A Rapid Image-based Bacterial Virulence Assay Using Amoeba
Summary
Here, we present a protocol to measure the virulence of planktonic or surface-attached bacteria using D. discoideum (amoeba) as a host. Virulence is measured over a period of 1 h and host killing is quantified using fluorescence microscopy and image analysis. We demonstrate this protocol using the bacterium P. aeruginosa.
Abstract
Traditional bacterial virulence assays involve prolonged exposure of bacteria over the course of several hours to host cells. During this time, bacteria can undergo changes in the physiology due to the exposure to host growth environment and the presence of the host cells. We developed an assay to rapidly measure the virulence state of bacteria that minimize the extent to which bacteria grow in the presence of host cells. Bacteria and amoebae are mixed together and immobilized on a single imaging plane using an agar pad. The procedure uses single-cell fluorescence imaging with calcein-acetoxymethyl ester (calcein-AM) as an indicator of host cell health. The fluorescence of host cells is analyzed after 1 h of exposure of host cells to bacteria using epifluorescence microscopy. Image analysis software is used to compute a host killing index. This method has been used to measure virulence within planktonic and surface-attached Pseudomonas aeruginosa sub-populations during the initial stage of biofilm formation and may be adapted to other bacteria and other stages of biofilm growth. This protocol provides a rapid and robust method of measuring virulence and avoids many of the complexities associated with the growth and maintenance of mammalian cell lines. Virulence phenotypes measured here using amoebae have also been validated using mouse macrophages. In particular, this assay was used to establish that surface attachment upregulates virulence in P. aeruginosa.
Introduction
Bacterial infection is one of the leading causes of mortality in human and animals1,2. The ability to measure virulence of bacteria in cultures or biofilms is important in healthcare and research settings. Here, we describe a versatile, rapid, and relatively simple method to quantify bacterial virulence. The eukaryotic organism Dictyostelium discoideum (amoeba) is used as the model host organism. D. discoideum has been used as a host to identify virulence factors in Pseudomonas aeruginosa (P. aeruginosa)3,4,5 and other bacteria6,7,8 and is susceptible to largely the same virulence factors that kill mammalian cells including type III secretion9,10. Previous virulence assays using D. discoideum have involved prolonged exposure of bacteria with D. discoideum cells over the course of hours3,4,5. The protocol, here, presents a rapid method of determining virulence using this amoeba. This protocol (Figure 1) describes how to: (1) grow the amoebae axenically (in the absence of bacteria), (2) grow bacteria for the assay, (3) prepare bacteria and host cells for microscopy, (4) perform epifluorescence microscopy, and (5) analyze amoeba fluorescence.
Amoebae are initially streaked out from frozen stocks and grown on a lawn of Escherichia coli (E. coli), where the amoebae produce spores. These spores are picked and inoculated into an enriched medium for axenic growth. The amoebae are maintained through axenic growth in nutrient-rich conditions until they are ready to be mixed with bacteria for the assessment of bacterial virulence. The survival or the death of the amoebae is quantified by measuring the fluorescence of calcein-acetoxymethyl (calcein-AM), which is cleaved by intracellular esterases and, thereby, activated for fluorescence11,12. Live amoebae exhibit little or no fluorescence whereas stressed and dying cells fluoresce intensely. This result is due to a little or no incorporation of calcein-AM into healthy amoebae and incorporation and cleavage of the substrate in stressed amoebae13. This behavior is notably distinct from calcein-AM fluorescence in mammalian cells11,14,15,16.
Bacteria that will be assessed for virulence are grown separately. Here, we describe how to measure the virulence of the opportunistic pathogen P. aeruginosa and detail how to quantify the virulence of planktonic (swimming) and surface-attached sub-populations. This protocol may be adapted to test the virulence of other bacteria. In the Representative Results section, we show that virulence is activated in surface-attached cells and is low in planktonic cells, which was reported previously13. Virulence-activated surface-attached P. aeruginosa kills amoebae while non-virulent planktonic cells are consumed by the amoebae. If the virulence of planktonic bacteria is solely being assayed, bacteria can be cultured in ordinary culture tubes rather than using Petri dishes as described in the protocol.
The growth of amoebae and P. aeruginosa cultures must be coordinated such that P. aeruginosa cultures reach the intended growth phase while the amoebae are growing at steady state in nutrient-rich conditions. This condition typically requires amoebae cultures to be diluted at least 1 day prior to when they are mixed with bacteria. Amoebae and bacteria are immobilized using agar pads, are co-incubated for 1 h, and imaged using a low resolution (10X, numerical aperture 0.3) objective, green fluorescence protein (GFP) filters, and an imaging camera. Analysis can be performed using freely-available ImageJ software or customized image analysis software. Our analysis was performed using our own software written using a scientific analysis package13. The software should create a mask using the phase contrast image and extract fluorescence values from the masked areas in the fluorescence image. Fluorescence values are averaged over at least 100 cells, resulting in a numerical host killing index.
Protocol
All experimental procedures were carried out at the University of California, Irvine.
1. Buffers and Solutions
- Prepare Lysogeny Broth (LB) in a 1 L glass bottle by adding 25 g of LB-Miller mix in 1 L of double-distilled water (ddH2O). For Petri dishes, add an additional 20 g of agar. Autoclave to sterilize. Pour 25 mL of molten agar medium into 10 cm-diameter Petri dishes and allow to solidify at room temperature. Store liquid media at room temperature and agar plates at 4 °C.
- Prepare Glucose Yeast Peptone (GYP) plates in a 1 L glass bottle by mixing 1 g of D-glucose, 2 g of peptone, 0.25 g of yeast extract, 4.2 g of KH2PO4, 5.1 g of Na2HPO47H2O, and 25 g of agar in 1 L of ddH2O. Autoclave to sterilize. Pour 25 mL of molten agar into 10 cm-diameter Petri dishes and allow to solidify at room temperature. Store at 4 °C.
- Prepare Peptone S (PS) medium in a 1 L glass bottle by mixing 10 g of peptone, 7 g of yeast extract, 15 g of D-glucose, 0.12 g of Na2HPO47H2O, 1.4 g of KH2PO4, 40 µg of vitamin B12, and 80 µg of Folic acid in 800 mL of ddH2O. Calibrate an electronic pH meter using pH 4 and pH 7 standards if the pH meter has not been used within a day.
- Measure the pH of the PS medium using the pH meter. Add either 5 M KOH or 5 M H3PO4 to adjust the pH to 6.5. Adjust the final volume to 1 L using ddH2O. Filter sterilize using 0.22 µm vacuum filter and store at 4 °C.
NOTE: Proceed cautiously with the pH adjustments by wearing protective equipment including gloves, protective clothing, eye protection, and face protection.
- Measure the pH of the PS medium using the pH meter. Add either 5 M KOH or 5 M H3PO4 to adjust the pH to 6.5. Adjust the final volume to 1 L using ddH2O. Filter sterilize using 0.22 µm vacuum filter and store at 4 °C.
- Prepare 10x Development Buffer Prime (DB') buffer in a 1 L glass bottle by adding 3.4 g of KH2PO4, 13.4 g of Na2HPO4 7H2O, and ddH2O to a total volume of 800 mL. Calibrate an electronic pH meter using pH 4 and pH 7 standards if the pH meter has not been used within a day.
- Measure the pH of the buffer using the electronic pH meter. Adjust the pH to 6.5 by gently adding either 5 M KOH or 5 M H3PO4 as needed. Adjust the final volume to 1 L using ddH2O and sterilize by autoclaving. Store at room temperature.
NOTE: Proceed cautiously with the pH adjustments by wearing protective equipment including gloves, protective clothing, eye protection, and face protection.
- Measure the pH of the buffer using the electronic pH meter. Adjust the pH to 6.5 by gently adding either 5 M KOH or 5 M H3PO4 as needed. Adjust the final volume to 1 L using ddH2O and sterilize by autoclaving. Store at room temperature.
- Make 1x Development buffer (DB) by mixing 100 mL of 10x DB' buffer with 1 mL of sterilized 2 M MgCl2, and 1 mL of sterilized 1 M CaCl2. Adjust the final volume to 1 L in a 1 L glass bottle using an autoclaved sterilized ddH2O. Store at room temperature.
- Prepare PS:DB medium in a 1 L glass bottle by mixing 100 mL of sterilized PS medium, 100 mL of sterilized 10x DB' buffer, and sterilized ddH2O to a total volume of 1 L. Supplement with 1 mL of sterilized 2 M MgCl2 and 1 mL of sterilized 1 M CaCl2. Store at 4 °C.
- Prepare the calcein-AM stock solution by adding 50 µL of DMSO to 50 µg of calcein-AM in the plastic container supplied by the manufacturer. Store at -20 °C.
2. Growth and Maintenance of Amoebae
- Grow E. coli strain B/r17 as a food source for amoebae during the initial growth period. Streak E. coli B/r from a frozen stock stored at -80 °C onto an LB agar plate and incubate at 37 °C overnight to obtain single colonies.
- Inoculate a single colony of E. coli B/r into 2 mL of LB medium and incubate in a roller drum or a shaker at 37 °C overnight.
- Bring the overnight E. coli culture to the room temperature. Using a wooden stick, inoculate amoebae from a frozen stock kept at -80 °C into 750 µL of the overnight culture.
NOTE: Use D. discoideum strain AX3, which is capable of axenic growth18. This step does not require axenic growth, but subsequent steps will have his requirement. - Immediately add 700 µL of the amoebae-E. coli mixture onto a Glucose Yeast Peptone (GYP) plate. Spread the culture evenly on the surface of the plate by tilting it back-and-forth and side-to-side as needed. Avoid the use of a spreader.
- Place the GYP plate with agar facing up in a box that maintains high humidity. Use two disposable plastic containers (183 cm x 183 cm x 91 cm) stacked on the top of each other. Fill the bottom container with approximately 25 mL of water and place the top container with several 0.5 cm-diameter holes drilled through the flooring of the container to allow moisture to move from the water reservoir to the top container.
- Incubate the GYP plate and box at 22 °C for 4 – 5 days until amoeba spores are observed growing near the surface of the plate (Figure 2). Use a refrigerated incubator to incubate plates rather than incubation at room temperature.
NOTE: After spores have grown, they remain viable on plates for several weeks when stored at 4 °C. Store the plates with agar side facing up. After 3 – 4 days of incubation, zones of clearing within the bacterial lawn can be observed, which indicate the onset of amoeba sporulation.- Observe small amoebae spores above the plate surface after 4 – 5 days of incubation (Figure 2).
NOTE: Do not grow the amoebae for longer than a week, as the quality of the spores decreases beyond this time period.
- Observe small amoebae spores above the plate surface after 4 – 5 days of incubation (Figure 2).
- Collect amoebae spores to prepare for the axenic growth of the amoebae by sweeping a sterile 1 mL pipette tip parallel to the surface of the plate. Collect spores from half of a 10 cm Petri dish.
NOTE: Avoid touching the tip to the surface of the agar plate as this motion will collect large numbers of E. coli. - Resuspend the amoebae on the tip of the pipette by immersing the tip into 1 mL of PS medium and gently pipetting up and down.
- Transfer the inoculated mixture into a 250-mL flask containing 25 mL of the PS medium supplemented with the antibiotic-antimycotic solution at 0.25x concentration.
NOTE: Store the antibiotic-antimycotic solution as 250 µL aliquots at -20 °C. Avoid repeated cycles of thawing and freezing of this solution as this may decrease its efficacy. - Grow amoebae axenically at 22 °C in a rotating shaker at 100 rpm. For the virulence assay in step 5, grow cells to an optical density measured at 600 nm (OD600) of 0.2 to 0.5.
NOTE: This step requires 3 – 4 days of growth from the time of inoculation (step 2.7). If cells have grown to a higher density, dilute the culture back in PS medium to an OD600 of 0.05 or lower and grow back to an OD600 of 0.2 to 0.5.
3. The Growth of P. aeruginosa
- Pick a single colony of P. aeruginosa from an LB plate, inoculate it into a 2 mL PS:DB culture, and grow overnight at 37 °C to saturation.
NOTE: Do not add the antibiotic-antimycotic solution to the PS:DB media. - Dilute the overnight culture 1:100 or 1:1,000 into a 6 cm-diameter Petri dish using PS:DB. If the virulence of planktonic cells is solely being assayed, grow the cultures using conventional culture tubes instead. Shake the culture on a benchtop rotator set at 100 rpm at 37 °C.
- Harvest cultures at specific time points to ensure reproducibility. At 30 min to 1 h prior to assessing virulence, prepare an agar pad as described in section 4.
NOTE: Virulence in P. aeruginosa is induced by approximately 8 h of growth on surfaces. - To isolate the surface-attached cells, remove the liquid medium from the Petri dish, wash immediately with 1 mL of DB buffer to remove planktonic cells, and proceed immediately to step 5.1.
CAUTION: Do not wash longer than 30 s as this will perturb and detach surface-attached cells and may affect the virulence measurement. - To isolate planktonic cells, transfer 10 µL of the culture from the Petri dish to a clean Petri dish and proceed immediately to step 5.1.
4. Microscopy – Preparation of the Agar Pad
- Prepare agar pads about 30 min to 1 h before amoebae are ready to be mixed with P. aeruginosa.
- Microwave 50 mL of 1% (w/v) agar in DB buffer in a 250 mL flask. Mix every 20 s until clumps of agar disappear.
- Cool the molten agar by placing it in preheated 55 °C water bath for 15 min.
- Add 15 µL of the calcein-AM stock solution to 15 mL of the molten agar in a conical tube. Mix by gently inverting the tube 4 – 5 times and proceed immediately to the next step.
NOTE: Perform this step in a polypropylene 15 mL centrifuge tube. Do not use a thick glass container as this will cause the agar to solidify too quickly. - Pour the molten agar mixture on top of a 12 cm x 10 cm glass plate and allow the agar to solidify for 10 minutes at room temperature. Cut the solidified agar pad into smaller 1.5 cm x 1.5 cm sections by placing the glass plate over a printed grid and cutting along the grid using a metal ruler.
- Keep the gel hydrated in a humid box until it is ready for use.
5. Microscopy – Preparation of the Bacteria-amoeba Sample for Imaging
- To assay the virulence of surface-attached bacterial cells, add 10 µL of amoeba cells from step 2.10 to surface-attached bacteria from step 3.4.
NOTE: This step describes the mixing of amoebae with bacteria. - To assay the virulence of planktonic cells, mix 10 µL of amoeba cells from step 2.10 with 10 µL of planktonic bacteria from step 3.5.
NOTE: Do not wash the axenically-grown amoeba cells (at an OD600 of 0.2 to 0.5) before mixing with the bacteria. No E. coli growth will be observed in the amoeba culture due to the use of the antibiotic-antimycotic solution. - Immediately place the 1.5 cm x 1.5 cm calcein-AM agar pad from step 4.5 on top of the bacteria-amoeba mixture on the Petri dish surface.
NOTE: Place the agar pad on top of the mixture using a quick smooth motion. Do not lower the pad slowly onto the mixture as this motion tends to push cells towards the periphery and away from the underside of the pad. This step will ensure that bacteria and amoebae are evenly mixed, immobilized and that both cell types are confined to the same plane. - Remove excess liquid by gently pipetting out any remaining liquid surrounding the agar pad. Allow the Petri dish to dry for 20 min at room temperature.
- Cover with the Petri dish with a lid and incubate the immobilized amoeba-bacteria mixture at room temperature for an additional 40 min. Proceed to step 6.1.
NOTE: It is important to incubate all samples for the same amount of time, as the background fluorescence increases over time. An incubation time of 1 h is recommended. Incubations for longer than 1 h are less reproducible as amoebae cells may begin to burst.
6. Microscopy – Image Acquisition
- Image amoebae using a fluorescence microscope capable of acquiring brightfield and green fluorescence images. For green fluorescence, use green fluorescence protein (GFP) filters with 474/27 nm and 525/45 nm for excitation and emission, respectively. Use a 10X (≥0.3 numerical aperture) objective to image the amoebae.
NOTE: The calcein-AM in the agar will cause dying amoebae to fluorescence in the green fluorescence channel. Healthy amoebae are otherwise not fluorescent. - Adjust acquisition times for the phase contrast, GFP channels to limit phototoxicity, and optimize the dynamic range of the image intensities. Substitute the differential interference contrast (DIC) phase contrast if necessary.
NOTE: Use 100 – 200 ms exposures for phase contrast and GFP fluorescence if possible. Keep the exposure and instrument settings unchanged throughout the entire experiment. This is critical for computing the host killing index. - Acquire images for at least 100 amoeba cells in order to obtain good statistics for calculating the host killing index.
7. Image Analysis and Host Killing Index
- Perform image analysis using ImageJ.
- Open the phase contrast image file in ImageJ by clicking on File | Open and selecting the image. Adjust the threshold by clicking on Image | Analysis | Threshold. Adjust the lower and upper threshold to select only the intensity peak (Figure 3A).
- Convert the image to binary by clicking Process | Binary | Make Binary. Fill holes by selecting Process | Binary | Fill Holes.
NOTE: Using the image in Figure 1 as the source, steps 7.1 – 7.2 will produce an image in which the amoebae and surface-attached bacteria appear as dark regions (Figure 3B). - Ensure that the Area and Mean gray value boxes are checked in Analyze | Set Measurements. Measure the approximate size of the amoebae by selecting the oval tool on the main toolbar. Click Analyze | Measure and note the area of representative amoebae.
- Create masks for the amoebae by clicking Analyze | Analyze Particles. In the size box, enter the area that will include the amoebae but exclude the bacteria. Select Masks in the dropdown menu bar for the Show option. Ensure that the Add to Manager box is checked.
- Click OK and an image showing only the amoebae and an ROI Manager dialog will appear (Figure 3C).
- Open the fluorescence image using File | Open and selecting the appropriate file. Select all the regions in the ROI Manager by holding down the Shift Key and Clicking on each region in the list (Figure 3D).
- Click the Measure button in the ROI Manager. This will open a Results window displaying the mean intensity value of each amoeba.
- Copy the fluorescence values for each amoeba into a spreadsheet program and calculate the host killing index by taking the average of all fluorescence values.
Representative Results
We grew wild-type P. aeruginosa strain PA1419 or a ΔlasR strain20 in the same PA14 background in 6 cm-diameter Petri dishes and assayed the virulence of planktonic and surface-attached cells. Cultures were inoculated from single-colonies into PS:DB cultures, grown overnight in culture tubes in a roller drum at 37 °C to saturation, diluted 1:100 into PS:DB, grown for 8 h in 6 cm-diameter Petri dishes shaking at 100 rpm, and planktonic and surface-attached P. aeruginosa sub-populations were isolated as described in section 3.
Surface-attached wild-type P. aeruginosa killed amoebae, which was indicated by a round amoeba cell shape and the observation of calcein fluorescence (Figure 4A, top-left panel). This resulted in a high host killing index (Figure 4B). Planktonic cells were consumed by the amoebae, which was indicated by amorphous amoeba cell shapes and the observation of little or no calcein fluorescence above background (Figure 4A, top-right panel). This resulted in a relatively low host killing index (Figure 4B). Both surface-attached and planktonic cells of the ΔlasR strain were consumed by the amoebae, which was indicated by low host killing indexes (Figure 4B, bottom panel). The results thus show that virulence is upregulated in surface-attached populations and the LasR is required for the surface-activated virulence, which was described previously13. These experiments thus establish robust positive and negative controls for future experiments.
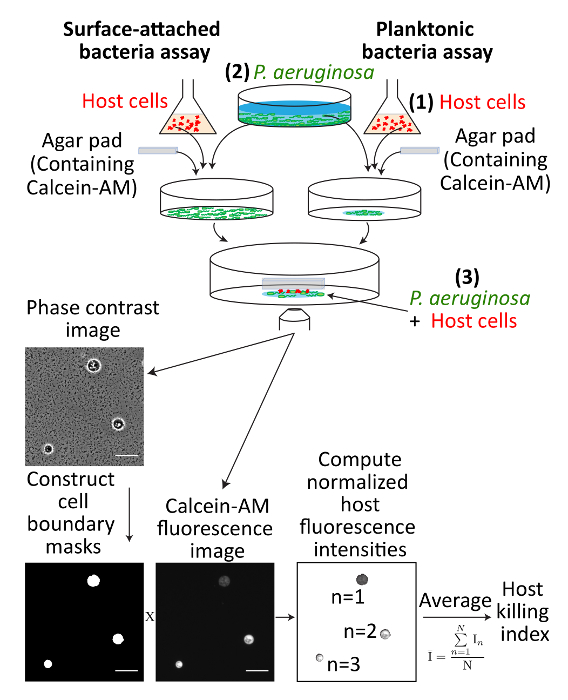
Figure 1: Schematic describing an overview of the virulence assay. (1) Amoeba host cells are grown, (2) bacterial cultures are grown, and (3) planktonic or surface-attached bacterial cells are mixed with amoebae and immobilized onto the same imaging plane using an agar pad. Host cells are quantified for health using calcein-AM, fluorescence microscopy, and image analysis. Scale bars represent 50 µm. Please click here to view a larger version of this figure.
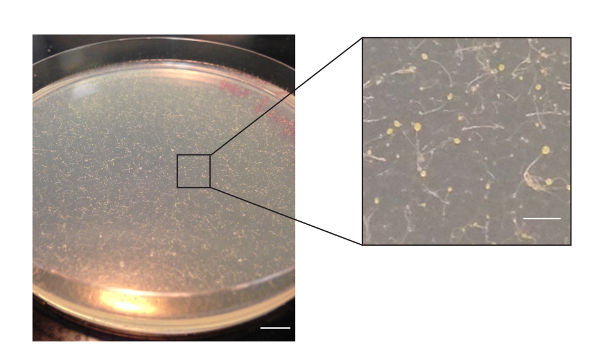
Figure 2: Amoeba spores on a GYP 10 cm-diameter Petri dish plate after 5 days of growth. Spores form above the Petri dish surface. The inset shows a magnified view of a single section of the dish surface. The surface may contain bacteria which cannot be discerned at this resolution. Scale bars in main and inset images represent 1 cm and 2 mm, respectively. Please click here to view a larger version of this figure.
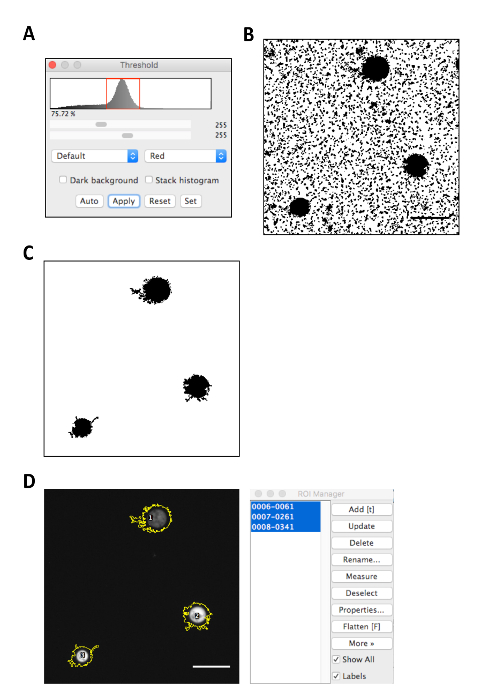

Figure 3: Image analysis of microscopy data using ImageJ. (A) Selection of the peak intensity using the threshold tool. (B) Resulting image after the phase contrast source image from Figure 1 is converted into a binary image. (C) Resulting image after the image is subsequently converted to a mask. (D) Screenshots of the selection of regions of interest using the ROI Manager. Scale bars represent 50 µm. Please click here to view a larger version of this figure.
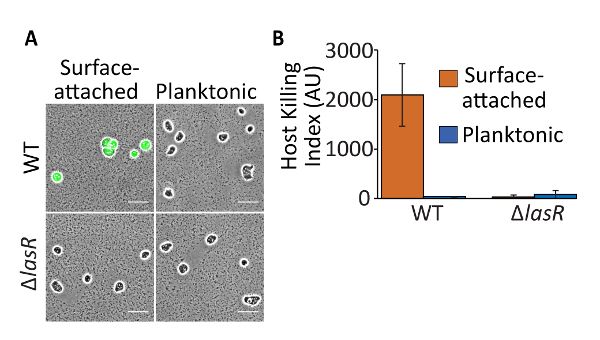
Figure 4: Representative results of amoeba killing by P. aeruginosa cells. (A) Calcein-AM fluorescence overlaid on phase microscopy images and (B) host killing indexes of surface-attached or planktonic wild-type or ΔlasR P. aeruginosa. Wild-type surface-attached P. aeruginosa kills amoebae, which exhibit significant calcein fluorescence and hence a significant host killing index. Planktonic cells are consumed by amoebae, which exhibit little calcein fluorescence and produce a low host killing index. Both surface-attached and planktonic ΔlasR are consumed by amoebae and result in a low host killing index. Scale bars represent 50 µm. Bars indicate the average of three independent experiments and errors bars indicate standard deviation. Please click here to view a larger version of this figure.
Discussion
This protocol describes a rapid and quantitative method to assay virulence in P. aeruginosa. This protocol may be tested with other bacteria. However, it is important to keep in mind that the growth medium should be compatible with the amoeba growth conditions. In particular, we have optimized the protocol using PS:DB as the bacterial growth medium. If other media are used, it may be necessary to perform a growth media-only control in which there are no bacterial cells present to verify that the medium is compatible with the amoebae.
This method can be expanded to assay the growth phase dependence of virulence. In particular, bacterial cultures can be grown for a wide range of times instead of a fixed time point. In our experience, it is important to harvest bacterial cultures at specific time points as virulence in P. aeruginosa appears to be dependent on the growth phase. We observed that virulence in P. aeruginosa was induced between 6 – 8 h of growth in Petri dishes.
Many variables associated with the growth environment affect host health and bacterial virulence. Thus, it is important to use proper positive and negative controls for each experiment. We have used wild-type surface-attached P. aeruginosa as a positive control and ΔlasR as a negative control. We suggest performing these controls each time the virulence assay is performed. These controls are important for verifying that host cell deaths are not due to any external factors such as the age of the amoebae, variations in temperature, etc. Furthermore, we suggest performing all experiments in biological replicate to establish the reproducibility of the virulence phenotypes.
We have performed our experiments using amoebae at an optical density of 0.2 to 0.5 after a dilution of at least 1:10 and have regulated the temperature of amoebae culture precisely to 22 °C. Outside of these growth conditions, we have found that the susceptibility of amoebae to virulent bacteria and the reproducibility of the virulence assay are altered. Amoeba cells pellet relatively quickly. Ensure that cultures are resuspended by pipetting up and down immediately before the optical density measurement is made. Do not grow the amoeba cultures higher than an OD600 density of 1 as growth to high densities affects the reproducibility of the experiment. Propagate axenic cultures for no longer than 1 week. In addition, it is important to verify that amoebae are grown axenically (beginning in step 2.7) through 20X or high magnification microscopy such that other microbes are not present in the culture. If cultures are turbid in step 2.10 after 2 days of growth following initial inoculation, this would likely indicate the presence of a microbial contaminant. If bacterial contamination is suspected, we suggest growing 1 mL of the amoebae culture at 37 °C for 4 – 8 h. The observation of a turbid culture under these conditions would suggest bacterial contamination. If repeated bacterial contamination is observed, the antibiotic-antimycotic solution should be replaced.
The host killing index is a reliable indicator of whether a bacterial population is virulent or avirulent. This assay has not been optimized for comparisons between different intermediate levels of virulence (i.e., low virulence compared to medium-low virulence) and over-interpretation of the results should be avoided. Potential methods to address this issue are the repetition of the virulence assay over multiple days using different batches of host cells to establish the confidence in the results, performing additional replicate experiments, and performing appropriate statistical analyses.
The multiplicity of infection (MOI) of planktonic cells can be controlled by normalizing the optical density of the bacterial culture. However, this assay does not control the MOI of surface-attached bacteria cells. We have observed that bacterial surface density increases with time. Thus, adjusting growth time in the Petri dish may result in a corresponding change in surface density. In addition, the surface density depends on the material of the surface. P. aeruginosa cells attach to polystyrene surfaces in the assay described here. However, other surfaces including glass, agar, and polyacrylamide may also be assayed13. The surface density of single-layered bacterial populations can be measured using 100X magnification phase microscopy. If surface densities are multi-layered, confocal microscopy may be appropriate.
Here, we have described a rapid and robust method to measure bacterial virulence. By modifying individual parameters such as incubation times, the fluorescent dye, the bacterial strain, the host cell type, or growth media, this method may be extended to quantify virulence across a wide range of organisms and growth conditions.
Disclosures
The authors have nothing to disclose.
Acknowledgements
KP and AS wrote and revised the manuscript. KP performed the experiments and the analysis. This work was supported by the National Institutes of Health (NIH) Career Transition Award (K22AI112816) to AS.
Materials
| Reagents | |||
| Bacto agar, dehydrated | BD Difco | 214010 | |
| Antibiotic-Antimycotic (100X) | Life Technologies | 15240062 | Aliquot < 1 mL and store at -20 °C |
| Calcein-acetoxymethyl ester (calcein-AM) | Life Technologies | C34852 | Calcein Acetoxymethyl (AM) |
| Calcium chloride, anhydrous | Sigma-Aldrich | C1016 | |
| D-Glucose | Fisher Chemical | D16500 | Dextrose |
| Dimethyl sulfoxide | Sigma-Aldrich | D5879 | |
| Folic acid | Sigma-Aldrich | F8758 | |
| LB-Miller | BD Difco | 244620 | |
| Magnesium chloride | Sigma-Aldrich | M8266 | |
| Yeast extract | Oxoid | LP0021 | |
| Special peptone | Oxoid | LP0072 | |
| Potassium hyroxide | Fisher Chemical | P250 | |
| Potassium phosphate monobasic | Sigma-Aldrich | P0662 | |
| Sodium phosphate dibasic heptahydrate | Fisher Chemical | S373 | |
| Vitamin B12 | Sigma-Aldrich | V2876 | |
| Strains | |||
| Dictyostelium discoideum | Siryaporn lab | Strain AX318 | |
| Escherichia coli | Siryaporn lab | Strain B/r17 | |
| Pseudomonas aeruginosa | Siryaporn lab | PA14 | PA14 strain19 |
| Pseudomonas aeruginosa ΔlasR | Siryaporn lab | AFS20.1 | PA14-derived strain20 |
| Supplies | |||
| 0.22 µm filter | Millipore | SCGPT01RE | For filter sterilization |
| Conical tube, 15 mL | Corning | 352097 | |
| Glass storage bottles | Pyrex | 13951L | 250 mL, 500 mL, 1000 mL |
| Petri dish, 6 cm diameter | Corning | 351007 | 60 x 15 mm polystyrene plates |
| Petri dish, 10 cm diameter | Fisher | FB0875712 | 100 x 15 mm polystyrene plates |
| Plastic containers with lid | Ziploc | 2.57E+09 | Square 3-cup containers |
| Glass plates | Bio-Rad | 1653308 | For preparing agar pads. Other glass plates may be used with similar dimensions. |
| Wooden sticks | Fisher | 23-400-102 | |
| Equipment | |||
| Eclipse Ti-E microscope | Nikon | MEA53100 | Microscope setup |
| 10X Plan Fluor Ph1 objective 0.3 NA | Nikon | MRH20101 | Microscope setup |
| Fluorescence excitation source | Lumencor | Sola light engine | Microscope setup |
| Fluorescence filter set | Semrock | LED-DA/FI/TX-3X3M-A-000 | Microscope setup |
| Orca Flash 4.0 V2 Camera | Hamamatsu | 77054098 | Microscope setup |
| ImageJ | NIH | v. 1.49 | Software for image analysis |
| MATLAB | Mathworks | R2013 | Software for image analysis |
| Orbital shaker incubator | VWR | 89032-092 | For growth of bacteria at 37 °C |
| Platform shaker | Fisher | 13-687-700 | For growth of amoebae at 22 °C |
| Spectrophotometer | Biochrom | Ultrospec 10 | |
| Undercounter refrigerated incubator | Fisher | 97990E | For growth of amoebae at 22 °C |
| Isotemp waterbath | Fisher | 15-462-21Q | For cooling media to 55 °C |
References
- Rasko, D. A., Sperandio, V. Anti-virulence strategies to combat bacteria-mediated disease. Nature Reviews Drug Discovery. 9 (2), 117-128 (2010).
- Coburn, B., Grassl, G. A., Finlay, B. B. Salmonella, the host and disease: a brief review. Immunology and Cell Biology. 85 (2), 112-118 (2007).
- Alibaud, L., et al. Pseudomonas aeruginosa virulence genes identified in a Dictyostelium host model. Cellular Microbiology. 10 (3), 729-740 (2008).
- Pukatzki, S., Kessin, R. H., Mekalanos, J. J. The human pathogen Pseudomonas aeruginosa utilizes conserved virulence pathways to infect the social amoeba Dictyostelium discoideum. Proceedings of the National Academy of Sciences U S A. 99 (5), 3159-3164 (2002).
- Cosson, P., et al. Pseudomonas aeruginosa Virulence Analyzed in a Dictyostelium discoideum Host System. Journal of Bacteriology. 184 (11), 3027-3033 (2002).
- Pukatzki, S., et al. Identification of a conserved bacterial protein secretion system in Vibrio cholerae using the Dictyostelium host model system. Proceedings of the National Academy of Sciences U S A. 103 (5), 1528-1533 (2006).
- Steinert, M., Heuner, K. Dictyostelium as host model for pathogenesis. Cellular Microbiology. 7 (3), 307-314 (2005).
- Hagele, S., Kohler, R., Merkert, H., Schleicher, M., Hacker, J., Steinert, M. Dictyostelium discoideum: a new host model system for intracellular pathogens of the genus Legionella. Cellular Microbiology. 2 (2), 165-171 (2000).
- Matz, C., et al. Pseudomonas aeruginosa uses type III secretion system to kill biofilm-associated amoebae. ISME Journal. 2 (8), 843-852 (2008).
- Hauser, A. R. The type III secretion system of Pseudomonas aeruginosa: infection by injection. Nature Reviews Microbiology. 7 (9), 654-665 (2009).
- Moore, P. L., MacCoubrey, I. C., Haugland, R. P. A rapid pH insensitive, two color fluorescence viability (cytotoxicity) assay. Journal of Cell Biology. 111, 58 (1990).
- Kaneshiro, E. S., Wyder, M. A., Wu, Y. -. P., Cushion, M. T. Reliability of calcein acetoxy methyl ester and ethidium homodimer or propidium iodide for viability assessment of microbes. Journal of Microbiological Methods. 17 (1), 1-16 (1993).
- Siryaporn, A., Kuchma, S. L., O’Toole, G. A., Gitai, Z. Surface attachment induces Pseudomonas aeruginosa virulence. Proceedings of the National Academy of Sciences U S A. 111 (47), 16860-16865 (2014).
- Decherchi, P., Cochard, P., Gauthier, P. Dual staining assessment of Schwann cell viability within whole peripheral nerves using calcein-AM and ethidium homodimer. Journal of Neuroscience Methods. 71 (2), 205-213 (1997).
- Braut-Boucher, F., Pichon, J., Rat, P., Adolphe, M., Aubery, M., Font, J. A non-isotopic, highly sensitive, fluorimetric, cell-cell adhesion microplate assay using calcein AM-labeled lymphocytes. Journal of Immunological Methods. 178 (1), 41-51 (1995).
- Grieshaber, P., Lagrèze, W. A., Noack, C., Boehringer, D., Biermann, J. Staining of fluorogold-prelabeled retinal ganglion cells with calcein-AM: A new method for assessing cell vitality. Journal of Neuroscience Methods. 192 (2), 233-239 (2010).
- Witkin, E. M. Inherited Differences in Sensitivity to Radiation in Escherichia Coli. Proceedings of the National Academy of Sciences U S A. 32 (3), 59-68 (1946).
- Loomis, W. F. Sensitivity of Dictyostelium discoideum to nucleic acid analogues. Experimental Cell Research. 64 (2), 484-486 (1971).
- Rahme, L. G., Stevens, E. J., Wolfort, S. F., Shao, J., Tompkins, R. G., Ausubel, F. M. Common virulence factors for bacterial pathogenicity in plants and animals. Science. 268 (5219), 1899-1902 (1995).
- O’Loughlin, C. T., Miller, L. C., Siryaporn, A., Drescher, K., Semmelhack, M. F., Bassler, B. L. A quorum-sensing inhibitor blocks Pseudomonas aeruginosa virulence and biofilm formation. Proceedings of the National Academy of Sciences U S A. 110 (44), 17981-17986 (2013).

