In Situ Hybridization Techniques for Paraffin-Embedded Adult Coral Samples
Instructor Prep
concepts
Student Protocol
1. Removal of Paraffin
Caution: Perform the following steps under a fume hood.
- Dewax the thin-sectioned paraffin-embedded slides with 100% xylene under the hood in glass Coplin jars for 10 min. Do not use plastic Coplin jars, as xylene melts plastic. Sterilize the Coplin jars in an autoclave before use.
- Prepare four sterile glass Coplin jars with the following: 100% ethanol, 80% ethanol, 70% ethanol and 60% ethanol. Dilute the ethanol with RNase-free water.
- Transfer the slides to the sterile glass Coplin jar with 100% ethanol and soak them for 10 min. After 10 min, empty the glass Coplin jar and replace with new 100% ethanol. Soak the slides again for 10 min.
- Transfer the slides to the sterile glass Coplin jar with 80% ethanol for 1 min.
- Transfer the slides to the sterile glass Coplin jar with 70% ethanol for 1 min.
- Transfer the slides to the sterile glass Coplin jar with 60% ethanol for 1 min.
2. Pretreatment of Slides for Preparation of RNA Probe Hybridization
- Perform the following washes at room temperature, unless specified otherwise. Perform room temperature washes on an orbital shaker at a slow speed. Use sterile slide mailers with room for up to five slides for the washing of the slides.
- Transfer the slides to a sterile slide mailer with 18 mL of 1x phosphate buffered saline (PBS). Wash for 5 min at room temperature.
- Pour off the 1x PBS, and immediately treat the tissue on the slides with proteinase K to enable probe penetration of the tissue. Add approximately 18 mL of proteinase K solution to the slide mailer (see Table 1 for working solution concentration). Incubate at 37 °C for 15 min. Do not shake.
- While the slides are incubating, prepare the prehybridization buffer (Prehybe Buffer). Place the Prehybe Buffer in a boiling water bath for 10 min, then cool on an ice bath for 5 min. For a recipe of Prehybe Buffer, see Table 1.
- Stop digestion of the proteinase K reaction by pouring out the proteinase K solution, and immediately add 18 mL of 0.2% glycine-PBS solution to the slide mailer. Let the slide mailer sit at room temperature for 5 min.
- Pour out the 0.2% glycine-PBS solution and add 18 mL of 2x saline sodium citrate (SSC) solution to each slide mailer. Wash the slides in the 2x SSC for 10 min at room temperature, with shaking at 100-150 rpm.
3. Prehybridization of Slides in Preparation for RNA Probe Hybridization
- While the slides are being washed with 2x SSC, obtain a new sterile slide mailer and put approximately 18 mL of Prehybe Buffer into the slide mailer. Place the slide mailer into the hybridization oven at hybridization temperature.
NOTE: The hybridization temperature will depend on the probe being used but usually ranges from 50-60 °C. - Once the 2x SSC wash is completed, pour out the 2x SSC, and place the slides within the slide mailer in the Prehybe Buffer in the hybridization oven for 1 h.
4. Hybridization of the RNA Probe
- While the slides are incubating with the Prehybe Buffer in the hybridization oven, prepare hybridization-probe solution.
- Dilute each probe in hybridization buffer (0.5 µL of probe with 24.5 µL of hybridization buffer).
NOTE: The recipe for hybridization buffer is found in Table 1. An example of probe preparation and concentrations can be found in Traylor-Knowles et al.11. - Place the diluted probe on an 86-90 °C heat block for 12 min, then cool for 1 min on ice.
- Dilute each probe in hybridization buffer (0.5 µL of probe with 24.5 µL of hybridization buffer).
- Remove the slide mailer from the hybridization oven and individually remove slides from the slide mailer using sterile tweezers. Lay the slides flat on a paper towel, and carefully wipe off excess Prehybe Buffer around the tissue samples. Be careful not to touch the samples, and work quickly to prevent drying of the tissue samples.
- Encircle the tissue with a PAP pen. Apply 25 µL of diluted probe solution with a pipette and cover the sample with a plastic coverslip.
- Place the samples in the slide moisture chamber with 4x SSC + 50% formamide solution in the bottom of the chamber.
- Once probes have been added to all the slides, place the slide moisture chamber into the hybridization oven for at least 24 h at a hybridization temperature of 50-60 °C. The incubation time can vary depending on the concentration of the probe but should be for a minimum of 24 hours.
- After overnight incubation with the diluted probes, remove the slide moisture chamber from the hybridization oven.
- Remove coverslips from each of the slides, being careful not to displace the tissue. In a 1000 µL pipette, add 1000 µL of 2x SSC solution and gently wash the slide off.
- Place the slide in a sterile slide mailer with 18 mL of 2x SSC solution for 5 min at room temperature. Pour out the solution and replace it with 18 mL of fresh 2x SSC. Repeat the incubation again for 5 min at room temperature with gentle shaking.
- Pour out the 2x SSC solution. Add 18 mL of 1x SSC solution and wash for 5 min at room temperature. Pour out the 1x SSC solution and replace it with 18 mL of fresh 1x SSC. Repeat the incubation for 5 min at room temperature with gentle shaking.
- Pour out the 1x SSC. Add 18 mL of 0.5x SSC and wash for 10 min at 42 °C without shaking. Pour out the 0.5x SSC and replace it with 18 mL of fresh 0.5x SSC. Wash again for 10 min at 42 °C without shaking.
5. Visualization of the RNA Probe
NOTE: For visualizing the probe, BM purple will be used during the development process. However, before this step, several washes are required to prepare the samples for staining.
- Wash the slides with 18 mL of alkaline phosphatase buffer (AP-buffer) without MgCl2 for 1 min. Pour out the AP-Buffer without MgCl2, and add 18 mL of 1x Boehringer-Mannheim blocking buffer diluted with maleic acid buffer. Incubate for at least 1 h at room temperature in a slide mailer with gentle shaking.
NOTE: The Boehringer-Mannheim blocking buffer and maleic acid buffer used were premade and purchased in the DIG Wash and Block Buffer Set (available commercially).- Alternatively, incubation overnight at 4 °C can be performed. A longer blocking period will lessen the appearance of non-specific binding.
- Prepare the 20 mL of diluted DIG anti-digoxigenin-AP Fab fragments (anti-DIG antibody = 2 µL of anti-DIG antibody + 20 mL 1x Boehringer-Mannheim blocking buffer). This is enough for use in 1 plastic slide mailer. More of this solution must be prepared if more than one slide mailer is to be used.
- Add the anti-DIG antibody to a new sterile slide mailer, and transfer the slides to the slide mailer with the anti-DIG antibody solution. Incubate at room temperature for 3 h with gentle shaking.
- After incubation, pour out the anti-DIG antibody and wash the slides in the slide mailer with 18 mL of AP-Buffer without MgCl2 for 5 min with gentle shaking.
- Pour out the AP-Buffer without MgCl2, and add 18 mL of AP-Buffer. Wash for 5 min with gentle shaking. After the 5 min incubation, pour off the AP-Buffer, replace it with 18 mL of fresh AP-Buffer, and wash for 5 min with gentle shaking.
- In the dark, pour out the AP-Buffer and add 18 mL of BM purple to the slide mailer. Incubate the slides at room temperature in the dark, checking for purple color development every ½ h. Reaction times for visualization will vary based on the probe that is being developed.
NOTE: Instead of BM purple visualization, development solution can be made using 5-bromo-4-chloro-3-indolyl-phosphate (BCIP) and nitro blue tetrazolium (NBT). See Table 1 for instructions. - Stop color development by transferring the slides to a new sterile slide mailer with 18 mL of Tris-EDTA (TE) buffer for 5 min at room temperature, in the dark.
- Pour off the TE buffer and add 18 mL of RNase-free water to the slide mailer and wash for 1 min, in the dark.
- Remove the slides from the water, and dry them around the edges of the tissue. Add glycerol mounting medium and place the coverslips. Store the slides at 4 °C until pictures are ready to be taken.
In Situ Hybridization Techniques for Paraffin-Embedded Adult Coral Samples
Learning Objectives
After completing this protocol, identification of cells and tissues that are expressing the RNA probe of interest will be achieved. The representative results for this protocol are for AP-1, FosB, and TNFR41. These results, previously published by Traylor-Knowles et al.11, show spatial expression of RNA probes on adult corals that were exposed to heat stress. Two examples of different staining types are presented in Figure 1. Figure 1A is an example of diffuse tissue staining. The stain is found throughout the tissue, not only in specific cells. Here, the expression of anti-sense AP-1 is found throughout the epidermis and the oral gastrodermis11 (Figure 1A). The staining is diffuse and not segregated to one specific cell type. For this type of staining, it is particularly critical to run a sense control at the same time, as some of the results may be due to non-specific staining.
The second type of staining is cell-specific, in which the stain is only found in specific cell types. Both FosB (Figure 1B) and TNFR41 (Figure 1C) show this type of staining. Both of these probes were used on tissue samples of adult corals that had been exposed to a heat stress11. In the anti-sense panel, purple staining is observed. TNFR41 was expressed in different types of cnidocytes, a family of cell types that is only found within cnidarians (Figure 1). Specifically, it stained nematocytes that produce the microbasic mastigophore organelle (nematocysts) and spirocytes that produce spirocysts, all found within the epidermis of coral tissue. Nematocytes are also found in other tissue layers of the coral, such as the calicodermis; however, due to the sectioning performed on this particular sample, that information could not be gathered. FosB also stained cnidocytes and was specific to both spirocytes and nematocytes. For both of these probes, the sense control showed no staining, indicating that the staining for the anti-sense probe was indeed specific. These are just two types of representative results (diffuse tissue staining and cell-specific staining) of the staining techniques that may be present with different types of probes. Due to this variability, it is critical to use a sense control to determine non-specific staining.
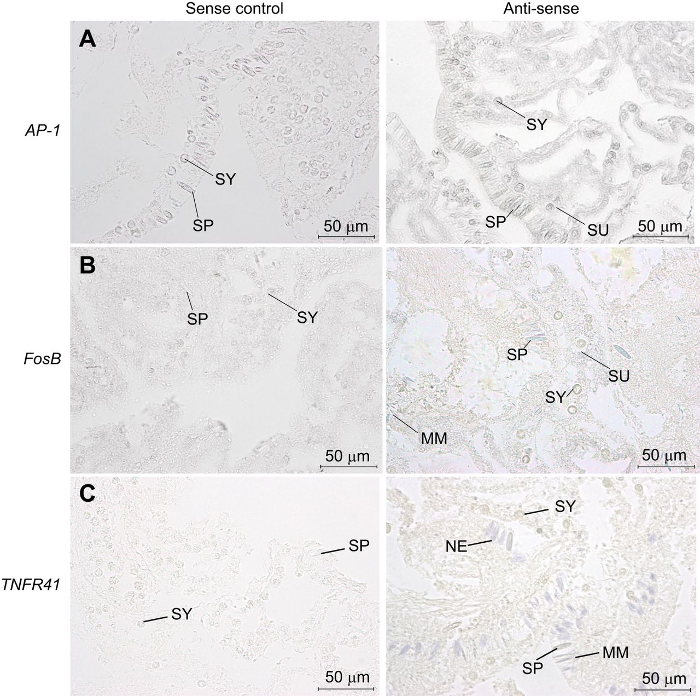
Figure 1: Representative outcome of ISH that stained specific cells. (A) In the first panel, sense control for the AP-1 probe shows that little staining is present within the slide. In the second panel, the anti-sense probe for AP-1 shows that staining with this probe is diffuse throughout the tissue. The stain is specific to the oral gastrodermis and epidermis. With this type of staining, it is critical to run a sense control to ensure that the observed staining is specific. (B) A sense control for the FosB probe in the first panel shows little staining present. In the second panel, the anti-sense probe for FosB shows staining specific to the spirocytes and nematocytes, with diffuse staining throughout the epidermis and oral gastrodermis. (C) In the first panel, the sense control for the TNFR 41 probe shows that little staining is present on the slide. In the second panel, the anti-sense probe for TNFR 41 shows specific staining within the spirocytes, nematocytes, and microblastic mastigophores. Abbreviations: spirocytes (SP), symbiodium (SY), nematocyte (NE), microbasic mastigophores (MM), symbiont-containing gastrodermal cell (SU). This figure was reproduced with permission11. Please click here to view a larger version of this figure.
| Note: All dilutions should be done with sterile, RNase Free water in autoclaved glassware. | |||
| Prehybe Buffer (50 mL) | ADD | ||
| Formamide | 25 mL | ||
| 20X SSC (pH 4.5) | 12.5 mL | ||
| 20 mg/mL heparin | 0.1 mL | ||
| 50X Denhardts | 5 mL | ||
| 20% Tween-20 | 0.5 mL | ||
| 20% SDS | 0.5 mL | ||
| 10 mg/mL Salmon Sperm DNA (denature before adding) | 2 mL | ||
| RNase Free water | 4.4 mL | ||
| Note: Solution can be made in a disposible 50 mL tube. Salmon Sperm should be denatured by boiling on a heat block for 5 minutes before adding to solution. | |||
| Hybridization Buffer (47 mL) | ADD | ||
| Formamide | 25 mL | ||
| 20X SSC (pH 4.5) | 12.5 mL | ||
| 20 mg/mL heparin | 0.1 mL | ||
| 50X Denhardts | 5 mL | ||
| 20% Tween-20 | 0.5 mL | ||
| 20% SDS | 0.5 mL | ||
| 10 mg/mL Salmon Sperm DNA (denature before adding) | 2 mL | ||
| RNase Free water | 1 mL | ||
| Note: Solution can be made in a disposible 50 mL tube. Salmon Sperm should be denatured by boiling on a heat block for 5 minutes before adding to solution. | |||
| 10X PBS (1.0 L) | ADD | ||
| 18.6 mM NaH2PO4 | 2.56 g | ||
| 84.1 mM Na2HPO4 | 11.94 g | ||
| 1,750 mM NaCl | 102.2 g | ||
| Notes: Mix phosphates in about 800 mL of water for a 1.0 L volume. Check pH. It should be 7.4 ± 0.4. Otherwise adjust pH to 7.4 with NaOH of HCl. Add NaCl and rest of water. After pH is adjusted autoclave. | |||
| 20X SSC (1.0 L) | ADD | ||
| 3 M NaCl | 175.3 g | ||
| 0.3 M Na Citrate | 88.2 g | ||
| Notes: Mix in about 800 mL of RNase free water to bring to a 1.0 L volume. Adjust pH to 4.5 and autoclave. | |||
| Alkaline Phosphatase (AP) buffer w/o MgCl2 (50 mL) | ADD | ||
| 1 M NaCl | 5.0 mL | ||
| 1 M Tris, pH 9.5 | 5.0 mL | ||
| 20% Tween-20 | 1.25 mL | ||
| RNase Free water | 38.75 mL | ||
| Notes: Prepare just prior to use in a 50 mL tube. | |||
| Alkaline Phosphatase (AP) buffer (100 mL) | ADD | ||
| 1 M NaCl | 10.0 mL | ||
| 1 M MgCl2 | 5 mL | ||
| 1 M Tris, pH 9.5 | 10 mL | ||
| 20% Tween-20 | 2.5 mL | ||
| RNase Free water | 72.5 mL | ||
| Notes: Prepare just prior to use in a 50 mL tube. The solution will become cloudy after a few hours and will no longer work for the enzymatic reaction. | |||
| AP Substrate Solution | ADD | ||
| AP Buffer | 25 mL | ||
| NBT | 82.5 uL | ||
| BCIP | 82.5 uL | ||
| Notes: This can be used instead of BM Purple. Prepare just prior to use in a 50 mL tube. Keep this solution in the dark by covering tube in foil, and preparing in low light. | |||
| Proteinase K Stock Solution (10 mg/mL) | ADD | ||
| Proteinase K | 10 mg | ||
| RNase Free water | 10 mL | ||
| Notes: Aliquot and store at -20 °C. | |||
| Proteinase K Working Solution | ADD | ||
| Proteinase K stock solution | 90 ul | ||
| 1X PBS | 18 mL | ||
| Notes: Make just prior to use for best results. | |||
| 0.2% glycine-PBS solution | ADD | ||
| 10X PBS | 45 mL | ||
| RNase Free Water | 405 mL | ||
| Glycine | 1 g | ||
| Notes: Prepare in autoclaved glass container. Mix at room temperature on a stirer plate until glycine is fully dissolved. | |||
| Boehringer-Mannheim Blocking Buffer (30 mL) (Part of the DIG Wash and Block Buffer Set) | ADD | ||
| 10x Blocking solution | 3 mL | ||
| 1x maleic acid buffer | 27 mL | ||
| Notes: Make just prior to use for best results. Stock Blocking solution and maleic acid buffer were used from the “DIG Wash and Block -Buffer Set". | |||
| 4X SSC—50% formamide (30 mL) | ADD | ||
| 20X SSC | 6 mL | ||
| 50% formamide | 24 mL | ||
| Notes: Prepare just prior to use in 50 mL tube. Prepare under laboratory hood. | |||
| Glycerol Mounting Media | ADD | ||
| 50 mM Tris, pH 8.4 | 80 µL | ||
| Glycerol | 20 µL | ||
Table 1: Stock solutions for performing in situ hybridization. The table is a list of the different stock solutions needed for this protocol. Total amounts are estimated for performance of approximately 2 slide mailers. It is advised to adjust total amounts based on the amount of slides that are being processed.
List of Materials
| Denhardt's solution | Affymetrix | 70468 50 ML | |
| Bioworld Alkaline phosphatase buffer | Fisher | 50-198-724 | |
| Slide mailers | Fisher | 12-587-17B | |
| Bioworld Alkaline phosphatase buffer | Fisher | 50-198-724 | |
| 50 mL Falcon tubes | Fisher | 14-959-49A | |
| UltraPure Salmon Sperm DNA solution | Invitrogen | 15632-011 | |
| PBS – Phosphate-Buffered Saline (10X) pH 7.4 | Invitrogen | AM9625 | |
| UltraPure DNase/RNase-Free Distilled Water, 10 x 500 mL | Invitrogen | 10977-023 | |
| UltraPure DNase/RNase-Free Distilled Water, 10 x 500 mL | Invitrogen | 10977-023 | |
| UltraPure Salmon Sperm DNA solution | Invitrogen | 15632-011 | |
| Slide white apex superior adhesive | Leica Biosystems | 3800080 | |
| PBS solution, pH 7.4 | Life Technologies | 10010072 | |
| Proteinase K, Molecular Grade, 2 mL | New England Biolabs | P8107S | |
| Super Pap Pen Liquid Blocker | Promega | 22309 | |
| DIG Anti-Digoxigenin-AP Fab fragments | Roche | 11093274910 | |
| BM Purple, 100 mL | Roche | 11442074001 | |
| DIG Wash and Block Buffer Set | Roche | 11585762001 | |
| NBT/BCIP | Roche | 11681451001 | |
| Formaldehyde solution, 500 mL size | Sigma-Aldrich | 252549-500ML | |
| SSC Buffer 20X concentration | Sigma-Aldrich | S6639-1L | |
| Acetic Anhydride | Sigma-Aldrich | 320102-100ML | |
| Formamide | Sigma-Aldrich | 47670-250ML-F | |
| Triethanolamine | Sigma-Aldrich | 90279-100ML | |
| Heparin sodium salt from porcine intestinal mucosa | Sigma-Aldrich | H3149-10KU | |
| Xylenes, AR (ACS), For Histological Use | VWR | MK866806 | |
| Ethanol | VWR | EM-EX0276-4S | |
| TE buffer | VWR | PAV6232 | |
| hybridization oven | VWR | 97005-252, 97005-254 | |
| Orbital shaker | VWR | 89032-088 |
Lab Prep
Corals are important ocean invertebrates that are critical for overall ocean health as well as human health. However, due to human impacts such as rising ocean temperatures and ocean acidification, corals are increasingly under threat. To tackle these challenges, advances in cell and molecular biology have proven to be crucial for diagnosing the health of corals. Modifying some of the techniques commonly used in human medicine could greatly improve researchers' ability to treat and save corals. To address this, a protocol for in situ hybridization used primarily in human medicine and evolutionary developmental biology has been adapted for use in adult corals under stress.
The purpose of this method is to visualize the spatial expression of an RNA probe in adult coral tissue that has been embedded in paraffin and sectioned onto glass slides. This method focuses on removal of the paraffin and rehydration of the sample, pretreatment of the sample to ensure permeability of the sample, pre-hybridization incubation, hybridization of the RNA probe, and visualization of the RNA probe. This is a powerful method when using non-model organisms to discover where specific genes are expressed, and the protocol can be easily adapted for other non-model organisms. However, the method is limited in that it is primarily qualitative, because expression intensity can vary depending on the amount of time used during the visualization step and the concentration of the probe. Furthermore, patience is required, as this protocol can take up to 5 days (and in many cases, longer) depending on the probe being used. Finally, non-specific background staining is common, but this limitation can be overcome.
Corals are important ocean invertebrates that are critical for overall ocean health as well as human health. However, due to human impacts such as rising ocean temperatures and ocean acidification, corals are increasingly under threat. To tackle these challenges, advances in cell and molecular biology have proven to be crucial for diagnosing the health of corals. Modifying some of the techniques commonly used in human medicine could greatly improve researchers' ability to treat and save corals. To address this, a protocol for in situ hybridization used primarily in human medicine and evolutionary developmental biology has been adapted for use in adult corals under stress.
The purpose of this method is to visualize the spatial expression of an RNA probe in adult coral tissue that has been embedded in paraffin and sectioned onto glass slides. This method focuses on removal of the paraffin and rehydration of the sample, pretreatment of the sample to ensure permeability of the sample, pre-hybridization incubation, hybridization of the RNA probe, and visualization of the RNA probe. This is a powerful method when using non-model organisms to discover where specific genes are expressed, and the protocol can be easily adapted for other non-model organisms. However, the method is limited in that it is primarily qualitative, because expression intensity can vary depending on the amount of time used during the visualization step and the concentration of the probe. Furthermore, patience is required, as this protocol can take up to 5 days (and in many cases, longer) depending on the probe being used. Finally, non-specific background staining is common, but this limitation can be overcome.
Procedure
Corals are important ocean invertebrates that are critical for overall ocean health as well as human health. However, due to human impacts such as rising ocean temperatures and ocean acidification, corals are increasingly under threat. To tackle these challenges, advances in cell and molecular biology have proven to be crucial for diagnosing the health of corals. Modifying some of the techniques commonly used in human medicine could greatly improve researchers' ability to treat and save corals. To address this, a protocol for in situ hybridization used primarily in human medicine and evolutionary developmental biology has been adapted for use in adult corals under stress.
The purpose of this method is to visualize the spatial expression of an RNA probe in adult coral tissue that has been embedded in paraffin and sectioned onto glass slides. This method focuses on removal of the paraffin and rehydration of the sample, pretreatment of the sample to ensure permeability of the sample, pre-hybridization incubation, hybridization of the RNA probe, and visualization of the RNA probe. This is a powerful method when using non-model organisms to discover where specific genes are expressed, and the protocol can be easily adapted for other non-model organisms. However, the method is limited in that it is primarily qualitative, because expression intensity can vary depending on the amount of time used during the visualization step and the concentration of the probe. Furthermore, patience is required, as this protocol can take up to 5 days (and in many cases, longer) depending on the probe being used. Finally, non-specific background staining is common, but this limitation can be overcome.
