Optical Photothermal Infrared – Fluorescence In Situ Hybridization (OPTIR-FISH)
Summary
Here, we present a protocol using optical photothermal infrared-fluorescence in situ hybridization (OPTIR-FISH), also known as mid-infrared photothermal-FISH (MIP-FISH), to identify individual cells and understand their metabolism. This methodology can be applied broadly for diverse applications, including mapping cellular metabolism with single-cell resolution.
Abstract
Understanding the metabolic activities of individual cells within complex communities is critical for unraveling their role in human disease. Here, we present a comprehensive protocol for simultaneous cell identification and metabolic analysis with the OPTIR-FISH platform by combining rRNA-tagged FISH probes and isotope-labeled substrates. Fluorescence imaging provides cell identification by the specific binding of rRNA-tagged FISH probes, while OPTIR imaging provides metabolic activities within single cells by isotope-induced red shift on OPTIR spectra. Using bacteria cultured with 13C-glucose as a test bed, the protocol outlines microbial culture with isotopic labeling, fluorescence in situ hybridization (FISH), sample preparation, optimization of the OPTIR-FISH imaging setup, and data acquisition. We also demonstrate how to perform image analysis and interpret spectral data at the single-cell level with high throughput. This protocol's standardized and detailed nature will greatly facilitate its adoption by researchers from diverse backgrounds and disciplines within the broad single-cell metabolism research community.
Introduction
Cellular metabolism stands as a foundational pillar in cell biology, steering many processes that determine cell health, function, and interaction with the environment. Analyzing metabolism at the individual cell level, particularly within the native environments, provides invaluable insights to reveal the heterogeneous and complex activities in biological systems1. This is especially crucial in the study of microorganisms, as many microbes exhibit unique growth requirements or environmental dependencies that challenge traditional cultivation methods2. For instance, some species may require specific nutrient compositions or symbiotic relationships not easily replicated in a laboratory setting, thus rendering them non-culturable by standard techniques3. Furthermore, the extended periods needed for the cultivation of certain species pose significant challenges for microbiological research, often extending beyond practical time frames for study and analysis. To circumvent these limitations, alternative methods such as polymerase chain reaction and fluorescence in situ hybridization (FISH) allow for the identification of microbial species without necessitating cultivation4, which can achieve a more accurate and holistic view of microbial ecosystems. However, these analytical technologies lack the ability to elucidate cellular metabolism. This gap highlights the ongoing challenges in the microbiology field: the concurrent task of differentiating cellular identity and elucidating metabolism at the single-cell level. Advancements in techniques such as imaging mass spectrometry (IMS) coupled with stable isotopes have emerged as powerful tools for single-cell metabolic analysis5. In these experiments, cells were incubated with substrates containing isotopes such as 13C or 15N. The newly anabolized biomolecules carry these isotopes, making them distinguishable on the m/z spectra. However, IMS suffers from expensive instrumentation, complicated sample preparation, relatively low throughput, and expertise required for analyzing the m/z spectra. When combined with molecular marker-specific fluorescence imaging, advancements have been made in elucidating cellular metabolism with increased specificity6. Nevertheless, challenges persist in bridging these two modalities. The difference in resolution between IMS and fluorescence imaging, combined with different operational setups, makes it difficult to align and correlate findings7.
The integration of vibrational spectroscopic imaging with stable isotope labeling offers a novel solution for the study of single-cell metabolism. The incorporation of heavier isotopes slows down the vibration of chemical bonds, leading to red-shifted peaks in the vibrational spectra8. Notably, vibrational imaging provides a spatial resolution comparable to fluorescence imaging, and both metabolic quantification and cell identification can be performed in a single setup, simplifying the image registration and correlation. Our recent work has demonstrated the combination of an advanced vibrational imaging platform: optical photothermal infrared (OPTIR) with fluorescence in situ hybridization (FISH) to probe glucose metabolism in bacterial communities9 (Figure 1). OPTIR is a vibrational spectroscopic microscopy system that harnesses the photothermal effect of mid-IR absorption by detecting the visible light change, which provides sub-micrometer resolution as in optical microscopy but with the additional vibrational spectroscopic information originating from the mid-IR absorption. FISH is a commonly used technique to determine the microorganism identity at the single-cell level. The sequence of oligonucleotides could be designed to target specific 16S sequences of different taxa, and different fluorophores could be attached. Specific hybridization of the designed oligonucleotide probes with the target rRNA leads to strong fluorescence signals of target species within individual cells, and multi-channel fluorescence imaging could be performed to identify multiple species within the population. As both OPTIR and fluorescence imaging are based on optical detection, combining OPTIR and FISH fluorescence is straightforward to implement. The two modalities share the same optical resolution for single-cell analysis and can be switched conveniently without requiring additional alignment or co-registration.
This protocol presents a detailed guide to leveraging the OPTIR-FISH platform for advanced single-cell structure-function analysis. We used the bacterial samples grown in 13C-glucose-containing media as a test bed and quantified de novo protein synthesis from 13C-glucose. In order to demonstrate the ability of the platform to identify cells, we used bacterial mixtures in which each species was labeled using rRNA-targeted FISH probes. This approach facilitated precise single-cell identification of specific bacterial strains, such as Escherichia coli (E. coli) and Bacteroides thetaiotaomicron (B. theta), and their metabolism. This protocol offers researchers a powerful tool for simultaneous metabolic profiling and species identification at the single-cell level, promising to advance our understanding of cellular interactions, physiology, and their roles in complex environments.
Protocol
The use of bacterial specimens in this study is in accordance with the guidelines of the Institutional Review Board (IRB) of Boston University and the National Institute of Health.
NOTE: The general workflow followed in this protocol is summarized in Figure 2.
1. Bacterial culture and isotope labeling (Figure 2A)
NOTE: The example given here is for labeling a pure bacterial culture. If applying this protocol to polymicrobial communities, the medium and the labeling time must be adjusted according to the community and the physiology of the organisms of interest.
- Inoculate the bacterial strain of interest from a single colony and preculture in a nutrient-rich medium such as tryptic soy broth (TSB) or Luria-Bertani broth (LB) for 3 h to reach the exponential growth (log) phase. E. coli BW25113 is used as an example strain in this protocol.
NOTE: The time to reach the log phase will vary according to the strain in use. This time needs to be adapted for each strain. - Measure the optical density at 600 nm to estimate the bacterial concentration.
- Prepare a minimal growth medium supplemented with isotopically-labeled substrates. For E. coli BW25113, supplement the M9 minimal medium with 13C-glucose at a final concentration of 0.2% (w/v).
NOTE: The 13C-glucose used (D-glucose U−13C6, 99%) was universally labeled, where all carbon atoms were replaced with 13C atoms. The choice of minimal growth medium will depend on the bacterial strain. The choice of isotope substrate will depend on the specific metabolic process of interest. The overall goal is that the isotopic substrate would serve as the major nutrition source compared with their unlabeled counterparts. - Dilute bacteria in the minimal growth medium containing the isotopically-labeled substrate to a final concentration of 5 × 105 CFU/mL.
NOTE: Usually, a high dilution ratio like 1:100 is used, and the unlabeled nutrition from the rich medium becomes negligible. Alternatively, to ensure complete removal of the nutrient-rich medium, a wash with 1x phosphate-buffered saline (PBS) could be performed by centrifugation at 14,000 x g for 10 min at 4 °C and resuspension in PBS followed by a second centrifugation and final resuspension in the minimal growth medium. - Grow bacterial samples in an orbital shaking incubator under optimal growth conditions. In general, set the orbital shaking incubator to a speed of 220 rpm and a temperature of 37 °C, and place the culture tubes tilted with a loose cap for aerobic incubation. For E. coli, 24 h of aerobic incubation at 37 °C is usually enough to reach maximum labeling of 13C.
NOTE: The incubation time depends on the metabolic process of interest. Longer incubation time generally leads to higher isotopic labeling. A growth curve measurement under the same condition with an unlabeled substrate could provide insight into determining the appropriate collection time points. - Collect bacterial cells at the desired time points by centrifugation at 14,000 x g for 10 min at 4 °C.
- Remove the supernatant and resuspend in an equal volume of 4% paraformaldehyde (PFA) in PBS. Fix the cells in PFA solution for 10 min at room temperature (RT) or 2 h at 4 °C. For long-term storage, wash the cells twice with PBS, resuspend the cells in 1 mL of 50% (v/v) of PBS and 96% ethanol, and store at -20 °C until further use.
2. Fluorescence in situ hybridization (FISH) (Figure 2B)
- Hybridize E. coli cells with the Gam42a10 oligonucleotide probe labeled with a cyanine5 (Cy5) fluorophore (Gam42a-Cy5) and B. theta cells with the Bac30311 oligonucleotide probe labeled with a cyanine3 (Cy3) fluorophore (Bac303-Cy3), following a standard FISH protocol.
NOTE: Fundamentals of rRNA-FISH for identification: rRNA molecules are ideal targets of FISH as they are ubiquitously distributed and naturally amplified within microbial cells. rRNA also contains both conserved and variable sequence regions so that the phylogenetic depth of the target group could be determined by the degree of conservation of the oligonucleotide probe. By specific binding of the designed probe to the target sequence, the tagged fluorophore shows fluorescent signals for individual cells. - Refer to previously published literature12,13,14 for the detailed protocol for fluorescence in situ hybridization. The steps below briefly describe the standard protocol for microbial cells.
- Dehydration:
- Centrifuge the fixed cells (100 µL) at 14,000 x g for 10 min to pellet, resuspend in 100 µL of 96% analytical grade ethanol, and incubate for 1 min at RT.
- Subsequently, centrifuge the cells again at 14,000 x g for 5 min, remove the ethanol, and let the cell pellet dry in air.
- Hybridization: Hybridize the cells in solution (100 µL) for 3 h at 46 °C.
NOTE: The hybridization buffer consists of 900 mM NaCl, 20 mM Tris-(hydroxymethyl)-amino methane HCl, 1 mM ethylenediamine tetraacetic acid, 0.01% sodium dodecylsulphate, 100 ng of the respective fluorescently labeled oligonucleotide and required formamide concentration to obtain stringent conditions. - Centrifuge: Immediately after hybridization, transfer the samples into a centrifuge with a rotor preheated at 46 °C and centrifuge at 14,000 x g for 15 min at the maximum allowed temperature (usually 40 °C).
- Wash and store:
- Wash the samples in a buffer of suitable stringency for 15 min at 48 °C. Then, centrifuge the cells at 14,000 x g for 15 min at the maximum allowed temperature in a centrifuge with a rotor preheated at 46 °C.
- Finally, wash the cells with 500 µL of ice-cold PBS, resuspend them in 20 µL of 1x PBS, and store them at 4 °C until further use.
NOTE: The specific binding of the probe is ensured by the correct hybridization stringency. High stringency could be achieved by increasing hybridization temperature, decreasing salt concentration, or adding denaturant formamide. Formamide interferences with the hydrogen bonds that stabilize nucleic acid duplexes. Here, the temperature and salt concentration are not modified in hybridization while tuning formamide concentration. The optimal formamide concentration for newly designed probes is determined experimentally using formamide dissociation curves10. The required formamide concentration for published probes can be found in probeBase15 or by a literature search. Alternatively, predicted formamide concentrations can be obtained from de–silico analysis using mathFISH16. In the washing step, a slightly higher temperature leads to a slightly more stringent condition, which leads to a higher binding specificity.
- Dehydration:
3. Sample preparation (Figure 2C)
- Slide cleaning and coating:
- Use standardized CaF2 slides (10 mm in diameter and 350 µm thick for compatibility with the instrument sample holder). Immerse the CaF2 in 70% ethanol for 15 min and rinse twice with DI water.
- Transfer the cleaned slides to 0.1% poly-L-lysine solution and incubate overnight at 4 °C. Rinse the slides once with DI water and dry in air.
- Thaw the prepared cell samples at RT while protected from light.
- To avoid excessive salt crystals from PBS during sample drying, follow steps 3.3.1-3.3.2.
- Wash the cells twice with deionized water (DI). Centrifuge at 14,000 x g for 10 min and remove the supernatant.
- Add back the original volume of DI water and vortex gently. Repeat the centrifugation and add the DI water wash again.
- (Optional) For mixture samples, transfer the same volume from each sample and mix into one centrifuge tube.
- Concentrate the bacterial cell samples by centrifugation at 14,000 x g for 10 min and remove the supernatant.
- Gently vortex the remaining solution for ~30 s and add 1-2 µL of solution on the prepared CaF2 slide. Air dry the sample for 30 min while protected from light.
NOTE: A small round dot should appear on the slide after air drying. Due to the "coffee ring" effect, the edges of the dot would have a higher density of bacterial cells. This centrifuge step is to reach an appropriate density for efficient single-cell bacterial imaging. The ideal concentration should lead to a uniform and dense layer of individual bacterial cells under the microscope. If there is a clear pellet that can be seen by the naked eye after centrifuge, remove the supernatant so that after the vortex, the solution looks almost clear. A quick test could be performed on the control sample to estimate the supernatant removal volume. An array of droplets with different amounts of remaining supernatant could be deposited onto the microscope glass slide to examine the optimal dilution volume under the bright-field microscope. If there is no visible pellet after centrifugation, remove most of the supernatant to concentrate the samples to the greatest extent. The trick is positioning the centrifuge tube with the hinge on the outside in a fixed-angle rotor so that the pellet appears on the same side of the tube hinge. Additionally, grade down pipette sizes (P1000, P200, P10) for increased precision during supernatant removal to keep the pellet intact. - Mount the slide in a supplied sample holder and use small dots of polyester tape to secure the slide during measurement.
4. OPTIR-FISH imaging
NOTE: The fluorescence imaging and OPTIR imaging will be performed on the OPTIR system.
- System geometry:
- Use counter-propagation of mid-IR pump and visible probe, backward (epi) detection of visible light with this protocol.
- To focus the visible light and collect back-reflected and scattered visible light, use an air objective (50x 0.8NA). To focus the mid-IR light, use a Cassegrain objective (40x 0.78NA).
- Ensure that the system is also equipped with a widefield epi-fluorescence module with multiple fluorescence filter sets. For the fluorophores used in this protocol, select the green fluorescent protein (GFP) filter set for Cy3 detection and the Alexa Fluor 647 filter set for Cy5 detection.
- Prepare for measurements:
- Power up the system and open the instrument control software. After initialization, toggle the Counterprop and Fluorescence slider to the on position.
- Set the objective to be 50x, and then perform Auto-background to acquire the IR power spectrum. Use the power spectrum to normalize acquired sample spectra or normalize OPTIR images at different wavenumbers. This will exclude the effect of mid-IR laser power differences on data analysis and quantification.
NOTE: Acquiring IR power spectra for normalization is imperative for the downstream quantitative analysis as the measured intensity is linearly related to the IR power. After performing Auto-background, the software will normalize the OPTIR spectra and OPTIR images to the IR power spectra automatically for users. Purging the system with N2 can minimize the influence of water vapor and increase reproducibility between experiments.
- To load the sample, use the Unload button to move the sample stage out. Use a low magnification objective such as 10x to locate the sample area and navigate to the region that has optimal sample density. Focus the sample by adjusting the objective height in the control software. Change the objective selection to 50x.
- Image acquisition
NOTE: The samples used are: i) E. coli (13C-glucose incubated); ii) E. coli (12C-glucose incubated); iii) B. theta (12C-glucose incubated); iv) mixture of E. coli (13C-glucose incubated) and B. theta (12C-glucose incubated).- Bright-field Images (Figure 2D): Bring the sample into focus by looking at the bright field images.
- Fluorescence images (Figure 2E): Change the filter set by clicking the filter name button (GFP for Cy3 and Alexa Fluor 647 for Cy5) to detect fluorophores associated with the designed FISH probes and acquire fluorescence imaging sequentially using the fluorescence module in the software. Adjust the light intensity and exposure time to get adequate signal contrast before bleaching the fluorophores.
NOTE: Since the probe used in OPTIR measurements may photobleach fluorophores, always acquire fluorescence images before OPTIR images. For concern about the effect of fluorescent markers on OPTIR detection, refer to a previously published work that demonstrated that FISH with Cy5 fluorophore does not affect the following quantification based on OPTIR images as Cy5 is small in size and low in abundance compared to target bacterial protein9. - OPTIR images (Figure 2F):
- Determine the normal peak and isotope-induced red-shifted peak position. In the case of protein synthesis from 13C-glucose, the normal amide I peak is detected at around 1656 cm-1, and the red-shifted peak is at around 1612 cm-1 (Figure 3A).
NOTE: The normal peak position depends on the biochemical component and/or metabolic product of interest. The shifted peak position also depends on which isotope is used for labeling. These two wavenumbers could also be experimentally determined by measuring the complete spectra of unlabeled and maximally labeled cells (see step 4.5) - Tune the IR pump wavelength to 1656 cm-1 using the software. Modify the probe power and detector gain accordingly to avoid saturation of the DC signal (8 V). Fine-tune the bottom Cassegrain objective height to maximize the AC signal.
NOTE: Typical IR power at the sample is around 5 mW, and the probe power at the sample is around 3-15 mW, depending on the sample. Both the IR and probe power can be adjusted using the control software to optimize the power level to ensure high image contrast while avoiding damaging the cells. - Set OPTIR images to contain 200 x 200 pixels and set the scan rate to 100 µm/s. Set the step size in the X and Y directions to 150 nm so the image size is about 30 x 30 μm2.
- Acquire the image.
- Tune the IR pump wavelength to 1612 cm-1 and acquire an image from the same field of view at this wavenumber.
- For each sample, image at least three fields of view for statistical analysis.
- Determine the normal peak and isotope-induced red-shifted peak position. In the case of protein synthesis from 13C-glucose, the normal amide I peak is detected at around 1656 cm-1, and the red-shifted peak is at around 1612 cm-1 (Figure 3A).
- (Optional) Spectral measurement: After image acquisition, if a spatial feature of interest is located, perform the spectral measurement.
- Select locations: Choose a single point or specify an array of points for analysis in the Spectral module of the software.
- Set spectral scanning parameter. Select the wavenumber coverage range, scan speed, and data output spacing based on requirements.
NOTE: if the scan speed and data output spacing are different compared to the ones used in the background acquisition step (step 4.2), a new background based on current spectral scanning parameters is needed for correct normalization. - Set the co-averaging number and start spectral measurement. Increasing co-averaging also increases the signal-to-noise of spectra but requires longer acquisition times.
NOTE: If significant fluctuation is observed during co-averaging, it may indicate laser-heating-induced sample instability. This can often be addressed by lowering the IR power and/or lowering the visible probe power.
5. Data processing and analysis at the single-cell level
- Quantify metabolic synthesis from isotopic substrates (Figure 2G)
NOTE: The following quantification of metabolic incorporation from isotopic substrates is based on two-wavenumber OPTIR images corresponding to the normal and shifted peaks of the target macromolecules. Compared to a full hyperspectral stack, two-wavenumber imaging could greatly reduce the time required for each FOV and improve the throughput while fully recapturing the isotopic effect critical for metabolism quantification.- Subtract the shifted peak OPTIR image from the normal peak OPTIR image generates a visual map of synthesis activities from isotope substrates. As a demonstration (Figure 3C), for E. coli cultured with 13C-glucose, the positive pixels in the (1656 cm-1-1612 cm-1) image indicate active incorporation of 13C into proteins, therefore, a higher protein synthesis level.
NOTE: The subtraction could be performed using the ImageJ Image Calculator function. Alternatively, the subtraction could easily be performed in any preferred programming language for batch processing. - Quantification of single-cell metabolic activity
- Obtain coefficients (represented by h de Figure 2G) from unlabeled and fully labeled reference samples. As the isotope-induced red-shifted peak would overlap partially with the normal peak, both unlabeled and labeled (newly synthesized from the isotopically-labeled substrate) samples will contribute to the OPTIR signals at the normal and shifted peak. Quantify these different contributions using the normalized OPTIR signal intensity at the normal and shifted peak from unlabeled and fully labeled reference samples.
NOTE: For reference samples, unlabeled reference samples are grown (or incubated) without any isotopically labeled substrate. Fully labeled reference samples are usually grown with the highest concentration of isotopically labeled substrate with the longest cultivation time. - For single-cell analysis, obtain single-cell masks based on the bright-field images.
NOTE: Various techniques in image processing could be applied here to generate this mask, including different particle detection like blob analysis and cell segmentation methods like watershed. - Measure the OPTIR signal intensity from individual cells. Apply the acquired single-cell mask to the two OPTIR images (at normal peak and shifted peak), and for each single cell, average the pixel values in the single-cell mask region. The averaged value from individual cells will be used in quantification.
- For each cell, calculate the relative contribution of unlabeled and labeled bio components based on the coefficients from step 5.1.2.1 and signals from step 5.1.2.3.
NOTE: An isotopic replacement ratio is defined as the relative contribution of labeled bio components over the sum of both labeled and unlabeled bio components (Figure 2G). This ratio should fall in the range of 0 to 1. The isotopic replacement ratio is linearly proportional to the 13C-glucose to 12C-glucose ratio (with a fixed total glucose amount)9. A high detection sensitivity of 5% 13C-glucose has been demonstrated for E. coli cells.
- Obtain coefficients (represented by h de Figure 2G) from unlabeled and fully labeled reference samples. As the isotope-induced red-shifted peak would overlap partially with the normal peak, both unlabeled and labeled (newly synthesized from the isotopically-labeled substrate) samples will contribute to the OPTIR signals at the normal and shifted peak. Quantify these different contributions using the normalized OPTIR signal intensity at the normal and shifted peak from unlabeled and fully labeled reference samples.
- Subtract the shifted peak OPTIR image from the normal peak OPTIR image generates a visual map of synthesis activities from isotope substrates. As a demonstration (Figure 3C), for E. coli cultured with 13C-glucose, the positive pixels in the (1656 cm-1-1612 cm-1) image indicate active incorporation of 13C into proteins, therefore, a higher protein synthesis level.
- Identify bacterial species from multi-channel fluorescence images (Figure 2H)
- Merge the multi-color fluorescence channels to create visual guidance for different bacterial species using ImageJ (Image > Color > Merge Channels).
- For single-cell analysis, for each fluorescence channel, generate a set of regions of interest (ROIs) for individual cells.
NOTE: Various techniques in image processing could be applied here to generate the mask, including different auto-thresholding methods and cell segmentation methods.
- Correlate FISH identities and OPTIR metabolic activities (Figure 2I)
- Link the single-cell level bacterial identities from FISH fluorescence images directly to the metabolic activities from OPTIR images by correlating the positions of the ROIs.
- Perform statistical analysis to compare the metabolic activities of different species.
NOTE: Masks are generated separately from fluorescence images and OPTIR images because metabolic activities could be analyzed for all the cells from OPTIR images, while unknown cells would not show up on any of the fluorescence channels. This ensures that all the cells could be included in the metabolic analysis. If there are a large number of unidentified cells, different FISH probes targeting the rRNA of different taxa of interest may need to be considered.
Representative Results
The general workflow for single-cell microbial metabolic analysis with genetic identification by OPTIR-FISH is summarized in Figure 2. The representative results demonstrating the single-cell metabolic imaging capability of OPTIR are shown in Figure 3. This example used E. coli cells incubated with 12C- or 13C-glucose for 24 h. The incorporation of 13C into proteins has been shown to cause a significant red shift of protein amides I and II (Figure 3A)17,18. Therefore, these two key wavenumbers representing the 12C-protein (1656 cm-1, normal amide I band) and 13C-protein (1612 cm-1, shifted amide I band) were selected, and OPTIR images at these two wavenumbers were acquired. The corresponding bright-field images are also shown to demonstrate the morphology of the cells (Figure 3B,E). Rod-shaped bacteria can be clearly resolved at the single cell level in both bright field and OPTIR images, confirming the high spatial resolution of the OPTIR technique (Figure 3B–G). For cells incubated with 12C-glucose (Figure 3B–D), a higher intensity at 1656 cm-1 was observed, while a higher 1612 cm-1 intensity was observed for cells incubated with 13C-glucose (Figure 3E–G). This peak shift indicates the incorporation of heavier carbon isotopes into protein mass17. The evaluation was extended to assess OPTIR-FISH's capability in discriminating bacterial taxa and their metabolic activities in multi-species samples. We artificially mixed two bacterial species that are prevalent in the human gut microbiome: E. coli and B. thetaiotaomicron (B. theta). E. coli cells incubated with 13C-glucose were hybridized with a Gam42a-Cy5 probe, targeting the 23S rRNA of Gammaproteobacteria10. B. theta cells incubated with 12C-glucose were hybridized with the Bac303-Cy3 probe, targeting the 16S rRNA of most Bacteroidaceae species11. It is very hard to differentiate the two species based on the bright-field images since they are both rod-shaped (Figure 4A). Thus, multi-color fluorescence imaging of hybridized probes was essential to assign a taxonomic identity to each analyzed bacterial cell (Figure 4B). Upon subtracting OPTIR images acquired at 1612 cm-1 from that of 1656 cm-1, we noticed that a segment of cells in this bi-species sample displayed positive subtraction values (Figure 4C), indicating the incorporation of 13C isotopes. Based on the culture conditions, we assigned the cells with positive subtraction value to be E. coli, and negative subtraction value to be B. theta. The assignment of bacterial species is confirmed by the fluorescence imaging results: the cells producing positive subtraction values produce Cy5 contrast, which is specific to E. coli, and the cells producing negative subtraction values produce B. theta-specific Cy3 contrasts. Then, the isotopic replacement ratio was calculated, in this case, newly synthesized 13C-protein from 13C-glucose (Figure 4D). As expected, a significant difference in isotopic replacement ratio between the two bacterial species (pairwise t-test, p = 9.74 x 10-33) was observed. This dataset underscores the feasibility of applying the OPTIR-FISH platform to study metabolism in a complex environment where multiple species of samples are present.
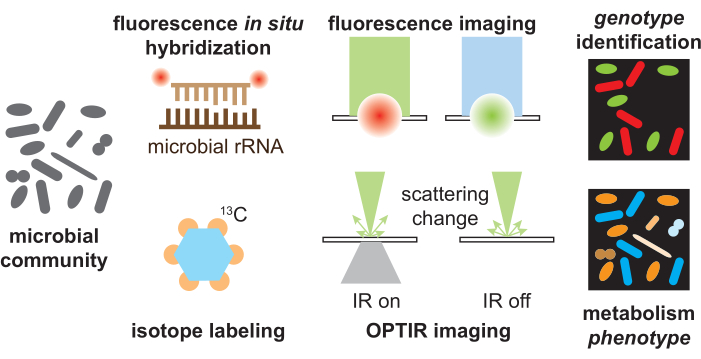
Figure 1: Schematic for the OPTIR-FISH platform for simultaneous species identification and metabolism analysis. Different members of the microbial community could be identified by specific binding of the microbial ribosomal RNA probes through fluorescence in situ hybridization and detected by fluorescence imaging of the attached fluorophores. Metabolic activity could be analyzed by culturing with isotopic substrates and detected by OPTIR imaging. OPTIR imaging detects the scattering change induced by the IR absorption, and OPTIR is inherently compatible with widely used fluorescent tools. Please click here to view a larger version of this figure.
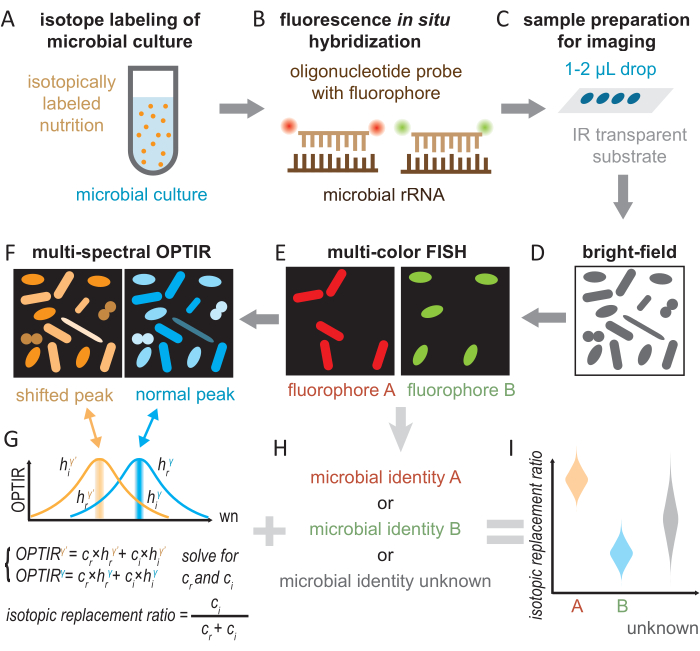
Figure 2: General workflow for single-cell analysis of metabolic activity with genetic identification by OPTIR-FISH. (A) Bacterial cells were cultured with isotopically labeled substrates. (B) Fluorescence in situ hybridization (FISH) was then performed. (C) Sample preparation on IR-transparent slide. (D) Bright-field images identify the regions with optimal cell density. (E,H) Multi-color fluorescence imaging of different FISH fluorophores reveals the genetic identity of individual cells as specified by FISH probes or remains unknown. (F,G) Subsequent multi-spectral OPTIR imaging at the normal and shifted peak reveals the isotopic incorporation into the microbial biomass. Referencing the OPTIR spectra for unlabeled and fully labeled samples (coefficient h), the relative contribution of regular (cr) and isotopic (ci) bio components could be quantified. Protein synthesis from metabolized isotopically-labeled substrates was quantified using the "isotopic replacement ratio". (I) By OPTIR-FISH, a high-throughput analysis of the metabolic activity of microbial species in a complex population can be obtained. Please click here to view a larger version of this figure.
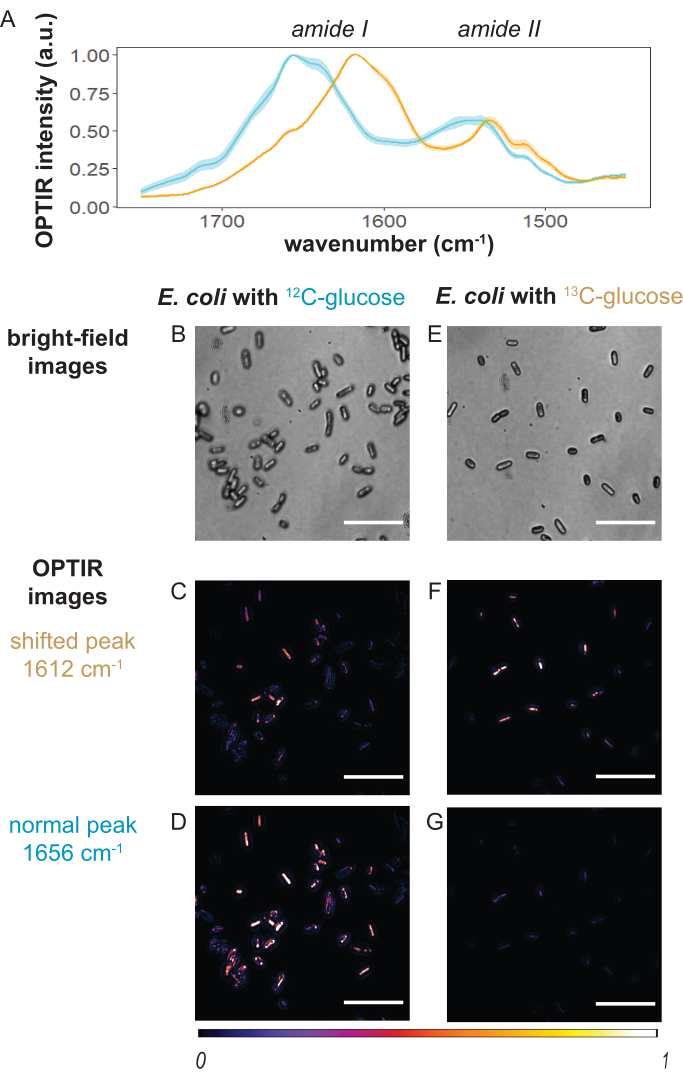
Figure 3: Representative OPTIR spectra and imaging results. (A) OPTIR spectra for E. coli cells cultured with 12C-glucose (cyan) and 13C-glucose (orange) covering the amide I and amide II region. A clear red-shifted amide I peak (1656 cm-1 to 1612 cm-1) is observed for 13C-glucose cultured cells. Representative bright-field and OPTIR images of E. coli cells at normal amide I and shifted amide I band under (B–D) 12C-glucose and (E–G) 13C-glucose culturing conditions. Scale bars: 10 µm. This figure was modified with permission from Bai et al.9. Please click here to view a larger version of this figure.
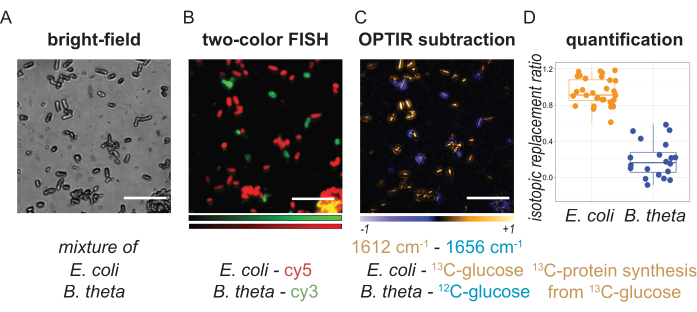
Figure 4: Representative images and quantification results from bacterial mixtures. E. coli cells were incubated with 13C-glucose, followed by hybridization with oligonucleotide probe Gam42a-Cy5. B. theta cells were incubated with 12C-glucose, followed by hybridization with oligonucleotide probe Bac303-Cy3. (A) Bright-field image shows the morphology of the cell mixture. (B) The two-color fluorescence image shows the distribution of E. coli (red) and B. theta (green). (C) OPTIR subtraction result (1612 cm-1-1656 cm-1) shows the protein metabolism of the bacteria cells. (D) Quantification of isotopic replacement ratio representing newly synthesized 13C-protein from 13C-glucose. (Pairwise t-test: p = 9.74 x 10-33). Scale bars: 10 µm. This figure was modified with permission from Bai et al.9. Please click here to view a larger version of this figure.
Discussion
Here, we described a detailed protocol for applying the OPTIR-FISH platform for simultaneous identification of microbial species and quantification of metabolic activities at the single-cell resolution. The critical steps include culture with stable isotope labeling for studying specific metabolic activities and fluorescence in situ hybridization for identifying target microbial species. Multi-channel fluorescence imaging and OPTIR imaging at selected wavenumbers could be performed sequentially on the same microscope. We showcased how to quantitatively analyze these images to reveal the metabolic activity levels of different species within the complex community. The protocol could be especially attractive for the metabolic study of diverse species within the complex community in their native environment.
We used bacteria as an example here, but this protocol can be adapted for other organisms, such as fungi and mammalian cells. One key point is to add the vibrational probes of the target metabolic process in the culture medium free of the normal counterpart. For example, if studying lipid metabolism from fatty acids, the azido-labeled fatty acids should be supplemented to their standard culture medium composed of de-lipid fetal bovine serum (FBS) as the fatty acids originally come from FBS. Additionally, the incubation time also needs to match with the growth rate of different fungi or mammalian cells. This ensures the normal growth of cells while maximizing the labeling efficiency of vibrational probes in the target macromolecules.
The protocol described here can also be adapted to study other metabolic processes beyond protein synthesis from glucose. All major biomolecules, including protein, lipids, nucleic acids, and carbohydrates, could be imaged by OPTIR19,20. Furthermore, there is a vast selection of labeling strategies, including isotopic labeling such as 13C, 15N, 18O, and 2H, and the addition of vibrational tags such as C≡C and C≡N to small molecules21. Due to the small label sizes and high biocompatibility, it is a preferred method for metabolism study compared with fluorescence analogs. To study other metabolic process using different vibrational probes, this protocol can be modified, including cell culture and metabolic labeling steps, as well as the wavenumber selection based on different probes and metabolic products, which can be found in corresponding references21,22,23. Some of the examples include mapping newly synthesized lipids from azide-palmitic acid in human-derived two-dimensional (2D) and three dimensional (3D) culture systems24, imaging newly synthesized protein from azidohomoalanine in macrophage cells23, imaging glucose metabolism with deuterium-glucose in PC3 cells25, and analysis of deuterium incorporation from heavy water as an activity marker in human gut microbiome26.
A potential limitation of the protocol is the detection limit of 13C in complex communities. We have shown that for the pure culture used here, the detection limit is 5% of 13C in total carbon9. We also demonstrate that by mixing fully 13C-labeled E. coli cells with unlabeled human complex gut microbiome samples, we can confidently differentiate the E. coli based on metabolic profiles despite the potential spectral variation associated with different cellular chemical compositions originating from various unlabeled gut microbiome species9. However, complex microbial communities, influenced by the unique physiologies of the microbial species present, can exhibit inconsistent 13C assimilation rates. In such cases, it is worth further testing the metabolic differentiation capability of 13C-labeled and 12C-labeled cells of OPTIR-FISH in the context of a complex community. The performance of the platform can be further improved. For example, by combining the widefield OPTIR setup, the imaging speed can be significantly increased27. Advanced denoising methods based on machine learning can be incorporated into the platform to further boost imaging speed28,29. To better visualize the detailed metabolite distribution in microbial cells, the recently developed super-resolution technologies can be adopted30,31.
Critical steps to this protocol include optimizing the efficiency of FISH labeling for target microbes, which could be achieved by rational design of the oligonucleotide probe, optimization of the formamide concentration, and careful control of the hybridization environment. Preparing samples at an appropriate single-cell density for high-throughput analysis is important for high throughput in practice. It is also critical to optimize the system to get adequate signal for quantification. This is done automatically in the instrument control software during the auto-background step. If there is a deviation in sample preparation, such as using a different type of slide instead of the standard CaF2 slide, manual optimization can be performed in the software. During the manual optimization step, the overlap between mid-IR and visible light is optimized by scanning an array of mid-IR positions. Major challenges include choosing the most suitable vibrational substrates and the corresponding OPTIR wavenumbers to quantify the metabolic activities of interest.
In conclusion, this study demonstrates how metabolic activity and microbial identification could be achieved simultaneously at the single-cell level by the OPTIR-FISH platform. We believe this detailed protocol will provide useful guidance to promote the widespread adoption of this new vibrational imaging platform with high compatibility with widely used fluorescence tools, enabling applications in diverse fields of life science and medicine and unlocking opportunities for new discoveries.
Declarações
The authors have nothing to disclose.
Acknowledgements
This work was supported by the National Institute of Health R35GM136223, R01AI141439 to J.X.C.
Materials
| 96% Ethanol | ThermoScientific | T032021000 | |
| Calcium Fluoride | Crystran | CAFP10-0.35 | |
| D-(+)-Glucose | Sigma-Aldrich | G7021-1KG | |
| D-Gluocose (U-13C6, 99%) | Cambridge Isotopic Laboratories | CLM-1396-1 | |
| Ethylenediaminetetraacetic acid | Sigma-Aldrich | E9884-100G | |
| formamide | ThermoScientific | 17899 | |
| Luria-Bertani broth | Sigma-Aldrich | L3522-250G | |
| M9 Minimal Salts 5x | SIGMA | M6030-1KG | |
| OPTIR instrument | Photothermal Spectroscopy Corp. | mIRage LS | |
| Paraforaldehyde Solution, 4% in PBS | ThermoScientific | J19943-K2 | |
| poly-L-lysine solution 0.1% (w/v) | Sigma-Aldrich | P8920-500ML | |
| Sodium Chloride | Sigma-Aldrich | S9888-25G | |
| Sodium dodecyl sulfate | Sigma-Aldrich | L3771-25G | |
| Tris(hydroxymethyl)aminomethane hydrochloride, 99+% | ThermoScientific | A11379.18 | |
| Trypic Soy Broth | Sigma-Aldrich | 22092-500G |
Referências
- Evers, T. M. J., et al. Deciphering metabolic heterogeneity by single-cell analysis. Anal Chem. 91 (21), 13314-13323 (2019).
- Overmann, J., Abt, B., Sikorski, J. Present and future of culturing bacteria. Annu Rev Microbiol. 71, 711-730 (2017).
- Epstein, S. The phenomenon of microbial uncultivability. Curr Opin Microbiol. 16 (5), 636-642 (2013).
- Su, C., Lei, L., Duan, Y., Zhang, K. -. Q., Yang, J. Culture-independent methods for studying environmental microorganisms: methods, application, and perspective. Appl Microbiol Biotechnol. 93 (3), 993-1003 (2012).
- Musat, N., Foster, R., Vagner, T., Adam, B., Kuypers, M. M. Detecting metabolic activities in single cells, with emphasis on nanoSIMS. FEMS Microbiol Rev. 36 (2), 486-511 (2012).
- Jiang, H., Kilburn, M. R., Decelle, J., Musat, N. NanoSIMS chemical imaging combined with correlative microscopy for biological sample analysis. Curr Opin Biotechnol. 41, 130-135 (2016).
- Porta Siegel, T., et al. Mass spectrometry imaging and integration with other imaging modalities for greater molecular understanding of biological tissues. Mol Imaging Biol. 20 (6), 888-901 (2018).
- Lagunov, O., Drenchev, N., Chakarova, K., Panayotov, D., Hadjiivanov, K. Isotopic labelling in vibrational spectroscopy: a technique to decipher the structure of surface species. Top Catal. 60, 1486-1495 (2017).
- Bai, Y., Guo, Z., Pereira, F. C., Wagner, M., Cheng, J. -. X. Mid-infrared photothermal-fluorescence in situ hybridization for functional analysis and genetic identification of single cells. Anal Chem. 95 (4), 2398-2405 (2023).
- Manz, W., Amann, R., Ludwig, W., Wagner, M., Schleifer, K. -. H. Phylogenetic oligodeoxynucleotide probes for the major subclasses of proteobacteria: problems and solutions. System Appl Microbiol. 15 (4), 593-600 (1992).
- Manz, W., Amann, R., Ludwig, W., Vancanneyt, M., Schleifer, K. -. H. Application of a suite of 16S rRNA-specific oligonucleotide probes designed to investigate bacteria of the phylum cytophaga-flavobacter-bacteroides in the natural environment. Microbiology. 142, 1097-1106 (1996).
- Daims, H., Stoecker, K., Wagner, M. Fluorescence In Situ Hybridization for the Detection of Prokaryotes. Molecular Microbial Ecology. , (2004).
- Bridger, J. M., Volpi, E. V. . Fluorescence In Situ Hybridization (FISH): Protocols and Applications. , (2010).
- Wang, D. O., et al. A quick and simple FISH protocol with hybridization-sensitive fluorescent linear oligodeoxynucleotide probes. RNA. 18 (1), 166-175 (2012).
- Greuter, D., Loy, A., Horn, M., Rattei, T. probeBase-an online resource for rRNA-targeted oligonucleotide probes and primers: new features 2016. Nucleic Acids Res. 44, D586-D589 (2016).
- Yilmaz, L. S., Parnerkar, S., Noguera, D. R. mathFISH, a web tool that uses thermodynamics-based mathematical models for in silico evaluation of oligonucleotide probes for fluorescence in situ hybridization. Appl Environ Microbiol. 77 (3), 1118-1122 (2011).
- Lima, C., Muhamadali, H., Xu, Y., Kansiz, M., Goodacre, R. Imaging isotopically labeled bacteria at the single-cell level using high-resolution optical infrared photothermal spectroscopy. Anal Chem. 93 (6), 3082-3088 (2021).
- Yin, J., et al. Nanosecond-resolution photothermal dynamic imaging via MHZ digitization and match filtering. Nat Commun. 12 (1), 7097 (2021).
- Bai, Y., Yin, J., Cheng, J. -. X. Bond-selective imaging by optically sensing the mid-infrared photothermal effect. Sci Adv. 7 (20), (2021).
- Xia, Q., Yin, J., Guo, Z., Cheng, J. -. X. Mid-infrared photothermal microscopy: principle, instrumentation, and applications. J Phys Chemi B. 126 (43), 8597-8613 (2022).
- Du, J., Wang, H., Wei, L. Bringing vibrational imaging to chemical biology with molecular probes. ACS Chem Biol. 17 (7), 1621-1637 (2022).
- Miao, Y., Shi, L., Hu, F., Min, W. Probe design for super-multiplexed vibrational imaging. Phys Biol. 16 (4), 041003 (2019).
- Shi, L., et al. Mid-infrared metabolic imaging with vibrational probes. Nat Methods. 17 (8), 844-851 (2020).
- Bai, Y., et al. Single-cell mapping of lipid metabolites using an infrared probe in human-derived model systems. Nat Commun. 15 (1), 350 (2024).
- Zhang, L., et al. Spectral tracing of deuterium for imaging glucose metabolism. Nat Biomed Eng. 3 (5), 402-413 (2019).
- Ge, X., et al. SRS-FISH: A high-throughput platform linking microbiome metabolism to identity at the single-cell level. Proc Natl Acad Sci U S A. 119 (26), e2203519119 (2022).
- Bai, Y., et al. Ultrafast chemical imaging by widefield photothermal sensing of infrared absorption. Sci Adv. 5 (7), 7127 (2019).
- Zhang, J., Zhao, J., Lin, H., Tan, Y., Cheng, J. -. X. High-speed chemical imaging by dense-net learning of femtosecond stimulated Raman scattering. J Phys Chem Lett. 11 (20), 8573-8578 (2020).
- Lin, H., et al. Microsecond fingerprint stimulated Raman spectroscopic imaging by ultrafast tuning and spatial-spectral learning. Nat Commun. 12 (1), 3052 (2021).
- Fu, P., et al. Super-resolution imaging of non-fluorescent molecules by photothermal relaxation localization microscopy. Nat Photon. 17, 330-337 (2023).
- Tamamitsu, M., et al. Mid-infrared widefield nanoscopy. arXiv. , (2023).

.
