Universal Molecular Retention with 11-Fold Expansion Microscopy
Summary
Presented here is a new version of expansion microscopy (ExM), Magnify, that is modified for up to 11-fold expansion, conserving a comprehensive array of biomolecule classes, and is compatible with a broad range of tissue types. It enables the interrogation of the nanoscale configuration of biomolecules using conventional diffraction-limited microscopes.
Abstract
The nanoscale imaging of biological specimens can improve the understanding of disease pathogenesis. In recent years, expansion microscopy (ExM) has been demonstrated to be an effective and low-cost alternative to optical super-resolution microscopy. However, it has been limited by the need for specific and often custom anchoring agents to retain different biomolecule classes within the gel and by difficulties with expanding standard clinical sample formats, such as formalin-fixed paraffin-embedded tissue, especially if larger expansion factors or preserved protein epitopes are desired. Here, we describe Magnify, a new ExM method for robust expansion up to 11-fold in a wide array of tissue types. By using methacrolein as the chemical anchor between the tissue and gel, Magnify retains multiple biomolecules, such as proteins, lipids, and nucleic acids, within the gel, thus allowing the broad nanoscale imaging of tissues on conventional optical microscopes. This protocol describes best practices to ensure robust and crack-free tissue expansion, as well as tips for handling and imaging highly expanded gels.
Introduction
Biological systems exhibit structural heterogeneity, from the limbs and the organs down to the levels of proteins at the nanoscale. Therefore, a complete understanding of the operation of these systems requires visual examination across these size scales. However, the diffraction limit of light causes challenges in visualizing structures smaller than ~200-300 nm on a conventional fluorescence microscope. In addition, optical super-resolution methods1,2,3, such as stimulated emission depletion (STED), photo-activated localization microscopy (PALM), stochastic optical reconstruction microscopy (STORM), and structured illumination microscopy (SIM), though powerful, present their own challenges, as they require expensive hardware and reagents and often have slow acquisition times and a poor ability to image large volumes in 3D.
Expansion microscopy4 (ExM) provides an alternative means of circumventing the diffraction limit of light by covalently anchoring biomolecules into a water-swellable polymer gel and physically pulling them apart, thus rendering them resolvable on conventional optical microscopes. A multitude of ExM protocol variants have been developed since the original publication of ExM less than a decade ago, and these protocols allow the direct incorporation of proteins5,6,7, RNA8,9,10, or lipids11,12,13 into the gel network by altering the chemical anchor or expanding the sample further (thus improving the effective resolution) either in a single step14 or multiple iterative steps15,16. Until recently, no single ExM protocol could retain these three biomolecule classes with a single commercially available chemical anchor while providing a mechanically sturdy gel that could expand ~10-fold in a single expansion round.
Here, we present Magnify17, a recent addition to the ExM arsenal that uses methacrolein as the biomolecule anchor. Methacrolein forms covalent bonds with tissue like that of paraformaldehyde, ensuring that multiple classes of biomolecules can be retained within the gel network without requiring various specific or custom anchoring agents. Additionally, this technique can expand a broad spectrum of tissues up to 11-fold, including notoriously challenging samples such as formalin-fixed paraffin-embedded (FFPE) clinical samples. Previous methods for expanding such mechanically rigid samples required harsh protease digestion, rendering the antibody labeling of proteins of interest impossible after the sample had been expanded. In contrast, this technique achieves the expansion of FFPE clinical samples using a hot denaturing solution, thus preserving whole protein epitopes within the gel, which can be targeted for post-expansion imaging (Figure 1).
Protocol
All the experimental procedures involving animals were conducted in accordance with the National Institutes of Health (NIH) guidelines and were approved by the Institutional Animal Care and Use Committee at Carnegie Mellon University. Human tissue samples were commercially obtained.
1. Preparation of the stock reagents and solutions
NOTE: Refer to the Table of Materials for a list of the reagents used.
- Prepare the gelling stock solutions. These will be combined immediately before the gelling step.
- Prepare the monomer stock solution (4 g/100 mL N,N- dimethylacrylamide acid [DMAA], 50 g/100 mL sodium acrylate [SA], 66.7 g/100 mL acrylamide [AA], 0.10 g/100 mL N,N′-methylenebisacrylamide [Bis], 33 g/100 mL sodium chloride [NaCl], 1x phosphate-buffered saline; PBS) using the components listed in Table 1. Dissolve or dilute all the stock solutions in water. The monomer stock solution can be stored at 4 °C for at least 3 months.
- Make the following stock solutions separately in water: 0.5% (w/w) 4-Hydroxy-TEMPO (4HT), which inhibits gelation during monomer diffusion into the tissues, and 10% (w/w) tetramethylethylenediamine (TEMED), which accelerates the radical generation initiated by ammonium persulfate (APS) and is prepared at 10% (w/w).
NOTE: The stock solutions can be distributed into 1-2 mL aliquots and stored at 4 °C for up to 3 months; however, APS is made immediately before preparing the gelling solution.
- Prepare the homogenization buffer (1% w/v S.D.S., 8 M urea, 25 mM EDTA, 2x PBS, pH 8, at room temperature [R.T.]) by combining 1 g of SDS, 48.048 g of urea, 5 mL of EDTA (0.5M, pH 8), 20 mL of 10x PBS, 25 mL of Tris (2 M), and 0.75 g of glycine. Add water for a total volume of 100 mL. The solution can be scaled up or down as needed and can be stored at R.T.
2. Tissue preparation for archived and freshly prepared clinical tissue slides
NOTE: The tissue pre-processing steps differ based on how the specimens are prepared.
- For formaldehyde-fixed paraffin-embedded (FFPE) clinical samples, follow the steps below.
- Place the samples in a sequential series of solutions (a glass slide staining jar or 50 mL conical tube may be used): xylene, 95% ethanol, 70% ethanol, and 50% ethanol, followed by doubly deionized water. Use each solution twice for at least 3 min each time. Use forceps to place the slide into the container, and add 15 mL of the respective solution (enough to cover the sample). Place the capped conical tube horizontally on a shaker at room temperature.
- For samples stained and mounted on permanent slides, follow the steps below.
- Place the slide briefly in xylene, and carefully remove the coverslips using an appropriate tool, such as a razor blade. If the coverslip remains stuck, return the slides in xylene to room temperature until the coverslip loosens.
- Then, process the samples as FFPE samples (step 2.1.1).
NOTE: For hematoxylin and eosin (H&E)-stained slides, the hematoxylin and eosin stain is lost during the expansion process.
- For unfixed frozen tissue slides in optimum cutting temperature (OCT) solution, fix the tissues in acetone at −20 °C for 10 min before washing the samples with 1x PBS solution three times for 10 min each time at room temperature.
- For previously fixed frozen clinical tissue slides, melt the OCT by incubating the slides for 2 min at room temperature, and then wash with 1x PBS solution three times for 5 min each time at room temperature.
3. Tissue preparation for paraformaldehyde-fixed mouse brain
- Follow this protocol for mouse brain sections that are 30-50 µm thick.
NOTE: Success may be found with larger sections, but longer times may be required for monomer solution diffusion and homogenization. - If the sections are not currently stored in a 1x PBS solution (for instance, in a 30% [v/v] glycerol and 30% [v/v] ethylene glycol solution in 1x PBS), wash three times for at least 10 min each time at room temperature in 1x PBS in a 6-to-24-well plate.
- Obtain an uncoated glass microscopy slide. If one side of the glass slide is coated, use a wet paintbrush to check for the uncoated side (often, the label of the coated side will be made of ground glass and will become less opaque when wet).
- Use a paintbrush to wet the uncoated slide. Remove the mouse brain tissue from the well plate with the paintbrush, and carefully place it onto the slide, using a rolling motion to ensure the tissue is flat. Allow excess PBS to remain on the tissue until all the sections are mounted.
NOTE: While a single glass slide may be used for multiple tissue sections, when more tissue sections are gelled at once (even on separate slides), more care must be taken to ensure they do not completely dry out. - Using a laboratory paper cloth, remove the majority of the excess PBS from the slide. Leave a small liquid ring around each tissue section, but leave the rest of the slide mostly dry. This remaining liquid will cover the tissue while the gel monomer solution is being prepared.
4. Gelling
NOTE: This protocol is suitable for all tissue types prepared for use with this technique.
- Apply spacers on either side of the tissue section (Figure 2A) to prevent tissue compression. To do so, make the spacers by thinly cutting pieces of cover glass using a diamond-tipped pen, and then secure them to the slide using small amounts of water or 1x PBS, or fix them using a small amount of super glue. Next, gently place the spacers on either side of the tissue.
- Prepare the final gelling solution. Generally, ~200 µL is sufficient per tissue section, but the volume should not exceed 800 µL per slide. Any more gelling solution will result in spillover.
NOTE: Methacrolein is used to anchor biomolecules into the gel. However, no separate anchoring step is required, and methacrolein may be added directly to the final gelling solution.- Per section, combine the following in order: 200 µL of monomer solution, 0.5 µL of 0.5% 4HT stock solution (1:400 dilution), 0.2-0.5 µL of methacrolein (1:400-1:1,000 dilution depending on the tissue type; generally, use a 1:1,000 dilution for paraformaldehyde [PFA]-fixed samples and 1:400 for FFPE clinical specimens), 2 µL of 10% TEMED stock solution (1:100 dilution), and 5 µL of 10% A.P.S. stock solution (1:40 dilution).
NOTE: Prepare the final gelling solution immediately before use. To prevent premature gelling, add the APS solution last.
- Per section, combine the following in order: 200 µL of monomer solution, 0.5 µL of 0.5% 4HT stock solution (1:400 dilution), 0.2-0.5 µL of methacrolein (1:400-1:1,000 dilution depending on the tissue type; generally, use a 1:1,000 dilution for paraformaldehyde [PFA]-fixed samples and 1:400 for FFPE clinical specimens), 2 µL of 10% TEMED stock solution (1:100 dilution), and 5 µL of 10% A.P.S. stock solution (1:40 dilution).
- Remove the excess solution from the tissue section, and place the slide in a Petri dish. Add all the freshly prepared, cold gelling solution to the sample, and incubate the mixture on the tissue for 30 min at 4 °C to allow diffusion into the tissue.
- Place a second uncoated glass slide carefully over the tissue and gel solution. Air bubbles should be avoided (Figure 2B).
NOTE: If air bubbles become trapped over the tissue, the glass lid may be carefully lifted and placed back down. Additionally, any monomer solution that spills out may be trimmed off after polymerization and will not cause any problems for the tissue. - Ensure that polymerization is completed. This polymerization can be done in two ways, each producing similar results. First, incubate the samples at 37 °C in a humidified environment (such as a closed Petri dish with a damp paper towel) overnight. Alternatively, for more rapid polymerization, incubate the samples in the humidified atmosphere for 2 h at 37 °C followed by 1 h at 60°C.
5. Sample digestion and tissue expansion
NOTE: This protocol is suitable for all tissue types.
- Wearing eye protection, slide a razor blade between the two glass slides of the gelling chamber to separate them.
NOTE: The glass slides should separate very easily. If different parts of the gel are stuck to both slides, a paintbrush may be used to guide the gel to one or the other. If the gel has become particularly dry during polymerization, 1x PBS may be applied to the outside of the gel chamber, but do not submerge the gel fully, as this will cause it to expand before the tissue is homogenized, which will result in cracking. - Trim the blank gel around the tissue to minimize the volume. Cutting the gel asymmetrically allows for the tracking of the orientation of the gel after homogenization, as the sample will be completely transparent.
- Gently lift the tissue-containing gel from the slide with a razor blade, and gently transfer it into a 2 mL centrifuge tube with a paintbrush.
- Fill the tube to the top with homogenization buffer (Table 2) pre-heated to 80 °C.
- Incubate the sample with shaking at 80 °C for 8 h (for PFA-fixed mouse brain) or 60 h (most FFPE clinical samples; see Table 3).
NOTE: The homogenization time is dependent on the tissue type and fixation method. Many of these conditions have been validated already, but for others, optimization may be necessary. - Pour the contents of the centrifuge tube into a single well of a 6-well plastic cell culture plate.
- Remove the denaturation buffer with a transfer pipet.
- To completely remove the remaining SDS from the hydrogel, wash the sample in a 1% non-ionic surfactant (C12E10) solution as follows: at least three times for 10 min each time at room temperature; for 60 min at 60 °C; and then another three washes for 10 min each time at room temperature.
- Store the sample in 1x PBS + 0.02% sodium azide at 4 °C until post-expansion staining and imaging need to be performed.
NOTE: At this point, the sample should be ~3-4.5-fold larger in each dimension, depending on the tissue type.
6. Post-expansion biomolecule profiling
- Antibody staining
NOTE: This follows a typical immunofluorescence (IF)/immunohistochemistry (I.H.C.) staining protocol. The amounts of primary and secondary antibodies used depend on the concentrations suggested by the manufacturer or by optimization for the specific experiment. Individual buffers may be substituted at the user's discretion.- With the sample in a single well of a 6-well cell culture plate, wash the expanded samples three times for 10 min each time in 1x PBS at R.T., transferring the liquid with a pipette.
- Perform an optional blocking step (for instance, 3% bovine serum albumin/0.1% Triton-X100 in 1x PBS) for at least 1 h at room temperature or overnight at 4 °C.
- Incubate with the primary antibodies in staining buffer (for instance, 0.1% Triton-X100 1x PBS or 9x PBS/1% TritonX/10 mg/L heparin) overnight at 37 °C. Depending on the sample size, 0.5-2 mL should adequately cover the gel.
- Wash the sample three times for at least 10 min each time in 1x PBS at room temperature.
- Incubate in the corresponding secondary antibodies for 3 h at 37 °C in the staining buffer of choice.
- Wash the sample three times for at least 10 min each time in 1x PBS at room temperature.
- NHS-ester pan-protein staining
- With the sample in a 6-well plate, pour 1-2 mL of polyethylene glycol (PEG 200) onto the sample (enough to fully cover it).
NOTE: The tissue should shrink and return to its original pre-expansion size (or close to it). - Stain with fluorophore-conjugated NHS-ester diluted to 13.3-80 µg/mL for 3 h at R.T. in 1-2 mL of staining buffer (for instance, 1x PBS, 2x S.S.C., or 100 mM sodium bicarbonate solution).
- Wash the sample three times for at least 10 min each time in 1x PBS at room temperature.
- With the sample in a 6-well plate, pour 1-2 mL of polyethylene glycol (PEG 200) onto the sample (enough to fully cover it).
- Lipid staining
NOTE: Post-expansion lipid staining is not effective on FFPE clinical samples, as lipids are lost during the fixation process.- Wash the sample three times for at least 10 min each time in 1x PBS at room temperature.
- Apply a conventional lipophilic dye (DiO, DiI, or DiD) diluted 200-fold in 2 mL of 0.1% TritonX-100/1x PBS for 72-96 h at room temperature.
- Wash at least three times with 1x PBS
- Fluorescence in situ hybridization (FISH)
- Place homogenized gel samples in hybridization buffer pre-heated to 60 °C for 30 min.
- Prepare the hybridization buffer for FISH: Combine 2x S.S.C. (300 mM NaCl, 30 mM sodium citrate, pH 7.0), 10% (v/v) dextran sulfate, 20% (v/v) ethylene carbonate, and 0.1% (v/v) Tween20.
- Incubate the gel samples with a hybridization buffer containing 10 pM (per oligo) of FISH probes against the target gene (at least 20 probes per 10 kb) at 45 °C overnight.
- Wash with stringency wash buffer (2x S.S.C. and 0.1% Tween20) two times for 15 min each time at 45 °C.
- Wash with stringency wash buffer another two times for 10 min each time at 37 °C.
- Finally, wash with 1x PBS at least three times for 10 min each time at room temperature.
- The samples are ready for imaging or storage in 1x PBS + 0.02% sodium azide at 4 °C.
- Full tissue expansion and fluorescence imaging
NOTE: The expansion factor attainable with Magnify varies with the tissue type and expansion method (a full summary can be found in Table 3). However, as a general estimate, a Magnify sample will expand 3-4.5-fold in 1x PBS, 5-7-fold in PBS diluted to 1:50 in ddH2O, and 8-11-fold in ddH2O. The optimal expansion factor for imaging depends on each experiment's needs. The expansion factor is highly tunable, such that a single sample may be expanded and shrunk many times for imaging at various scales.- Move the samples expanded 4-fold in PBS to a glass-bottom 6-well imaging plate using a paintbrush. Move any sample expanded further by cutting it into smaller pieces or placing gently into a large glass-bottom imaging plate with a thin piece of flexible plastic.
- Perform fluorescence imaging using a conventional wide-field microscope, a confocal microscope, or another imaging system of choice. Removing excess liquid around the gel with a paper laboratory cloth may prevent gel movement during imaging. The gels can also be immobilized by applying 0.1% poly-L-lysine to the glass surface before imaging.
NOTE: The application of poly-L-lysine will make the gel entirely immobile upon contact. For this reason, caution must be taken that the gel is applied to the glass without folding or stretching. Additionally, the gel should not be allowed to dry completely on poly-L-lysine-coated glass, as this could make removal difficult, and the gel should not be shrunk in PBS while it is still attached to the glass with poly-L-lysine, as this could cause gel or tissue damage. - After imaging, shrink the samples in 1x PBS with at least three washes for 10 min each time, as it is not recommended to store the samples long-term in ddH2O due to the degradation of the fluorescence signal. In particular, fully expanded samples stained with lipophilic dyes should be imaged as close to the time of expansion as possible to prevent dissociation of the dye in water.
- Store the samples in 1x PBS + 0.02% sodium azide at 4 °C.
Representative Results
If the protocol has been successfully completed (Figure 1), the sample will appear clear and flat after heat denaturation; any folding or wrinkling indicates incomplete homogenization. A successfully expanded sample will be 3-4.5-fold larger than before expansion in 1x PBS and 8-11-fold larger when fully expanded in ddH2O. Figure 3 shows example pre- and post-expansion images of 5 µm thick FFPE human kidney sample processed using this protocol and successfully expanded over 8-fold. The tissue was first stained with antibodies for ACTN4 along with DAPI to visualize the nuclear DNA. The specimen was then imaged using a spinning disk confocal microscope (Figure 3A). The tissue was treated following the above protocol, including the post-expansion staining of the same targets, and fully expanded in water (Figure 3B) before re-imaging. Following heat denaturation, biomolecules other than proteins may be stained for and imaged as well, as shown in Figure 4. Lipophilic dyes can be applied to reveal the cell and mitochondria membrane structures in the mouse brain (Figure 4A). Additionally, nucleic acids may be imaged through FISH, as in the FFPE normal human lymph node tissue shown in Figure 4B,C.
Figure 5 demonstrates a likely outcome if the sample is not properly homogenized. Following expansion, the tissue was stained with antibodies for ACTN4 and vimentin, along with DAPI to visualize the nuclear DNA and wheat germ agglutinin (WGA) to label the carbohydrates. The tissue was expanded 3.5-fold in PBS before imaging using a spinning disk confocal microscope. In one kidney section (Figure 5A,C), the crack-free expansion of a glomerulus can be seen. In a separate section (Figure 5B,D), cracking can be clearly seen in the tissue.
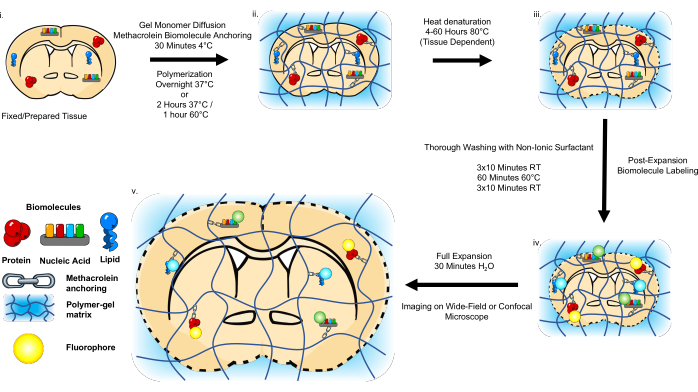
Figure 1: Schematic of the Magnify workflow. The pre-processing of clinically archived tissue slides is first performed based on the storage format. Free-floating paraformaldehyde-fixed sections need only be washed in PBS. The samples are then incubated in a gel monomer solution, including methacrolein to anchor the biomolecules to the hydrogel. In situ polymerization is performed prior to heat denaturation with urea, SDS, and EDTA. The samples are then thoroughly washed and stained using conventional immunostaining protocols, FISH protocols, or lipophilic dyes. The samples are then expanded in pure water before imaging. This figure has been modified from Klimas et al.17. Please click here to view a larger version of this figure.
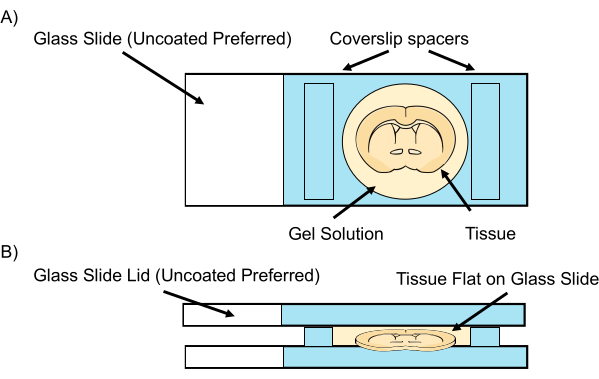
Figure 2: Tissue sample gelation chamber. (A) On each side of the tissue, two spacers, such as two pieces of #1.0 cover glass, are placed before the gel monomer solution is allowed to diffuse at 4 °C. To prevent compression, the spacers should be thicker than the tissue slices. (B) A lid, such as a second glass slide, is used to cover the sample before polymerization at 37 °C. Please click here to view a larger version of this figure.
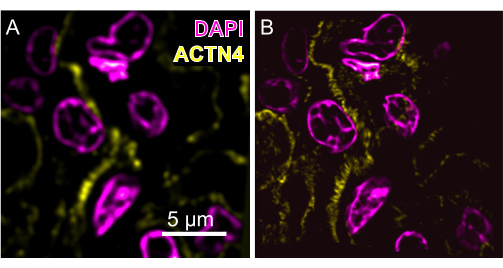
Figure 3: Example pre-expansion images of a human kidney tissue section. (A) An image taken at 60x magnification (1.4 NA) compared to (B) an image of the same field of view post-expansion with Magnify taken at 40x magnification (1.15 NA, expansion factor: 8.15x). Magenta, DAPI; Yellow, ACTN4. The maximum intensity of the post-expansion images is projected over 25 frames. Please click here to view a larger version of this figure.
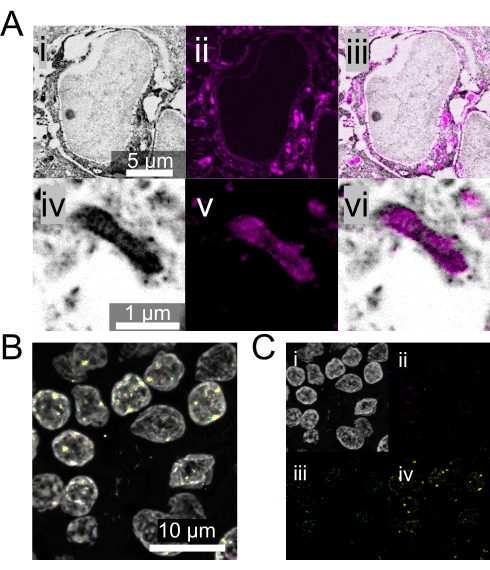
Figure 4: Alternative biomolecule staining strategies with Magnify. (A) (i and iv) NHS-ester pan-protein labeling of fully-expanded mouse brain tissue. (ii and v) Lipophilic dye (DiD) labeling of the same mouse brain tissue. (iii and vi) The channels merged. (B) DNA FISH with Magnify using FFPE normal human lymph node tissue. Expansion factor: 3.5x in 1x PBS. White, DAPI; Magenta, serine/threonine kinase 1 gene; Blue, APC/C activator protein CDH1 gene; Yellow, human satellite sequence S4. (C) Individual channels associated with B. All the images were obtained using a spinning disk confocal microscope at 40x magnification (1.15 NA). Please click here to view a larger version of this figure.
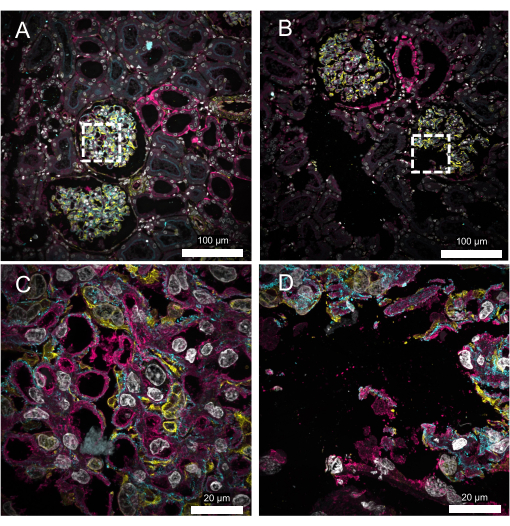
Figure 5: Representative results of 5 µm thick FFPE kidney samples. (A) Crack-free expansion. White, DAPI; Yellow, vimentin; Cyan, alpha-actinin 4; Magenta, wheat germ agglutinin. (B) Expanded kidney section exhibiting cracking. Cracking, distortions, and the loss of labeled targets can be the result of inadequate anchoring and/or homogenization. (C,D) Zoomed-in images of the boxed regions in A and B, respectively. All the images were obtained using a spinning disk confocal microscope at 10x (A,B; 0.45 NA) or 60x (C,D; 1.2 NA) magnification. Please click here to view a larger version of this figure.
Table 1: Gel monomer solution composition. Please click here to download this Table.
Table 2: Homogenization buffer composition. Please click here to download this Table.
Table 3: Summary of the methacrolein and homogenization conditions for validated tissues. Please click here to download this Table.
Discussion
Here, we present the Magnify protocol17, an ExM variant that can retain multiple biomolecules with a single chemical anchor and expand challenging FFPE clinical specimens up to 11-fold with heat denaturation. The key changes in this protocol that distinguish it from other ExM protocols include the use of a reformulated gel that remains mechanically robust even when fully expanded, as well as the use of methacrolein as the biomolecule anchor. The most critical steps in this protocol are as follows: 1) the composition of the final gel solution; 2) the timing of the gelation steps; 3) the setup of the gelation chamber; 4) the parameters for sample homogenization; and 5) the sufficient washing of SDS from the sample before the post-expansion staining.
The most critical parameter for this protocol is the composition of the final gelling solution, particularly the concentration of the biomolecule anchor methacrolein. Different methacrolein concentrations are required to expand different tissue types (Table 3), and care must be taken to ensure this value is well-matched to the sample to be expanded. Over-anchoring with methacrolein can result in a reduced expansion factor and a loss of epitopes available for post-expansion staining, while under-anchoring, especially in FFPE clinical samples, can result in tissue cracks or distortion. Therefore, the methacrolein concentration must be optimized for unvalidated tissue types, although the optimal concentration will likely fall near or within the range presented here. While we expect that this protocol will work for most tissue types, even those we have not yet validated, it is known not to work for samples containing bone.
Beyond methacrolein, using an incorrect concentration of a critical gel ingredient (4HT or TEMED) can result in the complete failure of the experiment, either due to incomplete or no gelling (excessive 4HT) or premature gelation (excessive TEMED, reduced 4HT, or adding APS before the other components). To prevent premature gelation, it is also necessary to keep the sample and gel at 4°C and exposed to air for 30 min and to only cover the sample and place it at 37°C in a humidified chamber after the diffusion process is complete.
When constructing the gel chamber, including spacers between the two uncoated glass slides is critical, especially for thicker tissue sections such as PFA-fixed mouse brain. A lack of spacers can cause tissue compression, resulting in distorted images and inaccurate data.
The sample homogenization depends on the digestion time with the digestion buffer, as well as the digestion buffer’s temperature, and composition. Inadequate homogenization can cause distortions and reduced expansion factors compared to the reported value for a given tissue type. If incomplete homogenization is suspected, the digestion time can be increased, particularly in the case of thicker tissues.
A simple yet crucial step is the adequate washing-out of the SDS from the sample with a non-ionic surfactant (such as C12E10) after the homogenization step. Any leftover SDS can result in suboptimal or entirely hindered antibody binding. Fortunately, if this is suspected, further washing and the re-application of the antibodies will often be a satisfactory solution.
The protocol provides a cost-effective alternative to current super-resolution imaging and electron microscopy techniques to interrogate nanoscale structures in various tissue samples, including in FFPE clinical specimens. The use of methacrolein and heat denaturation allows the post-expansion profiling of any biomolecules that are preserved during tissue fixation (this precludes the imaging of lipids in FFPE samples, for instance). Our protocol, as an extension of the ExM framework, is modular and likely compatible with other techniques such as optical super-resolution methods (STED18, STORM19) or iterative expansion microscopy (iExM)15. However, these have yet to be tested with the presented protocol, and the large expansion factors may present challenges, especially with fluorophore dilution. Additionally, the large size of the samples after full expansion in water necessitates care when handling (although the gel used here is more resilient than previous high-expansion factor ExM gel formulations, such as ten-fold robust expansion microscopy (TREx), to handling miscues17), and creative solutions are occasionally required to transfer and image these fully expanded gels. For instance, using wide-based implements such as a thin sheet of plastic to transfer the fully expanded gels rather than a paintbrush, or using custom-made large imaging plates (in our hands, this means laser-cut or 3D-printed plates with large pieces of #1.5 coverglass adhered to the bottom; these can be seen in the accompanying video). Most importantly, this method broadens the applicability of nanoscale imaging by allowing the nanoscale imaging of common biological and pathological sample preparations on conventional wide-field or confocal microscopes.
Declarações
The authors have nothing to disclose.
Acknowledgements
This work was supported by Carnegie Mellon University and the D.S.F. Charitable Foundation (Y.Z. and X.R.), the National Institutes of Health (N.I.H.) Director's New Innovator Award DP2 OD025926-01, and the Kauffman Foundation.
Materials
| 4-hydroxy-TEMPO (4HT) | Sigma Aldrich | 176141 | Inhibitor |
| 6-well glass-bottom plate (#1.5 coverglass) | Cellvis | P06-1.5H-N | |
| Acrylamide | Sigma Aldrich | A8887 | Gel Monomer component |
| Ammonium persulfate (APS) | Sigma Aldrich | A3678 | Initiatior |
| DAPI (1 mg/mL) | Thermo Scientific | 62248 | |
| Decaethylene glycol mono dodecyl ether (C12E10) | Sigma Aldrich | P9769 | Non-ionic surfactant |
| Diamond knife No. 88 CM | General Tools | 31116 | |
| Ethanol | Pharmco | 111000200 | |
| Ethanol | Pharmco | 111000200 | |
| Ethylenediaminetetraacetic acid (EDTA) 0.5 M |
VWR | BDH7830-1 | Homogenization Buffer Component |
| Forceps | |||
| Glycine | Sigma Aldrich | G8898 | Homogenization Buffer Component |
| Heparin | Sigma Aldrich | H3393 | |
| Methacrolein | Sigma Aldrich | 133035 | Anchoring Agent |
| Micro cover Glass #1 (24x60mm) | VWR | 48393 106 | |
| Micro cover Glass #1.5 (24x60mm) | VWR | 48393 251 | |
| N,N,N′,N′- Tetramethylethylenediamine (TEMED) |
Sigma Aldrich | T9281 | Accelerator |
| N,N′-Methylenebisacrylamide (Bis) | Sigma Aldrich | M7279 | Gel Monomer component |
| N,N-dimethylacrylamide (DMAA) | Sigma Aldrich | 274135 | Gel Monomer component |
| Nunclon 4-Well x 5 mL MultiDish Cell Culture Dish | Thermo Fisher | 167063 | |
| Nunclon 6-Well Cell Culture Dish | Thermo Fisher | 140675 | |
| Nunc™ 15mL Conical | Thermo Fisher | 339651 | |
| Nunc™ 50mL Conical | Thermo Fisher | 339653 | |
| Orbital Shaker | |||
| Paint brush | |||
| pH Meter | |||
| Phosphate Buffered Saline (PBS), 10x Solution | Fischer Scientific | BP399-1 | |
| Polyethylene glycol 200 | Sigma Aldrich | P-3015 | |
| Proteinase K (Molecular Biology Grade) | Thermo Scientific | EO0491 | |
| Razor blade | Fischer Scientifc | 12640 | |
| Safelock Microcentrifuge Tubes 1.5 mL | Thermo Fisher | 3457 | |
| Safelock Microcentrifuge Tubes 2.0 mL | Thermo Fisher | 3459 | |
| Sodium acrylate (SA) | AK Scientific | R624 | Gel Monomer component |
| Sodium azide | Sigma Aldrich | S2002 | |
| Sodium chloride | Sigma Aldrich | S6191 | |
| Sodium citrate tribasic dihydrate | Sigma Aldrich | C8532-1KG | |
| Sodium dodecyl sulfate (SDS) | Sigma Aldrich | L3771 | Homogenization Buffer Component |
| Tris Base | Fischer Scientific | BP152-1 | Homogenization Buffer Component |
| Triton X-100 | Sigma Aldrich | T8787 | |
| Urea | Sigma Aldrich | U5378 | Homogenization Buffer Component |
| Xylenes | Sigma Aldrich | 214736 | |
| 20x SSC | Thermo Scientific | AM9763 | |
| Tween20 | Sigma Aldrich | P1379 | |
| poly-L-lysine | Sigma Aldrich | P8920 |
Referências
- Hell, S. W. Far-field optical nanoscopy. Science. 316 (5828), 1153-1158 (2007).
- Combs, C. A., Shroff, H. Fluorescence microscopy: A concise guide to current imaging methods. Current Protocols in Neuroscience. 79, 1-25 (2017).
- Schermelleh, L., Heintzmann, R., Leonhardt, H. A guide to super-resolution fluorescence microscopy. Journal of Cell Biology. 190 (2), 165-175 (2010).
- Chen, F., Tillberg, P. W., Boyden, E. S. Expansion microscopy. Science. 347 (6221), 543-548 (2015).
- Tillberg, P. W., et al. Protein-retention expansion microscopy of cells and tissues labeled using standard fluorescent proteins and antibodies. Nature Biotechnology. 34 (9), 987-992 (2016).
- Chozinski, T. J., et al. Expansion microscopy with conventional antibodies and fluorescent proteins. Nature Methods. 13 (6), 485-488 (2016).
- Ku, T., et al. Multiplexed and scalable super-resolution imaging of three-dimensional protein localization in size-adjustable tissues. Nature Biotechnology. 34 (9), 973-981 (2016).
- Chen, F., et al. Nanoscale imaging of R.N.A. with expansion microscopy. Nature Methods. 13 (8), 679-684 (2016).
- Tsanov, N., et al. SmiFISH and FISH-quant – A flexible single R.N.A. detection approach with super-resolution capability. Nucleic Acids Research. 44 (22), 165 (2016).
- Wang, G., Moffitt, J. R., Zhuang, X. Multiplexed imaging of high-density libraries of R.N.A.s with MERFISH and expansion microscopy. Scientific Reports. 8 (1), 4847 (2018).
- Wen, G., et al. Evaluation of direct grafting strategies via trivalent anchoring for enabling lipid membrane and cytoskeleton staining in expansion microscopy. ACS Nano. 14 (7), 7860-7867 (2020).
- Sun, D., et al. Click-ExM enables expansion microscopy for all biomolecules. Nature Methods. 18, 107-113 (2021).
- White, B. M., Kumar, P., Conwell, A. N., Wu, K., Baskin, J. M. Lipid expansion microscopy. Journal of the American Chemical Society. 144 (40), 18212-18217 (2022).
- Truckenbrodt, S., et al. X10 expansion microscopy enables 25-nm resolution on conventional microscopes. EMBO Reports. 19 (9), e45836 (2018).
- Chang, J. -. B., et al. Iterative expansion microscopy. Nature Methods. 14 (6), 593-599 (2017).
- Park, J., et al. Epitope-preserving magnified analysis of proteome (eMAP). Science Advances. 7 (46), (2021).
- Klimas, A., et al. Magnify is a universal molecular anchoring strategy for expansion microscopy. Nature Biotechnology. , (2023).
- Gao, M., et al. Expansion stimulated emission depletion microscopy (ExSTED). ACS Nano. 12 (5), 4178-4185 (2018).
- Zwettler, F. U., et al. Molecular resolution imaging by post-labeling expansion single-molecule localization microscopy (Ex-SMLM). Nature Communications. 11, 3388 (2020).

