A 3D Quantification Technique for Liver Fat Fraction Distribution Analysis Using Dixon Magnetic Resonance Imaging
Summary
This study introduces a unique 3D quantification method for liver fat fraction (LFF) distribution using Dixon Magnetic Resonance Imaging (Dixon MRI). LFF maps, derived from in-phase and water-phase images, are integrated with 3D liver contours to differentiate LFF patterns between normal and steatotic livers, enabling precise assessment of liver fat content.
Abstract
This study presents a 3D quantification methodology for the distribution of liver fat fraction (LFF) through the utilization of Dixon MRI image analysis. The central aim is to offer a highly accurate and non-invasive means of evaluating liver fat content. The process involves the acquisition of In-phase and Water-phase images from a Dixon sequence. LFF maps are then meticulously computed voxel by voxel by dividing the Lipid Phase images by the In-phase images. Simultaneously, 3D liver contours are extracted from the In-phase images. These crucial components are seamlessly integrated to construct a comprehensive 3D-LFF distribution model. This technique is not limited to healthy livers but extends to those afflicted by hepatic steatosis. The results obtained demonstrate the remarkable effectiveness of this approach in both visualizing and quantifying liver fat content. It distinctly discerns patterns that differentiate between normal and steatotic livers. By harnessing Dixon MRI to extract the 3D structure of the liver, this method offers precise LFF assessments spanning the entirety of the organ, thereby holding great promise for the diagnosis of hepatic steatosis with remarkable effectiveness.
Introduction
Non-Alcoholic Fatty Liver Disease (NAFLD) encompasses a spectrum of pathological conditions, ranging from the abnormal accumulation of triglycerides in liver cells (hepatic steatosis) to the development of inflammation and damage to liver cells, known as non-alcoholic Steatohepatitis (NASH). In some cases, NAFLD can progress to more severe stages, including fibrosis, cirrhosis, end-stage liver disease, or even Hepatocellular carcinoma (HCC)1. Published data from the World Health Organization and the Global Burden of Disease suggest that approximately 1,235.7 million individuals worldwide are affected by NAFLD across all age groups2. NAFLD currently ranks as one of the most prominent causes of liver-related diseases globally and is expected to become the leading cause of end-stage liver disease in the coming decades3.
The accurate assessment of hepatic steatosis's extent holds substantial importance for precise diagnosis, appropriate treatment selection, and effective disease progression monitoring. The gold standard for assessing liver fat content continues to be liver biopsy. However, due to its invasive nature, the potential for pain, bleeding, and other postoperative complications, it is not a practical option for frequent follow-up examinations4,5,6. Consequently, there is a pressing need for noninvasive imaging techniques that can reliably quantify hepatic fat deposition. Magnetic resonance imaging (MRI) shows promise in this area due to its lack of ionizing radiation and its ability to sensitively detect fat content through chemical shift effects7,8.
Recent studies have outlined MRI techniques for quantifying hepatic fat, based on chemical shift gradient echo methods like Dixon imaging9,10. Nonetheless, the majority of these techniques rely on the analysis of two-dimensional regions of interest. The comprehensive evaluation of the three-dimensional distribution of liver fat fraction (LFF) has remained limited. In the present study, a unique 3D LFF quantification approach is introduced, combining Dixon MRI with liver structural imaging. The resulting 3D LFF model allows for precise visualization and measurement of the distribution of fat content throughout the entire volume of the liver. This technique demonstrates substantial clinical utility for the accurate diagnosis of hepatic steatosis.
Protocol
The study was approved, and the patient was recruited from the Department of Infectious Diseases at Dongzhimen Hospital, Beijing University of Chinese Medicine, in Beijing, China. The patient underwent a routine abdominal Dixon MRI scan after providing informed consent. In this investigation, a 3D distribution modeling approach is employed to reconstruct the liver fat fraction (LFF) in a standard patient with medically diagnosed hepatic steatosis. Furthermore, the study provides a quantitative assessment comparing the patient's liver with a healthy liver. The software tools utilized in this research are listed in the Table of Materials.
1. Preparation and data collection
NOTE: The variance in parameters remains unaffected by the research approach. In this investigation, genuine DICOM data were obtained from clinical imaging. The data were acquired using a MRI apparatus with a field strength of 1.5 Tesla. The dataset comprises four distinct phases derived from the Dixon sequence, specifically In-phase, Out-of-phase, Water, and Fat.
- Check the Dixon sequence for the upper abdomen.
- Copy all DICOM data to a customized working directory.
NOTE: The working directory is the same in both the operating system and MATLAB. - Go to the directory housing the data within MATLAB's current working directory (as shown in Figure 1).
- Confirm that the DIXON data displays a horizontal resolution of 520 pixels by 704 pixels, with a pixel spacing of 0.5682 mm and a slice thickness of 5 mm.
- Copy all DICOM data to a customized working directory.
- Quickly check images of Dixon.
- Change the directory to access folders for different phases, which include In-phase, Out-phase, Water, and Fat phases, each stored in distinct folders for Dixon imaging.
- Utilize the Slice_View function to visualize the images for each phase.
- See Figure 2 to view an image of the MRI-Dixon sequence's interactive graphical user interface (GUI). Employ the scroll bar situated at the bottom of the GUI to efficiently navigate through the various sequences.
- Employ the MRI-Dixon In-phase sequence to enhance descriptions of liver tissue boundaries.
NOTE: In Figure 2, one can find icons located in the upper right corner of the GUI. These icons offer functions such as zooming out, zooming in, returning to the global view, and marking the coordinates of the selected pixel. Make use of the zoom feature to closely examine local lesion features. - Use the Mark pixel coordinates feature to calculate the distance between two points, aiding in the measurement of nodule sizes.
- The initial color bar uses the jet colormap, which signifies a color shift from blue to red, signifying low to high values respectively. To switch to the standard gray colormap and reset the entire GUI, right-click on the Color Bar and select the option from the pop-up menu.
- If the default filter window is not suitable, adjust it by dragging up and down in the middle of the figure using the left mouse button to modify the window level. Dragging left and right adjusts the window width, and the precise filtering range will be shown on the color bar.
NOTE: Subsequent operations will focus on utilizing the DIXON In-phase sequence to represent the 3D region of the liver.
2. Extracting the 3D region of the liver
NOTE: To compute the Liver Fat Fraction (LFF), each voxel within the 3D liver region acts as a spatial carrier, with its fat fraction value obtained from MRI-Dixon data. Before calculating LFF, it's crucial to extract the 3D liver region. Although deep learning methods could achieve this more efficiently, the focus here is on using mature software tools like MIMICS for liver region extraction.
- Open the MIMICS software and choose New Project. In the subsequent dialog box, locate the folder housing the DIXON Out-phase images. Continue by clicking on NEXT, and then hit Convert to enter the sequence-editing mode.
- Generate an empty Mask by clicking on novo within the MASK dialog box situated on the right-hand side, and opt for the maximum threshold.
- Use the Edit Masks tool located under the "Segment" label to delineate the liver area in all horizontal views.
- Create a 3D spatial representation of the liver by choosing the liver mask depicted earlier and clicking on Calculate Part from Mask. The resulting 3D liver region will be presented (refer to Figure 3).
- Navigate to File, then select Export, and choose the DICOM option. In the pop-up dialog box, select the liver mask, specify the file path and names, and then click on OK to export the 3D liver region to the designated DICOM files.
3. Generating Fat Fraction Map (FF-Map)
NOTE: The fat fraction map (FF-Map) has a value range of 0-1. In this study, the FF of each voxel is calculated by dividing the voxel value of In-phase minus Water-only by that of In-phase using Dixon MRI.
- Change the directory to the folder of In-phase images and select the Volume_In function to generate the in-phase Volume.
- Change the directory to the folder of Water-only images and select the Volume_Water function to generate the water-only Volume.
- Select the FF_Volume function using the two Volumes generated in step 3.1 and step 3.2 as inputs to obtain the FF-Volume of the abdominal MRI, visualized as shown in Figure 4.
- The Fat Fraction (FF) map shown in Figure 4 is presented as an RGB true-color image with a data structure of 520 by 704 by 44 matrix. In this representation, each pixel is composed of three values, representing the red, green, and blue channels. Take note of the color bar on the left side, which provides a visual reference for the fat fraction values associated with different colored pixels.
NOTE: The GUI shown in Figure 4 allows closer observation of the patient's liver through the zooming tool in the top right corner and obtaining the exact FF values using the probing tool. By dragging the scroll bar at the bottom, users can observe the FF-maps of all scanning positions in the horizontal plane.
4. 3D-volume of liver fat fraction distribution
NOTE: Figure 4 shows the LFF map calculated based on the Dixon MRI images of the upper abdomen. Combined with the 3D liver region in Figure 3, the 3D-LFF volume of the entire liver can be computed separately.
- Utilize the LFF_Volume function, providing it with the 3D liver region (as depicted in Figure 3) and the Liver stiffness map (shown in Figure 4) as input parameters. This will result in the creation of the 3D-Volume representation of the liver fat fraction, as illustrated in Figure 5.
- To examine the stiffness map for each liver layer, simply use the scroll bar located beneath the GUI displayed in Figure 5.
NOTE: In this view, only liver tissue is accurately depicted. - Pay attention to the icons located in the upper right corner of the GUI (as shown in Figure 5). These icons provide functionalities for zooming out, zooming in, returning to the global view, and marking the coordinates of the selected pixel.
NOTE: The default color bar utilizes the "jet" colormap, where colors transition from blue to red to represent varying values (measured in units of kPa) from low to high. - Run the LFF_Distribution function using the identical input parameters as "LFF_Volume" to produce the spatial distribution of the 3D Liver Fat Fraction (3D-LFF), as depicted in Figure 6.
5. 3D-LFF quantitative analysis
NOTE: Normal liver voxels: LFF < 5%; Mild fatty liver voxels: 5%-10%; Moderate fatty liver voxels: 10%-20%; Severe fatty liver voxels: LFF ≥ 20%11,12,13,14,15. A key quantitative focus of this study is determining the proportion of voxels at different LFF stages in the patient's liver. Figure 6 demonstrates the uneven spatial distribution of liver fat fraction in the patient. The lack of distinct clinical symptoms is primarily attributed to a substantial proportion of normal liver tissue. Therefore, precise quantification of differences between patients and healthy individuals is imperative. This represents a vital quantitative concept herein.
- Calculate the 3D-LFF distribution for a healthy liver by repeating steps 1-4.
NOTE: The built-in MATLAB bar function enables the comparison of the 3D-LFF distributions for healthy and fatty livers (Figure 7).
Representative Results
This investigation utilizes actual patient datasets acquired using a commercially available MRI scanner to validate the 3D liver fat fraction quantification methodology (Figure 1). The MRI protocol included Dixon's four-phase imaging9,10: In-phase, Out-of-phase, Water-only, and Fat-only (Figure 2). The fat fraction (FF) of each voxel is computed by dividing the In-phase minus Water-only voxel signal by the In-phase voxel signal using Dixon MRI. This numeric model allows accurate calculation of the fat content percentage in each voxel.
Although deep learning methods can extract 3D liver anatomy, they have inherent algorithmic errors. To ensure precise quantification, mature software tools like MIMICS were used to extract an accurate 3D liver contour combined with expert guidance (Figure 3).
Fusing the 3D liver contour with the 2D FF map in Figure 4 generates an integrated 3D-FF distribution model in Figure 5. This overcomes the limitation of 2D FF maps and provides visualization of fat deposition in the entire liver volume. Doctors can now accurately locate fat content in 3D liver space instead of a vague impression.
As Figure 6 shows, the 3D-FF distribution reveals fat fraction values at different liver positions. By comparing this to standard FF thresholds, the percentage of voxels falling into different stages of hepatic steatosis can be quantified. This enables precise measurement of the proportion of liver at various steatosis levels.
Comparison between a normal and fatty liver (Figure 7) validates the technique's ability to discern different 3D-LFF distribution patterns. The proposed workflow demonstrates clinical value in the 3D visualization, quantification, and diagnosis of hepatic steatosis based on patient's Dixon MRI data.
Figure 1: MRI-Dixon sequence folders. A list of folder names corresponding to the Dixon MRI scan sequences used in the study. Please click here to view a larger version of this figure.
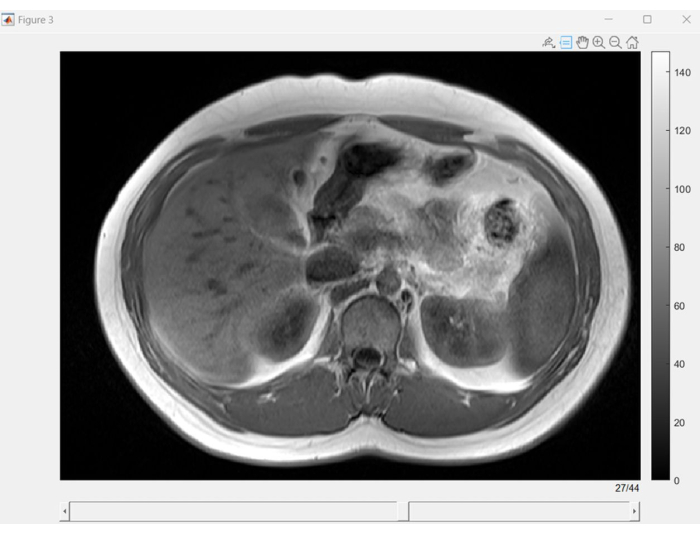
Figure 2: Dixon MRI slice browser. Graphical user interface (GUI) displaying slices from each phase sequence of Dixon MRI. Dixon MRI is valuable for enhancing image quality and interpretability, especially when precise fat and water separation is essential. Please click here to view a larger version of this figure.
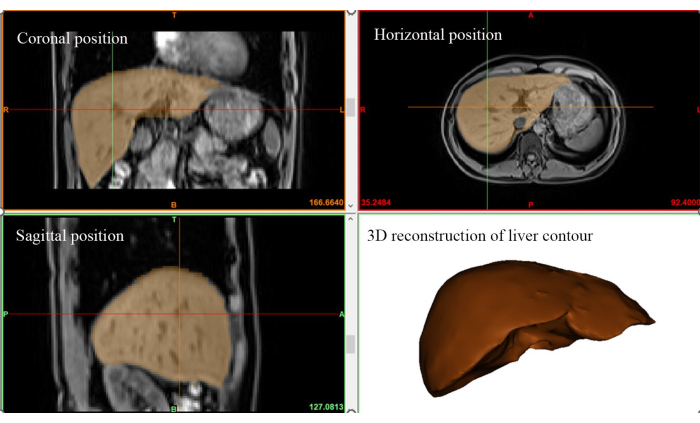
Figure 3: 3D liver region extraction. Visualization of the three-dimensional spatial extent of the liver based on In-Phase images acquired during the MRI scan. Please click here to view a larger version of this figure.
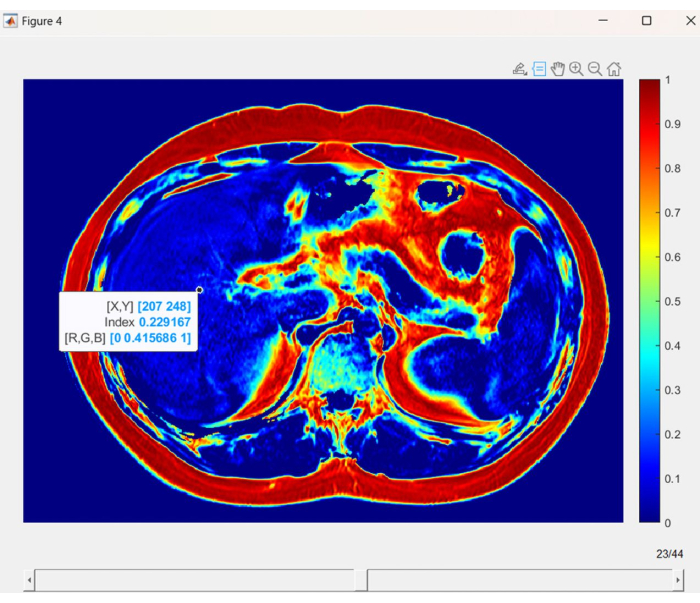
Figure 4: Liver fat fraction map. A visual representation of liver fat fraction (LFF) in each voxel, using distinct colors to indicate variations in fat content. Please click here to view a larger version of this figure.
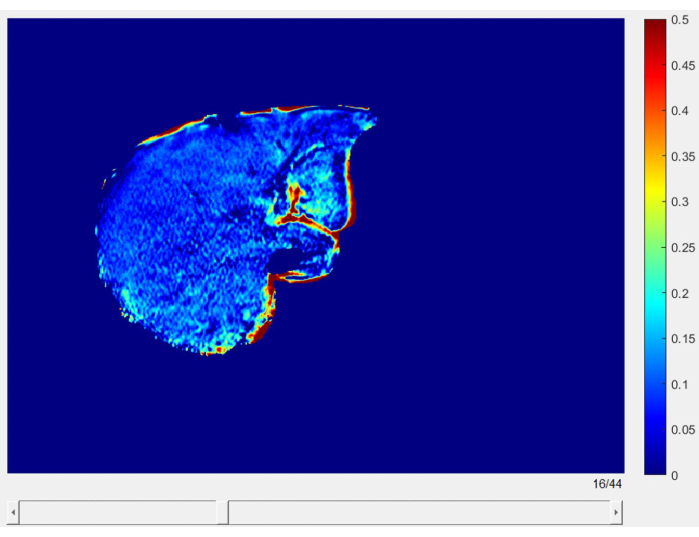
Figure 5: Liver fat fraction slices. High-resolution slices displaying the liver fat fraction map, providing a detailed view of LFF distribution throughout the entire liver. Please click here to view a larger version of this figure.
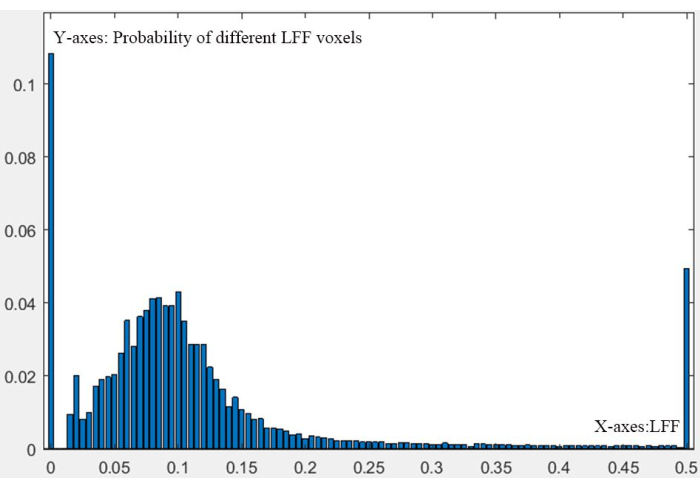
Figure 6: Whole liver 3D-LFF distribution. A figure depicting the numeric probability distribution of liver fat fraction (LFF) across the entire liver, presented in a three-dimensional format. Please click here to view a larger version of this figure.
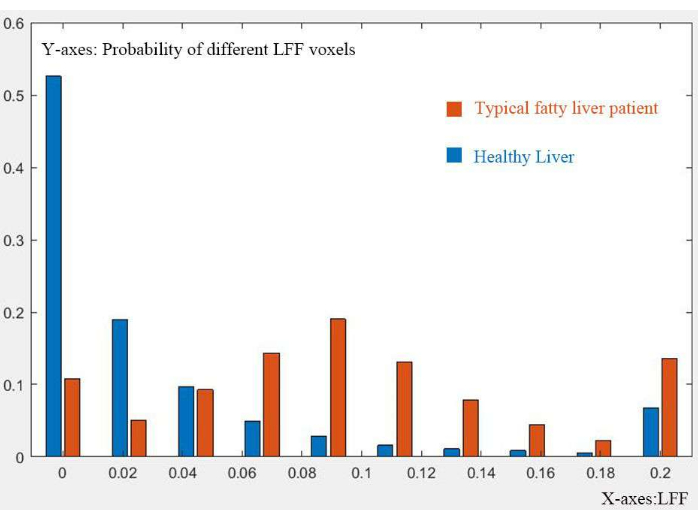
Figure 7: Comparison of 3D-LFF distribution. Comparison of the 3D-LFF distribution between a healthy liver and a typical fatty liver, highlighting differences in fat content and distribution. Please click here to view a larger version of this figure.
Discussion
This research presents an innovative 3D quantification technique for analyzing the distribution of liver fat fraction (LFF) using Dixon MRI9,10. By integrating LFF maps, which are generated from in-phase and water-phase images, with 3D liver contours, this method distinguishes between LFF patterns in normal and steatotic livers6. Consequently, it facilitates a precise evaluation of liver fat content.
Step 3 represents a vital stage in calculating the FF map to quantify fat content in each voxel. Step 4 integrates the FF data with the 3D liver contour to construct an integrated 3D-LFF distribution model. Step 5 validates the efficacy of the 3D-LFF approach for accurate quantification of hepatic steatosis13.
Regarding future modifications, machine vision could enhance the efficiency of 3D liver segmentation. Compiling an atlas of 3D-LFF distributions for healthy livers and different steatosis grades could facilitate clinical diagnosis and typing.
One limitation is that while the method can quantitatively stage early steatosis, it does not elucidate the mechanisms underlying disease progression. Variations in equipment and protocols may lead to inconsistent outcomes. Standardizing the computational workflow remains an ongoing challenge.
This technique introduces and implements the concept of 3D-LFF distribution, providing clinicians with comprehensive insights into fat deposition patterns and disease severity across the entire liver organ. This holds significant significance for precise diagnosis, treatment decisions, and monitoring of therapeutic efficacy. The approach also bears importance for health screening and prevention in the general population.
The method exhibits vast potential in multiple research domains, including: (1) large-scale validation of the technique across heterogeneous cohorts; (2) investigation of 3D-LFF variations among different steatosis etiologies; (3) correlation of 3D-LFF distribution with clinical parameters and risk factors; (4) applying 3D-LFF patterns to build diagnostic, prognostic, and treatment response models; (5) comparing 3D quantification with two-dimensional imaging assessments. Numerous research avenues exist to translate this methodology into clinical utility.
Declarações
The authors have nothing to disclose.
Acknowledgements
This publication received support from the fifth national program for the identification of outstanding clinical talents in traditional Chinese medicine, organized by the National Administration of Traditional Chinese Medicine. The official network link is'http://www.natcm.gov.cn/renjiaosi/zhengcewenjian/2021-11-04/23082.html.
Materials
| MATLAB | MathWorks | 2022B | Computing and visualization |
| Mimics | Materialise | Mimics Research V20 | Model format transformation |
| Tools for 3D_LFF | Intelligent Entropy | HepaticSteatosis V1.0 | Beijing Intelligent Entropy Science & Technology Co Ltd. Modeling for CT/MRI fusion |
Referências
- Pouwels, S., et al. Non-alcoholic fatty liver disease (NAFLD): a review of pathophysiology, clinical management and effects of weight loss. BMC Endocr Disord. 22 (1), 63 (2022).
- Younossi, Z. M., et al. The global burden of liver disease. Clin Gastroenterol Hepatol. 21 (8), 1978-1991 (2023).
- Younossi, Z., et al. Global burden of NAFLD and NASH. Trends, predictions, risk factors and prevention. Nat Rev Gastroenterol Hepatol. 15 (1), 11-20 (2018).
- Boyum, J. H., et al. Incidence and risk factors for adverse events related to image-guided liver biopsy. Mayo Clin Proc. 91 (3), 329-335 (2016).
- Khalifa, A., Rockey, D. C. The utility of liver biopsy in 2020. Curr Opin Gastroenterol. 36 (3), 184-191 (2020).
- Sumida, Y., Nakajima, A., Itoh, Y. Limitations of liver biopsy and non-invasive diagnostic tests for the diagnosis of nonalcoholic fatty liver disease/nonalcoholic steatohepatitis. World J Gastroenterol. 20 (2), 475-485 (2014).
- Ajmera, V., Loomba, R. Imaging biomarkers of NAFLD, NASH, and fibrosis. Mol Metab. 50, 101167 (2021).
- Castera, L., Friedrich-Rust, M., Loomba, R. Noninvasive assessment of liver disease in patients with nonalcoholic fatty liver disease. Gastroenterology. 156 (5), 1264-1281 (2019).
- Jiang, Y., et al. Application of multi-echo Dixon and MRS in quantifying hepatic fat content and staging liver fibrosis. Sci Rep. 13 (1), 12555 (2023).
- Yang, Y., et al. The accuracy and clinical relevance of the multi-echo dixon technique for evaluating changes to hepatic steatosis in patients with non-alcoholic fatty liver disease treated with formulated food. Magn Reson Med Sci. 22 (2), 263-271 (2023).
- Peng, H., et al. Fat-water separation based on Transition REgion Extraction (TREE). Magn Reason Med. 82 (1), 436-448 (2019).
- Hines, C. D. G., et al. T(1) independent, T(2)(*) corrected chemical shift based fat-water separation with multi-peak fat spectral modeling is an accurate and precise measure of hepatic steatosis. Magn Reason Imaging. 33 (1), 873-881 (2011).
- Tang, A., et al. Nonalcoholic fatty liver disease: MR imaging of liver proton density fat fraction to assess hepatic steatosis. Radiology. 267 (2), 422-431 (2013).
- Caussy, C., Reeder, S. B., Sirlin, C. B., Loomba, R. Noninvasive, quantitative assessment of liver fat by MRI-PDFF as an endpoint in NASH Trials. Hepatology. 68 (2), 763-772 (2018).
- Reeder, S. B., Cruite, I., Hamilton, G., Sirlin, C. B. Quantitative assessment of liver fat with magnetic resonance imaging and spectroscopy. Magn Reson Imaging. 34 (4), 729-749 (2011).

