Functional Surface-immobilization of Genes Using Multistep Strand Displacement Lithography
Instructor Prep
concepts
Student Protocol
NOTE: A time schedule for the steps in the different sections is given in the supplementary information (section 1).
1. Chip Fabrication
NOTE: As substrates, use silicon wafers (100 mm diameter, 0.525 mm thickness) with a 50 nm thick layer of silicon dioxide or glass slides (24 mm x 24 mm, no. 1.5; 22 mm x 50 mm, no. 4). Depending on the application, other sizes and thicknesses may be more suitable.
- Cleaning the substrates via an RCA clean procedure (water, ammonia solution NH3(aq) 30% and hydrogen peroxide H2O2 30%, at a ratio 5:1:1)
- Mix water and ammonia solution in a beaker glass and heat the mixture to 70 °C on a hot plate while stirring.
- Add the hydrogen peroxide and the substrate. For higher throughput, place multiple substrates (especially the small glass slides) in a polytetrafluoroethylene (PTFE) holder.
- After 30 min, take out the substrate, rinse it thoroughly with water from a wash bottle, and dry it with a nitrogen gun. Immediately proceed with the PEGylation of the substrate.
CAUTION: Wear skin and eye protection and work in a fume hood. Do not close the waste container tightly in order to allow for gas release from the mixture.
- PEGylation of the substrate
NOTE: Repeated freezing and thawing of the Biotin-PEG-Silane stock reduces the reactivity of the silane due to condensation of air moisture. Therefore, prepare 5 mL-tubes with appropriate amounts of Biotin-PEG-Silane (e.g. 10-20 mg) and fill them with dry argon before freezing them until use.- Dissolve Biotin-PEG-Silane in dry toluene by vortexing (5 mg/mL).
- Place the substrate in a glass petri dish (120 cm diameter, 2 cm height) in a fume hood.
- Pipette the solution (≈200 µL for a single glass cover slip, a few mL for a large silicon wafer) onto the substrate, covering the whole surface, but avoid the solution flowing over the edge of the substrate.
- Close the petri dish. To reduce drying of the substrate, add an additional cover with a wet paper towel.
- After 30 min, add approximately 40 ml of isopropyl alcohol, then take out the substrate, rinse it again thoroughly with isopropyl alcohol and dry it with a nitrogen gun (for glass slides, remember the PEGylated side). Store the substrate in the dark until use.
CAUTION: Wear skin and eye protection and work in a fume hood.
2. Preparation of Genes for Immobilization
NOTE: Primer sequences, DNA modifications and an exemplary PCR protocol are given in the supplementary information (sections 2-4).
- Use modified primers to amplify the gene of interest by polymerase chain reaction (PCR). One primer carries a fluorophore for visualization in fluorescence microscopy, and the other primer is separated from the DIS sequence by a triethylene glycol spacer in order to keep the DIS sequence single-stranded throughout the PCR.
- Purify the DNA using a spin column purification kit and measure its concentration using an absorption photometer. The concentration should not be much lower than 100 nM.
- Adjust the concentration of NaCl to 1 M using a concentrated 5 M NaCl solution. The high salt concentration facilitates the DNA brush assembly process22.
3. Photolithography
NOTE: The photocleavable DNA (PC) should be handled only in a yellow-light environment. Yellow foil for cleanrooms can be used to filter the light of conventional white light lamps.
- Preparation of the substrate
- Prepare the Bephore-mix (1 µM streptavidin, 1.5 µM PC DNA, 7.5 µM PH DNA in 1x PBS (phosphate-buffered saline)) and the passivation mix (10 µM unlabeled DIS DNA, 1 mg/mL BSA (bovine serum albumin), in 1x PBS). Both can be stored in the dark in the fridge for several weeks.
- Cut the substrate into smaller pieces (a few cm2) either by using a glass cutter or by breaking a silicon wafer along a crystal line by pressing onto an edge with a scalpel. As a simple alignment mark, create a short scratch from the center towards an edge of the substrate using the glass cutter. Blow small particles off the chip with a nitrogen gun.
- Mix two droplets of two-component silicone glue, stirring it e.g. with a pipette tip, and apply the glue to the surface of the chip, leaving the area around the tip of the scratch blank ( Figure 2A). The glue provides a hydrophobic barrier that facilitates washing steps and reduces the amount of DNA required for incubations. In a later step, the glue can be easily peeled off.
- Incubate 10 µL (or as much as necessary to cover the chip) of the Bephore-mix at room temperature (RT) on the chip. During the incubation, place the chip in a box, e.g. a pipette tip box partially filled with water to reduce evaporation.
- After 1 h, wash the chip several times (5x with 50 µL) by pipetting with 1x PBS to remove unbound Bephore-mix.
- Take off as much buffer as possible without drying the chip and incubate 10 µL of the passivation-mix on the chip at RT.
- After 2 h, wash the chip several times (10x with 50 µL) by pipetting with 1x PBS to remove unbound passivation agents.
- Projection lithography
NOTE: For lithography, an upright microscope with a 60x water immersion objective can be used. Illumination times depend on the objective, the light source, the mask, the substrate (silicon chip or glass slide) and the desired DNA surface density. With a 60x objective, typical exposure times ranged from below one minute for a two-step lithography with overlapping regions (15 s for silicon chips, 45 s for glass slides) to 2-3 min for a full exposure. Do not use a cover slip (except fused quartz) on top of the substrate, since it blocks UV light.- Cut a printed photomask (see supplementary file with exemplary lithography masks and supplementary section 5) to an appropriate size to fit a suitable mask holder, which can be inserted at the position of the field stop (Figure 2B). For precise multi-step lithography, align the mask with alignment marks on the holder.
- Place the substrate on a microscopy slide and move it to the microscope stage. For patterning of Bephore glass slides, insert black foil (e.g. spare foil from the printed photomasks) between microscopy slide and Bephore slide to provide a dark background for lithography.
- Insert a red filter into the illumination path, focus onto the substrate surface and navigate to the region of the substrate, which will be exposed, e.g. the region at the tip of the scratch.
- Insert the mask holder, block the illumination and change to the UV filter (Figure 2C). At a low light intensity, open the shutter and use the camera to quickly focus the mask on the substrate.
- Block the light path to the camera and illuminate the substrate at a high light intensity for the desired exposure time.
- Take the sample from the microscope and carefully remove as much buffer as possible from the substrate without drying it.
- Add 10-20 µL of the DNA with DIS-sequence. Place the substrate in a wet box to keep it from drying. Incubate the DNA on the substrate for 2 h (oligonucleotides at a micromolar concentration) or several hours/overnight (kbp-long DNA at approximately 100 nM concentration) at RT.
- Remove the DNA from the chip and wash it several times with 1x PBS. In order to reduce the waste of DNA (especially for longer fragments), store the DNA taken from the substrate and reuse it.
- Multi-step lithography
NOTE: For precise alignment of two or more exposures in a single area, keep the sample on the stage to avoid angular misalignment. Make sure the chip does not dry up. For the positioning of several DNA brushes in different regions on a substrate, use a motorized stage to drive to the designated area or use a substrate with appropriate alignment marks.- After the first exposure (section 3.2), incubate DNA with DIS-sequence on the substrate. It is important that all binding sites resulting from the first exposure are occupied. Therefore, incubate oligonucleotides (10 µM) for approximately 3 h at RT.
- To immobilize DIS-labeled genes, place them on the substrate for several hours or overnight at RT, then wash the sample and add the passivation mix for another 2 h at RT. Proceed with the next exposure.
- For additional exposures, repeat the previous steps (3.2.3 – 3.3.2).
4. PDMS Devices
NOTE: Preferably, work in a clean-room. The fabrication of a PDMS device follows a standard protocol such as described by McDonald et al.30
- Fabrication of master molds via photolithography
- Clean a silicon wafer (76.2 mm diameter, 525 µm thickness) by sonication in acetone for 5 min at high power and rinse with isopropyl alcohol and de-ionized water. Afterwards place the wafer on a hot plate at 150 °C for 15 min.
- Optionally, in order to improve the adhesion of resist to the wafer, add 2-3 mL adhesion promoter and spin at 3000 rpm for 30 s. Heat the wafer to 120 °C for 2 min.
- Add about 2-3 mL photoresist (see Table of Materials). Spin the wafer for 15 s at 500 rpm, then at 3000 rpm for 30 s. This should result in a 20 µm thick layer of photoresist. Let the photoresist layer relax at room temperature for 10 min.
- For the pre-exposure bake, place the wafer on a hot plate at 50 °C for 2 min, then at 85 °C for 5 min.
- Place the wafer under the photomask with the blueprint of the compartments. Preferably, use a mask-aligner. The photomask should have a resolution of 64,000 dpi (see supplementary file with lithography masks and supplementary section 5).
- Expose the wafer for 1 min with UV-light (I-line 5-10 mW/cm2) through the photomask.
- For the post-exposure bake, place the wafer at 50 °C for 2 min and then at 85 °C for 5 min. Let the wafer cool down and relax for 1 h at room temperature.
- Place the wafer in a dish with developer for 3 min while gently agitating the beaker. Rinse the wafer with isopropyl alcohol and dry it with a nitrogen gun. Place the wafer on a hot plate at 130 °C for 45 min.
- Preparation of the PDMS device
- Mix PDMS base and PDMS curing agent in a 10:1 ratio and pour it onto the wafer in a closed container until a layer of about 5 mm has formed above the wafer. To obtain a PDMS device of only 1 mm thickness, instead spin coat the wafer with PDMS at 100 rpm for 2 min.
- Place the container in a desiccator and apply a vacuum (about 85 kPa) for about 30 min. Then, heat the PDMS to about 70°C for 1-2 h in an oven.
- Separate the PDMS device from the wafer. Cut the PDMS into slabs so that each piece contains one set of compartments. In case the PDMS will be connected to a microfluidic setup, punch holes (1 mm diameter) as an inlet and outlet at the ends of the supply channel.
- Clean the PDMS with isopropyl alcohol and de-ionized water and dry it with a nitrogen gun. Store the PDMS dust-free.
5. Compartmentalized Gene Expression
NOTE: The following procedure describes the assembly of a sample holder (Figure 3) for the observation of compartmentalized gene expression on an inverted microscope with a cage incubator for temperature control. The holder was built using readily available materials and tools (3.5-5 mm thick polyvinyl chloride (PVC) plastics, screws and nuts, drill) and can be customized to fit different types of microscopes. The steps described in 5.1 and 5.2 should be performed such that both parts of the holder are ready at the same time.
- Bottom holder assembly
- Prepare a Bephore chip with alignment mark (e.g. a scratch) and glue it onto the chip holder using double-sided adhesive tape (Figure 3B). The chip should be larger than the PDMS device which will cover it.
- Immobilize one or multiple genes on the chip (see sections 2 and 3).
- Add some vacuum grease around the central hole of the bottom holder, then plug in the chip holder.
NOTE: The chip holder now adheres to the bottom holder by means of the grease, while retaining the possibility for re-positioning of the chip in the x-y-plane.
- Top holder assembly
- Prepare a thin PDMS chip with compartments using the spin coating procedure in step 4.2.1. Cut the PDMS as small as possible, leaving a channel open to one side to allow for the exchange of waste and precursor molecules by diffusion.
- Clean a glass cover slip (24 mm x 24 mm, no. 1.5) and the backside of the PDMS chip (side without compartments) with oxygen plasma for 42 s (operated at 200 W and 0.8 mbar with the sample in a Faraday cage).
- Place the PDMS chip at the center of the glass slide with the compartments pointing upwards. Bake the glass with the PDMS for 1 h at 70 °C.
- Add some vacuum grease around the large hole of the top holder.
- Before starting the experiment, clean the glass slide and the PDMS chip with oxygen plasma for 42 s to render the PDMS hydrophilic.
- Place the glass slide with the PDMS onto the top holder ( Figure 3C) and gently press the glass onto the grease (Figure 3C).
- Holder assembly
- Prepare 100 µL of a cell-free expression system and keep it on ice.
- Carefully remove the buffer from the chip and wash it once with 10 µL of cell-free expression system. Next, add 60 µL of cell-free expression system onto the chip and remove the two-component silicone glue from the edges of the chip (already removed in Figure 3B).
- Add 20 µL of expression system onto the PDMS. After placing the droplet onto the PDMS, quickly check in the stereoscopic microscope that the compartments are well wet and without air bubbles. If there are air bubbles, try to wash them off with the rest of the cell-free expression system.
- To assemble the two pieces of the holder and to align chambers and DNA brushes, work under a stereoscopic microscope and immobilize the bottom holder with a gripper arm, so the two screws and wingnuts are easily accessible with both hands (Figure 3D-F).
- Insert the top holder into the bottom holder and lower it until the cell-free expression system droplets fuse (Figure 3D). Check through the stereoscopic microscope whether the compartments and the alignment mark are in a similar region in the x-y-plane, otherwise pull up the top holder and re-position the glass slide on the top holder.
- From the bottom, screw up the wingnuts until they touch the bottom side of the holder. Carefully tighten the wingnuts, while aligning the compartments and the chip in the x-y-plane via the handle at the bottom of the chip holder (Figure 3E). Once in contact, tighten the wingnuts only very gently (Figure 3F).
NOTE: This step requires some experience and depends on the experimental setup. Under the microscope, interference patterns arising from liquid between PDMS and glass can indicate that the applied force is too small, while a lower fluorescence intensity (from a DNA brush or expressed proteins) in the center of a compartment hints at a too high pressure (see also supplementary information, section 6). - Spray the sponge in the anti-evaporation enclosure with water and insert the holder into the box. Fill a 5 mL syringe with two-component silicone glue and use it to seal the box (Figure 3G).
- Transfer the box to a temperature-controlled microscope and image the DNA brush and the reaction in fluorescence microscopy. Even without illumination, the fluorescence of the DNA brush fades in a cell-free expression system. Nevertheless, the brush retains its activity, as can be shown by repeated gene expression from the same brush.
NOTE: When using a Bephore glass slide instead of a silicon chip, the position (top/bottom) of PDMS and Bephore chip can be exchanged, allowing for the use of objectives with smaller working distances.
6. Sustained Expression in Microfluidic Devices
NOTE: The experimental setup is assembled from the parts shown in Figure 4A. Details on the assembly of the temperature-controlled stage are given in the supplementary information (section 7).
- Cell-free expression system and tubing
- Prepare the cell-free expression system in a sample tube (about 200 µL). Polystyrene beads labeled with fluorescent dyes can be added to later monitor the flow rate through a supply channel.
- Connect the tube to a pressure controller. Place the tube in a large ice reservoir that keeps cold for about 12 hours. Refill ice if necessary.
NOTE: As an alternative to a pressure controller, a syringe pump provides a constant flow rate of cell-free expression system even if the flow resistance changes, e.g. due to air bubbles, which might help with long term experiments. - Connect the tube containing the cell-free expression system to PTFE tubing (inner diameter of 0.8 mm), which reaches to the heated stage. Break a syringe needle (diameter of 0.9 mm) into 1-2 cm pieces with blunt ends and insert one piece into the PTFE tubing. It will later serve as connector to the PDMS. Try to minimize the length of the tubing between the stage and the tube (Figure 4C).
- To cool the PTFE tubing, wrap it around larger rubber tubing that is connected to an ice water reservoir and a peristaltic pump. Tape them together with Scotch tape (Figure 4C).
- Top holder assembly
- Clean the flat side of the PDMS chip with oxygen plasma (42 s) and place it on a microscopy slide with holes fitting inlet and outlet of the PDMS (Figure 4B). The holes in the glass can be drilled beforehand with a glass drilling head. Make sure that the micro-chambers are not facing the glass slide.
- Apply double-sided adhesive tape to the flat side of the PDMS holder. Stick a piece of black foil (e.g. from a lithography mask) in the region between the two holes of the holder to avoid background fluorescence from the adhesive tape.
- Stick the glass slide with the PDMS chip to the holder.
- Treat the PDMS and the PDMS holder with the attached glass slide with oxygen plasma in a plasma cleaner (42 s).
- Apply vacuum grease around the large hole in the top holder and insert the PDMS holder (Figure 4B). The PDMS holder now adheres to the top holder, while retaining the possibility for re-positioning in x-y directions.
- Connect the PTFE tubing with the syringe needle connector to the inlet of the PDMS chip. To facilitate access to the PDMS through the top holder, the upper plastic plate can be briefly removed for this and the following step.
- Insert a second piece of PTFE tubing (about 5 cm) with a syringe needle connector at the outlet (Figure 4D).
- Position the PDMS holder in the top holder so it is free to move in all directions. Place the top holder loosely onto the heated stage like in Figure 4E without tightening any wingnuts.
- With the microscope, navigate to the position of the PDMS compartments and center them in the camera's field of view. Do not move the stage from this point on.
- Remove the top holder with the PDMS from the stage.
- Heated stage and Bephore glass slide
- Set the temperature of the heated stage to 37 °C
- Glue a Bephore glass slide (no. 4) with the gene brushes to the temperature-controlled stage of the inverted fluorescence microscope using double sided adhesive tape, with the alignment mark approximately at the center of the microscope's field of view. This positioning of the alignment mark ensures that the compartments will be lowered roughly to the same region.
- Take an image in the corresponding fluorescence channel of the gene brush and mark its position in the camera image.
- Remove the buffer from the gene brush, but do not let it go dry. Add 20 µL of the cell- free expression system to the gene brush and use tweezers to remove the silicone glue frame.
- Assembly of the microfluidic setup
- Add pressure to the sample tube until the cell-free expression system has filled the PTFE tubing and appears at the bottom of the PDMS device. Additionally, pipette 10 µL of cell-free expression system onto the micro-chambers on the PDMS. Make sure the compartments are free of air bubbles.
- Carefully lower the top holder onto the glass slide and align the micro-chambers with the gene brush. Before both PDMS and alignment mark reach the same focal plane, use the small screws at the back of the PDMS holder to precisely align the x-y-position and orientation of the PDMS device with the gene brush (Figure 4E).
- Hold the position and press the PDMS onto the glass slide by tightening the wingnuts along the screws that connect the top holder to the microscope stage (Figure 4E).
- Increase the pressure in the tube containing the cell-free expression system and monitor the flow of the fluorescent beads through the supply channel (flow rates between 0.5 to 5 µl per hour are recommended) (Figure 4F). If necessary, increase the pressure of the PDMS on the glass by tightening the wingnuts.
Functional Surface-immobilization of Genes Using Multistep Strand Displacement Lithography
Learning Objectives
Two-step lithography: Figure 5 shows the result of a two-step lithographic process on a glass slide with overlapping patterns of fluorescently labeled DIS strands.
Expression of a fluorescent protein from a gene brush: Figure 6 demonstrates the expression of the fluorescent protein YPet from immobilized DNA. At several points in time we assessed the gene expression rate by partly bleaching the fluorescent protein and observing the recovery of the fluorescence signal, disregarding the immediate recovery, which does not result from protein expression. After the first bleaching at two hours of expression, the fluorescence intensity recovered quickly and rose above its value before the bleaching. After four and six hours, the fluorescence did not recover to its previous intensity, indicating that without the supply of fresh expression mix, the reaction terminated after approximately 3-4 h.
Coupling to microfluidics: Gene expression can be sustained over longer periods of time by supplying the expression compartments with additional precursor molecules via microfluidics. Figure 7 shows such a system, enabling the expression of YPet over 10 h.
TIRF observation: Bephore can also be applied to study the interaction of fluorescently labeled proteins with a DNA brush at the single molecule level. Especially in a noisy environment, lithography helps to distinguish between specific and unspecific interaction with the brush or the surface, respectively. Figure 8 gives such an example, with fluorescently labeled T7 RNA polymerase binding or adhering preferentially to the DNA brush compared to the surrounding surface.
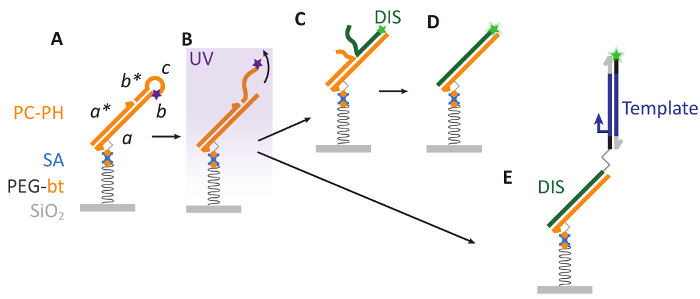
Figure 1: Bephore photolithography. A. A substrate with a surface of silicon dioxide (SiO2) is covered with a layer of biotinylated polyethylene glycol (PEG-bt), which is biocompatible and allows for the attachment of a photocleavable DNA hairpin via biotin-streptavidin interactions. Here, PC contains sequence domains abcb* and a photocleavable modification (purple star) and is hybridized to the strand PH with domain a*. B. Ultraviolet (UV) illumination cleaves PC, releasing a DNA fragment (cb*) into solution. C-D. The resulting single-stranded region on PC (b) aids as a toehold for the displacement of PH by a fluorescently labeled (green star) DIS strand. E. Also longer, double-stranded DNA ("Template") can be attached to the patterned surface. Such DNA is prepared by PCR with a primer carrying a DIS sequence at its 5' end, where primer and DIS are separated by a triethylene glycol spacer to keep DIS single-stranded during PCR (see also supplementary information, sections 2-4). Please click here to view a larger version of this figure.
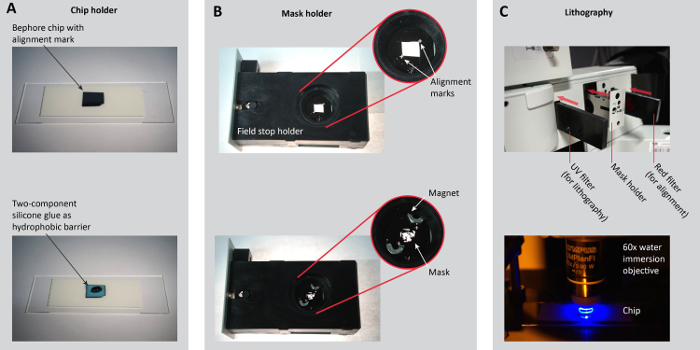

Figure 2: Sample preparation for photolithography. A. Place a Bephore chip with an alignment mark on a microscopy slide or another chip holder (e.g. the holder in Figure 3B). Apply two-component silicone glue as a hydrophobic barrier along the edges of the chip (steps 3.1.2-3). B. Place the mask onto a mask holder. Here, we removed the iris of the field stop and modified its holder so the mask can be held by small magnets. For precise (angular) alignment in multi-step lithography, ensure that alignment marks on the holder and the mask match (step 3.2.1). C. To navigate on the chip and to align the mask with the alignment marks on the chip, slide in the red filter. For the UV exposure, insert the UV filter (steps 3.2.2-5). Figure reproduced from the Supporting Information of previous work27. Please click here to view a larger version of this figure.
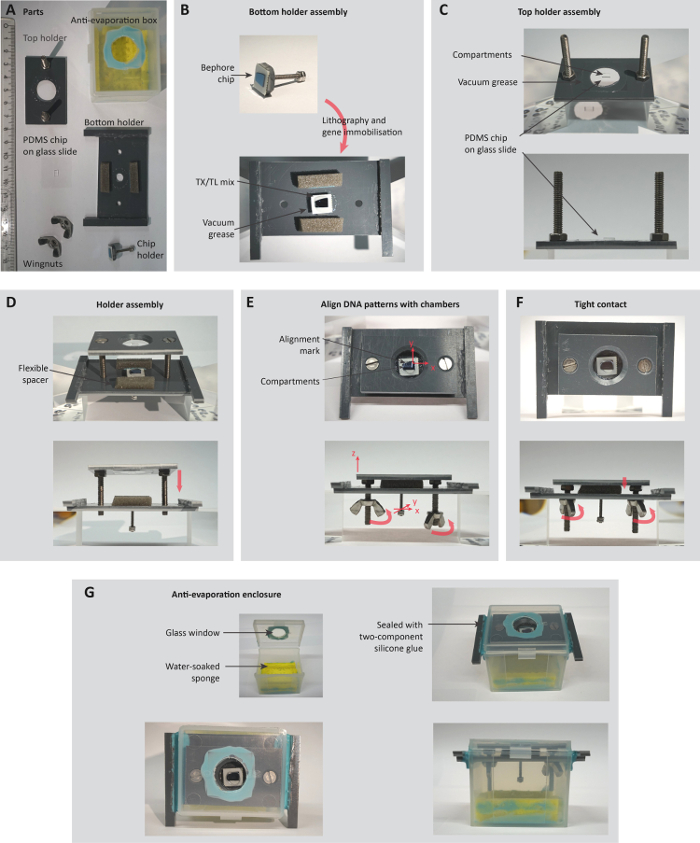

Figure 3: Assembly of a holder for the observation of compartmentalized gene expression. A. Parts of the holder (ruler unit: cm). B-C. Top and bottom part of the holder are assembled separately, with the bottom holder carrying a patterned Bephore chip (section 5.1) and the top holder carrying a PDMS chip with compartments (section 5.2). D-F. Chip and PDMS are carefully brought into tight contact, while simultaneously observing and aligning compartments and DNA brushes in a stereoscopic microscope (steps 5.3.1-6). G. Before transferring the holder to an inverted, temperature-controlled microscope, the whole system is encapsulated in an anti-evaporation box (steps 5.3.7-8). Figure reproduced from the Supporting Information of previous work27. Please click here to view a larger version of this figure.
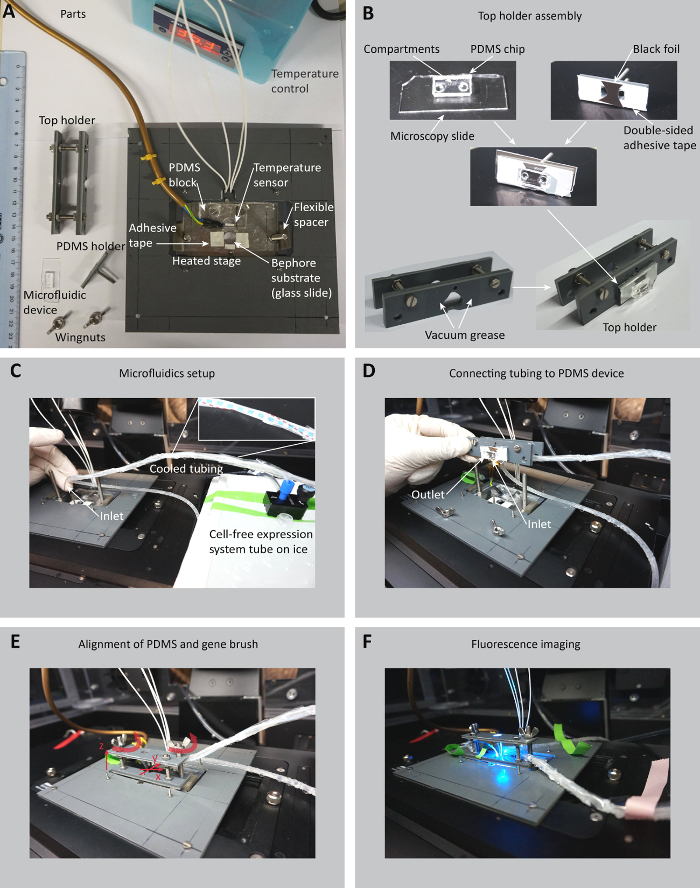

Figure 4: Microfluidic setup and sample holder for the compartmentalized gene expression. A. Parts of the sample holder, the PDMS device, the temperature-controlled microscope stage and the Bephore glass slide (ruler unit: cm). B. A microscopy slide carrying the PDMS with compartments is glued to the PDMS holder and exposed to oxygen plasma together with the PDMS in a plasma cleaner. The PDMS is then inserted into the top holder (steps 6.2.2-5). C. The cell-free expression system tube is connected to a pressure controller and placed on ice. The tubing (red dashed line in the inset) for the cell-free expression system is cooled by rubber tubing (blue dashed line) through which ice water is pumped by a peristaltic pump (section 6.1). D. The tubing is connected to the inlet position on the PDMS device. Another piece of tubing is connected to the outlet (steps 6.2.6-7). E. The top holder is placed on the microscope stage and carefully lowered towards the Bephore slide. The PDMS holder can still be moved in the x-y-plane to align the compartments in the PDMS with the gene brush on the Bephore chip. The wingnuts are used to press the PDMS onto the Bephore chip and fix the top holder to the microscope stage (steps 6.4.1-3). F. Cell-free expression system is pumped through the micro-channels in the PDMS and gene expression from DNA brushes can be monitored in epifluorescence microscopy (step 6.4.4). Please click here to view a larger version of this figure.
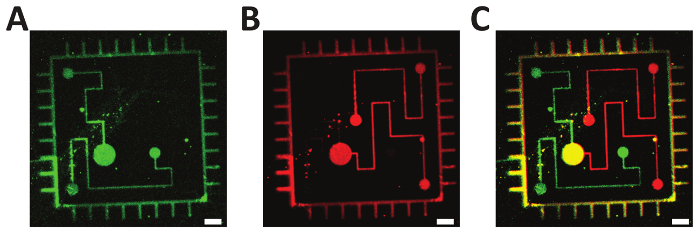
Figure 5: Two-step photolithography. A-B. Fluorescently labeled oligonucleotides (green: ATTO 532; red: Alexa Fluor 647) were consecutively immobilized on a Bephore glass slide via mask projection lithography with exposure times of 45 s. C. Overlay of subfigures A and B, demonstrating the precise alignment of the single exposures. Scale bars: 10 µm. Please click here to view a larger version of this figure.
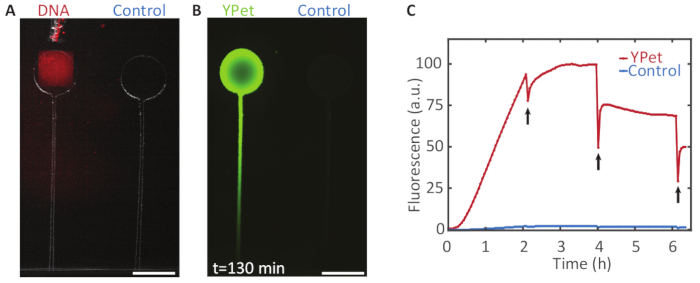
Figure 6: Compartmentalized gene expression. A. A DNA brush on a Bephore glass slide (UV exposure time: 2 min), coding for the fluorescent protein YPet was aligned with a compartment on a PDMS chip (see section 5 and Figure 3). B. Gene expression in the chamber with DNA yielded a strong fluorescence signal with a protein concentration gradient forming along a channel, which connected the chamber to the expression mix outside of the PDMS device. The control chamber without immobilized genes remained relatively dark. C. Fluorescence intensity profile over time for both chambers. Every two hours the fluorescent protein was partly bleached (black arrows) to check whether the expression was still active. After 4 h, the fluorescence did not recover to its previous intensity, indicating that expression had terminated. Scale bars: 300 µm. Please click here to view a larger version of this figure.
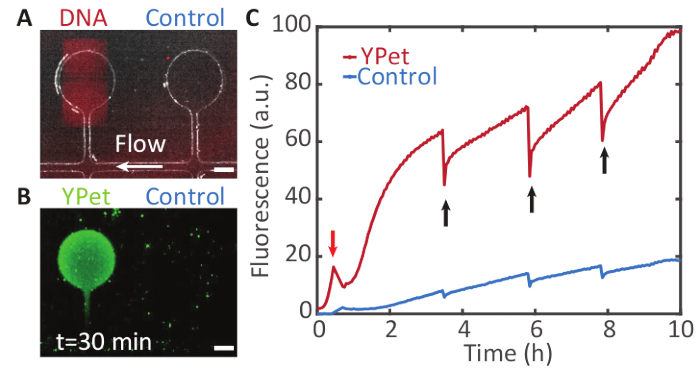
Figure 7: Sustained compartmentalized gene expression. A. A DNA brush on a Bephore glass slide (UV exposure time: 2 min), coding for YPet was aligned with a compartment on a PDMS chip. The compartments are connected to a supply channel (white arrow) through which cell-free expression system is pumped (see section 6 and Figure 4). B. The compartment containing the gene brush shows a fluorescence signal from YPet expression (in green) by the cell-free gene expression system. The neighboring compartment without a gene brush remains relatively dark. Fresh components of the cell-free expression system flow through the supply channel and diffuse into the compartments, while waste products are transported away. C. Fluorescence intensity profile over time for both chambers. The fluorescent proteins were partly bleached at different points in time (black arrows) to check whether expression was still active. Due to flow in the supply channel, gene expression was maintained for at least 10 h. (The peak in the red trace marked by the red arrow was caused by an air bubble that temporarily drained the solution from the compartments). Scale bars: 50 µm. Please click here to view a larger version of this figure.
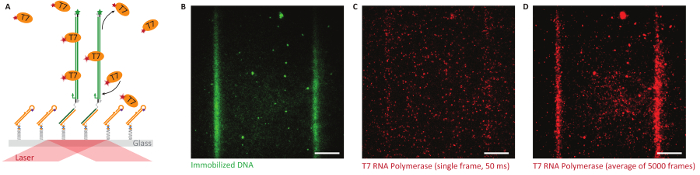
Figure 8: Single-molecule studies on Bephore glass slides in total internal reflection fluorescence microscopy (TIRFM). A. Objective-type TIRFM enables single-molecule imaging close to the glass-water interface. We immobilized fluorescently labeled genes (green, ATTO 532, UV exposure time: 2 min) with a T7 promoter along lithographically defined stripes and observed the behavior of T7 RNA polymerase (orange, labeled with Alexa Fluor 647) interacting with the surface. B. Fluorescence image showing two stripes of immobilized genes. C. T7 RNA polymerases attach specifically and non-specifically to the surface (single image, 50 ms exposure time). D. An average image obtained from all frames of a fluorescence video (5,000 frames like in subfigure C) exposes the specific interaction of the RNA polymerase with the DNA brush. Scale bars: 10 µm. Please click here to view a larger version of this figure.
Supplemental File 1. Please click here to download this file.
Supplemental File 2. Please click here to download this file.
List of Materials
| Silicon wafer with 50 nm silicon dioxide (Bephore substrate) | Siegert Wafer | Thickness (µm): 525 ±25, Diameter (mm): 100 | |
| Silicon wafer (for PDMS master mold) | Siegert Wafer | Thickness (µm): 525 ±25, Diameter (mm): 76.2 (3”) | |
| Glass slides no. 4 | Menzel | 22 mm x 50 mm | |
| Glass slides no. 1.5 | Assistent | 24 mm x 24 mm | |
| Biotin-PEG-Silane | Laysan Bio | MW 5,000 | |
| Anhydrous toluene | Sigma Aldrich (Merck) | 244511 | |
| Streptavidin | Thermo-Fisher Scientific | S888 | |
| DNA | Integrated DNA Technologies (IDT) | ||
| Phusion High-Fidelity PCR Master Mix with HF Buffer | New England Biolabs | M0531S | PCR kit |
| Wizard SV Gel and PCR Clean-Up System | Promega | A9281 | Spin-column PCR clean-up kit |
| PURExpress | New England Biolabs | E6800S | Cell-free expression system |
| PDMS | Dow Corning | Slygard 184 | |
| FluoSpheres | Thermo-Fisher Scientific | F8771 | |
| PTFE tubing (ID: 0.8mm, OD: 1.6 mm) | Bola | S 1810-10 | |
| EpoCore 20 | micro resist technology GmbH | Photoresist | |
| mr-Dev 600 | micro resist technology GmbH | Photoresist developer | |
| Ti-Prime | MicroChemicals | Adhesion promoter | |
| Two-component silicon glue | Picodent | Twinsil | |
| UV-protection yellow foil | Lithoprotect (via MicroChemicals) | Y520E212 | |
| Equipment | |||
| Masks for photolithography | Zitzmann GmbH | 64.000 dpi, 180×240 mm | |
| Upright microscope | Olympus | BX51 | Photolithography and fluorescence imaging |
| 60x water immersion objective | Olympus | LumPlanFl | Used with Olympus BX51, NA 0.9 |
| 20x water immersion objective | Olympus | LumPlanFl | Used with Olympus BX51, NA 0.5 |
| Camera | Photometrics | Coolsnap HQ | Used with Olympus BX51 |
| Ligtht source | EXFO | X-Cite 120Q | Used with Olympus BX51 |
| Inverted microscope | Nikon | Ti2-E | Fluorescence imaging of gene expression |
| 4x objective | Nikon | CFI P-Apo 4x Lambda | Used with Nikon Ti2-E |
| Camera | Andor | Neo5.5 | Used with Nikon Ti2-E |
| Light source | Lumencor | SOLA SM II | Used with Nikon Ti2-E |
| Cage incubator | Okolab | bold line | Used with Nikon Ti2-E |
| Pressure Controller | Elveflow | OB1 MK3 | |
| NanoPhotometer | Implen | DNA concentration measurement | |
| Plasma cleaner | Diener | Femto | 200 W, operated at 0.8 mbar with the sample in a Faraday cage |
Lab Prep
Immobilization of genes on lithographically structured surfaces allows the study of compartmentalized gene expression processes in an open microfluidic bioreactor system. In contrast to other approaches towards artificial cellular systems, such a setup allows for a continuous supply with gene expression reagents and simultaneous draining of waste products. This facilitates the implementation of cell-free gene expression processes over extended periods of time, which is important for the realization of dynamic gene regulatory feedback systems. Here we provide a detailed protocol for the fabrication of genetic biochips using a simple-to-use lithographic technique based on DNA strand displacement reactions, which exclusively uses commercially available components. We also provide a protocol on the integration of compartmentalized genes with a polydimethylsiloxane (PDMS)-based microfluidic system. Furthermore, we show that the system is compatible with total internal reflection fluorescence (TIRF) microscopy, which can be used for the direct observation of molecular interactions between DNA and molecules contained in the expression mix.
Immobilization of genes on lithographically structured surfaces allows the study of compartmentalized gene expression processes in an open microfluidic bioreactor system. In contrast to other approaches towards artificial cellular systems, such a setup allows for a continuous supply with gene expression reagents and simultaneous draining of waste products. This facilitates the implementation of cell-free gene expression processes over extended periods of time, which is important for the realization of dynamic gene regulatory feedback systems. Here we provide a detailed protocol for the fabrication of genetic biochips using a simple-to-use lithographic technique based on DNA strand displacement reactions, which exclusively uses commercially available components. We also provide a protocol on the integration of compartmentalized genes with a polydimethylsiloxane (PDMS)-based microfluidic system. Furthermore, we show that the system is compatible with total internal reflection fluorescence (TIRF) microscopy, which can be used for the direct observation of molecular interactions between DNA and molecules contained in the expression mix.
Procedure
Immobilization of genes on lithographically structured surfaces allows the study of compartmentalized gene expression processes in an open microfluidic bioreactor system. In contrast to other approaches towards artificial cellular systems, such a setup allows for a continuous supply with gene expression reagents and simultaneous draining of waste products. This facilitates the implementation of cell-free gene expression processes over extended periods of time, which is important for the realization of dynamic gene regulatory feedback systems. Here we provide a detailed protocol for the fabrication of genetic biochips using a simple-to-use lithographic technique based on DNA strand displacement reactions, which exclusively uses commercially available components. We also provide a protocol on the integration of compartmentalized genes with a polydimethylsiloxane (PDMS)-based microfluidic system. Furthermore, we show that the system is compatible with total internal reflection fluorescence (TIRF) microscopy, which can be used for the direct observation of molecular interactions between DNA and molecules contained in the expression mix.
