Using a Chemical Biopsy for Graft Quality Assessment
Summary
The protocol presents utilization of the chemical biopsy approach followed by comprehensive metabolomic and lipidomic analysis for quality assessment of kidney grafts allocated for transplantation.
Abstract
Kidney transplantation is a life-saving treatment for a large number of people with end-stage renal dysfunction worldwide. The procedure is associated with an increased survival rate and greater quality of patient’s life when compared to conventional dialysis. Regrettably, transplantology suffers from a lack of reliable methods for organ quality assessment. Standard diagnostic techniques are limited to macroscopic appearance inspection or invasive tissue biopsy, which do not provide comprehensive information about the graft. The proposed protocol aims to introduce solid phase microextraction (SPME) as an ideal analytical method for comprehensive metabolomics and lipidomic analysis of all low molecular compounds present in kidneys allocated for transplantation. The small size of the SPME probe enables performance of a chemical biopsy, which enables extraction of metabolites directly from the organ without any tissue collection. The minimum invasiveness of the method permits execution of multiple analyses over time: directly after organ harvesting, during its preservation, and immediately after revascularization at the recipient’s body. It is hypothesized that the combination of this novel sampling method with a high-resolution mass spectrometer will allow for discrimination of a set of characteristic compounds that could serve as biological markers of graft quality and indicators of possible development of organ dysfunction.
Introduction
According to the United States’ Organ Procurement and Transplantation Network, there were 94,756 patients waiting for kidney transplants in the US in 2019; while in Europe in 2018, that number was 10,791. Every ten minutes, someone is added to the national transplant waiting list in the U.S., and it is estimated that 20 people die each day waiting for a transplant1,2. Kidney transplantation is a life-saving treatment for a large number of people suffering with end-stage renal dysfunction worldwide. The procedure is associated with increased survival rate and greater quality of life when compared to conventional dialysis.
However, transplantation faces many serious problems, such as organ shortage or lack of effective tools for organ quality assessment. The standard protocols are limited to macroscopic appearance inspection or invasive tissue biopsy, which do not provide comprehensive information regarding the quality of the graft. While a visual appraisal allows for identification of tumors visible to the eye, anatomical abnormalities, or extensive damage to the grafts, this approach is very subjective, varying in its effectiveness according to the experience of the observers. Biopsy, on the other hand, can provide valuable information regarding pre-existing renal disorders, and is thus considered to be a method of objective and evidenced value in determining graft outcomes. However, the biopsy procedure is not free of flaws; there is the risk of potential complications such as bleeding and an additional 4-5 hours of sample preparation is required, which significantly prolongs the cold ischemic time. Therefore, especially in Europe, the use of direct tissue analysis is limited to expanded criteria donors (ECD) and donors after circulatory death (DCD)3,4.
Metabolomics and lipidomics have been recently recognized as promising approaches to attaining a better understanding of the changes in biochemical pathways occurring during organ preservation. Metabolomic and lipidomic profiling enables monitoring of immediate responses of the system to sudden environmental changes related to organ removal with subsequent consequences: ischemia, oxidative stress, or inflammatory responses5,6,7,8. The kidney is an organ that is largely associated with metabolic processes, thus measurements of metabolites and lipids concentrations may permit identification of potential organ quality biomarkers and enable better predictions of graft outcome.
Given the above complications and limitations associated with current organ quality assessment methods, a less invasive diagnostic solution is needed for quick and complex organ quality assessment. Solid phase microextraction (SPME) complies with these requirements as a minimally invasive analytical method that enables coverage of a broad spectrum of metabolites and lipids. The technique is based on the insertion of a thin (~200 µm), biocompatible, titanium-nickel alloy probe covered with a selective extraction phase into the examined organ for a short time. It should be emphasized that SPME prevents protein extraction, and hence enables metabolism inhibition already at the stage of sample collection, which is a significant advantage over alternative methods. Moreover, the miniaturization of the device allows for the execution of repetitive and simultaneous analyses of few structures of the organ9,10,11.
Protocol
All animals received humane care in compliance with the ‘‘Principles of Laboratory Animal Care’’ formulated by the National Society for Medical Research and the ‘‘Guide for the Care of Laboratory Animals’’ published by the National Institute of Health, Ontario, Canada. The Animal Care Committee of the Toronto General Research Institute approved all studies. The research involved human subjects was approved by the Bioethical Committee at Collegium Medicum in Bydgoszcz Nicolaus Copernicus University in Torun.
NOTE: Obtain approval from appropriate ethical boards. Remember to always wear safety gloves. Do not touch the extraction phase of SPME probes. The use of deactivated glass vials is recommended for lipidomic analyses.
1. Preparation of probes
- Prepare titanium-nickel alloy probes (40 mm length; 0.2 mm diameter) coated with 7 mm mix-mode sorbent. The number of probes depends on the time points targeted during the entire procedure and the number of replicates (3 is recommended per time point).
NOTE: The length and type of extraction phase may be adjusted based on the mode of study, the polarity of metabolites, and the sample matrix. - Prepare a cleaning mixture composed of 2:1 chloroform:methanol (v/v). Pipet 1.0 mL of the solution to each 2.0 mL glass vial and place one probe, previously pierced through the cap, in each vial.
NOTE: Before use, clean all probes to remove contaminating particles. - Put the vials on the agitator and set the agitation speed at 1,200 rpm. After 45 min, stop the device and rinse the coatings with LC-MS grade water.
- As coatings must undergo a preconditioning step to activate them, prepare a preconditioning mixture composed of 1:1 methanol:water (v/v). Pipet 1.0 mL of the solution to each 2.0 mL glass vial and place one probe, previously pierced through the cap, in each vial.
- Put the vials on the vortex agitator and set the agitation speed at 1,200 rpm.
- After 60 min, stop the agitator and rinse the coatings with LC-MS grade water.
- Sterilize probes according to the standard surgical equipment sterilization protocol.
2. Extraction
- Open the sterile package right before sampling to ensure a sterile environment.
- Insert two probes directly into the kidney cortex for 10 min at each time point. The entire length of the coating must be covered by the tissue matrix; no specific angle is required, but ca. 90 deg is usually used.
NOTE: The entire medical procedure follows standard protocols in given institution. No modification concerning SPME sampling is considered. The procedure involves the six following sampling time points:
a) before kidney resection, in vivo from donor;
b)-e) after 1 h, 3 h, 5 h, 7 h of kidney perfusion, ex vivo in organ chamber;
f) after reperfusion, in vivo from recipient. - Retract the probes by pulling it out from the tissue and then rinse coatings with LC-MS grade water to clear any remaining blood from the coating surface. Rinse away from the surgical site, and immediately after removal of the probes.
3. Transport and storage
- Place probes in separate vials and close them.
- Place vials in a Styrofoam box filled with dry ice or in liquid nitrogen for transport.
- Store samples in a freezer (-80 °C), or immediately commence the desorption step.
4. Desorption
- Prepare desorption solutions composed of 80:20 acetonitrile:water (v/v) for metabolomic analysis, and 1:1 isopropanol:methanol (v/v) for lipidomic analysis.
- Pipet 100 μL of the solution to inserts placed into 2.0 mL labeled vials and place one probe, previously pierced through the cap, in each vial.
- Put the vials on the vortex agitator and set the agitation speed at 1,200 rpm for 120 min.
- Remove probes from the vials. Obtained extracts are now ready for instrumental analysis.
5. LC-MS analysis
- Place vials containing extracts in the autosampler of the LC-MS system.
NOTE: Liquid chromatography (RP, HILIC) coupled with high resolution mass spectrometry and an orbitrap mass analyzer were used for this study. For metabolomic analysis parameters, go to step 5.2. For lipidomic analysis parameters, go to step 5.6. - Use a pentafluorophenyl (PFP) column (2.1 mm x 100 mm, 3 μm) for reversed phase separation.
- Set the flow rate to 300 µL/min, and autosampler and column temperatures to 4 °C and 25 °C, respectively.
- Prepare mobile phases according to the following proportions: mobile phase A: water:formic acid (99.9:0.1, v/v), and mobile phase B: acetonitrile:formic acid (99.9:0.1, v/v). Set the mobile phase flow according to the following parameters: starting mobile phase flow (0-3 min): 100% A followed by a linear gradient to 10% A (3-25 min), ending with isocratic flow of 10% A until 34 min, followed by 6 min of column re-equilibrium time.
- For HILIC separation, use a HILIC column (2.0 mm x 100 mm, 3 μm, 200A). Set the flow rate to 400 μL/min.
- Prepare mobile phases according to the following proportions: mobile phase A: acetonitrile:ammonium acetate buffer (9:1, v/v, effective salt concentration 20 mM), mobile phase B: acetonitrile:ammonium acetate buffer (1:1, v/v, effective salt concentration 20 mM).
- Set the mobile phase flow according to the following parameters: starting mobile phase flow (0-3 min) at 100% A, hold for 3.0 min and then ramp to 100% B within 5 min. Hold with 100% B until 12 min, followed by 8 min of column re-equilibrium time.
- Set the scan range to 85-1000 m/z. Set HESI ion source parameters in positive ionization mode to: spray voltage 1,500 V, capillary temperature 300 °C, sheath gas 40 a.u., aux gas flow rate 15 a.u., probe heater temperature 300 °C, S-Lens RF level 55%.
- Run QC samples composed of 10 µL of each analyzed sample (regularly, every 10-12 samples) to monitor instrument performance.
- For reversed phase separation, use a C18 column (2.1 mm x 75 mm, 3.5 μm).
- Set the flow rate to 200 µL/min, and autosampler and column temperatures to 4 °C and 55 °C, respectively. Prepare mobile phases according to the following proportions: mobile phase A: H2O:MeOH (60:40, v/v), 10 mM ammonium acetate and 1 mM acetic acid; mobile phase B: IPA:MeOH (90:10, v/v), 10 mM ammonium acetate and 1 mM acetic acid.
- Set the mobile phase flow according to the following parameters: 0-1 min (20% B), 1-1.5 min (20-50% B), 1.5-7.5 min (50-70% B), 7.5-13 min (70-95% B), 13-17 min (95% B), 17-17.1 min (95-20% B), 17.1-23 min (20%).
- For HILIC separation, use a HILIC column (100 x 2.1 mm, 3 μm).
- Set the flow rate to 400 µL/min, and autosampler and column temperatures to 4 °C and 40 °C, respectively.
- Prepare mobile phases according to the following proportions: mobile phase A: ACN; mobile phase B: 5 mM ammonium acetate in water. Set the mobile phase flow according to the following parameters: 0-2 min (96% B), 2-15 min (96-80% B), 15-15.1 min (80-96% B), 15.1-21 min (96% B).
- Set HESI ion source parameters in positive ionization mode to spray voltage 3,500 V, capillary temperature 275 °C, sheath gas 20 a.u., aux gas flow rate 10 a.u., probe heater temperature 300 °C, S-Lens RF level 55%.
- Run QC samples composed of 10 µL of each analyzed sample (every 10-12 samples) to monitor instrument performance.
- Perform data acquisition in software compatible with the instrument.
6. Data analysis
- Perform data processing, putative identification, and statistical analysis with the use of software dedicated to untargeted metabolomics and lipidomics analysis.
NOTE: Principal components analysis (PCA) and box-whisker plots can be obtained to visualize data structure.
Representative Results
Sample collection was performed according to the SPME method described above, using probes coated with a 7 mm mixed-mode extraction phase. Sampling was conducted directly from the graft tissue (in vivo before transplantation), ex vivo in kidney chamber after 1 h, 3 h, 5 h, and 7 h of perfusion, and in vivo after revascularization in the recipient. Untargeted metabolomic and lipidomic analyses were carried out with the use of liquid chromatography (RP, HILIC) coupled with high resolution mass spectrometry, with the mass analyzer set in positive ionization mode. To assess data quality and attain general insights regarding the results, data was subjected to principal component analysis (PCA). As shown in Figure 1, QC samples formed a tight cluster, confirming the quality of the analyses. The studied groups exhibit relatively good separation, allowing for visualization of differences in metabolic and lipidomic profiles before and after transplantation, as well as during organ perfusion (Figure 2). SPME sampling permits metabolic profiling of the organ over time. Box-whisker plots of selected metabolites serve to exemplify alterations in metabolite levels throughout the experiment (Figure 3). The wide spectrum of extracted features separated on both RP and HILIC columns is shown in Figure 4.
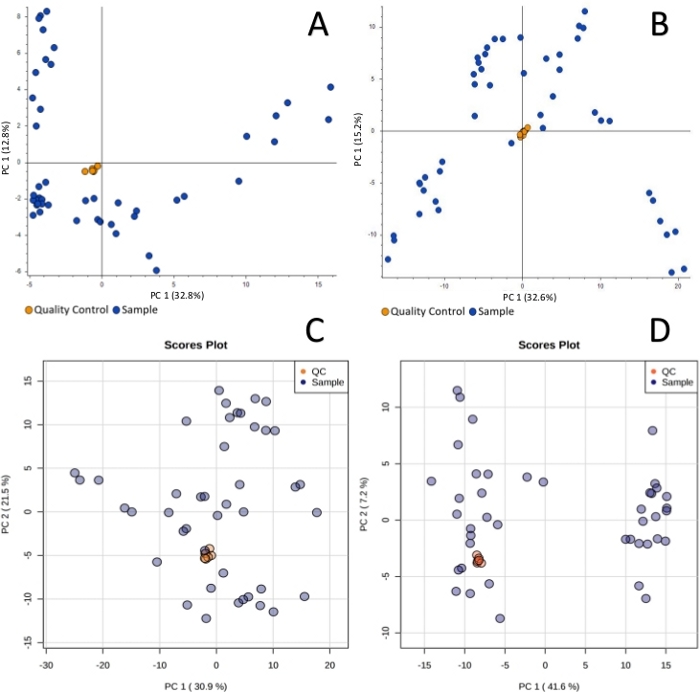
Figure 1: Principal component analysis showing clustering of quality control samples for metabolomic reversed-phase (A), HILIC (B) and lipidomic reversed-phase (C), HILIC (D) analyses. The quality control is necessary in untargeted analysis to monitor any possible changes induced by instrumental instability. Tight cluster of QC samples ensures that all changes observed are of biological origin. Please click here to view a larger version of this figure.
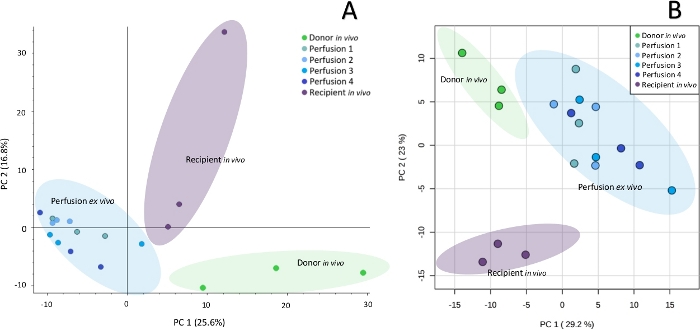
Figure 2: Principal component analysis showing metabolomic (A) and lipidomic (B) differences between particular sampling points. Untargeted profiling of kidney with SPME-LC-HRMS enables to observe biochemical differences in organs at subsequent steps of their preservation. Please click here to view a larger version of this figure.
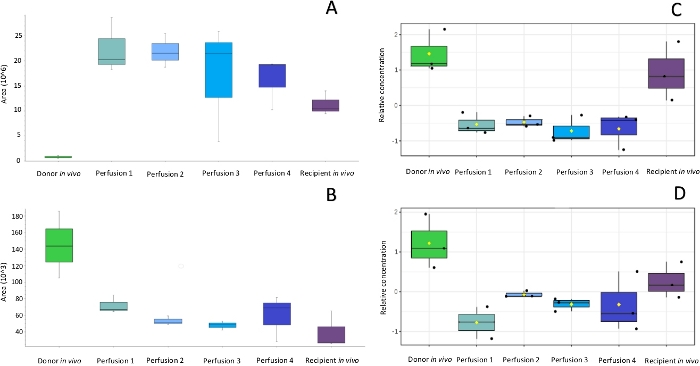
Figure 3: Box-whisker plots of selected compounds showing alterations in metabolites (A, B) and lipids (C, D) during the peritransplant period. After selection and identification of discriminating compounds box-whisker plots enables to monitor changes in levels of these compounds at subsequent steps of the protocol and compare the metabolites levels in organs subjected to different preservation protocols or harvested from different donors e.g. heart beating donors or donors after cardiac death. Please click here to view a larger version of this figure.
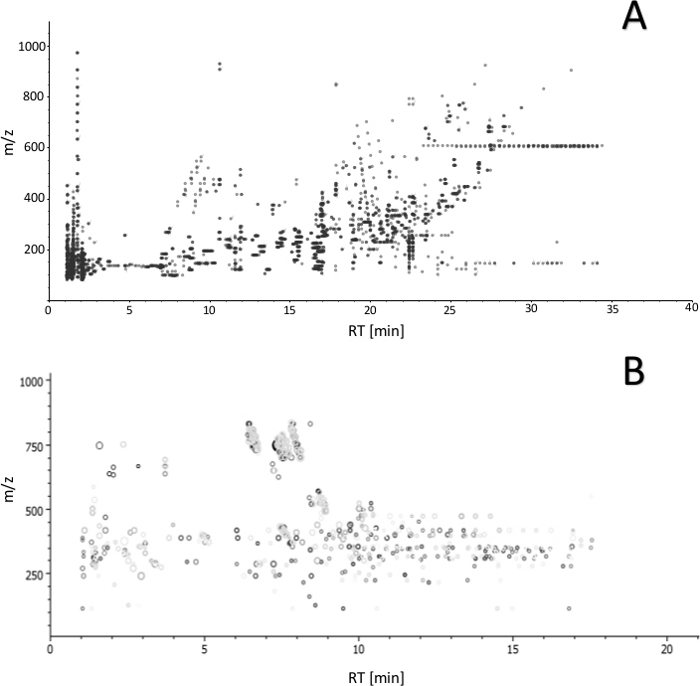
Figure 4: Ion map (m/z versus retention time) of features obtained by LC-MS analysis with (A) reversed-phase and (B) HILIC separation. The plot enables to estimate overall number of features obtained with the proposed protocol (from extraction to detection) and assess analyte coverage based on the polarity. Please click here to view a larger version of this figure.
Discussion
Evaluation of organ quality remains a large challenge for physicians, who must make quick informed decisions regarding whether a given organ is viable for transplantation or whether it must be discarded. Multiple factors, such as donor age, duration of ischemia, and infections and inflammatory processes, can affect the long-term graft outcome. While diverse methods have been developed to date to diagnose kidney allograft function, histopathological inspection remains the gold standard in that matter3,4,12. Although the biopsy procedure can yield significant information regarding pre-existing donor disease and vascular changes, it is not free of flaws. Sampling errors associated to interobserver variability and sampling of insufficient glomeruli for comprehensive information regarding organ function remain typical concerns in this regard. Moreover, specimen preparation brings some issues such as an incomplete appraisal of the graft in case of frozen sections, and extension of procedure time for paraffin sectioning. However, the increased risk of hemorrhage, which may appear acutely as microscopic or gross hematuria, is the major life-threatening complication associated with the biopsy procedure. For this reason, the number of allowable biopsies is strictly limited in transplantation procedures, a factor that hampers the capture of dynamic changes and time series analyses via this method12,13,14. The benefits of a histological analysis must be weighed against the risks associated with the methodology. The value of histological findings is indisputable, but they do not explain the molecular mechanisms of the aberrations.
Metabolomics and lipidomics are the youngest domains of the “-omics” scientific family. The complete set of low molecular (<1,200 Da) human metabolites and lipids connected within a metabolic network is defined as a human metabolome. The genome remains relatively constant throughout its lifetime, with slight modifications caused by mutations occurring infrequently. The metabolome is the product of gene expression, which is highly sensitive to changes in all biological processes as well as environmental factors. The dynamic nature of metabolites and lipids makes them perfect indicators of current organ condition7,8,15,16. The SPME method proposed in the abovementioned protocol enables detection of changes occurring in the organ during its preservation, starting from organ removal from the donor’s body till revascularization at the recipient’s. The small diameter of the probe (~200 µm) provides minimal invasiveness and allows for several samplings from the same organ without causing any damage to the tissue. Conducting studies using kidney, as the most frequently transplanted organ, allows for a better understanding and further characterization of the metabolic pathways responsible for declining the quality and function of grafts. The possibility of monitoring modifications over time certainly is an important advantage of the technique as compared to conventional invasive methods such as biopsy. The currently presented analysis identified altered concentrations of various groups of lipids and metabolites, especially of essential amino acids, purines, purine nucleosides, and glycerophospholipids. These results are consistent with previous tissue analysis reports5,6,17,18,19,20. To date, the majority of scientific reports utilizing metabolomics or lipidomics to explain processes inducing complications after transplantation or ischemia/reperfusion injury (IRI) phenomena have been limited to analysis of biofluids21,22,23.
Each clinical application requires optimization of the sampling protocol to ensure that the performance of the analytical method meets expected criteria. In this regard, the benefit of utilizing SPME is the possibility of adjusting conditions for various experimental designs. The variety of accessible extraction phases provides a broad spectrum of extracted metabolites with diversified polarities. At the same time, this might be considered as a limitation of the method due to the fact that each sorbent provides selectivity towards specific features and does not extract all compounds present in the sample matrix. It should be noted that SPME coatings extract only via free molecules, and simply do not interact with a bound fraction of the analyte. The biocompatibility of the coatings does not introduce toxicity to the tissue while restraining the extraction of large molecules such as proteins; as a consequence, the enzymatic processes are inhibited already at the stage of sample collection and the presence of artefacts is minimized, which is a great advantage over alternative sampling methods. The length of the coating influences the efficiency of extraction (i.e., the length of the coating designates the surface area and the extraction phase volume); thus, longer coatings yield higher recoveries. On the other hand, shorter coatings enable higher spatial resolution. For reliable results, it is crucial to submerge the probe to the exact same depth of kidney cortex. Insertion too deep causes the risk of entering the kidney medulla. The time of the extraction is also proportional to the extraction efficiency. Therefore, selection of optimum extraction time is one of the most critical steps in SPME method development. The accuracy of time measurement provides the highest repeatability. In biological applications such as the one discussed, there is always a compromise between the sensitivity and repeatability of the analytical protocol and the restrictions of the medical procedure. While equilibrium extraction provides the highest sensitivity, for safety reasons, pre-equilibrium conditions are often used in such applications, as extraction time should not affect the total duration of the surgery. The efficiency of desorption is determined by the time of the process and the composition of the desorption solvent, which should be compatible with the mobile phase used for chromatographic separation9,10,11.
One of the major requirements for diagnostic instrumentation used for intra-surgical assessments is time of analysis. Current attempts are being made to develop a rapid tool for in vivo SPME extraction coupled directly to a mass spectrometer via microfluidic open interface (MOI)24 or coated blade spray (CBS)25. Such approaches would allow for the disclosure of analytical results in real or close to real time. The use of such methods for pre-intervention analyses of metabolic and lipidomic profiles could enhance the decision-making process during transplantation procedures, enabling the best possible personalized approach and fast response in case of organ failure.
As a summary, it is hypothesized that the proposed protocol will enable attainment of full metabolic and lipidomic profiles of kidney grafts, which in turn would provide a comprehensive assessment of organ quality and characterization of the processes responsible for ischemia-reperfusion injury. The novelty of the project includes the utilization of solid-phase microextraction (SPME), offering low invasive sampling of living systems, in combination with one of the most innovative technologies available for metabolomics and lipidomics analysis (e.g., the Orbitrap high resolution mass spectrometer). SPME combines sample collection, extraction, and quenching of metabolites in one step, therefore making it a perfect tool for rapid analysis. It is expected that this protocol will help to answer questions related to what pre-transplant conditions of the kidney are responsible for delayed organ function or its dysfunctions after transplantation, as well as to how the graft preservation protocol influences the biochemistry of the organ. Such knowledge would not only have significant impact on the prevention of possible complications related to transplantation, but may help to improve current graft preservation protocols, minimizing loss of viable transplant tissue as well as loss of life. The proposed solution will open the door to further investigations in this field, including the validation of specific potential biomarkers and improvement of therapeutic outcomes in transplantology.
Disclosures
The authors have nothing to disclose.
Acknowledgements
The study was supported by grant Opus UMO-2017/27/B/NZ5/01013 from National Science Centre. The authors would like to acknowledge MilliporeSigma, a business of Merck KGaA, Darmstadt, Germany for providing SPME devices. The life science business of Merck operates as MilliporeSigma in the U.S. and Canada. Also, the authors want to thank Thermo Fisher Scientific for the access to Q-Exactive Focus orbitrap mass spectrometer. The authors would like to thank Dr. Aleksandra Woderska-Jasińska and the personnel of the Department of Transplantology and General Surgery in Bydgoszcz for their kind assistance in the project. BB want to thank Prof. Janusz Pawliszyn for the opportunity of sample collection at Toronto General Hospital during her stay at University of Waterloo.
Materials
| Acetic acid | Merck | 5330010050 | Mobile phase additive |
| Acetonitrile | Alchem | 696-34967-4X2.5L | HPLC solvent |
| Ammonium acetate | Merck | 5330040050 | Mobile phase additive |
| BENCHMIXER XL MULTI-TUBE VORTEXER | Benchmark Scientific | BV1010 | Vortex mixer |
| Caps | Perlan Technologies | 5183-2076 | Blue scrw tp, pre-slit PTFE/Si spta, 100PK |
| Chloroform | Merck | 1024441000 | |
| Discovery HS F5 Supelguard Cartridge, 3 μm, L × I.D. 2 cm × 2.1 mm | Merck | 567570-U | HPLC guard column |
| Discovery HS F5, 2.1 mm x 100 mm, 3 μm | Merck | 567502-U | HPLC column |
| Formic acid | Alchem | 497-94318-50ML | Mobile phase additive |
| Glass vials | Perlan Technologies | 5182-0714 | |
| HILIC Luna 3 μm, 200A, 100 x 2.0 mm | Shim-Pol | PHX-00D-4449-B0 | HPLC column |
| HILIC SecurityGuard Cartridge, 3 μm, 4 x 2.0 mm | Shim-Pol | PHX-AJ0-8328 | HPLC guard column |
| Isopropanol | Alchem | 231-AL03262500 | HPLC solvent |
| Methanol | Alchem | 696-34966-4X2.5L | HPLC solvent |
| Nano-pure water | Merck | 1037281002 | HPLC solvent |
| Q Exactive Focus hybrid quadrupole-Orbitrap MS | Thermo Scientific | Q Exactive Focus | Mass Spectrometer |
| SeQuant ZIC-cHILIC 3µm,100Å 100 x 2.1 mm | Merck | 1506570001 | HPLC column |
| SeQuant ZIC-HILIC Guard Kit 20 x 2.1 mm | Merck | 1504360001 | HPLC guard column |
| SPME LC fiber probes, mixed mode | Supelco | prototype fibers | |
| UltiMate 3000 HPLC systems | Thermo Scientific | UltiMate 3000 | HPLC system |
| Vial inserts (deactivated) | Perlan Technologies | 5181-8872 | |
| XSelect CSH C18 3.5μm 2.1x75mm | Waters | 186005644 | HPLC column |
| XSelect CSH C18 VanGuard Cartridge 3.5μm, 2.1x5mm | Waters | 186007811 | HPLC guard column |
References
- . Organ Procurement and Transplantation Network Available from: https://optn.transplant.hrsa.gov (2019)
- Branger, P., Undine, S. . Annual Report 2018/Eurotransplant International Foundation. , (2018).
- Dare, A. J., Pettigrew, G. J., Saeb-Parsy, K. Preoperative assessment of the deceased-donor kidney: From macroscopic appearance to molecular biomarkers. Transplantation. 97, 797-807 (2014).
- Mueller, T. F., Solez, K., Mas, V. Assessment of kidney organ quality and prediction of outcome at time of transplantation. Seminars in Immunopathology. 33, 185-199 (2011).
- Xu, J., et al. Lipidomics comparing DCD and DBD liver allografts uncovers lysophospholipids elevated in recipients undergoing early allograft dysfunction. Scientific Reports. 5, 1-10 (2015).
- Rao, S., et al. Early lipid changes in acute kidney injury using SWATH lipidomics coupled with MALDI tissue imaging. American Journal of Physiology – Renal Physiology. 310, 1136-1147 (2016).
- Wishart, D. S. Metabolomics: The principles and potential applications to transplantation. American Journal of Transplantation. 5, 2814-2820 (2005).
- Kim, S. J., Kim, S. H., Kim, J. H., Hwang, S., Yoo, H. J. Understanding metabolomics in biomedical research. Endocrinology and Metabolism. 31, 7-16 (2016).
- Bojko, B., et al. Solid phase microextraction fills the gap in tissue sampling protocols. Analytica Chimica Acta. 803, 75-81 (2013).
- Filipiak, W., Bojko, B. SPME in clinical, pharmaceutical, and biotechnological research – How far are we from daily practice. TrAC – Trends in Analytical Chemistry. , 115-213 (2019).
- Bojko, B., et al. Low invasive in vivo tissue sampling for monitoring biomarkers and drugs during surgery. Laboratory Investigation. 94, 586-594 (2014).
- Ahmad, I. Biopsy of the transplanted kidney. Seminars in Interventional Radiology. 21, 275-281 (2004).
- Plattner, B. W., et al. Complications and adequacy of transplant kidney biopsies: A comparison of techniques. Journal of Vascular Access. 19, 291-296 (2018).
- Tapia-Canelas, C., et al. Complications associated with renal graft biopsy in transplant patients. Nefrologia. 34, 115-119 (2014).
- Afshinnia, F., et al. Lipidomics and Biomarker Discovery in Kidney Disease. Seminars in Nephrology. 38, 127-141 (2018).
- Zhao, Y. Y., Vaziri, N. D., Lin, R. C. Lipidomics: New insight into kidney disease. Advances in Clinical Chemistry. 68, 153-175 (2015).
- Kaminski, J., et al. Oxygen Consumption by Warm Ischemia-Injured Porcine Kidneys in Hypothermic Static and Machine Preservation. Journal of Surgical Research. 242, 78-86 (2019).
- Wijermars, L. G. M., et al. Defective postreperfusion metabolic recovery directly associates with incident delayed graft function. Kidney International. 90, 181-191 (2016).
- Huang, H., et al. Proteo-metabolomics reveals compensation between ischemic and non-injured contralateral kidneys after reperfusion. Scientific Reports. 8, 1-12 (2018).
- Solati, Z., Edel, A. L., Shang, Y., Karmin, O., Ravandi, A. Oxidized phosphatidylcholines are produced in renal ischemia reperfusion injury. PLoS One. 13, 1-24 (2018).
- Wishart, D. S. Metabolomics in monitoring kidney transplants. Current Opinion in Nephrology and Hypertension. 15, 637-642 (2006).
- Abbiss, H., Maker, G. L., Trengove, R. D. Metabolomics approaches for the diagnosis and understanding of kidney diseases. Metabolites. 9, (2019).
- Zhang, Z. H., et al. Metabolomics insights into chronic kidney disease and modulatory effect of rhubarb against tubulointerstitial fibrosis. Scientific Reports. 5, 1-17 (2015).
- Looby, N. T., et al. Solid phase microextraction coupled to mass spectrometry: Via a microfluidic open interface for rapid therapeutic drug monitoring. Analyst. 144, 3721-3728 (2019).
- Gómez-Ríos, G. A., Tascon, M., Pawliszyn, J. Coated blade spray: Shifting the paradigm of direct sample introduction to MS. Bioanalysis. 10, 257-271 (2018).

