A Microfluidic Technique to Probe Cell Deformability
Summary
We demonstrate a microfluidics-based assay to measure the timescale for cells to transit through a sequence of micron-scale constrictions.
Abstract
Here we detail the design, fabrication, and use of a microfluidic device to evaluate the deformability of a large number of individual cells in an efficient manner. Typically, data for ~102 cells can be acquired within a 1 hr experiment. An automated image analysis program enables efficient post-experiment analysis of image data, enabling processing to be complete within a few hours. Our device geometry is unique in that cells must deform through a series of micron-scale constrictions, thereby enabling the initial deformation and time-dependent relaxation of individual cells to be assayed. The applicability of this method to human promyelocytic leukemia (HL-60) cells is demonstrated. Driving cells to deform through micron-scale constrictions using pressure-driven flow, we observe that human promyelocytic (HL-60) cells momentarily occlude the first constriction for a median time of 9.3 msec before passaging more quickly through the subsequent constrictions with a median transit time of 4.0 msec per constriction. By contrast, all-trans retinoic acid-treated (neutrophil-type) HL-60 cells occlude the first constriction for only 4.3 msec before passaging through the subsequent constrictions with a median transit time of 3.3 msec. This method can provide insight into the viscoelastic nature of cells, and ultimately reveal the molecular origins of this behavior.
Introduction
Changes in cell shape are critical in numerous biological contexts. For example, erythrocytes and leukocytes deform through capillaries that are smaller than their own diameter1. In metastasis, cancer cells must deform through narrow interstitial gaps as well as tortuous vasculature and lymphatic networks to seed at secondary sites2. To probe the physical behavior of individual cells, microfluidic devices present an ideal platform that can be customized to study a range of cell behaviors including their ability to migrate through narrow gaps3 and to passively deform through micron-scale constrictions3-9. Polydimethylsiloxane (PDMS) microfluidic devices are optically transparent, enabling cell deformations to be visualized using light microscopy and analyzed using basic image processing tools. Moreover, arrays of constrictions can be precisely defined, enabling analysis of multiple cells simultaneously with a throughput that exceeds many existing techniques10,11.
Here we present a detailed experimental protocol for probing cell deformability using the ‘Cell Deformer’ PDMS microfluidic device. The device is designed so that cells passage through sequential constrictions; this geometry is common in physiological contexts, such as the pulmonary capillary bed12. To gauge cell deformability, transit time provides a convenient metric that is easily measured as the time required for an individual cell to transit through a single constriction4,6. To maintain a constant pressure drop across the constricted channels during cell transit, we use pressure-driven flow. Our protocol includes detailed instructions on device design and fabrication, device operation by pressure-driven flow, preparation and imaging of cells, as well as image processing to measure the time for cells to deform through a series of constrictions. We include both device designs and vision data processing code as supplemental files. As a representative sample of data, we show cell transit time through a series of constrictions as a function of the number of constrictions passaged. Analysis of the timescale for cells to transit though narrow constrictions of a microfluidic device can reveal differences in the deformability of a variety of cell types4,5,13. The device demonstrated here uniquely surveys cell transit through a series of micron-scale constrictions; this design emulates the tortuous path that cells experience in circulation and also enables probing additional physical characteristics of the cells such as relaxation time.
Protocol
1. Microfluidic Device Design
NOTE: The device design has four basic functional regions: entry port, cell filter, constriction array, and exit port (Figure 1). The overall design can be applied to a wide array of cell types, with minor adjustments to dimensions. Provided here are a few basic design recommendations along with device parameters that are effective for a selection of both primary and immortalized cells.
- Select the width of constriction array channels (Figure 1B) to be approximately 30-50% of the average cell diameter; this constriction-to-cell size ratio results in significant cell deformation but minimal clogging. Given a cell type that has not previously been tested, it is prudent to design a range of constriction widths from ~30-50% the cell diameter to find the optimum width. As with many types of microfluidic devices, over 120 device master molds can be patterned on a single 4 inch silicon wafer, enabling an array of different device designs to be fabricated in a single fabrication process.
- Select the channel height to be at least 50% of the cell diameter; this ensures the cell is confined in 2 dimensions. To prevent channel collapse during bonding, ensure that the channel aspect ratio is no less than 1:10 (height:width) throughout the device. For channels that must exceed this aspect ratio, add support posts to prevent collapse. Space support posts at a distance that is at least twice the cell diameter as to not impede cell flow.
- Include a filter at the entry ports to remove debris and disaggregate cell clusters (Figure 1B). Adjust the size and spacing of the filter posts so that the distance separating the posts is approximately equal to one cell diameter.
NOTE: The lower limit of feature size is set by the resolution at which the lithography mask is printed. Ensure that all features are within the resolution limit specified by the mask supplier.
2. Supplies and Preparation
NOTE: Before commencing any experiment, the following items must be prepared. A schematic of the entire setup is given in Figure 1.
- Fabricate the device master using standard techniques for lithographic micromachining14,15. Verify the height of the patterned features using a profilometer.
NOTE: While masters can be easily fabricated in a cleanroom facility, some labs have developed inexpensive lithography methods for use in a semi-clean environment16,17. - Prepare the air source for pressure-driven flow, using a tank of compressed air and sequence of air regulators and fittings (Figure 1A).
- Set up an air supply tank and manual regulator.
NOTE: A composition of 5% CO2 in air maintains a similar environment as a typical cell culture incubator, but other gas mixtures can also be used. - Set up an electronically controlled pressure regulator in-line with the manual regulator. Use an electro-pneumatic converter to regulate and maintain a stable pressure drop across the channels, with pressure fluctuations of less than 2 kPa. Use the simple code written in LabVIEW (Supplementary Information) to input the desired pressure.
NOTE: The electro-pneumatic converter uses an internal feedback loop to adjust the valve outlet pressure to match the specified pressure across the microfluidic device. - Set up a pressurized chamber to drive flow of the cell suspension. Assemble a cell suspension chamber out of a standard flow cytometer tube and a machined cap that creates a pressure-tight seal on the tube.
NOTE: The pressure chamber cap contains two orifices: an inlet that connects to the compressed air tank, and an outlet through which cells flow from the pressurized chamber into the device (Figure 2).
- Set up an air supply tank and manual regulator.
- Set up an inverted microscope fitted with a camera that has an acquisition rate of at least 100 frames per sec to capture images of the cells flowing through the constriction array. Interface the camera with a video capture card and a computer.
- Prepare a stock surfactant solution of 10% w/w Pluronic F-127 in phosphate buffered saline (PBS) as adding a small amount of F127 to the cell media solution will help to minimize cell adhesion to the PDMS walls. To prepare the F127 solution, vortex and heat gently at 37 °C to dissolve the F127. Prior to use, pass the solution through a 0.2 µm syringe filter and store at room temperature. Prepare the surfactant solution in advance as vortexing and filtering will generate bubbles that will persist for more than 24 hr.
3. Microfluidic Device Fabrication
- Fabricate PDMS block with microfluidic channels.
- Mix PDMS thoroughly at a ratio of 1:10 (w/w) curing agent to base.
- Pour the mixture over the device master (Step 2.1). Degas in a bell jar with applied vacuum until the entrapped bubbles disappear, or for about 10-20 min.
NOTE: After this time, there may still be bubbles at the air-PDMS interface, which is normal; these will typically dissipate during baking. It is most critical to ensure that there are no remaining bubbles adjacent to the channels. - Bake the degassed device at 65 °C for 4 hr. To prevent channels from collapsing, bake the device overnight to further crosslink the PDMS for a device with a higher elastic modulus that is more resistant to channel collapse. However, note that extended baking also embrittles the PDMS and may lead to cracking when punching holes (Step 3.3).
- Remove microfluidic devices from the master mold. Cut individual microfluidic devices out of the PDMS block with a razor blade and remove the device from the master. Start by gently lifting off one corner of the device; slow and gentle peeling, as opposed to lifting the PDMS block straight up, reduces the stress on both the PDMS and the master.
- Punch holes in the PDMS device to create connection ports for access between tubing and microchannels. Create holes using a biopsy punch, and punch from the channel side through to the exterior side of the device.
NOTE: A biopsy punch with a 0.75 mm bore size creates holes that interface perfectly with polyetheretherketone (PEEK) tubing (outer diameter = 1/32” or 0.79 mm) or polyethylene tubing (PE-20, outer diameter = 0.043” or 1.09 mm). Use a ring light to help visualize holes in devices with shallow channels18. Be sure the biopsy punch is sharp; after repeated use the cutting edge dulls and the biopsy punch should be discarded and replaced. - Rinse the PDMS device with isopropanol (HPLC grade) to remove dust and lodged chunks of PDMS. Make sure the punched holes are clear of debris by directing a steady stream of pure isopropanol from a squeeze bottle through the holes. Blow dry with filtered air. After cleaning, place PDMS blocks with channels face up in a clean Petri dish to maintain device cleanliness. If required, store devices in this form until bonding.
- Clean the glass substrate by rinsing with methanol (HPLC grade), blow dry with filtered air, and place on a 200 °C hotplate for 5-10 min to ensure the glass is completely clean and dry before plasma treatment.
NOTE: The glass substrate forms the bottom of each channel. Typically a glass slide suffices, since a 10X or 20X objective is ideal for imaging multiple channels in parallel. For higher resolution imaging, bond the device to a coverslip (#1.5 thickness). - Bond the PDMS device to the glass substrate.
- Oxygen plasma treat the channel side of the PDMS block and 1 side of a glass slide.
NOTE: With a hand-held discharge unit, use a treatment time of 1 min, but optimize the treatment time for each oxygen plasma apparatus. - Place the channel side of the PDMS on top of the treated side of the glass immediately after treatment.
- Hold together with light pressure for ~10 sec to promote bonding. Bake the entire device at an elevated temperature (60-80 °C) for at least 15 min to improve bond strength.
- Oxygen plasma treat the channel side of the PDMS block and 1 side of a glass slide.
4. Deforming Cells through Constricted Channels
- Inspect the microfluidic device under a microscope. Ensure that the channels are not collapsed or broken and that both inlet and outlet holes connect directly into the device channels by using a low-power objective (10X). Designate defective devices by scoring the surface of the PDMS with a razor blade and do not use them for experiments.
- Prepare the cell culture samples to be assayed. Culture cells using standard conditions.
- For example, culture HL-60 cells in RPMI-1640 medium supplemented with 1% penicillin-streptomycin solution and 10% fetal bovine serum in 5% CO2 incubator at 37 °C.
- For adherent cells, harvest them by trypsinization and resuspend in fresh growth medium. Use a standard trypsinization protocol; for example, add ~0.5 ml trypsin to a flask with 25 cm2 culture area, and resuspend in ~30 ml fresh medium. Differentiate HL-60 cells into neutrophil-type cells by all-trans retinoic (ATRA) at a final concentration of 5 µM per 1 x 105 cells/ml as previously described19,20.
- Centrifuge the cells at 300 x g for 3 min, carefully aspirate the supernatant and resuspend cells to the appropriate final density in fresh culture medium with 0.1% v/v F-127.
- Count the cells using a Coulter counter, hemacytometer, or flow cytometer. Adjust the cell suspension to a density of 1 x 106 to 5 x 106 cells/ml. Once the suspension density is set, supplement with approximately 0.1% v/v F-127.
NOTE: The cell density and surfactant concentration may need to be modified depending on the particular cell systems; Table 1 provides a record of previously used concentrations of cells and F-127 for different cell types. - Place cell suspension in a flow cytometer tube and connect to the pressure cap. Before connecting the tubing to the microfluidic device, adjust the pressure to about 14-21 kPa and flush until the cell suspension emerges from the tip of the tubing. This will help to minimize air bubbles in the tubing and device.
- Insert the tip of the tubing containing the cell suspension into the device inlet. If the device is bonded to a coverslip, place the device on a flat, solid surface such as the microscope stage before connecting the tubing; the coverslip is fragile and may otherwise break.
- Insert a piece of tubing into the exit port and route it into an empty tube for waste collection such as an empty Falcon tube taped to the side of the microscope stage. For consistency in the overall fluidic resistance of the device between experiments, ensure that the inlet and outlet tubing is of the same diameter and approximately the same length for each experiment.
- Gently ramp up the pressure to about 28 kPa or until cells flow through the channels. Position the device so that multiple channels are in the field of view and are perpendicular to the bottom of the screen. If the camera is limited by data size, acquire multiple videos to generate a sufficient number of independent data points. Adjust the frame rate as necessary for the desired data output. To capture changes in cell shape, use high speed imaging at 300 frames per sec (fps); to measure transit times of cells, use frame rates of approximately 100 fps.
5. Data Analysis
- Open the file Master_script.m (Downloadable Supplemental File) and run the program.
- Select the first video to be analyzed from the Windows Explorer window that appears. Specify the frame rate for the selected video and press enter.
NOTE: A figure will appear that prompts the user to select a rectangular cropping window for the first video. - Select a window that encompasses all the channels from left to right and intersects the channels just above the first bulb of the array of channels at the top and bottom (Figure 3).
- Select the constriction regions from the cropped image. Pay special attention to maintain a fixed pixel distance between the top and bottom window boundary for every video selected.
NOTE: The top window edge specifies the topmost constriction and bottom window edge specifies the bottommost constriction (Figure 4); intermediate constrictions are approximated from this user input. Next, the algorithm displays the segmentation of cells for the first 50 frames of each video (Figure 5). - Monitor the top left image (Figure 5A) closely to determine if the algorithm is accurately locating cells; a binarized image is superimposed on the source video to demonstrate the location of identified cells.
NOTE: The different windows, (Figures 5B-5D) are for code diagnosis and demonstrate the video frames after specific image processing operations. - Select the next video to be processed from the Windows Explorer window.
NOTE: The algorithm will repeat steps 5.2-5.5 for each video selected. - Select cancel once all desired videos have been added via the Windows Explorer window to instruct the function that the video list is complete.
NOTE: The algorithm will perform the remaining operations without user intervention. This will take approximately 1-2 hr depending on the number of videos processed and computer performance. Upon completion, the algorithm will produce a histogram of the transit time data and save the data in the directory as a MATLAB .mat file and a Microsoft Excel .xls file.
Representative Results
To investigate the deformability of different cell types, human myeloid leukemia cells (HL-60), differentiated neutrophil cells, mouse lymphocyte cells, and human ovarian cancer cell lines (OVCAR8, HEYA8) are evaluated using the ‘Cell Deformer’ microfluidic technique. Representative results for the transit time of HL-60 and neutrophil-type HL-60 cells show the timescale for a single cell to transit through a series of constrictions, as shown in Figure 6. Transit time is measured for a population of individual cells at each 7 µm constriction in a series of 7 constrictions at a driving pressure of 28 kPa (Figure 6).
As shown in Figure 6, HL-60 cells temporarily occlude the first constriction for a median time of 9.3 msec before passaging through the subsequent constrictions. By contrast, neutrophil-type HL-60 cells occlude the first constriction for only 4.3 msec before passaging. The shorter transit time of the HL-60 cells is consistent with their reduced elastic and viscous moduli, as determined by atomic force microscopy21 and micropipette aspiration22. These results are also consistent with the reduced levels of the mechanoregulating protein, lamin A, which determine transit times of cells through micron-scale gaps23. Once through the first constriction, cells transit more quickly through the remaining constrictions, from 2 to 7, with a median transit time of 4.0 msec for the HL-60 cells and 3.3 msec for the neutrophil-type cells (Figure 6). While the HL-60 cells still exhibit slightly longer but significant transit times (C2-C7, p = 1.7E-9), the distribution of transit times for constrictions 2-7 are nearly identical for each cell type. The observation of the longer transit time required for the first constriction may reflect that the viscoelastic cells do not fully relax to their initial shape on these ~msec timescales. This behavior may also be explained by irreversible structural changes that develop within the cell, and facilitate their transit through the subsequent micron-scale gaps. Importantly, by comparing transit time among cell populations, even for the first constriction, can reveal differences in their deformability23.
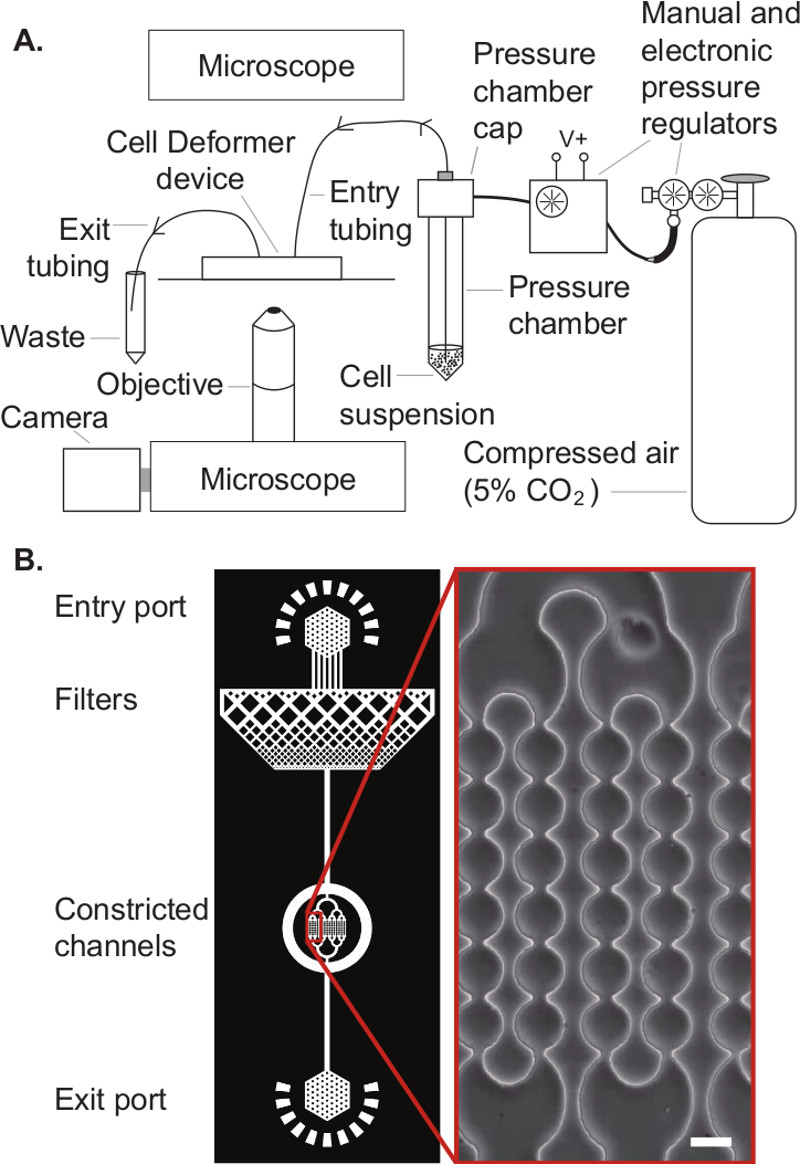
Figure 1. Schematic illustration of the experimental setup. A. ‘Cell Deformer’ device in the experimental setup showing the peripheral connections. B. The device design has 4 functional regions: entry port, cell filter, constriction array, and exit port. Architecture of the microfluidic device showing its main features; inset shows a transmitted light image of the constricted channels. Scale, 10 µm.
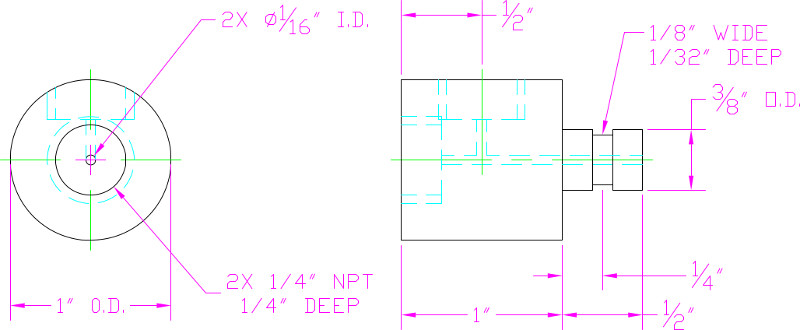
Figure 2. Engineering drawings for a custom cap to pressurize cell media contained in a flow cytometer tube.
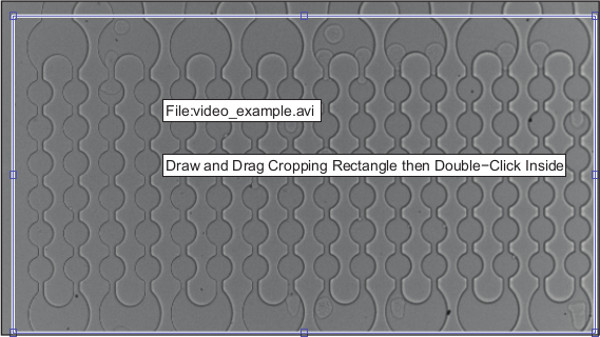

Figure 3. Demonstration of how to select the region of interest for video frame cropping.
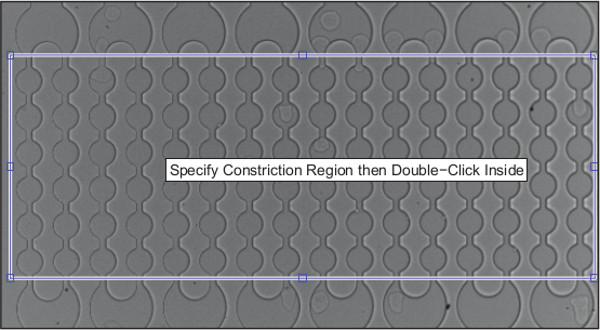

Figure 4. Demonstration of how to select constriction locations.
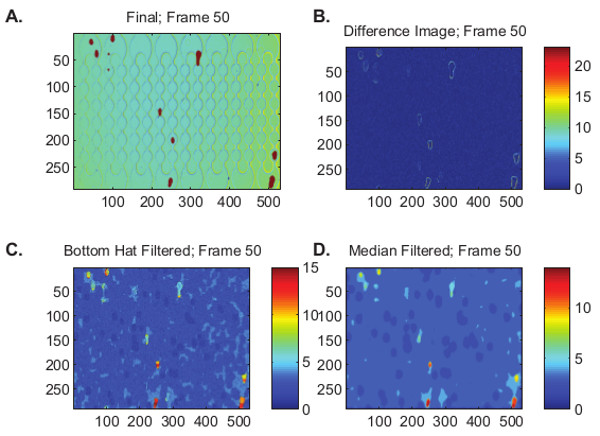
Figure 5. Observation window used to verify that the algorithm properly identifies cell locations as they process through the constriction array. A. Binarized image is superimposed on the source video to show the location of the cells. B. Difference image; C. Bottom hat filtered image; D. Median filtered.
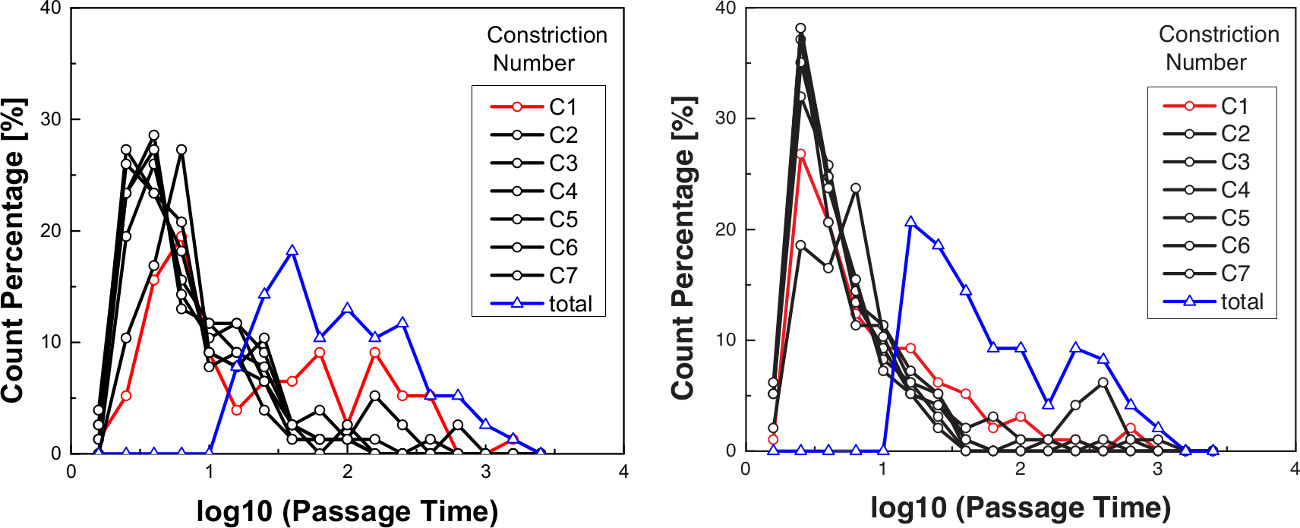
Figure 6. Representative transit time measurements as a function of constriction number. A. HL-60 cells exhibit a longer transit time through the first constriction than B. ATRA-treated HL-60 (neutrophil-type) cells. A comparison of the transformed passage time (log10) through the first constriction as evaluated using the nonparametric Mann-Whitney test reveals the difference is significant, p = 1.47E-5. Cells typically transit through the first constriction more slowly than the subsequent constrictions. Data is shown here for cells transiting through 7 μm-wide constrictions at a driving pressure of 28 kPa. Cell density, mean cell diameter, and surfactant concentration are given in Table 1. HL-60, N = 77; ATRA-treated HL-60, N = 97. Results were replicated in independent experiments over the course of 3 different days.
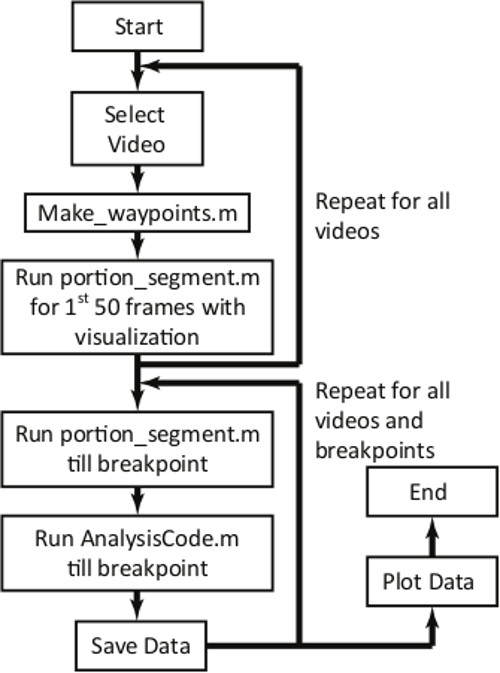
Figure 7. Overview of the MATLAB script for measuring the transit time. The first loop requires user intervention and observation for the first 50 frames for each video. After the first loop, the entire program will run without user intervention and automatically compiles and plots population data.
| Cell Type | Channel Height (µm) | Channel constriction (µm) | Mean Cell Diameter (µm) | Cell Concentration (cells/ml) | Surfactant Concentration (vol%) |
| HL-60 | 10.2 | 5, 7, 9 | 14 | ~1 x 106 | F127: 0.1 |
| Neutrophil-type HL-60 | 10.2 | 5, 7, 9 | 14 | ~1 x 106 | F127: 0.1 |
| OVCAR8 | 10.2 | 7, 9 | 16 | ~5 x 106 | F127: 0.1 |
| HEYA8 | 10.2 | 7, 9 | 17 | ~5 x 106 | F127: 0.1 |
| Mouse lymphocyte | 5.5 | 3, 5 | 8 | ~3 x 106 | F127: 0.33 |
Table 1. Previously studied cell systems and their operating conditions.
Discussion
Here we provide a comprehensive experimental procedure for analyzing the deformation of cells transiting through constricted microfluidic channels using pressure-driven flow. A MATLAB script enables automated data processing (Supplemental Material); an updated version of the code is maintained (www.ibp.ucla.edu/research/rowat). More broadly, the techniques presented here can be adapted in many cell-based microfluidic assays, including the effect of cytoskeletal and nuclear stiffening agents24,23 as well as assaying the deformability of cancer cell types4,5. Taken together with higher resolution microscopy to image subcellular deformations, this method provides a powerful approach to study cell mechanical properties and alterations that occur in physiological and pathological conditions.
Automatic Cell Processing Algorithm
The acquired movies can be analyzed with little user intervention using a MATLAB script. The algorithm is outlined in Figure 7, and the set of MATLAB functions is provided in the Supplementary Information. In brief, the user is instructed to select all the videos to analyze; crop each video for the region of interest; and specify a frame rate for the set of videos; thereafter, the software automatically processes video data to determine the transit time of each cell for each constriction. There are certain requirements for the script to operate: cells should flow from the top of the frame to the bottom of the frame in the video; the microfluidic device position and the lighting conditions should not change appreciably over the duration of the video; and videos should be in .avi format. Importantly, the tracking algorithm includes cells that both enter and passage in the time window of a single video recording, which is approximately 8 sec. Therefore, any objects that have a longer transit time are excluded from the analysis. By performing background subtraction, objects that are present throughout the entire video are removed from the analysis. The code also implements a lower size cutoff, which may be set by the user. The size cutoff for the video analysis was 5 pixels, equivalent to a cell diameter of 2.7 µm.
Since the high-speed video data files can be sizable (~500 MB), the program is computationally intensive. Code is most efficiently run using a desktop computer with at least 6 GB of RAM and a 64-bit chipset with either a local hard drive with at least a 1 TB capacity or interfaced with a large external hard drive or data server with a USB 3.0, FireWire, or Ethernet connection. Data is output in a histogram format for rapid visualization of results; it is also tabulated as data that is stored in a MATLAB (.mat) data file and in an Excel spreadsheet.
The code that is presented provides a simple way to compare cell populations by analysis of the transit time for cells to passage through each constriction. For more detailed analysis, the code could be extended to investigate other parameters including cell size, aspect ratio, as well as relaxation timescale. Previous studies reveal only a weak dependence of cell size on transit time6.
Device-related Pitfalls
Occluded channels are a major hindrance in any flow experiment. Cell adhesion to the device walls will impair the flow of cells through the constricted channels: cells may be observed to smear out along the channel walls. To prevent adhesion, proper surface treatment with F-127 is essential.
The surface properties of PDMS change with time after plasma treatment, which renders the PDMS temporarily hydrophilic25,26. Over the course of one week, hydrophilic PDMS surfaces slowly degenerate to their natural hydrophobic surface energy; this can challenge the removal of air bubbles through the channels. Devices should thus be used within seven days of plasma treatment. Most importantly, the time between plasma treatment and the deformability assay should be consistent across all experiments.
While we have successfully distinguished transit times between distinct cell types using devices fabricated with a glass floor and three PDMS walls, devices can also be fabricated so they have uniform surface properties on all four channel walls27. The PDMS device can be bonded to a glass substrate spincoated with a thin PDMS layer: mix curing agent to base at a ratio of 1:5 (w/w), pour onto the device mold, and cure for 20 min at 65 °C. Meanwhile, spincoat a thin layer of 1:20 (w/w) curing agent to base onto the glass substrate; bake at 85 °C for 4 min. Place the PDMS device on the PDMS-coated slide, and finish baking overnight to complete bonding. This method is also valuable if a plasma machine for glass-PDMS bonding is not available.
Cell-related pitfalls.
hen preparing the cell suspension, the density of cells is critical: if the cell density is too low, cell transit events will be infrequent; if the cell density is too high, there will be multiple cells passaging a single channel simultaneously, the pressure drop across a single cell will not be consistent, and it will be difficult to delineate individual cells with an automated script.
While experiments are typically performed within 30 min of cell suspensions being prepared, cells can settle to the bottom of the pressure chamber over longer times of >30 min. To avoid settling over longer time periods, the pressure chamber and connecting tubing can be periodically rotated. For longer experiments, a small magnetic stir bar can be added to the cell suspension, with a stir plate positioned beneath the pressure chamber to agitate the cells and prevent settling.
Disclosures
The authors have nothing to disclose.
Acknowledgements
The authors would like to acknowledge Lloyd Ung for constructive input in early versions of this technique, Dr. Jeremy Agresti for pressure cap design tips, and Dr. Dongping Qi for his help in fabricating the pressure cap. We are grateful to the laboratories of M. Teitell and P. Gunaratne for providing a variety of cell samples for testing. We are grateful to the National Science Foundation (CAREER Award DBI-1254185), the UCLA Jonsson Comprehensive Cancer Center, and the UCLA Clinical and Translational Science Institute for supporting this work.
Materials
| Name of Material/ Equipment | Company | Catalog Number | Comments/Description |
| Pluronic F-127 Block Copolymer Surfactant | Fisher Scientific | 8409400 | Produced by BASF, also available through Sigma |
| PDMS base and crosslinker | Essex Brownell | DC-184-1.1 | Product commonly named Sylgard 184 Elastomer |
| Oxygen plasma discharge unit | Enercon | Dyne-A-Mite 3D Treater | |
| Biopsy Punch, Harris Uni-Core (0.75 mm) | Ted Pella, Inc. | 15072 | |
| Fingertight Ferrule, 1/32" | Upchurch Scientific | UP-F-113 | |
| Fingertight III Fitting, 10-32 | Upchurch Scientific | UP-F-300X | |
| polyetheretherketone (PEEK) tubing, outer diameter = 1/32"or 0.79 mm | Valco | TPK.515-25M | |
| polyethylene (PE-20) tubing, 0.043" or 1.09 mm | Becton Dickinson | 427406 | |
| Pressure regulator | Airgas or Praxair | ||
| Polyurethane tubing, 5/32” OD | McMaster Carr | 5648K284 | |
| Push-to-connect fittings | McMaster Carr | 5111K91 | |
| Voltage to Pressure (E/P) Electropneumatic Converter | Omega | IP413-020 | |
| 16-bit,250 kS/S, 80 Analog Inputs Multifunction DAQ | National Instruments | NI PCI 6225-779295-01 | |
| Analog Connector Block-Screw Terminal | National Instruments | SCB-68-776844-01 | |
| LabView System Design Software | National Instruments | ||
| Matlab Software | The MathWorks, Inc. | Matlab R2012a | Code requires the Image Processing Toolbox |
| Shielded Cable | National Instruments | SHC68-68 |
References
- Doerschuk, C. M., Beyers, N., Coxson, H. O., Wiggs, B., Hogg, J. C. Comparison of neutrophil and capillary diameters and their relation to neutrophil sequestration in the lung. Journal of applied physiology. 74 (6), 3040-3045 (1993).
- Fidler, I. J. The pathogenesis of cancer metastasis: the `seed and soil’ hypothesis revisited. Nature Reviews Cancer. 3, 453-458 (2003).
- Jowhar, D., Wright, G., Samson, P. C., Wikswo, J. P., Janetopoulos, C. Open access microfluidic device for the study of cell migration during chemotaxis. Integrative biology: quantitative biosciences from nano to macro. 2 (11-12), 648-658 (2010).
- Hou, H. W., Li, Q. S., Lee, G. Y. H., Kumar, A. P., Ong, C. N., Lim, C. T. Deformability study of breast cancer cells using microfluidics. Biomedical microdevices. 11 (3), 557-564 (2009).
- Byun, S., et al. Characterizing deformability and surface friction of cancer cells. Proceedings of the National Academy of Sciences. 110 (19), 7580-7585 (2013).
- Rosenbluth, M. J., Lam, W. A., Fletcher, D. A. Analyzing cell mechanics in hematologic diseases with microfluidic biophysical flow cytometry. Lab on a chip. 8 (7), 1062-1070 (2008).
- Chen, J., et al. Classification of cell types using a microfluidic device for mechanical and electrical measurement on single cells. Lab on a Chip. 11 (18), 3174 (2011).
- Zheng, Y., Shojaei-Baghini, E., Azad, A., Wang, C., Sun, Y. High-throughput biophysical measurement of human red blood cells. Lab on a Chip. 12 (14), 2560 (2012).
- Zheng, Y., Nguyen, J., Wang, C., Sun, Y. Electrical measurement of red blood cell deformability on a microfluidic device. Lab on a Chip. 13 (16), 3275 (2013).
- Hogg, J. C. Neutrophil kinetics and lung injury. Journal of applied physiology. 67 (4), 1249-1295 (1987).
- Hochmuth, R. M. Micropipette aspiration of living cells. Journal of biomechanics. 33 (1), 15-22 (2000).
- Yap, B., Kamm, R. D. Cytoskeletal remodeling and cellular activation during deformation of neutrophils into narrow channels. Journal of applied physiology. 99 (6), 2323-2330 (2005).
- Bow, H., et al. A microfabricated deformability-based flow cytometer with application to malaria. Lab on a Chip. 11 (6), 1065-1073 (2011).
- Qi, D., Hoelzle, D. J., Rowat, A. C. Probing single cells using flow in microfluidic devices. The European Physical Journal Special Topics. 204 (1), 85-101 (2012).
- Doll, J. C., et al. SU-8 force sensing pillar arrays for biological measurements. Lab on a Chip. 9, 1449-1454 (2009).
- Huntington, M. D., Odom, T. W. A Portable, Benchtop Photolithography System Based on a Solid-State Light Source. Small. 7 (22), 3144-3147 (2011).
- Grimes, A., Breslauer, D. N., Long, M., Pegan, J., Lee, L. P., Khine, M. Shrinky-Dink microfluidics: rapid generation of deep and rounded patterns. Lab on a chip. 8 (1), 170-172 (2008).
- Rowat, A. C., Weitz, D. A. Chips & Tips: see where to punch holes easily in a PDMS microfluidic device. Lab on a Chip. 8, 1888-1895 (2008).
- Meyer, P., Kleinschnitz, C. Retinoic Acid Induced Differentiation and Commitment in HL-60 cells. Environmental Health Perspectives. 88, 179-182 (1990).
- Olins, A., Herrmann, H., Lichter, P., Olins, D. E. Retinoic Acid Differentiation of HL-60 Cells Promotes Cytoskeletal Polarization. Experimental Cell Research. 254 (1), 130-142 (2000).
- Rosenbluth, M. J., Lam, W. A., Fletcher, D. A. Force Microscopy of Nonadherent Cells: A Comparison of Leukemia Cell Deformability . Biophysical Journal. 90 (8), 2994-3003 (2006).
- Tsai, M., Waugh, R., Keng, P. Changes in HL-60 cell deformability during differentiation induced by DMSO. Biorheology. 33 (1), 1-15 (1996).
- Rowat, A. C., et al. Nuclear Envelope Composition Determines the Ability of Neutrophil-type Cells to Passage through Micron-scale Constrictions. Journal of Biological Chemistry. 288 (12), 8610-8618 (2013).
- Lam, W. A., Rosenbluth, M. J., Fletcher, D. A. Chemotherapy exposure increases leukemia cell stiffness. Blood. 109 (8), 3505-3508 (2007).
- Bhattacharya, S., Datta, A., Berg, J. M., Gangopadhyay, S. Studies on surface wettability of poly(dimethyl) siloxane (PDMS) and glass under oxgen-plasma treatment and correlation with bond strength. Journal of Microelectromechanical Systems. 14 (3), 590-597 (2005).
- Wu, M. H. Simple poly(dimethylsiloxane) surface modification to control cell adhesion. Surface and Interface Analysis. 41 (1), 11-16 (2009).
- Unger, M. A., Chou, H. P., Thorsen, T., Scherer, A., Quake, S. R. Monolithic Microfabricated Valves and Pumps by Multilayer Soft Lithography. Science. 288 (5463), 113-116 (2000).

