scales. Finally, a detection head is used to predict bounding boxes and the corresponding confidence scores. Data collected from 2,680 patients were used to conduct the experiments, and the results showed that this method achieved the best mAP score of 44.8%, outperforming CNN-based baselines. In addition, we gained better sensitivity (90.5%) than the competitors. This indicates that context modeling in this model is effective for thyroid nodule detection.
Introduction
The incidence of thyroid cancer has increased rapidly since 1970, especially among middle-aged women1. Thyroid nodules may predict the emergence of thyroid cancer, and most thyroid nodules are asymptomatic2. The early detection of thyroid nodules is very helpful in curing thyroid cancer. Therefore, according to current practice guidelines, all patients with suspected nodular goiter on physical examination or with abnormal imaging findings should undergo further examination3,4.
Thyroid ultrasound (US) is a common method used to detect and characterize thyroid lesions5,6. US is a convenient, inexpensive, and radiation-free technology. However, the application of US is easily affected by the operator7,8. Features such as the shape, size, echogenicity, and texture of thyroid nodules are easily distinguishable on US images. Although certain US features-calcifications, echogenicity, and irregular borders-are often considered criteria for identifying thyroid nodules, the presence of interobserver variability is unavoidable8,9. The diagnosis results of radiologists with different levels of experience are different. Inexperienced radiologists are more likely to misdiagnose than experienced radiologists. Some characteristics of US such as reflections, shadows, and echoes can degrade the image quality. This degradation in image quality caused by the nature of US imaging makes it difficult for even experienced physicians to locate nodules accurately.
Computer-aided diagnosis (CAD) for thyroid nodules has developed rapidly in recent years and can effectively reduce errors caused by different physicians and help radiologists diagnose nodules quickly and accurately10,11. Various CNN-based CAD systems have been proposed for thyroid US nodule analysis, including segmentation12,13, detection14,15, and classification16,17. CNN is a multilayer, supervised learning model18, and the core modules of CNN are the convolution and pooling layers. The convolution layers are used for feature extraction, and the pooling layers are used for downsampling. The shadow convolutional layers can extract primary features such as the texture, edges, and contours, while deep convolutional layers learn high-level semantic features.
CNNs have had great success in computer vision19,20,21. However, CNNs fail to capture long-range contextual dependencies due to the limited valid receptive field of the convolutional layers. In the past, backbone architectures for image classification mostly used CNNs. With the advent of Vision Transformer (ViT)22,23, this trend has changed, and now many state-of-the-art models use transformers as backbones. Based on non-overlapping image patches, ViT uses a standard transformer encoder25 to globally model spatial relationships. The Swin Transformer24 further introduces shift windows to learn features. The shift windows not only bring greater efficiency but also greatly reduce the length of the sequence because self-attention is calculated in the window. At the same time, the interaction between two adjacent windows can be made through the operation of shifting (movement). The successful application of the Swin Transformer in computer vision has led to the investigation of transformer-based architectures for ultrasound image analysis26.
Recently, Li et al. proposed a deep learning approach28 for thyroid papillary cancer detection inspired by Faster R-CNN27. Faster R-CNN is a classic CNN-based object detection architecture. The original Faster R-CNN has four modules-the CNN backbone, the region proposal network (RPN), the ROI pooling layer, and the detection head. The CNN backbone uses a set of basic conv+bn+relu+pooling layers to extract feature maps from the input image. Then, the feature maps are fed into the RPN and the ROI pooling layer. The role of the RPN network is to generate region proposals. This module uses softmax to determine whether anchors are positive and generates accurate anchors by bounding box regression. The ROI pooling layer extracts the proposal feature maps by collecting the input feature maps and proposals and feeds the proposal feature maps into the subsequent detection head. The detection head uses the proposal feature maps to classify objects and obtain accurate positions of the detection boxes by bounding box regression.
This paper presents a new thyroid nodule detection network called Swin Faster R-CNN formed by replacing the CNN backbone in Faster R-CNN with the Swin Transformer, which results in the better extraction of features for nodule detection from ultrasound images. In addition, the feature pyramid network (FPN)29 is used to improve the detection performance of the model for nodules of different sizes by aggregating features of different scales.
Protocol
This retrospective study was approved by the institutional review board of the West China Hospital, Sichuan University, Sichuan, China, and the requirement to obtain informed consent was waived.
1. Environment setup
- Graphic processing unit (GPU) software
- To implement deep learning applications, first configure the GPU-related environment. Download and install GPU-appropriate software and drivers from the GPU's website.
NOTE: See the Table of Materials for those used in this study.
- To implement deep learning applications, first configure the GPU-related environment. Download and install GPU-appropriate software and drivers from the GPU's website.
- Python3.8 installation
- Open a terminal on the machine. Type the following:
Command line: sudo apt-get install python3.8 python-dev python-virtualenv
- Open a terminal on the machine. Type the following:
- Pytorch1.7 installation
- Follow the steps on the official website to download and install Miniconda.
- Create a conda environment and activate it.
Command line: conda create –name SwinFasterRCNN python=3.8 -y
Command line: conda activate SwinFasterRCNN - Install Pytorch.
Command line: conda install pytorch==1.7.1 torchvision==0.8.2 torchaudio==0.7.2
- MMDetection installation
- Clone from the official Github repository.
Command line: git clone https://github.com/open-mmlab/mmdetection.git - Install MMDetection.
Command line: cd mmdetection
Command line: pip install -v -e .
- Clone from the official Github repository.
2. Data preparation
- Data collection
- Collected the ultrasound images (here, 3,000 cases from a Grade-A tertiary hospital). Ensure that each case has diagnostic records, treatment plans, US reports, and the corresponding US images.
- Place all the US images in a folder named "images."
NOTE: The data used in this study included 3,853 US images from 3,000 cases.
- Data cleaning
- Manually check the dataset for images of non-thyroid areas, such as lymph images.
- Manually check the dataset for images containing color Doppler flow.
- Delete the images selected in the previous two steps.
NOTE: After data cleaning, 3,000 images were left from 2,680 cases.
- Data annotation
- Have a senior doctor locate the nodule area in the US image and outline the nodule boundary.
NOTE: The annotation software and process can be found in Supplemental File 1. - Have another senior doctor review and revise the annotation results.
- Place the annotated data in a separate folder called "Annotations."
- Have a senior doctor locate the nodule area in the US image and outline the nodule boundary.
- Data split
- Run the python script, and set the path of the image in step 2.1.2 and the paths of the annotations in step 2.3.3. Randomly divide all the images and the corresponding labeled files into training and validation sets at a ratio of 8:2. Save the training set data in the "Train" folder and the validation set data in the "Val" folder.
NOTE: Python scripts are provided in Supplemental File 2.
- Run the python script, and set the path of the image in step 2.1.2 and the paths of the annotations in step 2.3.3. Randomly divide all the images and the corresponding labeled files into training and validation sets at a ratio of 8:2. Save the training set data in the "Train" folder and the validation set data in the "Val" folder.
- Converting to the CoCo dataset format
NOTE: To use MMDetection, process the data into a CoCo dataset format, which includes a json file that holds the annotation information and an image folder containing the US images.- Run the python script, and input the annotations folder paths (step 2.3.3) to extract the nodule areas outlined by the doctor and convert them into masks. Save all the masks in the "Masks" folder.
NOTE: The Python scripts are provided in Supplemental File 3. - Run the python script, and set the path of the masks folder in step 2.5.1 to make the data into a dataset in CoCo format and generate a json file with the US images.
NOTE: Python scripts are provided in Supplemental File 4.
- Run the python script, and input the annotations folder paths (step 2.3.3) to extract the nodule areas outlined by the doctor and convert them into masks. Save all the masks in the "Masks" folder.
3. Swin Faster RCNN configuration
- Download the Swin Transformer model file (https://github.com/microsoft/Swin-Transformer/blob/main/models/swin_transformer.py), modify it, and place it in the “mmdetection/mmdet/models/backbones/” folder. Open the “swin_transformer.py” file in a vim text editor, and modify it as the Swin Transformer model file provided in Supplemental File 5.
Command line: vim swin_transformer.py - Make a copy of the Faster R-CNN configuration file, change the backbone to Swin Transformer, and set up the FPN parameters.
Command line: cd mmdetection/configs/faster_rcnn
Command line: cp faster_rcnn_r50_fpn_1x_coco.py swin_faster_rcnn_swin.py
NOTE: The Swin Faster R-CNN configuration file (swin_faster_rcnn_swin.py) is provided in Supplemental File 6. The Swin Faster R-CNN network structure is shown in Figure 1. - Set the dataset path to the CoCo format dataset path (step 2.5.2) in the configuration file. Open the "coco_detection.py" file in the vim text editor, and modify the following line:
data_root = "dataset path(step 2.5.2)"
Command line:vim mmdetection/configs/_base_/datasets/coco_detection.py
4. Training the Swin Faster R-CNN
- Edit mmdetection/configs/_base_/schedules/schedule_1x.py, and set the default training-related parameters, including the learning rate, optimizer, and epoch. Open the "schedule_1x.py" file in the vim text editor, and modify the following lines:
optimizer = dict(type="AdamW", lr=0.001, momentum=0.9, weight_decay=0.0001)
runner = dict(type='EpochBasedRunner', max_epochs=48)
Command line:vim mmdetection/configs/_base_/schedules/schedule_1x.py
NOTE: In this protocol for this paper, the learning rate was set to 0.001, AdamW optimizer was used, the maximum training epoch was set to 48, and the batch size was set to 16. - Begin training by typing the following commands. Wait for the network to begin training for 48 epochs and for the resulting trained weights of the Swin Faster R-CNN network to be generated in the output folder. Save the model weights with the highest accuracy on the validation set.
Command line: cd mmdetection
Command line: python tools/train.py congfigs/faster_rcnn/swin_faster_rcnn_swin.py –work-dir ./work_dirs
NOTE: The model was trained on an "NVIDIA GeForce RTX3090 24G" GPU. The central processing unit used was the "AMD Epyc 7742 64-core processor × 128", and the operating system was Ubuntu 18.06. The overall training time was ~2 h.
5. Performing thyroid nodule detection on new images
- After training, select the model with the best performance on the validation set for thyroid nodule detection in the new images.
- First, resize the image to 512 pixels x 512 pixels, and normalize it. These operations are performed automatically when the test script is run.
Command line: python tools/test.py congfigs/faster_rcnn/swin_faster_rcnn_swin.py –out ./output - Wait for the script to automatically load the pretrained model parameters to the Swin Faster R-CNN, and feed the preprocessed image into the Swin Faster R-CNN for inference. Wait for the Swin Faster R-CNN to output the prediction box for each image.
- Finally, allow the script to automatically perform NMS postprocessing on each image to remove duplicate detection boxes.
NOTE: The detection results are output to the specified folder, which contains the images with the detection boxes and the bounding box coordinates in a packed file.
- First, resize the image to 512 pixels x 512 pixels, and normalize it. These operations are performed automatically when the test script is run.
Representative Results
The thyroid US images were collected from two hospitals in China from September 2008 to February 2018. The eligibility criteria for including the US images in this study were conventional US examination before biopsy and surgical treatment, diagnosis with biopsy or postsurgical pathology, and age ≥ 18 years. The exclusion criteria were images without thyroid tissues.
The 3,000 ultrasound images included 1,384 malignant and 1,616 benign nodules. The majority (90%) of the malignant nodules were papillary carcinoma, and 66% of the benign nodules were nodular goiter. Here, 25% of the nodules were smaller than 5 mm, 38% were between 5 mm and 10 mm, and 37% were larger than 10 mm.
All the US images were collected using Philips IU22 and DC-80, and their default thyroid examination mode was used. Both instruments were equipped with 5-13 MHz linear probes. For good exposure of the lower thyroid margins, all the patients were examined in the supine position with their backs extended. Both thyroid lobes and the isthmus were scanned in the longitudinal and transverse planes according to the American College of Radiology accreditation standards. All the examinations were carried out by two senior thyroid radiologists with ≥10 years of clinical experience. The thyroid diagnosis was based on the histopathological findings from fine needle aspiration biopsy or thyroid surgery.
In real life, as US images are corrupted by noise, it is important to conduct proper preprocessing of the US images, such as image denoising based on wavelet transform30, compressive sensing31, and histogram equalization32. In this work, we used histogram equalization to preprocess the US images, enhance image quality, and alleviate image quality degradation caused by noise.
In what follows, true positive, false positive, true negative, and false negative are referred to as TP, FP, TN, and FN, respectively. We used mAP, sensitivity, and specificity to evaluate the model's nodule detection performance. mAP is a common metric in object detection. Sensitivity and specificity were calculated using equation (1) and equation (2):
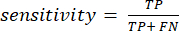 (1)
(1)
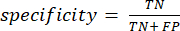 (2)
(2)
In this paper, TP is defined as the number of correctly detected nodules, which have an intersection over union (IoU) between the prediction box and the ground truth box of >0.3 and a confidence score >0.6. IoU is the intersection over union, which is computed by using equation (3):
 (3)
(3)
We compared several classic object detection networks, including SSD33, YOLO-v334, CNN backbone-based Faster R-CNN27, RetinaNet35, and DETR36. YOLO-v3 and SSD are single-stage detection networks, DETR is a transformer-based object-detection network, and Faster R-CNN and RetinaNet are two-stage detection networks. Table 1 shows that the performance of Swin Faster R-CNN is superior to the other methods, reaching 0.448 mAP, which is 0.028 higher than CNN backbone's Faster R-CNN and 0.037 higher than YOLO-v3. By using Swin Faster R-CNN, 90.5% of thyroid nodules can be detected automatically, which is ~3% higher than CNN backbone-based Faster R-CNN (87.1%). As shown in Figure 2, using Swin Transformer as the backbone makes boundary positioning more accurate.
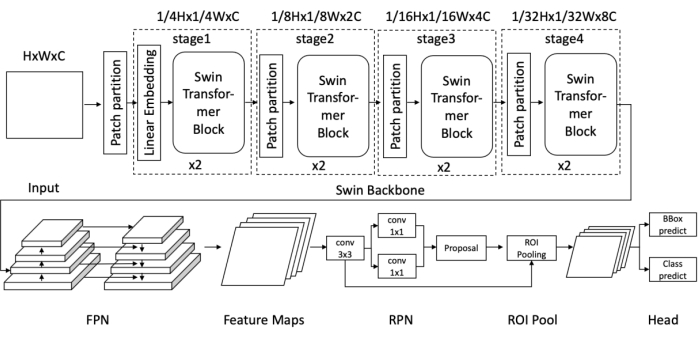
Figure 1: Diagram of the Swin Faster R-CNN network architecture. Please click here to view a larger version of this figure.
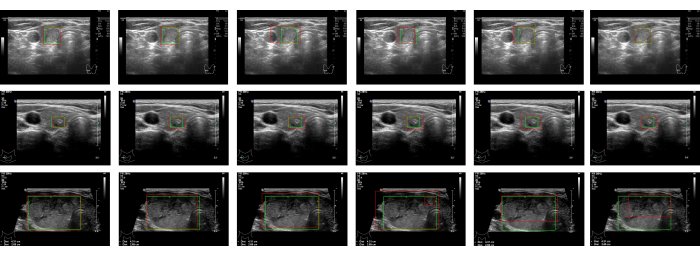
Figure 2: Detection results. The detection results for the same image are in a given row. The columns are the detection results, from left to right, for Swin Faster R-CNN, Faster R-CNN, YOLO-v3, SSD, RetinaNet, and DETR, respectively. The ground truths of the regions are marked with green rectangular boxes. The detection results are framed by the red rectangular boxes. Please click here to view a larger version of this figure.
| Method | Backbone | mAP | Sensitivity | Specificity |
| YOLO-v3 | DarkNet | 0.411 | 0.869 | 0.877 |
| SSD | VGG16 | 0.425 | 0.841 | 0.849 |
| RetinaNet | ResNet50 | 0.382 | 0.845 | 0.841 |
| Faster R-CNN | ResNet50 | 0.42 | 0.871 | 0.864 |
| DETR | ResNet50 | 0.416 | 0.882 | 0.86 |
| Swin Faster R-CNN without FPN | Swin Transformer | 0.431 | 0.897 | 0.905 |
| Swin Faster R-CNN with FPN | 0.448 | 0.905 | 0.909 |
Table 1: Performance comparison with state-of-the-art object detection methods.
Supplemental File 1: Operating instructions for the data annotation and the software used. Please click here to download this File.
Supplemental File 2: Python script used to divide the dataset into the training set and validation set, as mentioned in step 2.4.1. Please click here to download this File.
Supplemental File 3: Python script used to convert the annotations file into masks, as mentioned in step 2.5.1. Please click here to download this File.
Supplemental File 4: Python script used to make the data into a dataset in CoCo format, as mentioned in step 2.5.2. Please click here to download this File.
Supplemental File 5: The modified Swin Transformer model file mentioned in step 3.1. Please click here to download this File.
Supplemental File 6: The Swin Faster R-CNN configuration file mentioned in step 3.2. Please click here to download this File.
Discussion
This paper describes in detail how to perform the environment setup, data preparation, model configuration, and network training. In the environment setup phase, one needs to pay attention to ensure that the dependent libraries are compatible and matched. Data processing is a very important step; time and effort must be spent to ensure the accuracy of the annotations. When training the model, a "ModuleNotFoundError" may be encountered. In this case, it is necessary to use the "pip install" command to install the missing library. If the loss of the validation set does not decrease or oscillates greatly, one should check the annotation file and try to adjust the learning rate and batch size to make the loss converge.
Thyroid nodule detection is very important for the treatment of thyroid cancer. The CAD system can assist doctors in the detection of nodules, avoid differences in diagnosis results caused by subjective factors, and reduce the missed detection of nodules. Compared with existing CNN-based CAD systems, the network proposed in this paper introduces the Swin Transformer to extract ultrasound image features. By capturing long-distance dependencies, Swin Faster R-CNN can extract the nodule features from ultrasound images more efficiently. The experimental results show that Swin Faster R-CNN improves the sensitivity of nodule detection by ~3% compared to CNN backbone-based Faster R-CNN. The application of this technology can greatly reduce the burden on doctors, as it can detect thyroid nodules in early ultrasound examination and guide doctors to further treatment. However, due to the large number of parameters of the Swin Transformer, the inference time of Swin Faster R-CNN is ~100 ms per image (tested on NVIDIA TITAN 24G GPU and AMD Epyc 7742 CPU). It can be challenging to meet the requirements of real-time diagnosis with Swin Faster R-CNN. In the future, we will continue to collect cases to verify the effectiveness of this method and conduct further studies on dynamic ultrasound image analysis.
Declarações
The authors have nothing to disclose.
Acknowledgements
This study was supported by the National Natural Science Foundation of China (Grant No.32101188) and the General Project of Science and Technology Department of Sichuan Province (Grant No. 2021YFS0102), China.
Materials
| GPU RTX3090 | Nvidia | 1 | 24G GPU |
| mmdetection2.11.0 | SenseTime | 4 | https://github.com/open-mmlab/mmdetection.git |
| python3.8 | — | 2 | https://www.python.org |
| pytorch1.7.1 | 3 | https://pytorch.org |
Referências
- Grant, E. G., et al. Thyroid ultrasound reporting lexicon: White paper of the ACR Thyroid Imaging, Reporting and Data System (TIRADS) committee. Journal of the American College of Radiology. 12 (12 Pt A), 1272-1279 (2015).
- Zhao, J., Zheng, W., Zhang, L., Tian, H. Segmentation of ultrasound images of thyroid nodule for assisting fine needle aspiration cytology. Health Information Science and Systems. 1, 5 (2013).
- Haugen, B. R. American Thyroid Association management guidelines for adult patients with thyroid nodules and differentiated thyroid cancer: What is new and what has changed. Cancer. 123 (3), 372-381 (2017).
- Shin, J. H., et al. Ultrasonography diagnosis and imaging-based management of thyroid nodules: Revised Korean Society of Thyroid Radiology consensus statement and recommendations. Korean Journal of Radiology. 17 (3), 370-395 (2016).
- Horvath, E., et al. An ultrasonogram reporting system for thyroid nodules stratifying cancer risk for clinical management. The Journal of Clinical Endocrinology & Metabolism. 94 (5), 1748-1751 (2009).
- Park, J. -. Y., et al. A proposal for a thyroid imaging reporting and data system for ultrasound features of thyroid carcinoma. Thyroid. 19 (11), 1257-1264 (2009).
- Moon, W. -. J., et al. Benign and malignant thyroid nodules: US differentiation-Multicenter retrospective study. Radiology. 247 (3), 762-770 (2008).
- Park, C. S., et al. Observer variability in the sonographic evaluation of thyroid nodules. Journal of Clinical Ultrasound. 38 (6), 287-293 (2010).
- Kim, S. H., et al. Observer variability and the performance between faculties and residents: US criteria for benign and malignant thyroid nodules. Korean Journal of Radiology. 11 (2), 149-155 (2010).
- Choi, Y. J., et al. A computer-aided diagnosis system using artificial intelligence for the diagnosis and characterization of thyroid nodules on ultrasound: initial clinical assessment. Thyroid. 27 (4), 546-552 (2017).
- Chang, T. -. C. The role of computer-aided detection and diagnosis system in the differential diagnosis of thyroid lesions in ultrasonography. Journal of Medical Ultrasound. 23 (4), 177-184 (2015).
- Li, X. Fully convolutional networks for ultrasound image segmentation of thyroid nodules. , 886-890 (2018).
- Nguyen, D. T., Choi, J., Park, K. R. Thyroid nodule segmentation in ultrasound image based on information fusion of suggestion and enhancement networks. Mathematics. 10 (19), 3484 (2022).
- Ma, J., Wu, F., Jiang, T. A., Zhu, J., Kong, D. Cascade convolutional neural networks for automatic detection of thyroid nodules in ultrasound images. Medical Physics. 44 (5), 1678-1691 (2017).
- Song, W., et al. Multitask cascade convolution neural networks for automatic thyroid nodule detection and recognition. IEEE Journal of Biomedical and Health Informatics. 23 (3), 1215-1224 (2018).
- Wang, J., et al. Learning from weakly-labeled clinical data for automatic thyroid nodule classification in ultrasound images. , 3114-3118 (2018).
- Wang, L., et al. A multi-scale densely connected convolutional neural network for automated thyroid nodule classification. Frontiers in Neuroscience. 16, 878718 (2022).
- Krizhevsky, A., Sutskever, I., Hinton, G. E. Imagenet classification with deep convolutional neural networks. Communications of the ACM. 60 (6), 84-90 (2017).
- He, K., Zhang, X., Ren, S., Sun, J. Deep residual learning for image recognition. Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition. , 770-778 (2016).
- Hu, H., Gu, J., Zhang, Z., Dai, J., Wei, Y. Relation networks for object detection. Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition. , 3588-3597 (2018).
- Szegedy, C., et al. Going deeper with convolutions. Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition. , 1-9 (2015).
- Dosovitskiy, A., et al. An image is worth 16×16 words: Transformers for image recognition at scale. arXiv preprint arXiv:2010.11929. , (2020).
- Touvron, H., et al. Training data-efficient image transformers & distillation through attention. arXiv:2012.12877. , (2021).
- Liu, Z., et al. Swin Transformer: Hierarchical vision transformer using shifted windows. 2021 IEEE/CVF International Conference on Computer Vision (ICCV). , 9992-10002 (2021).
- Vaswani, A., et al. Attention is all you need. Advances in Neural Information Processing Systems. 30, (2017).
- Chen, J., et al. TransUNet: Transformers make strong encoders for medical image segmentation. arXiv. arXiv:2102.04306. , (2021).
- Ren, S., He, K., Girshick, R., Sun, J. Faster r-cnn: Towards real-time object detection with region proposal networks. Advances in Neural Information Processing Systems. 28, 91-99 (2015).
- Li, H., et al. An improved deep learning approach for detection of thyroid papillary cancer in ultrasound images. Scientific Reports. 8, 6600 (2018).
- Lin, T. -. Y., et al. Feature pyramid networks for object detection. Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition. , 2117-2125 (2017).
- Ouahabi, A. A review of wavelet denoising in medical imaging. 2013 8th International Workshop on Systems, Signal Processing and their Applications. , 19-26 (2013).
- Mahdaoui, A. E., Ouahabi, A., Moulay, M. S. Image denoising using a compressive sensing approach based on regularization constraints. Sensors. 22 (6), 2199 (2022).
- Castleman, K. R. . Digital Image Processing. , (1996).
- Liu, W., et al. Ssd: Single shot multibox detector. European Conference on Computer Vision. , 21-37 (2016).
- Redmon, J., Farhadi, A. Yolov3: An incremental improvement. arXiv. arXiv:1804.02767. , (2018).
- Lin, T. -. Y., Goyal, P., Girshick, R., He, K., Dollár, P. Focalloss for dense object detection. arXiv. arXiv:1708.02002. , (2017).
- Carion, N., et al. End-to-end object detection with transformers. Computer Vision-ECCV 2020: 16th European Conference. , 23-28 (2020).

