Optimizing Tear Collection in Mice for mRNA and Protein Analysis
Summary
The identification of RNA and protein biomarkers from tears in mouse models holds great promise for early diagnostics in various diseases. This manuscript provides a comprehensive protocol for optimizing the efficacy and efficiency of mRNA and protein isolation from mouse tears.
Abstract
The tear film is a highly dynamic biofluid capable of reflecting pathology-associated molecular changes, not only in the ocular surface but also in other tissues and organs. Molecular analysis of this biofluid offers a non-invasive way to diagnose or monitor diseases, assess medical treatment efficacy, and identify possible biomarkers. Due to the limited sample volume, collecting tear samples requires specific skills and appropriate tools to ensure high quality and maximum efficiency. Various tear sampling methodologies have been described in human studies. In this article, a comprehensive description of an optimized protocol is presented, specifically tailored for extracting tear-related protein information from experimental animal models, especially mice. This method includes the pharmacological stimulation of tear production in 2-month-old mice, followed by sample collection using Schirmer strips and the evaluation of the efficacy and efficiency of the protocol through standard procedures, SDS-PAGE, qPCR, and digital PCR (dPCR). This protocol can be easily adapted for the investigation of the tear protein signature in a variety of experimental paradigms. By establishing an affordable, standardized, and optimized tear sampling protocol for animal models, the aim was to bridge the gap between human and animal research, facilitating translational studies and accelerating advancements in the field of ocular and systemic disease research.
Introduction
Tears are considered a plasma ultrafiltrate and have also been described as an intermediate fluid between plasma serum and cerebrospinal fluid due to a significant overlap in the biomolecules they share1. It has been reported that human tears contain proteins, tear lipids, metabolites, and electrolytes2. Recently, other biomolecules such as mRNAs, miRNAs, and extracellular vesicles have also been identified3,4,5,6,7.
In humans, basal tears are located in the tear film, which consists of three layers: the outer lipid layer, which maintains the tear surface smooth to allow us to see through it and prevent tear evaporation; the middle aqueous layer, which keeps the eye hydrated, repels bacteria, protects the cornea, and constitutes 90% of the tear film; and finally, the mucin layer, a family of high molecular weight proteins that are in contact with the cornea and allow the tear to adhere to the eye8. Tear distribution across the ocular surface begins with secretion from the lacrimal gland. This fluid is then guided through tear ducts to pass over the eye's surface and flow into drainage channels. Each blink allows tears to disperse evenly over the entire eye, keeping it moist9.
The tear film is a highly dynamic biofluid capable of reflecting molecular changes that occur not only on the ocular surface but also in other tissues and organs. The analysis of differential expression in this biofluid represents a promising approach for the discovery of biomarkers in human diseases10,11. The utilization of tear film as a source of biomarkers for early diagnosis in various pathologies has been greatly facilitated by the presence of non-invasive collection methods. The most common method for collecting tears in human and veterinary clinics involves membrane-based support (Schirmer's strip), which operates on the principle of capillary action, allowing the water in tears to travel along the length of a paper test strip or capillary tubes placed in the subject's lower conjunctival sac12,13,14. Despite the inherent limitation of obtaining a small sample volume through this method, the biochemical analysis of tear composition using various sensitive techniques has facilitated the identification of potential biomarker molecules11,15. Protocols for optimizing and evaluating the elution of tear proteins from Schirmer strips and capillaries in human patients are well documented16,17. However, complete descriptions of optimized protocols specifically designed to extract molecular information related to tears from experimental animal models are available but scarce. Existing methods, such as tear induction through direct stimulation of the lacrimal gland18, while allowing for the collection of larger volumes, are invasive and can cause discomfort to the animals. Non-invasive methods, such as collecting tears from the ocular surface, have been described as a way to isolate DNA and miRNAs19,21.
This protocol aims to establish a cost-effective and optimized method for collecting and processing tears in mice. The method prioritizes non-invasiveness while obtaining sufficient tear volumes suitable for molecular analysis through techniques like SDS-PAGE, qPCR, and dPCR. The extracted protein and mRNA content information can then be utilized to identify potential biomarkers in existing experimental models of disease.
Protocol
All procedures described here were approved by the Animal Ethics Committee of Cinvestav (CICUAL, # 0354/23). The laboratory animals were treated and handled in strict accordance with the journal's animal use guidelines and according to the Association for Research in Vision and Ophthalmology (ARVO) Statement for the Use of Animals in Ophthalmic and Vision Research. Ocular health before and after the procedures was evaluated by assessing ocular discharges, swollen eyelids, ocular abnormalities, and behavior changes.
1. Animals and reagent preparation
- Use three 2-month-old C57BL/6 male mice per replicate for the procedure, with a starting weight of ~25 g (Figure 1). Process each mouse individually and place it on a heating pad to maintain warmth.
- To induce tear secretion, prepare a stock solution of pilocarpine at 20 mg/mL using sterile water.
- Cut the Schirmer's test strips to 5 mm in length and bend each strip at the notch to a 90° angle.
- Prepare 250 µL of sterile water with a protease inhibitor (1 tablet for 50 mL, according to manufacturer instructions).
2. Tear production stimulation and collection
- Weigh each mouse individually to calculate the appropriate dosage of treatments.
- To stimulate tear production, administer 30 mg/kg of pilocarpine intraperitoneally with a 6 mm insulin syringe (Figure 1B). Wait for 2 min post-drug injection for its effects to manifest.
NOTE: Data from unstimulated animals could not be obtained due to the limited availability of tear fluid. - Collect the tears by placing the end of each Schirmer strip fragment over the center of the lower eyelid with a pair of tweezers and keep it in place until the tear reaches the strip limit (Figure 1C). Then, replace the strip. Sequentially place 2 Schirmer strips in each eye, with a 2 min interval between strips for collection. The tear collection process requires around four strip fragments over 10 min per mouse.
- Place the Schirmer strip fragments in sterile tubes on ice to avoid degradation of components. As soon as the collection finishes, start tear processing.
3. Protein isolation
- Place the strip fragments in a 600 µL microcentrifuge tube containing 20 µL of sterile water with a protease inhibitor and centrifuge for 1 h at 600 x g at 4 °C, as previously shown17 (Figure 1D).
- Make a puncture using a 7.5 cm stainless-steel injector needle at the tip of the 600 µL tube containing the sample (Figure 1E). Then, transfer this punctured tube to a new sterilized 1.5 mL tube and centrifuge the samples at 12,000 x g for 10 min at 4 °C, as previously reported17, to recover the volume of diluted tears. The tear film volume (supernatant) will be at the bottom of the tube.
- Create a protein pool by combining the supernatant recovered from all the samples. In this experiment, a total volume of approximately 200 µL was obtained.
NOTE: Tear production can last over 1 hour. During this time, tears can still be collected. - Quantify the total protein content of the tear pool using the Bradford standard method22.
4. SDS-PAGE analysis
- Heat the protein samples at 99 °C for 4 min to denature the proteins. Vortex the samples briefly to mix the components.
- Load 30 µg of quantified protein content from each sample onto a 10% SDS-PAGE gel (Table 1) and run the samples at 90 V for 2 h using a standard method23.
- After electrophoresis, cover the gel with staining solution (Table 1) and incubate for 30 min to stain the proteins.
- Rinse the gel with the destaining solution several times until the background becomes clear and protein bands are visible.
- Acquire images with a gel documentation system.
5. mRNA isolation for qPCR and digital PCR
- Place the strips in a 600 µL microcentrifuge tube containing 20 µL of sterile water. Make a puncture at the tip of the tube containing the sample, put it in a new sterilized 1.5 mL tube, and centrifuge, as previously reported in human tears3, for 20 min at 2,000 x g at 4 °C (Figure 1E-H).
- Create a pool from all samples. The recovered volume of this experiment was approximately 22 µL of supernatant for each mouse, i.e., 66 µL of total volume. Place the tubes on dry ice to avoid biomolecule degradation.
- Add 200 µL of monophasic solution of phenol and guanidinium isothiocyanate (commercially obtained) and homogenize by pipetting. Let stand for 5 min at room temperature. At this step, samples can be stored at -20 °C for several weeks or -70 °C for several months.
- Add 40 µL of chloroform and mix in a vortex for 15 s. Let stand for 2 min at room temperature. Centrifuge at 10,000 x g for 15 min at 4°C.
- Carefully transfer the aqueous phase (without taking the interphase) to a new 1.5 mL tube.
- Add 0.5 µL of glycogen (0.05 µg/µL) to a 100 µL of isopropanol and mix by pipetting. Glycogen is co-precipitated with the RNA and does not interfere with subsequent steps.
- Let stand for 10 min at -20 °C. Centrifuge at 10,000 x g for 10 min at 4 °C. Quickly decant the supernatant.
- Add 200 µL of cold 75% ethanol and resuspend the RNA pellet by vortexing. Centrifuge at 8,000 x g for 5 min at 4 °C.
- Decant the ethanol. Without inverting the tube, let it air dry for 20-30 min. Add 20 µL of RNase-free water and resuspend.
- Proceed with reading the optical density in the microvolume spectrophotometer at 260 nm and 280 nm to assess the purity of RNA. Samples must be on ice.
NOTE: Ribosomal bands 28S and 18S cannot be observed using an agarose gel due to the nature of the sample. The presence of other RNAs, such as miRNAs or mRNAs, can be analyzed using a bioanalyzer, as previously shown18. - Synthesize the cDNA from tears RNA using a commercial kit as per the manufacturer's protocol.
- Start with an input RNA in a range from 10 ng-5000 ng; according to the kit used, mix it with 1 µL of Oligo (dT) and up to 12 µL of RNase-free water. Spin it at 500 x g for 30 s and incubate at 65 °C for 5 min.
- Add to the mixing tube 4 µL of 5x reaction buffer, 1 µL of RNase inhibitor, 2 µL of dNTP mix, and 1 µL of reverse transcriptase.
- Spin the tube at 500 x g for 30 s to pull down the mixture. Then, incubate the tube at 42 °C for 60 min. Finish the reaction by incubating the tube at 70 °C for 10 min.
- Due to the low concentration of mRNA in tears, it is recommended to use undiluted cDNA for qPCR24. For this experiment, 150 ng of cDNA were used. Perform qPCR using DNMT3a primers:
Forward: GCCGAATTGTGTCTTGGTGGATGACA, Reverse: CCTGGTGGAATGCACTGCAGAAGGA and GAPDH primers:
Forward: ACTGGCATGGCCTTCCGTGTTCTTA, Reverse: TCAGTGTAGCCCAAGATGCCCTTC following instructions from manufacturers and equipment procedures.- For qPCR cycle, use the following program: Initial denaturation: 95 °C for 10 min followed by 45 cycles of C 95°C – 15 s, 60 °C- 30 s, and 72 °C- 30 s, end with a final extension at 72 °C for 5 min.
- Perform a melt curve analysis to assess the presence of primer dimers or unspecific amplicons in the reaction. The temperature ramp started from 50 °C to a final 99 °C, with an initial hold of 90 s for the first step and a hold of 5 s between each step.
- For dPCR25, perform serial dilutions to optimize the amount of cDNA that is required to detect the mRNA of interest. In this experiment, use the following dilutions of cDNA in nuclease-free water: 150 ng, 75 ng, 37.5 ng, and 18.75 ng of cDNA. Primers for DNMT3a previously mentioned were used. For dPCR cycle, use the following program: initial denaturation: 95 °C for 2 min followed by 40 cycles of 95 °C – 15 s and 60 °C- 30 s.
Representative Results
The protocol described in this paper provides an easy and affordable method for obtaining molecular information from tear fluid using techniques commonly available in most molecular biology laboratories. Furthermore, the protocol can be scaled up by employing highly sensitive techniques such as ELISA for enzymatic activity detection.
After these procedures, the total protein yield was approximately 3-4 µg/µL. Coomassie-stained SDS page analysis of total protein extracts reveals patterns of protein expression in total retina and tears (Figure 2A).
Isolated RNA had a 260/280 UV absorption ratio between 1.9 and 2.0. The average yield was 134.8 ng/µL. qPCR was performed using primer sets for a housekeeping gene (glyceraldehyde 3-phosphate dehydrogenase, GAPDH)3 and a putative biomarker gene (DNA-methyl transferase 3a, DNMT3a)26. An undiluted cDNA sample 1:1 (150 ng) was used.
A melting curve analysis was performed, and Ct rates were calculated in the qPCR software. All the experiments were performed in duplicate. The analysis revealed Ct values between 20 and 24, which is a suitable range of starting amount of target mRNA in the sample (Figure 2B). However, the melt curve analysis in Figure 2C suggests the low presence of target mRNA. Nonetheless, the 2% agarose gel confirmed the presence of amplicons at the expected size (Figure 2D).
dPCR was performed using the same pair of primer sets for DNMT3a. The dilution of cDNA samples used were 1:1 (150 ng), 1:2 (75 ng), 1:4 (37.5 ng), 1:8 (18.75 ng). All experiments were performed in duplicate. dPCR was able to detect from 1.33 copies/µL in the undiluted sample, to 0.45 copies/µL in the 8-fold diluted sample (Figure 3A,3B). Thus, dPCR showed an increased sensitivity to the target mRNA present in tears compared to qPCR.
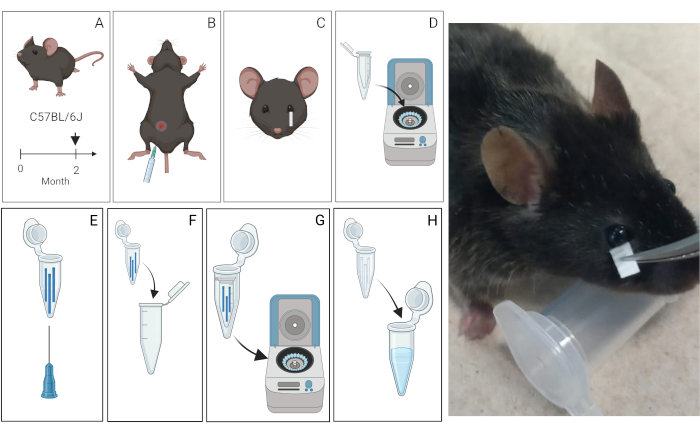
Figure 1: Schematic of the tear extraction protocol from mice. (A) Tear Collection. Three 2-month-old C57BL/6J mice were used. (B) Tear production was pharmacologically induced by intraperitoneal administration of pilocarpine. Photograph illustrating the placement of the Schirmer strip. (C) Tear Processing. Tears were then collected using Schirmer strips for 10 min and transferred into microcentrifuge tubes. (D) Tears collected in the Schirmer strips were centrifuged with sterile water and protease inhibitors or just sterile water. (E, F) These tubes were perforated at the bottom and subsequently transferred to a new sterilized 1.5 mL tube to facilitate fluid elusion. (G, H) The tears are collected in the 1.5 tube mL. (I) Representative photograph of the procedure to collect tears. Please click here to view a larger version of this figure.
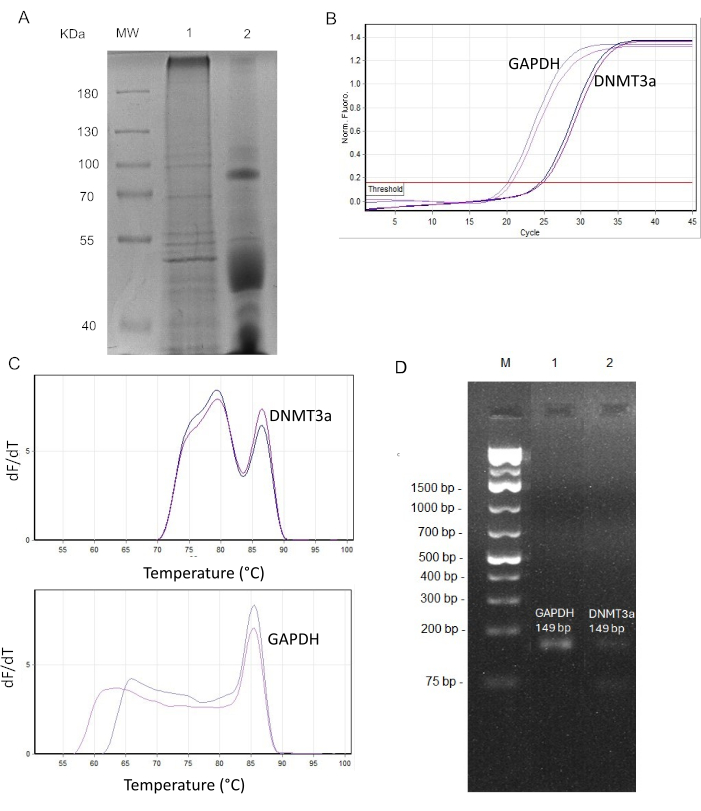
Figure 2: Downstream analysis of tear film for molecular applications. (A) SDS-PAGE of mouse tears and a retinal protein extract. MW: Molecular weight marker.; lane 1: Protein profiles of retina (30 µg); lane 2: protein profiles of tears (30 µg). (B) qPCR for GAPDH and DNMT3a in mice tears. Average CT and ΔCT are shown in Table 2. (C) Melting curve analysis for DNMT3a and GAPDH PCR products. The y-axis represents the rate of change of fluorescence in the amplification reaction (dF/dT), and the x-axis represents temperature in °C (D) Analysis of the PCR amplicon in 2% agarose gel electrophoresis containing 0.5 µg/mL ethidium bromide. Lane M: 1 kb Plus DNA Ladder. Lane 1,2: GAPDH and DNMT3a PCR amplicon, respectively. Please click here to view a larger version of this figure.
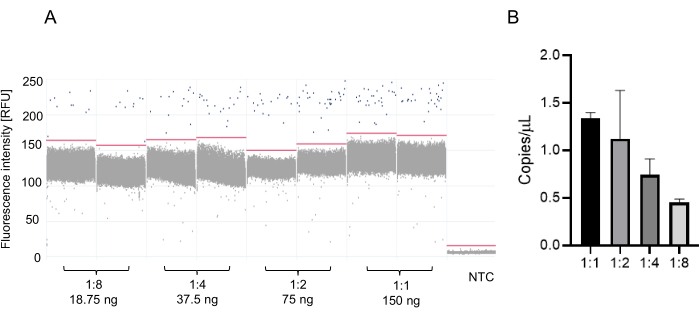
Figure 3: Digital PCR (dPCR) for DNMT3a expression in mice tears. (A) dPCR results are displayed as a 1D scatter plot showing in blue dots the DNMT3a transcript copies. Black dots are partitions without the DNMT3a transcript copies present. (B) A dilution series of cDNA was generated to determine the optimal concentration for quantifying DNMT3a transcript copies. Error bars represent standard deviation. Please click here to view a larger version of this figure.
| Name of Material | Description | ||||
| 10% SDS Gel | For one separating gel: 3.8 mL distilled H2O; 2.6 mL Tris; pH 8.8; 1.5 M 3.4 mL acrylamide; 100 µL 10% SDS; 100 µL 10% APS; 4 µL TEMED. | ||||
| For one collecting gel: 2.72 mL distilled H2O, 0.52 mL Tris; pH 6.8; 1.5 M 0.68 mL acrylamide, 40 µL 10% SDS, 40 µL 10% APS 4 µL TEMED. | |||||
| Electrophoresis Buffer | 250 mM Tris base, 2 M Glycine, 1% SDS | ||||
| Staining Solution | Dissolve the following reagents in 43 mL of water (store at room temperature in an amber bottle): Coomassie 0.25 g; acetic acid 7 mL; methanol 50 mL | ||||
| Destaining solution | Dissolve the following reagents in 63 mL of water (store at room temperature): acetic acid 7 mL; ethanol 30 mL | ||||
Table 1. Set up for SDS-Page.
| Gene name | CT1 | CT2 | Average with SD | ΔCT |
| GAPDH | 20.14 | 20.73 | 20.44±0.17 | |
| DNMT3a | 24.59 | 24.97 | 24.78±0.07 | 4.35 |
Table 2. Ct values of DNMT3a and GAPDH in tears of mice.
Discussion
Tear fluid is easily accessible, and the determination of biomarkers in tears can be employed as a successful complementary technique for the early diagnosis of various human diseases27. While the biochemical analysis of tear composition in experimental animal models complements this approach and promises significant progress in understanding the molecular basis of diseases, there is a scarcity of available data and protocols, which led us to develop one. The method described in this report is technically simple to perform and allows for the preliminary identification of the differential expression of different disease-associated candidate molecules.
Tear collection can be achieved through various methods such as capillary tubes and Schirmer strips16,17, swabs28and sponges29. The Schirmer strip method was preferred due to its ease of access, quantitative capabilities, adaptability during sample collection, cost, and simplicity in extracting tear components for laboratory processing. Additionally, this method is cost-effective, minimally invasive, and poses no risk of injury or damage to the cornea, thanks to the composition of the paper filter strip16,17. Critical steps of this protocol include the need for pharmacological stimulation of tear production using pilocarpine and the selection of an appropriate elution buffer.
Pilocarpine is a cholinergic agonist that binds to M3 muscarinic receptors and can cause pharmacological stimulation of exocrine glands, enhancing tear production, salivary secretion, and urination. Although studies in humans demonstrated that tear stimulation does not change the osmolarity of the collected fluid30, it clearly influences protein profiles31. The use of pilocarpine as a cholinergic pharmacological stimulator of exocrine glands and tear secretion enhancer has been successfully reported in rabbits and mice19,32,33, but the effect of stimulation on protein content has not been evaluated. To avoid this limitation and allow accurate comparison of tear profiles between experimental conditions, it is crucial to consider the specific collection settings. Similarly, the introduction of a rinsing step prior to tear collection could be beneficial to avoid prior contamination of the eye surface. In this protocol, a concentration of 30 mg/kg of pilocarpine was employed13. As the mice remained immobile and calm after the injection of this drug, the tears were collected without any anesthesia, making sample collection easy.
The choice of an appropriate elution buffer is closely related to the selected method for analyzing the biochemical composition of a tear sample and the recovery rate of biological material from the strip. For example, in a comparative study focusing on protein retention from Schirmer's strips in patients, it was established that a buffer comprising 100 mM ammonium bicarbonate along with 0.25% Nonidet P40 (NP40) yielded superior results for proteomics or multiplex ELISA analysis applications17. In our hands, the use of sterile water proved to be sufficient to visualize total protein extracts in SDS-PAGE and to detect the mRNA of the genes of interest by PCR.
To assess the presence of DNMT3a and GAPDH mRNA in mice tears, we compared the performance of qPCR and dPCR. The presence of primer dimers observed in the qPCR melting curve is attributed to the low abundance of the DNMT3a transcript. As demonstrated, dPCR has significantly greater sensitivity and efficacy in detecting the target mRNA in these samples in undiluted and diluted samples. This precise detection capability makes dPCR the preferred method for tear analysis in mice, offering researchers and clinicians a reliable tool for future studies and potential clinical applications.
The development of a standardized protocol for tear sample collection in animal models represents a significant advancement in research. This protocol not only facilitates the systematic and precise collection of tear samples but also opens a wide range of possibilities for their application in various areas of study. The ability to obtain tear samples from animal models allows for the investigation of a broad spectrum of diseases and conditions, as well as the exploration of new biomarkers and potential therapies. Furthermore, by establishing a standardized protocol, the reproducibility of results across different studies and laboratories is promoted, thereby contributing to the validity and reliability of research. Ultimately, this advancement in tear sample collection methodology in animal models has the potential to drive important discoveries and improve the understanding of the pathophysiology of various diseases, opening new avenues for diagnosis, treatment, and prevention.
Disclosures
The authors have nothing to disclose.
Acknowledgements
This work was supported by VELUX STIFTUNG [project 1852] to M.L. and postgraduate fellowship grants from CONAHCYT to M.B. (836810), E.J.M.C. (802436) and A.M.F (CVU 1317418). Sincere gratitude to all the members of the laboratory, Centro de Investigación sobre el Envejecimiento, and Departamento de Farmacobiología (Cinvestav) for their contributions to the stimulating discussions.
Materials
| 2-mercaptoethanol | Gibco | 1985023 | |
| 2x Laemmli buffer | Bio-Rad | 16-0737 | |
| Acetic Acid | Quimica Meyer | 64-19-7 | |
| Acrylamide | Sigma-Aldrich | A4058 | |
| Bradford Reagent | Sigma-Aldrich | B6916 | |
| Chloroform | Sigma-Aldrich | 1003045143 | |
| Coomassie Blue R 250 | US Biological | 6104-59-2 | |
| Ethanol | Quimica Rique | 64-17-5 | |
| GeneRuler 1kb plus DNA Ladder | Thermofisher scientific | SM1331 | |
| Glycerol | US Biological | G8145 | |
| Glycine | SANTA CRUZ | SC- 29096 | |
| Glycogen | Roche | 10901393001 | |
| HCl | Quimica Rique | 7647-01-0 | |
| Isopropyl alcohol | Quimica Rique | 67-63-0 | |
| Methanol | Quimica Meyer | 67-56-1 | |
| Micro tubes 1.5 ml | Axygen | MCT-150-C | |
| Micro tubes 600 µl | Axygen | MCT-060-C | |
| NaCl | Sigma-Aldrich | S3014 | |
| PCR tubes & strips | Novasbio | PCR 0104 | |
| Pilocarpine | Sigma-Aldrich | P6503-10g | |
| Protease inhibitor | Roche | 11873580001 | |
| QIAcuity EvaGreen PCR Kit (5mL) | Qiagen | 250112 | |
| QIAcuity Nanoplate 26k 24-well (10) | Qiagen | 250001 | |
| Real qPlus 2x Master Mix Green | Ampliqon | A323402 | |
| RevertAid First Strad cDNA Synthesis Kit | Thermofisher scientific | K1622 | |
| Schirmer's test strips | Laboratorio Santgar | SANT1553 | |
| SDS | Sigma-Aldrich | L3771 | |
| TEMED | Sigma aldrich | 102560430 | |
| TRI reagent | Sigma-Aldrich | T9424-200ML | monophasic solution of phenol and guanidinium isothiocyanate |
| Tris | US Biological | T8650 | |
| Tris base | Chem Cruz | sc-3715A |
References
- Ravishankar, P., Daily, A. Tears as the next diagnostic biofluid: A comparative study between ocular fluid and blood. Appl Sci. 12 (6), 2884 (2022).
- Masoudi, S. Biochemistry of human tear film: A review. Exp Eye Res. 220, 109101 (2022).
- Dara, M. Novel RNA extraction method from human tears. Mol Biol Res Commun. 11 (4), 167-172 (2022).
- Amorim, M., et al. Putative biomarkers in tears for diabetic retinopathy diagnosis. Front Med (Lausanne). 9, 873483 (2022).
- Arslan, M. A., et al. Expanded biochemical analyses of human tear fluid: Polyvalent faces of the Schirmer strip. Exp Eye Res. 237, 109679 (2023).
- Liu, R., et al. Analysis of th17-associated cytokines and clinical correlations in patients with dry eye disease. PloS one. 12 (4), e0173301 (2017).
- Cross, T., et al. Rna profiles of tear fluid extracellular vesicles in patients with dry eye-related symptoms. Int J Mol Sci. 24 (20), 15390 (2023).
- Paranjpe, V., Phung, L., Galor, A. The tear film: Anatomy and physiology. Ocular Fluid Dyn: Anat, Physiol, Imaging Tech, and Math. , 329-345 (2019).
- Jung, J. H., et al. Proteomic analysis of human lacrimal and tear fluid in dry eye disease. Sci Rep. 7 (1), 13363 (2017).
- Di Zazzo, A., Micera, A., De Piano, M., Cortes, M., Bonini, S. Tears and ocular surface disorders: Usefulness of biomarkers. J Cell Physiol. 234 (7), 9982-9993 (2019).
- Von Thun Und Hohenstein-Blaul, N., Funke, S., Grus, F. H. Tears as a source of biomarkers for ocular and systemic diseases. Exp Eye Res. 117, 137-126 (2013).
- Pieczynski, J., Szulc, U., Harazna, J., Szulc, A., Kiewisz, J. Tear fluid collection methods: Review of current techniques. Eur J Ophthalmol. 31 (5), 2245-2251 (2021).
- Shiraki, T., Shigeta, M., Tahara, N., Furukawa, H., Ohtsuka, H. A tear production assessment by using Schirmer tear test strips in mice, rats and dogs. Animal Eye Res. 24 (1-2), 1-2_1-1-2_5 (2005).
- Zhao, J., Nagasaki, T. Lacrimal gland as the major source of mouse tear factors that are cytotoxic to corneal keratocytes. Exp Eye Res. 77 (3), 297-304 (2003).
- Khanna, R. K., et al. Metabolomics and lipidomics approaches in human tears: A systematic review. Surv Ophthalmol. 67 (4), 1229-1243 (2022).
- Bachhuber, F., Huss, A., Senel, M., Tumani, H. Diagnostic biomarkers in tear fluid: From sampling to preanalytical processing. Sci Rep. 11 (1), 10064 (2021).
- Koduri, M. A., et al. Optimization and evaluation of tear protein elution from Schirmer’s strips in dry eye disease. Indian J Ophthalmol. 71 (4), 1413-1419 (2023).
- Kakan, S. S., et al. Tear miRNAs identified in a murine model of sjogren’s syndrome as potential diagnostic biomarkers and indicators of disease mechanism. Front Immunol. 13, 833254 (2022).
- Moore, M., Ma, T., Yang, B., Verkman, A. Tear secretion by lacrimal glands in transgenic mice lacking water channels aqp1, aqp3, aqp4 and aqp5. Exp Eye Res. 70 (5), 557-562 (2000).
- Balafas, E., et al. A noninvasive ocular (tear) sampling method for genetic ascertainment of transgenic mice and research ethics innovation. OMICS. 23 (6), 312-317 (2019).
- Nakagawa, A., Nakajima, T., Azuma, M. Tear miRNA expression analysis reveals mir-203 as a potential regulator of corneal epithelial cells. BMC Ophthalmol. 21 (1), 377 (2021).
- Bradford, M. M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem. 72, 248-254 (1976).
- Sambrook, J., Fritsch, E. F., Maniatis, T. . Molecular cloning: A laboratory manual. , (1989).
- . RealQPlus2x Master Mix Green without ROX Available from: https://ampliqon.com/en/pcr-enzymes/pcr-enzymes/real-time-pcr/realq-plus-2x-master-mix-green (2023)
- Qiagen. . Quick-Start Protocol QIAcuity EG PCR Kit. , (2023).
- Anier, K., Malinovskaja, K., Aonurm-Helm, A., Zharkovsky, A., Kalda, A. DNA methylation regulates cocaine-induced behavioral sensitization in mice. Neuropsychopharmacology. 35 (12), 2450-2461 (2010).
- Hagan, S., Martin, E., Enriquez-De-Salamanca, A. Tear fluid biomarkers in ocular and systemic disease: Potential use for predictive, preventive, and personalized medicine. EPMA J. 7 (1), 15 (2016).
- Gasparini, M. S., et al. Identification of sars-cov-2 on the ocular surface in a cohort of covid-19 patients from Brazil. Exp Biol Med. (Maywood). 246 (23), 2495-2501 (2021).
- Sebbag, L., Harrington, D. M., Mochel, J. P. Tear fluid collection in dogs and cats using ophthalmic sponges. Vet Ophthalmol. 21, 249-254 (2018).
- Abusharha, A. A., et al. Analysis of basal and reflex human tear osmolarity in normal subjects: assessment of tear osmolarity. Ther Adv Ophthal. 10, 2515841418794886 (2018).
- Fullard, R. J., Snyder, C. Protein levels in nonstimulated and stimulated tears of normal human subjects. Inves Opht Vis Sci. 31 (6), 1119-1126 (1990).
- Longwell, A., Birss, S., Keller, N., Moore, D. Effect of topically applied pilocarpine on tear film pH. J Pharm Sci. 65 (11), 1654-1657 (1976).
- Dartt, D. A. Neural regulation of lacrimal gland secretory processes: relevance in dry eye diseases. Prog Ret Eye Res. 28 (3), 155-177 (2009).

.
