The ChIP-exo Method: Identifying Protein-DNA Interactions with Near Base Pair Precision
Summary
Here, we present a protocol to achieve near base pair resolution of protein-DNA interactions. This is obtained by exonuclease treatment of DNA fragments selectively enriched by chromatin immunoprecipitation (ChIP-exo) followed by high throughput sequencing.
Abstract
Chromatin immunoprecipitation (ChIP) is an indispensable tool in the fields of epigenetics and gene regulation that isolates specific protein-DNA interactions. ChIP coupled to high throughput sequencing (ChIP-seq) is commonly used to determine the genomic location of proteins that interact with chromatin. However, ChIP-seq is hampered by relatively low mapping resolution of several hundred base pairs and high background signal. The ChIP-exo method is a refined version of ChIP-seq that substantially improves upon both resolution and noise. The key distinction of the ChIP-exo methodology is the incorporation of lambda exonuclease digestion in the library preparation workflow to effectively footprint the left and right 5' DNA borders of the protein-DNA crosslink site. The ChIP-exo libraries are then subjected to high throughput sequencing. The resulting data can be leveraged to provide unique and ultra-high resolution insights into the functional organization of the genome. Here, we describe the ChIP-exo method that we have optimized and streamlined for mammalian systems and next-generation sequencing-by-synthesis platform.
Introduction
Chromatin immunoprecipitation (ChIP) is a powerful method to study mechanisms of gene regulation by selectively enriching for DNA fragments that interact with a given protein in living cells. Detection methods of ChIP-enriched DNA fragments have evolved as technology improves, from detection of a single locus (standard ChIP-qPCR) to hybridization on oligonucleotide microarrays (ChIP-chip) to high-throughput sequencing (ChIP-seq) 1. Although ChIP-seq has seen widespread application, chromatin heterogeneity and nonspecific DNA interactions have hampered data quality leading to false positives and imprecise mapping. To circumvent these limitations, Dr. Frank Pugh developed the ChIP-exo method 2. The salient feature of ChIP-exo is that it incorporates a 5' to 3' exonuclease, effectively footprinting transcription factor binding locations. As a result, the ChIP-exo methodology achieves higher resolution, greater dynamic range of detection, and lower background noise.
Although ChIP-exo is more technically challenging to master than ChIP-seq, it is being widely adopted as studies aim to gain unique ultra high-resolution insights using diverse biological systems 3-8. Indeed, ChIP-exo has been successfully applied to bacteria, yeast, mouse, rat, and human cell systems. As proof of principle, ChIP-exo was originally used to identify the precise binding motif for a handful of yeast transcription factors 2. The technique was also used in yeast to study the organization of the transcription pre-initiation complex, and to decipher subnucleosomal structure of various histones 9,10. More recently, we leveraged ChIP-exo to resolve adjacent TFIIB and Pol II binding events at human promoters, and showed that widespread divergent transcription arises from distinct initiation complexes 11.
The workflow presented here is optimized and streamlined for mammalian ChIP-exo (Figure 1). First, living primary or tissue culture cells are treated with formaldehyde to preserve in vivo protein-DNA interactions through a covalent crosslink. The cells are lysed and chromatin sheared to ~ 100 – 500 base pair size fragments. ChIP then selectively enriches for DNA fragments crosslinked to the protein of interest. At this point, ChIP-seq libraries are typically prepared, which inherently limits the detection resolution to the average fragment size of a few hundred base pairs. However, ChIP-exo overcomes this limitation by trimming the left and right 5' DNA borders of the protein-DNA crosslink site with lambda exonuclease. Sequencing libraries are then constructed from exonuclease digested DNA as detailed below. The resulting nested 5' borders represent an in vivo footprint of the protein-DNA interaction (Figure 1, step 14), and are detected by high throughput sequencing. Although the ChIP-exo methodology is more involved than ChIP-seq, transitions between most steps requires simple bead washing, which minimizes sample loss and experimental variability. Importantly, since ChIP-exo is a refined version of ChIP-seq, any sample that is successful with ChIP-seq should also be successful with ChIP-exo.
The in vivo footprinting of protein-DNA interactions with ChIP-exo results in a fundamentally distinct data structure from ChIP-seq. Although common ChIP-seq callers may be applied to ChIP-exo data, to obtain the most precise peak calls we recommend bioinformatics tools specifically designed with the unique structure of ChIP-exo data in mind. These include Genetrack, GEM, MACE, Peakzilla, and ExoProfiler 12-15.
Protocol
Note: Double distilled H2O or molecular grade equivalent is recommended for all buffers and reactions mixes.
Day 0: Material Preparation and Cell Harvest
1. Buffer Preparation
- Prepare Lysis Buffers 1 – 3 (Tables 1 – 3) and add 100 µl complete protease inhibitor stock (CPI) to 50 ml of each buffer just prior to use. Prepare CPI stock by dissolving one tablet in 1 ml molecular grade H2O.
- Prepare ChIP Buffers (Tables 4 – 7). Add 100 µl CPI stock to 50 ml of Blocking and RIPA buffers. Do not add CPI to Tris and ChIP Elution buffers.
2. Annealing of Adapter Oligonucleotides
Note: Specific oligonucleotide sequences may be found in the Additional Information Section.
- Prepare Adapter Annealing mixes (Tables 8 – 9). Vortex to mix and briefly spin tubes to collect contents. Aliquot 100 µl of each mix into 0.5 ml tubes.
- Hybridize the oligonucleotides by running the program (Table 10) in the Thermocycler.
Store annealed oligonucleotides at -80 °C.
3. In Vivo Chromatin Crosslinking with Formaldehyde
NOTE: For a typical ChIP experiment, starting material should contain approximately 50 million cells.
- Add fresh 37% methanol-free formaldehyde stock solution to phosphate buffered saline (PBS) washed cell culture to a final concentration of 1% (v/v), mix thoroughly, and let sit at room temperature for 10 min.
NOTE: For example, we typically add 1.4 ml of 37% methanol-free formaldehyde to cells in 50 ml of room temperature PBS. Also, avoid formaldehyde crosslinking in media since proteins in the media will quench some of the formaldehyde. - Quench crosslinking reaction by adding 2.5 M glycine for a final concentration of 125 mM. Mix thoroughly.
- Transfer cells to 50 ml tube on ice. Spin cells for 5 min at 1,000 x g at 4 °C. Decant supernatant.
- Add 1 ml ice-cold 1x PBS to cell pellet and resuspend by pipetting. Transfer to 1.5 ml tube.
- Spin cells for 3 min at 2,000 x g at 4 °C. Aspirate supernatant.
- Immediately flash freeze cell pellets in 1.5 ml tubes with liquid nitrogen. Store at -80 °C. Crosslinked cells are stable indefinitely at -80 °C.
Day 1: Cell Lysis, Sonication, and ChIP
4. Cell Lysis
NOTE: During all cell lysis sections, the samples MUST be kept on ice or at 4 °C to minimize crosslink reversal.
- Briefly thaw crosslinked cell pellet. Thoroughly resuspend each pellet in 0.5 ml Lysis Buffer 1 and combine with 4.5 ml Lysis Buffer 1 in a 15 ml polystyrene tube.
- Rock tubes for 10 min at 4 °C for on a rocking platform. Spin for 4 min at 2,000 x g at 4 °C. Decant supernatant.
- Thoroughly resuspend each pellet in 0.5 ml Lysis Buffer 2 and add 4.5 ml Lysis Buffer 2 once resuspended.
- Rock for 5 min at 4 °C on a rocking platform. Spin for 5 min at 2,000 x g at 4 °C. Decant supernatant and gently tap excess on paper towel.
- Thoroughly resuspend each pellet in 0.5 ml Lysis Buffer 3 and add 1 ml Lysis Buffer 3 once resuspended. Keep nuclear lysates on ice and immediately proceed to sonication.
5. Sonication of Nuclear Lysates
NOTE: During all sonication sections, the samples MUST be kept on ice to minimize crosslink reversal. The specific model of sonicator used in association with the protocol below may be found in the Table of Materials. Details on specific sonicator usage and guidelines for other instruments may be found in the Additional Information Section.
- Place resonance adapters in 15 ml polystyrene tubes containing nuclear extracts (the metallic bar should not touch the wall of the tube).
- Sonicate nuclear lysates in an ice cold water bath for 2 x 15 min sessions with 30 sec ON/30 sec OFF at medium power. Only sonicate two 15 ml tubes at a time. These settings work on a broad range of cell lines and primary cell types, but further optimization may be needed if chromatin shearing is incomplete.
- To check sonication results, transfer 2 x 10 µl samples from lysate into 1.5 ml tubes.
- To reverse crosslinks, combine the first 10 µl aliquot with 10 µl TE-RNase A and 0.2 µl of Proteinase K. Incubate at 37 °C for 30 min. Add 4 µl 6x xylene DNA dye.
- To preserve crosslinks, combine the second 10 µl aliquot with 2 µl 6x xylene DNA dye.
- Run both samples on a 1.5% agarose gel at 140 V for approximately 30 – 45 min until the bromophenol dye in the ladder is ¾ of the way down the gel.
- If sonication was successful (most DNA fragments sheared to 100 – 500 bp), transfer sonicated lysates to 2 ml tubes containing 150 µl of 10% Triton X-100. Vortex to mix.
- To pellet insoluble chromatin and debris, spin sonicated nuclear lysate for 10 min at 20,000 x g at 4 °C.
- Transfer supernatant to a new 2 ml tube. Proceed immediately to ChIP or store samples at -80 °C indefinitely. Sonicated extracts should not be frozen and thawed more than once.
6. Coupling Antibody to Beads
NOTE: Never vortex or freeze/thaw magnetic beads since they will shatter and increase background signal.
- After thoroughly mixing stock, aliquot Y µl bead slurry into 1.5 ml tube, where Y = 1.1 x 2.5 µl x (number of ChIP samples). Binding capacity for 2.5 µl bead slurry is up to 13 µg IgG. Wash pooled beads 3x with 1 ml Blocking Buffer.
- For more consistent aliquotting of beads after washes, resuspend beads in 10x original slurry volume (25 µl/ChIP) with Blocking Buffer. Aliquot 25 µl per ChIP sample into 1.5 ml tubes.
- Add 5 – 10 µg of antibody to corresponding aliquot of resuspend beads. Bring final volume up to 250 µl with Blocking Buffer. Incubate samples for 4 hr (alternatively, overnight for 16 hr) on a rotating platform at 4 °C.
- Aspirate supernatant. To remove unbound or excess antibody, wash beads once with 1 ml Blocking Buffer.
- After aspirating last wash, immediately add 50 million cell equivalents of sonicated extracts (~ 1.6 ml) to antibody:bead conjugates. If sonicated extracts are not ready, resuspend beads in 100 µl Blocking Buffer until they are ready. It is critical to never let the beads dry.
7. Chromatin Immunoprecipitation (ChIP)
- For each ChIP sample, combine 1.5 ml of sonicated extracts with antibody:bead conjugates in 1.5 ml or 2 ml tubes.
- Incubate tubes on a mini-tube rotator overnight (approximately 16 hr) at speed setting of 9 at 4 ˚C. Check after a few min to ensure samples are not leaking and are mixing properly.
Day 2: ChIP Washes and On-resin Enzymatic Reactions
8. ChIP Washes
NOTE: To minimize cross contamination, briefly spin tubes between each wash. For the first aspiration, change tips between each sample. During subsequent washes, the same tip can be used as long as it did not touch the beads. See Additional Information Section for directions on proper ChIP washing.
- Very briefly spin tubes using a microcentrifuge (~ 3 sec at 500 x g) to collect liquid in caps and place on the magnetic rack for 1 min. While still on the magnetic rack, aspirate extract.
- Add 0.75 ml RIPA Buffer to each tube. Remove tubes from magnetic rack and invert several times to mix. Replace tubes on magnetic rack and aspirate supernatant. Repeat 7x.
- Add 0.75 ml 10 mM Tris HCl (pH 7.5) to each tube. Remove tubes from magnetic rack and invert several times to mix. Replace tubes on magnetic rack and aspirate supernatant. Repeat 2x.
- After aspirating last wash, proceed to Polishing.
Note: After each incubation reaction in sections 9.3, 10.3, 11.3, 12.3, 13.3, 14.3, and 15.3, spin tubes briefly and place against magnetic rack for 1 min and aspirate supernatant. Wash 2x with 0.75 ml RIPA Buffer and 2x with 0.75 ml Tris HCl (pH 7.5).
9. Polishing Reaction
- Fill out Polishing master mix calculations (Table 11). Make Polishing mix in 1.5 ml tube on ice. Pipet to mix.
- Immediately after the last ChIP wash is aspirated, add 50 µl of Polishing mix to each sample resin while still on the magnetic rack. Incubate the samples for 30 min at 3 x g at 30 ˚C in a thermomixer.
- After aspirating last wash as described in the note after Section 8.4, proceed to A-tailing.
10. A-tailing Reaction
- Fill out A-tailing master mix calculations (Table 12). Make A-tailing mix in a 1.5 ml tube on ice. Pipet to mix.
- Immediately after Polishing section, add 50 µl of A-tailing mix to each sample resin while still on the magnetic rack. Incubate samples for 30 min at 3 x g at 37 ˚C in a thermomixer.
- After aspirating last wash as described in the note after Section 8.4, proceed to P7 Adapter Ligation.
11. P7 Adapter Ligation Reaction
- Fill out Ligation master mix calculations (Table 13). Make Ligation mix in 1.5 ml tube on ice. Pipette to mix.
- Immediately after A-tailing section, add 48 µl P7 Ligation master mix and 2 µl of a different Adapter Index to each sample resin while still on the magnetic rack. Incubate samples for 2 hr at 3 x g at 25 ˚C in a thermomixer.
Note: The use of different indexes for each sample will allow more samples to be sequenced in a single flow cell. - After aspirating last wash as described in the note after Section 8.4, proceed to Φ-29 Nick Repair.
12. Phi-29 Nick Repair Reaction
- Fill out Φ-29 master mix calculations (Table 14). Make Φ-29 mix in 1.5 ml tube on ice. Pipet to mix.
- Immediately after P7 Ligation section, add 50 µl of Φ-29 mix to each sample resin while still on the magnetic rack. Incubate samples for 20 min at 3 x g at 30 ˚C in a thermomixer.
- After aspirating last wash as described in the note after Section 8.4, proceed to Kinase reaction.
13. Kinase Reaction
- Fill out Kinase master mix calculations (Table 15). Make Kinase mix in 1.5 ml tube on ice. Pipet to mix.
- Immediately after Φ-29 section, add 50 µl of Kinase mix to each sample resin while still on the magnetic rack. Incubate samples for 20 min at 3 x g at 37 ˚C in a thermomixer.
- After aspirating last wash as described in the note after Section 8.4, proceed to Lambda Exonuclease reaction.
14. Lambda Exonuclease Reaction
- Fill out Lambda Exonuclease master mix calculations (Table 16). Make Lambda Exonuclease mix in 1.5 ml tube on ice. Pipet to mix.
- Immediately after Kinase section, add 50 µl of Lambda Exonuclease mix to each sample resin while still on the magnetic rack. Incubate samples for 30 min at 3 x g at 37 ˚C in a thermomixer.
- After aspirating last wash as described in the note after Section 8.4, proceed to RecJf reaction.
15. RecJf Nuclease Reaction
- Fill out RecJf master mix calculations (Table 17). Make RecJf mix in 1.5 ml tube on ice. Pipet to mix.
- Immediately after Lambda Exonuclease section, add 50 µl of RecJf mix to each sample resin while still on the magnetic rack. Incubate samples for 30 min at 3 x g at 37 ˚C in a thermomixer.
- After aspirating last wash as described in the note after Section 8.4, proceed to ChIP Elution.
16. Elution and Crosslink Reversal
- Prepare master mix: number of samples x 1.1 x (200 µl ChIP Elution Buffer + 1 µl 20 mg/ml Proteinase K) in 1.5 ml tube. Add 200 µl ChIP Elution Buffer + Proteinase K master mix to each sample resin.
- Incubate samples overnight (approximately 16 hr) at 3 x g at 65 ˚C in a thermomixer with a heated lid to prevent condensation.
Day 3: DNA Extraction and Adapter Ligation
17. Elution and Crosslink Reversal (continued)
- After overnight incubation at 65 ˚C, briefly spin samples to collect condensate. Place samples on magnetic rack for 1 min.
- Transfer 200 µl supernatant to new 1.5 ml tube containing 200 µl TE Buffer (pH 7.5).
18. Phenol Chloroform Isoamyl Alcohol (PCIAA) Extraction
- Add 400 µl phenol chloroform isoamyl alcohol to each sample. Vortex to mix for 20 sec.
- Centrifuge for 10 min at 20,000 x g at room temperature (RT). Carefully transfer 325 µl of the upper aqueous layer to a fresh tube.
Note: Take care not to transfer any of the organic (lower layer) into the new tube. - Add 1/10 volume of 3 M NaOAc (pH 5.5) and 1 µl 20 mg/ml glycogen to each sample. For multiple samples, prepare a master mix.
- Add 3 volumes of ice cold 100% ethanol to each sample. Vortex to mix. Incubate for 15 min at -80 ˚C.
- Centrifuge for 15 min at 20,000 x g at 4 ˚C. Carefully decant supernatant, making sure to not disturb the pellet.
- Gently add 500 µl freshly made ice cold 70% ethanol, making sure to not disturb the pellet. Centrifuge for 5 min at 20,000 x g at 4 ˚C. Carefully decant supernatant.
- Dry pellet for approximately 20 min (or until dry) in a speed vacuum at 45 ˚C.
NOTE: This is a pause point in the protocol. Dry DNA pellets can be stored at -20 ˚C. - Resuspend pellets in 10 µl ddH2O. Pipet repeatedly over the area where the DNA pelleted, even though the dried pellet cannot be seen.
- Transfer 10 µl of each sample to a fresh 0.3 ml PCR tube or 8-rack, depending on number of samples.
19. P7 Primer Extension Reaction
- Fill out P7 Primer Extension master mix calculations (Table 18). Make Extension mix in 1.5 ml tube on ice. Pipet to mix.
- Add 10 µl of Extension mix (minus Φ-29 polymerase) to each 10 µl sample. Pipet to mix.
- Run samples in the Thermocycler using the program (Table 19) to anneal primer to the template until the 30 ˚C "hold" step.
- Add 1 µl Φ-29 polymerase during the 30 ˚C "hold" step in the program. Pipet to mix. Resume the remainder of the program (Table 19).
20. A-tailing Reaction
- Fill out A-tailing master mix calculations (Table 20). Make A-tailing mix in a 1.5 ml tube on ice. Pipet to mix.
- Add 10 µl of A-tailing mix to each sample. Pipet to mix.
- In the Thermocycler, incubate samples for 30 min at 37 ˚C, and then heat inactivate for 20 min at 75 ˚C.
21. P5 Adapter Ligation Reaction
- Fill out P5 Adapter Ligation master mix calculations (Table 21). Make Ligation mix in 1.5 ml tube on ice. Pipet to mix.
- Add 20 µl of P5 Ligation mix to each sample. Pipet to mix.
- In the Thermocycler, incubate samples for 2 hr at 25 ˚C.
22. Bead Clean-up
Note: Please see Materials section for manufacturer details on bead clean up.
- Transfer each 50 µl sample to a new 1.5 ml tube.
- Resuspend clean up beads on a mini-tube rotator at speed setting of 15 for 15 min at RT. Do not let beads settle before pipetting.
- Add 60 µl clean up beads to sample (1.2:1 clean up beads to sample ratio is critical to remove DNA that is shorter than 200 base pairs). Pipet to mix for 20 sec.
- Resuspend clean up beads on a mini-tube rotator at speed setting of 15 for 3 min at RT. Briefly spin tubes.
- Place tubes on the magnetic rack for 1 min. Pipet off and discard the supernatant.
NOTE: DO NOT use vacuum aspiration in this section because it will suck up the clean-up beads. - Add 400 µl of freshly made RT 70% ethanol to each sample while they are on magnetic rack. DO NOT mix. Aspirate supernatant. Repeat 2x.
- Dry the clean-up beads for 10 min at RT. The color of the beads will turn dark to light brown and very small cracks will be visible in the resin pellet when beads are dry.
- To elute DNA, add 40 µl 10mM Tris (pH 7.5). Thoroughly pipet to mix for 20 sec.
- Place the samples on the magnetic rack for 1 min. Slowly pipet and transfer 36 µl eluate directly to 0.3 ml PCR tubes.
- Proceed directly to PCR reaction or store eluates at -20 ˚C.
Day 4: PCR and Gel Analysis
23. PCR
- Fill out PCR master mix calculations (Table 22). Make PCR mix. Pipet very gently to mix to avoid bubbles.
- Add 14 µl of PCR mix to each 36 µl DNA sample. Pipet to mix. Keep the samples on ice until ready to put in Thermocycler.
- For the positive PCR control, include a previously prepared library. For the negative PCR control, include water. Run samples in the Thermocycler using PCR program (Table 23).
24. Gel Preparation, DNA Excision, and Visualization
- Thoroughly clean and rinse an appropriate size gel box with deionized water. Prepare a 1.5% agarose gel (containing 0.5 mg/ml ethidium bromide) with thick, wide-well comb that can hold 60 µl.
NOTE: Ethidium bromide (EtBr) is a carcinogen and must be handled with care. - Spin down PCR sample tubes briefly to collect condensation.
- Add 1/5 volume of 6x xylene DNA dye to combined sample. Do not use bromophenol blue in DNA dye since it migrates at the same location as the sample DNA.
- Load entire sample into each well, preferably with empty wells in between samples.
Note: Libraries with the same indexes must NOT be run in the same gel. - Load 7 µl 100 bp ladder on either side of samples. Run gel at 140 V for approximately 30-45 min until bromophenol dye in ladder is ¾ way down the gel.
- Image and visualize gel on a transilluminator at a low UV setting. Excise the sections of agarose containing DNA fragments 200 – 500 bp. Be very careful to avoid cutting out adapter dimer band that runs at 125 bp.
- Place each excised gel slice into a 15 ml tube. Record net weight of each gel slice. Write this weight directly on tube.
- Image, save, and annotate excised gel to confirm correct size range selected.
25. Gel Purification
- Purify DNA from excised gel slice according to the manufacturer's instructions, with the following modifications.
- Dissolve the gel slice by rocking at RT on rocking platform. It will take approximately 20 min to dissolve.
- To obtain highly purified DNA, wash columns with 0.5 ml Buffer QG after dissolved gel has passed through column.
- After Buffer PE wash, let columns sit at RT for 2 – 5 min. Spin for 1 min at 13,000 x g at RT.
- To allow room for PCR reaction master mix, elute samples with 40 µl EB Buffer into a fresh 1.5 ml tube.
26. DNA Quantification
- Prepare samples according to the manufacturer's instructions (Table 24). Measure samples in specified instrument with optical tubes.
- Once ChIP-exo libraries are quantified, submit for sequencing on a platform that uses sequencing-by-synthesis chemistry 16 that is compatible with DNA adapters in this protocol.
NOTE: Typically, 2 µl of sample is sufficient for quantification, but more may be necessary if sample concentration is lower. DNA yields typically range between 50 and 200 ng. - After quantification, store samples at -20 ˚C.
Representative Results
The following figures illustrate representative results from the ChIP-exo protocol presented here. In contrast to traditional ChIP-seq methodologies with few enzymatic steps, ChIP-exo requires eleven sequentially dependent enzymatic reactions (Figure 1). Thus, care must be taken at each step to ensure that each reaction component is added to its respective reaction master mix. We recommend generating a formulaic spreadsheet based on the reaction Tables to automatically perform the reaction master mix calculations, printing the resulting tables, and then checking off each item after it is added to the master mix.
We employ a number of quality control measures throughout the protocol to ensure high quality sequencing results (Figure 2). Each ChIP reaction contains three basic components: 1) sonicated chromatin extract from cells of interest, 2) an antibody directed against the protein of interest, and 3) ProteinG (or ProteinA) resin to immobilize the precipitated immune complexes. Obtaining high quality sonicated chromatin extracts (Figure 2A) can be quite challenging since sonication conditions must be optimized for each cell type and sonication instrument. It is worth spending time to optimize this step because the highest quality libraries start with a sonication result that yields DNA fragments between 100 – 500 bp (Figure 2A, "+" lane). Since formaldehyde crosslinks are labile and formaldehyde itself has a limited shelf-life, we use an electrophoretic mobility shift assay (Figure 2A, "-" lane) to verify that the sonicated extracts contain intact protein-DNA crosslinks, evident as super-shift of the DNA fragments. To assess background signal we routinely perform a "mock" ChIP that omits antibody from the ChIP reaction. A high quality ChIP-exo library preparation will have very little, if any, background signal in the "mock" ChIP ("-") relative to the specific ChIP antibody, in this case directed against Pol II. As seen in Figure 2B, traditional ChIP-seq libraries have substantially more background signal than ChIP-exo. Lastly, prior to sequencing, ChIP-exo library quality is assessed on the bioanalyzer (Figure 2C – D). Analysis accurately measures the library size distribution and detects contaminating adapter dimers (denoted by arrow) that run at 125 bp. If adapter dimers are present, they will reduce the sequencing bandwidth. Therefore, we recommend an additional bead cleanup that will efficiently remove DNA fragments less than 200 bp.
ChIP-exo is a powerful functional genomic technique because it is the only method capable of spatially resolving divergent, initiating, paused, and elongating RNA polymerase II on a genome-wide scale (Figure 3) 11. Since these adjacent binding events are tens of base pairs apart, ChIP-seq is unable to distinguish these binding events with resolving power of several hundred base pairs.
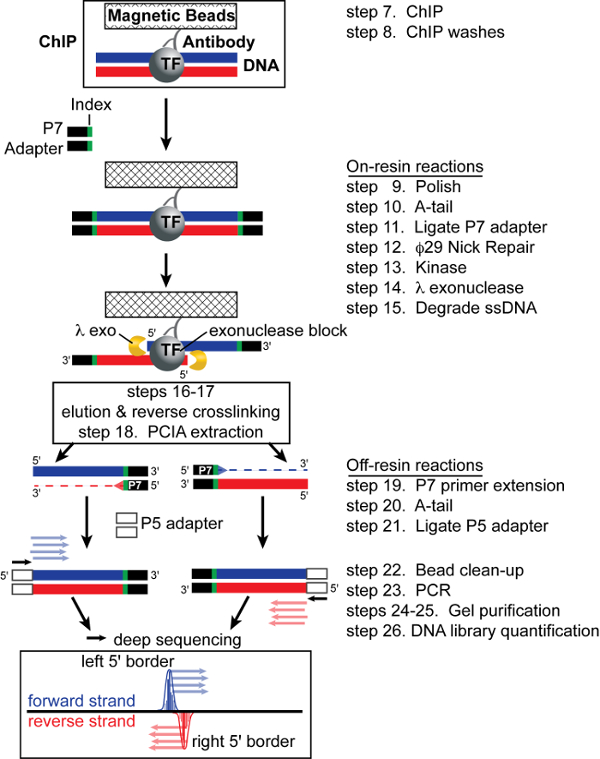
Figure 1: ChIP-exo Schematic. After ChIP, the P7 adapter is ligated to the sonication borders. Lambda exonuclease then trims DNA 5' to 3' to the crosslink point, thereby footprinting the protein-DNA interaction. After elution and crosslink reversal, primer extension synthesizes duplex DNA. Lastly, ligation of the P5 adapter marks the left and right exonuclease borders and the resulting library is subjected to high-throughput sequencing. Mapping the 5' ends of the sequence tags to the reference genome demarcates the exonuclease barrier and thus the precise site of protein-DNA crosslinking. Figure modified from Rhee and Pugh 17. Please click here to view a larger version of this figure.
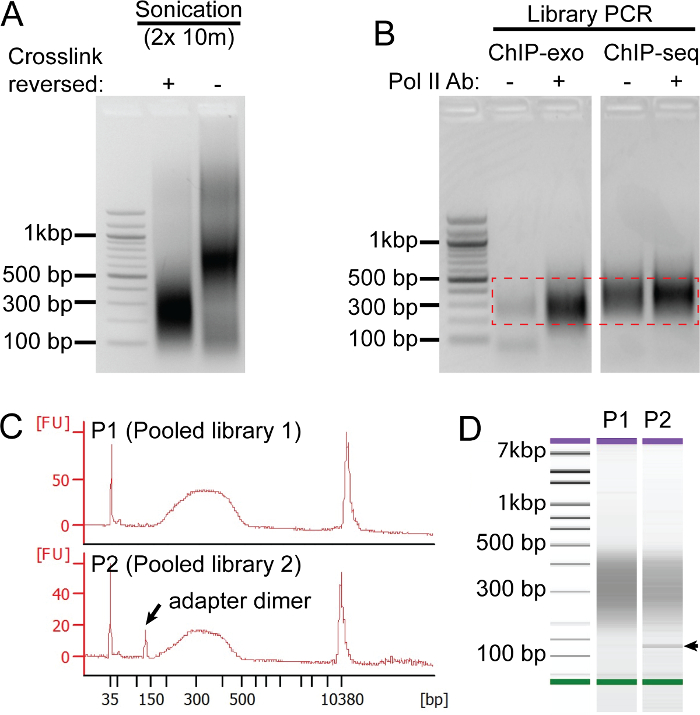
Figure 2: ChIP-exo Quality Controls. Panels A-C show representative results from unrelated experiments. (A) The quality of the sonicated chromatin extract (from the human HCC1806 breast cancer cell line) is assessed by agarose gel electrophoresis on extracts with (+) and without (-) the crosslinks reversed. Intact crosslinks will cause protein-DNA complexes to migrate slower. Thus, running extract without crosslinks reversed (-) allows for the quality of the formaldehyde crosslinks to be assessed, which are critical for a successful ChIP. (B) Comparison of a mock IP (-) and Pol II (+) ChIP-exo and ChIP-seq library preparations after 21 cycles of PCR amplification. After PCR, DNA fragments 200-500 bp are excised (denoted by red hashed box) and purified using a gel extraction kit. (C-D) In panel C, the top trace represents an ideal trace from a pooled library (P1), and the bottom trace shows a pooled library (P2) that contains adapter dimers. Panel D shows the DNA density plot corresponding to the P1-2 library traces from panel C (arrow denotes adapter dimer band). Please click here to view a larger version of this figure.
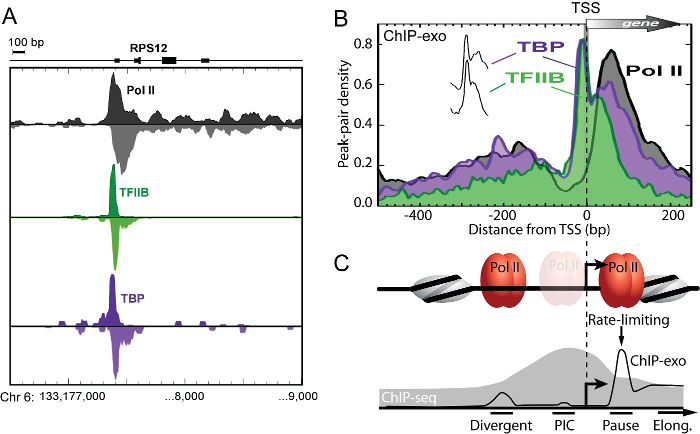
Figure 3: ChIP-exo Spatially Resolves Distinct Bidirectional Transcription Initiation Complexes. (A) Smoothed distribution of strand-separated ChIP-exo tag 5' ends for Pol II, TFIIB, and TBP at the human RPS12 gene in proliferating K562 cells. (B) Averaged ChIP-exo patterns around the closest RefSeq TSS. Peak-pair tags were aligned to the TSS gene-by-gene, binned in non-overlapping 10bp intervals relative to the TSS, and then the average peak-pair density value across all TFIIB-occupied (n = 6,511) genes was plotted as a percent of the total. The "spikes" of TBP and TFIIB are indiscernible (vertically offset in inset). (C) Model based on panel B data, illustrating distinct transcription initiation complexes resolved by ChIP-exo (black trace). Pol II occupied two separate resolvable locations that coincided with sites of divergent transcription initiation ("Divergent") and "Pause" sites. This clear spatial separation of Pol II complexes indicates that divergent transcripts arise from distinct initiation complexes. The vast majority Pol II crosslinked about 50 bp downstream of the TSS at the "Pause" site, where it is expected to pause after initiating transcription. Pol II was most depleted 20 – 60 bp upstream of the TSS where the pre-initiation complex ("PIC") forms, indicating that on average it likely spends less time there than at the paused sites. This suggests that in most (but not necessarily all) cases, once Pol II is recruited, it rapidly clears the promoter and assumes a paused-state approximately 30 – 50 bp downstream of the TSS, consistent with the observation that Pol II pause release is a rate-limiting step in transcription. These adjacent initiation complexes are unresolvable by ChIP-seq (illustrated by gray fill trace) since its resolution is limited to a few hundred base pairs. Figure modified from Pugh and Venters 11. Please click here to view a larger version of this figure.
Supplemental File 1: Additional Information. Please click here to download this file.
| Reagent | Volume (ml) | [Final] |
| 1 M HEPES-KOH (pH 7.5) | 50 | 50 mM |
| 5 M NaCl | 28 | 140 mM |
| 0.5 M EDTA | 2 | 1 mM |
| 100% Glycerol | 100 | 10% |
| 10% NP40 | 50 | 0.50% |
| 10% Triton X100 | 25 | 0.25% |
| ddH2O | Fill to 1 L |
Table 1. Recipe for Lysis Buffer 1. Filter using 0.22 μm filter. Store in 50 ml tubes at 4 ˚C. Add 100 μl CPI stock to 50 ml of buffer just prior to use.
| Reagent | Volume (ml) | [Final] |
| 1 M Tris-HCl (pH 8) | 10 | 10 mM |
| 5 M NaCl | 40 | 200 mM |
| 0.5 M EDTA | 2 | 1 mM |
| 0.5 M EGTA | 1 | 0.5 mM |
| ddH2O | Fill to 1 L |
Table 2. Recipe for Lysis Buffer 2. Filter using 0.22 μm filter. Store in 50 ml tubes at 4 ˚C. Add 100 μl CPI stock to 50 ml of buffer just prior to use.
| Reagent | Volume (ml) | [Final] |
| 1 M Tris-HCl (pH 8) | 10 | 10 mM |
| 5 M NaCl | 20 | 100 mM |
| 0.5 M EDTA | 2 | 1 mM |
| 0.5 M EGTA | 1 | 0.5 mM |
| 10% Deoxycholate | 10 | 0.10% |
| N-lauroylsarcosine | 5 g | 0.5% (w/v) |
| ddH2O | Fill to 1 L |
Table 3. Recipe for Lysis Buffer 3. Filter using 0.22 μm filter. Store in 50 ml tubes at 4 ˚C. Add 100 μl CPI stock to 50 ml of buffer just prior to use.
| Reagent | Volume (ml) | [Final] |
| 10x PBS | 50 | 1x |
| Bovine Serum Albumin | 2.5 g | 0.50% |
| ddH2O | Fill to 500 |
Table 4. Recipe for Blocking Buffer. Filter using 0.22 μm filter. Store in 50 ml tubes at 4 ˚C. Add 100 μl CPI stock to 50 ml of buffer just prior to use.
| Reagent | Volume (ml) | [Final] |
| 1 M HEPES (pH 7.5) | 25 | 50 mM |
| 0.5 M EDTA (pH 8) | 1 | 1 mM |
| 10% Sodium Deoxycholate | 35 | 0.70% |
| 10% NP40 | 50 | 1% |
| 1 M LiCl | 250 | 500 mM |
| ddH2O | Fill to 500 |
Table 5. Recipe for RIPA Buffer. Filter using 0.22 μm filter. Store in 50 ml tubes at 4 ˚C. Add 100 µl CPI stock to 50 ml of buffer just prior to use.
| Reagent | Volume (ml) | [Final] |
| 1 M Tris-Cl (pH 7.5) | 2.5 | 50 mM |
| 0.5 M EDTA | 1 | 10 mM |
| 20% SDS | 2.5 | 1% |
| ddH2O | Fill to 50 |
Table 6. Recipe for ChIP Elution Buffer. Filter using 0.22 μm filter. Store at RT.
| Reagent | Volume (ml) | [Final] |
| 1 M Tris-Cl (pH 7.5) | 0.5 | 10 mM |
| ddH2O | Fill to 50 |
Table 7. Recipe for TE Buffer. Filter using 0.22 μm filter. Store at 4 ˚C.
| Volume (μl) | [Final] | |
| 100 μM ExA2-iX | 75 | 15 μM |
| 100 μM ExA2-33 | 75 | 15 μM |
| 1 M Tris (pH 7.5) | 50 | 100 mM |
| 5 M NaCl | 5 | 50 mM |
| ddH2O | 295 | – |
| Total volume | 500 |
Table 8. P7 Adapter Annealing mix.
| Volume (μl) | [Final] | |
| 100 μM ExA1-58 | 75 | 15 μM |
| 100 μM ExA1-13 | 75 | 15 μM |
| 1 M Tris (pH 7.5) | 50 | 100 mM |
| 5 M NaCl | 5 | 50 mM |
| ddH2O | 295 | – |
| Total volume | 500 |
Table 9. P5 Adapter Annealing mix.
| Temp (˚C) | Time |
| 95 | 5 min |
| 72 | 5 min |
| 65 to 60 ramp decline | 5 min |
| 55 to 50 ramp decline | 3 min |
| 45 to 40 ramp decline | 3 min |
| 30 | 3 min |
| 20 | 3 min |
| 10 | 3 min |
| 4 | Forever |
Table 10. Adapter Annealing Program.
| 1x (μl) | [Final] | |
| ddH2O | 39.8 | |
| 10x Reaction Buffer 2 | 5 | 1x |
| 100 µM ATP | 0.5 | 1 mM |
| 3 mM dNTPs | 1.7 | 100 μM |
| 3 U/μl T4 polymerase | 1 | 3 U |
| 5 U/μl Klenow | 1 | 5 U |
| 10 U/μl T4 Polynucleotide Kinase | 1 | 10 U |
| Total reaction volume | 50 |
Table 11. Polishing master mix.
| 1x (μl) | [Final] | |
| ddH2O | 42.3 | |
| 10x Reaction Buffer 2 | 5 | 1x |
| 3 mM dATP | 1.7 | 100 μM |
| 5 U/μl Klenow 3'-5' exo minus | 1 | 5 U |
| Total reaction volume | 50 |
Table 12. A-tailing master mix.
| 1x (μl) | [Final] | |
| ddH2O | 41 | |
| 100 mM ATP | 0.5 | 1 mM |
| 10x Reaction Buffer 2 | 5 | 1x |
| 400 U/μl T4 DNA Ligase | 1.5 | 600 U |
| Reaction mix volume | 48 | |
| 15 mM Index Adapter | 2 | 30 picomoles |
| Total reaction volume | 50 |
Table 13. P7 Adapter Ligation master mix.
| 1x (μl) | [Final] | |
| ddH2O | 41 | |
| 10x Φ-29 Buffer | 5 | 1x |
| 3 mM dNTPs | 2.5 | 150 μM |
| 10 U/μl Φ-29 polymerase | 1.5 | 15 U |
| Total reaction volume | 50 |
Table 14. Φ-29 Nick Repair master mix.
| 1x (μl) | [Final] | |
| ddH2O | 43.5 | |
| 100 mM ATP | 0.5 | 1 mM |
| 10x Reaction Buffer 2 | 5 | 1x |
| 10 U/μl T4 Polynucleotide Kinase | 1 | 10 U |
| Total reaction volume | 50 |
Table 15. Kinase Reaction master mix.
| 1x (μl) | [Final] | |
| ddH2O | 43 | |
| 10x Lambda Buffer | 5 | 1x |
| 5 U/μl Lambda exonuclease | 2 | 10 U |
| Total reaction volume | 50 |
Table 16. Lambda Exonuclease Reaction master mix.
| 1x (μl) | [Final] | |
| ddH2O | 44 | |
| 10x Reaction Buffer 2 | 5 | 1x |
| 30 U/μl RecJf exonuclease | 1 | 30 U |
| Total reaction volume | 50 |
Table 17. RecJf Nuclease Reaction master mix.
| 1x (μl) | [Final] | |
| ddH2O | 6.45 | |
| 10x Φ-29 Buffer | 2 | 1x |
| 3 mM dNTP | 1.3 | 200 μM |
| 20 μM P7 Primer | 0.25 | 0.25 μM |
| Reaction mix volume | 10 | |
| EtOH precipitated sample | 10 | |
| Total reaction volume | 20 |
Table 18. P7 Primer Extension Reaction master mix.
| Temp (˚C) | Time |
| 95 | 5 min |
| 65 | 5 min |
| 30 | 2 min |
| 30 | Hold until Φ-29 is added |
| 30 | 20 min |
| 65 | 10 min |
| 4 | Forever |
Table 19. P7 Primer Extension program.
| 1x (μl) | [Final] | |
| ddH2O | 5 | |
| 10x Reaction Buffer 2 | 3 | 1x |
| 3 mM dATP | 1 | 0.1 mM |
| 5 U/μl Klenow 3’ to 5’ exo minus | 1 | 5 U |
| Reaction mix volume | 10 | |
| Primer extended sample | 20 | |
| Total reaction volume | 30 |
Table 20. A-tailing master mix.
| 1x (μl) | [Final] | |
| ddH2O | 11.5 | |
| 10x T4 Ligase Buffer | 5 | 1x |
| 15 μM ExA1-58/13 adapter | 2 | 30 picomoles |
| 400 U/μl T4 DNA Ligase | 1.5 | 600 U |
| Reaction mix volume | 20 | |
| A-tailed sample | 30 | |
| Total reaction volume | 50 |
Table 21. P5 Adapter Ligation master mix.
| 1x (μl) | [Final] | |
| 5x PCR Buffer | 10 | 1x (2 mM MgCl2) |
| 10 mM of each dNTP | 1 | 200 μM each |
| 20 μM P1.3 Primer | 1.25 | 0.5 μM |
| 20 μM P2.1 Primer | 1.25 | 0.5 μM |
| 2 U/μl Hot Start polymerase | 0.5 | 1 U |
| Reaction mix volume | 14 | |
| Bead eluted sample | 36 | |
| Total reaction volume | 50 |
Table 22. PCR.
| Temp (˚C) | Time | Cycles |
| 98 | 30 sec | 1 |
| 98 | 10 sec | 15-21 (depending on ChIP efficiency) |
| 52 | 30 sec | |
| 72 | 20 sec | |
| 72 | 2 min | 1 |
| 4 | Forever | Hold |
Table 23. PCR program.
| Per DNA Standard (μl) | Per Sample (μl) | |
| 1:200 diluted buffer/dye mix | 190 | 198 |
| DNA standards | 10 | |
| ChIP-exo library | 2 | |
| Total volume | 200 | 200 |
Table 24. Quantification.
Discussion
We present a functional genomic protocol to determine the precise binding location for chromatin interacting proteins in an unbiased, genome-wide manner at near base pair resolution. The most critical step to achieve near base pair mapping resolution is the exonuclease treatment of the ChIP-enriched DNA while the immunoprecipitate remains on the magnetic resin. Ostensibly, protein complexes could potentially block in vivo footprinting of any given subunit (e.g., chromatin remodeling complexes or the nucleosome core particle). However, as reported previously 10, since formaldehyde is an inefficient crosslinker, it becomes increasingly unlikely that multiple subunits of a complex would crosslink to DNA and each other in the same cell at the same locus. Thus, in vivo footprinting of individual subunits of a protein complex, such as individual histone subunits of a nucleosome, is possible with ChIP-exo.
The most notable advantages of ChIP-exo are its near base pair resolution and low background. The ultra-high resolution permits detailed structural and spatial insights to be made on a genome-wide scale that are not currently possible with any other method. On the other hand, the primary limitation of ChIP-exo is that it is a technically challenging molecular biology methodology to master. In addition, the general limitations of the ChIP step also apply to ChIP-exo (e.g., commercial antibody availability and specificity, epitope accessibility, and relatively large number of cells required). Common pitfalls include poor quality sonicated extracts, using a non-ChIP grade antibody, and not keeping samples on ice as much as possible. Thus, sonication conditions must be carefully optimized and each antibody validated as previously described 18 to avoid the detrimental effects these parameters can have on the experimental outcome.
As far as sequencing depth for a transcription factor target, we typically aim for about 20 million uniquely aligned reads. Since Pol II and histone modifications are more broadly distributed, we aim for 30 to 50 million reads. It is important to note that since ChIP-exo has substantially less background than ChIP-seq, fewer reads are required to achieve similar sequencing depth.
The ChIP-exo technology is now being widely adopted despite its technical challenges, as robust variations on the original protocol continue to be published 17,19,20. In particular, one variation that may prove useful for difficult to ChIP proteins is called ChIP-nexus, which uses a single ligation step to increase the efficiently of the library preparation 20. In summary, ChIP-exo is an increasingly employed and powerful methodology for ultra-high resolution mapping of chromatin interacting proteins on a global scale. As the list of commercially available ChIP-grade antibodies continues to grow, future applications of the ChIP-exo methodology will be directed at mapping uncharted gene regulatory networks to understand the molecular circuitry of the cell in ultra-high resolution. In addition, ChIP-exo will likely be further refined and adapted to footprint in vivo protein-RNA interactions at near base pair resolution.
Disclosures
The authors have nothing to disclose.
Acknowledgements
We thank the Venters Lab and members of the Molecular Physiology and Biophysics Department for helpful discussions. Special thanks to Frank Pugh for his guidance, mentoring, and many insightful discussions on the nuances of ChIP-exo while I was a post-doctoral fellow in his laboratory.
Materials
| 37% formaldehyde, methanol free, 10x10ml ampules | ThermoFisher Scientific | 28908 | Section 3 |
| Complete Protease Inhibitor cocktail (CPI) | Roche Life Science | 11873580001 | Sections 4, 8 |
| Bioruptor sonicator | Diagenode | UCD200 | Section 5 |
| 15 ml polystyrene tubes | BD Falcon | 352095 | Section 5 |
| MagSepharose Protein G Xtra beads | GE Healthcare | 28-9670-66 | Section 6 |
| DynaMag-1.5ml Side Magnetic Rack | Invitrogen | 12321D | Sections 6-17, 22 |
| Mini-Tube Rotator | Fisher Scientific | 05-450-127 | Sections 7, 22 |
| T4 DNA Polymerase | New England BioLabs | M0203 | Section 9 |
| DNA Polymerase I, Klenow | New England BioLabs | M0210 | Section 9 |
| 10x NEBuffer 2 (10x Reaction Buffer 2) | New England BioLabs | B7002 | Sections 9-11, 13, 15, 20 |
| Thermomixer C | Eppendorf | 5382 | Sections 9-16 |
| ATP | Roche Life Science | 010419979001 | Sections 9, 11, 13 |
| dNTPs | New England BioLabs | N0447 | Sections 9, 12, 19, 23 |
| T4 Polynucleotide Kinase | New England BioLabs | M0201 | Sections 9, 13 |
| dATP | New England BioLabs | N0440 | Sections 10, 20 |
| Klenow 3'-5' Exo Minus | New England BioLabs | M0212 | Sections 10, 20 |
| T4 DNA ligase | New England BioLabs | M0202 | Sections 11, 21 |
| Φ-29 DNA Polymerase | New England BioLabs | M0269 | Sections 12, 19 |
| 10x Φ-29 Buffer | New England BioLabs | B0269 | Sections 12, 19 |
| Lambda Exonuclease | New England BioLabs | M0262 | Section 14 |
| 10x Lambda Buffer | New England BioLabs | B0262 | Section 14 |
| RecJf Exonuclease | New England BioLabs | M0264 | Section 15 |
| Proteinase K | Roche Life Science | 03115828001 | Section 16 |
| Glycogen | Roche Life Science | 010901393001 | Section 18 |
| 10x T4 Ligase Buffer | New England BioLabs | B0202 | Section 21 |
| AMPure XP (clean up) beads | Beckman Coulter | A63881 | Section 22 |
| Q5 Hot Start DNA Polymerase | New England BioLabs | M0493 | Section 23 |
| 5x Q5 Buffer (5x PCR Buffer) | New England BioLabs | B9027 | Section 23 |
| Qubit Fluorometer | Invitrogen | Q33216 | Section 25 |
| QIAquick Gel Extraction Kit | Qiagen | 28704 | Section 25 |
| Optical Clear Qubit tubes | Invitrogen | Q32856 | Section 26 |
| Qubit dsDNA High Sensitivity Assay kit | Invitrogen | Q32851 | Section 26 |
References
- Venters, B. J., Pugh, B. F. How eukaryotic genes are transcribed. Crit Rev Biochem Mol Biol. 44, 117-141 (2009).
- Rhee, H. S., Pugh, B. F. Comprehensive genome-wide protein-DNA interactions detected at single-nucleotide resolution. Cell. 147, 1408-1419 (2011).
- Chen, J., et al. Single-molecule dynamics of enhanceosome assembly in embryonic stem cells. Cell. 156, 1274-1285 (2014).
- Wales, S., Hashemi, S., Blais, A., McDermott, J. C. Global MEF2 target gene analysis in cardiac and skeletal muscle reveals novel regulation of DUSP6 by p38MAPK-MEF2 signaling. Nucleic acids research. 42, 11349-11362 (2014).
- Cho, S., et al. The architecture of ArgR-DNA complexes at the genome-scale in Escherichia coli. Nucleic Acids Res. 43, 3079-3088 (2015).
- Katainen, R., et al. CTCF/cohesin-binding sites are frequently mutated in cancer. Nat Genet. 47, 818-821 (2015).
- Murphy, M. W., et al. An ancient protein-DNA interaction underlying metazoan sex determination. Nat Struct Mol Biol. 22, 442-451 (2015).
- Zere, T. R., et al. Genomic Targets and Features of BarA-UvrY (-SirA) Signal Transduction Systems. PLoS One. 10, e0145035 (2015).
- Rhee, H. S., Pugh, B. F. Genome-wide structure and organization of eukaryotic pre-initiation complexes. Nature. 483, 295-301 (2012).
- Rhee, H. S., Bataille, A. R., Zhang, L., Pugh, B. F. Subnucleosomal structures and nucleosome asymmetry across a genome. Cell. , 1377-1388 (2014).
- Pugh, B. F., Venters, B. J. Genomic Organization of Human Transcription Initiation Complexes. PLoS One. 11, e0149339 (2016).
- Albert, I., Wachi, S., Jiang, C., Pugh, B. F. GeneTrack–a genomic data processing and visualization framework. Bioinformatics. 24, 1305-1306 (2008).
- Guo, Y., Mahony, S., Gifford, D. K. High resolution genome wide binding event finding and motif discovery reveals transcription factor spatial binding constraints. PLoS computational biology. 8, e1002638 (2012).
- Bardet, A. F., et al. Identification of transcription factor binding sites from ChIP-seq data at high resolution. Bioinformatics. 29, 2705-2713 (2013).
- Wang, L., et al. MACE: model based analysis of ChIP-exo. Nucleic Acids Res. 42, e156 (2014).
- Bentley, D. R., et al. Accurate whole human genome sequencing using reversible terminator chemistry. Nature. 456, 53-59 (2008).
- Rhee, H. S., Pugh, B. F. ChIP-exo method for identifying genomic location of DNA-binding proteins with near-single-nucleotide accuracy. Curr Protoc Mol Biol. Chapter 21, Unit 21.24 (2012).
- Landt, S. G., et al. ChIP-seq guidelines and practices of the ENCODE and modENCODE consortia. Genome Res. 22, 1813-1831 (2012).
- Serandour, A. A., Brown, G. D., Cohen, J. D., Carroll, J. S. Development of an Illumina-based ChIP-exonuclease method provides insight into FoxA1-DNA binding properties. Genome Biol. 14, R147 (2013).
- He, Q., Johnston, J., Zeitlinger, J. ChIP-nexus enables improved detection of in vivo transcription factor binding footprints. Nat Biotechnol. 33, 395-401 (2015).

