Inducible and Reversible Dominant-negative (DN) Protein Inhibition
Instructor Prep
concepts
Student Protocol
Generation of the transgenic CBRb mouse and all animal care and experiments associated with the study were approved by the Creighton University Institutional Animal Care and Use Committee (IACUC) and performed by their guidelines.
1. Transgenic CB-Myc6-Rb1 Construct
NOTE: The cloning of CBRb into a pTet_Splice vector was done in a multi-step process (Figure 1A and 1B).
- Perform the first stage cloning into the pCS2+CB-Myc6 vector.
- To create the CBRb transgene construct, amplify a 1583-bp-long Rb1 cDNA fragment (528 amino acids corresponding to the amino acid region 369–896) of the RB1 protein using the following primer set: for EcoRI+RB 1243F, use GGGGAATTCATTAAATTCAGCAAGTGATCAACCTTC, and for XbaI+EcoRV+RB 2826R, use CCCTCTAGATATCTATTTGGACTCTCCTGGGAGATGTTTACTTCC.
- Clone the fragment generated (1546 bp) between the EcoR1 and XbaI restriction sites of the pCS2+CB-Myc6 vector as previously described1,12.
NOTE: The 1012-bp-long CB-Myc6 construct (337 amino acids) fused to the Rb1 fragment resulted in a protein of approximately 865 amino acids (about 108 kDa), which is slightly smaller than the endogenous RB1 protein (~110 kDa) (Figure 1A), with a CB-Myc6 tag at the N-terminus and Rb1 at the C-terminus.
- Perform the second stage subcloning into the pTet-Splice vector.
- To amplify the CB-RB-Myc6 fusion fragment and to gain the Tet-promoter, amplify the cassette, consisting of the CB-myc6-Rb1 using primers containing the EcoRV (XbaI+EcoRV+RB 2826R) and SalI sites: for SalI+CBF, use CCAGTCGACAGGATGTGGTGGTCCTTGATCCTTC.
- Subclone the fragment generated (2567 bp) into the pTet-Splice vector, between the SalI and EcoRV restriction sites, in such a way that the entire TetO-DN-CB-myc6-Rb1 transgene, including the SV40 intron and the PolyA signal, can be isolated through a single digestion with NotI (Figure 1B).
NOTE: The NotI fragment was used to generate the transgenic funders.
2. In Vitro Testing of the TetO-DN-CB-myc6-Rb1 Transgene
- Dox-regulation: NIH3T3 cell line
NOTE: Unless otherwise stated, all volumes have been adjusted for a 6-well plate.- Grow NIH3T3 cells following the vendor's specifications, using the recommended cell culture media containing 10% fetal bovine serum at 37 °C with 10% CO2.
- Cotransfect the cells with pTet-Splice and pCMV-Tet3G vectors (from step 1) for 24 h. Follow the steps mentioned below for cotransfection.
- Mix 2–4 µL of the lipid-based transfection reagent/well and 1 mL of incomplete (without serum) Dulbecco's Modified Eagle Medium (DMEM) for 5 min at room temperature. For a 12- or 24-well plate, use 250 or 500 µL of culture media, respectively, and adjust the transfection reagent volume accordingly.
- Add 2–3 µg of plasmid DNA/well to the transfection reagent-DMEM and incubate for 20 min at room temperature.
- Add the mix containing DNA and the transfection reagent in DMEM to each well. After 3–4 h of incubation, add 1 mL of complete media (so that the final volume is 2 mL/well). Keep aliquots of Dox stock (1 mg/mL) and add 2 µL in each of the wells (+ Dox). Incubate the cells at 37 °C with 5% CO2.
NOTE: Control samples should consist of cotransfected cells not treated with Dox (- Dox), as well as NIH3T3 cells transfected only with the transactivator pCMV-Tet3G vector and treated with Dox for a 24 h period. For pCMV-Tet3G, concentrations of 0.1–1 µg/mL Dox should be sufficient to induce the transgene expression.
- Test the reversibility of the construct using the HEK293 cell line.
- To test the reversibility of the TetO-DN-CB-myc6-Rb1 transgene, culture HEK293 cells in Eagle's Minimum Essential Medium (EMEM) following the vendor's specifications and cotransfect with both pTet-Splice and pCMV-Tet3G vectors for 24 h, as described above (step 2.1.2).
- For the transgene inactivation, remove the Dox-containing media after 24 h, wash the cells 2x–3x with phosphate buffered saline (PBS) and incubate them in Dox-free media for an additional 24 h.
NOTE: Upon Dox-removal, the transgene inactivation and regular protein expression should be resumed within that 24 h period.
- To assess the functionality of the TetO-DN-CB-myc6-Rb1 construct in promoting unscheduled cell proliferation, perform the functional analyses using an HEI-OC1 cell line, as described below.
- Culture HEI-OC1 cells in 10% DMEM under permissive conditions (33 °C with 10% CO2). For cell proliferation studies, count the cells using a cell counter, and plate 10,000 HEI-OC1 cells on a 96-well plate in a 200 µL volume. Incubate the cells overnight.
- On the following day, perform a transient cotransfection of pTet-Splice and pCMV-Tet3G vectors using a lipid-based transfection reagent (see steps 2.1.2). In a separate well, perform pmR-ZsGreen1 transfection at2-µg using the lipid-based transfection reagent (steps 2.1.2) to calculate the transfection rate. Use non-transfected cells as controls.
- Detect the presence of green fluorescence under a fluorescence microscope (excitation = 485 nm, emission = 530 nm) for the transfected cells 24 h after transfection. Record the presence or absence of green fluorescence and calculate the rate of transfection as the total number of GFP-positive cells (transfected cells) over the cells labeled with DAPI (total cells).
NOTE: We estimated a transfection rate of ~70%. - To induce transgene expression, add 1 µg/mL Dox to a subset of transfected cells while using the other subset as control (-Dox). Use untreated HEI-OC1 cells as additional controls.
- To assess cell proliferation, use a cell proliferation kit according to the manufacturer's protocol.
- In brief, 48 h after transfection, remove the cell culture medium and add 100 µL of 1x dye-binding solution to each well of the microplate. Incubate for 1 h at 37 °C.
- Thereafter, measure the fluorescence intensity of each sample using a fluorescence microplate reader (excitation = 485 nm, emission = 530 nm).
- To further assess cell proliferation using immunocytochemistry, plate the HEI-OC1 cells on a cover glass in a 12-well plate and incubate overnight at 37 °C.
- On the following day, cotransfect with the pTet-Splice and pCMV-Tet3G and perform the Dox treatment (+ Dox). Use untransfected cells as controls. Process HEI-OC1 cells for Ki-67 labeling.
- Fix the cells with 4% paraformaldehyde (PFA) in PBS, pH 7.4, for 10 min at room temperature. Wash the cells 3x with ice-cold PBS and incubate the cells in 0.25% non-ionic detergent in PBS for 10 min.
- Wash the cells again, 3x in PBS for 5 min, and block using 10% serum in a humidified chamber for 1 h at room temperature.
- Incubate the cells in 500 µL of Ki-67 primary antibody (1:200 dilution) for 3 h at room temperature or overnight at 4 °C.
- Wash the cells 3x for 5 min with PBS. After that, incubate the cells with the secondary antibody for 1 h at room temperature in the dark.
- Remove the secondary antibody solution and wash the cells 3x with PBS for 5 min each in the dark.
- For phalloidin labeling, incubate the HEI-OC1 cells in 1:200 phalloidin for 30 min. To label the nuclei of cells, incubate the cells with 5 µg/mL DAPI for 10 min at room temperature.
NOTE: Ensure that the phalloidin dye conjugate is different than the secondary antibody conjugate.
3. Generation of CBRb+/ROSA-CAG-rtTA+ (CBRb) Transgenic Mice and In Vivo Testing of the DN-CBRb Approach
- Upon confirmation of the effective Dox regulation of the TetO-DN-CB-myc6-Rb1 transgene, purify the NotI fragment and microinject them into mouse zygotes, which are later transferred into pseudo-pregnant females.
NOTE: These procedures were carried out at the University of Nebraska Medical Center (UNMC) Mouse Genome Engineering Core Facility. - After delivery, genotype the pups using a primer set specific to the CB-Rb1 fusion region: for CBRb F, use 5' CTGTGGCATTGAATCAGAAATTGTGGCTGG 3′, and for CBRB R, use 5′ GTACTTCTGCTATATGTGGCCATTACAACC 3′.
NOTE: The PCR product size observed on agarose gel is 401 bp (60-bp CB region + 341-bp Rb1 region (Table 1). Ten independent founder lines were identified, based on the presence of the TetO-DN-CB-myc6-Rb1 transgene. In five of these lines, progeny inherited the transgene, which confirmed the germline transmission of the transgene. One of the transgenic lines was ultimately used to establish and maintain the line and generate experimental TetO-DN-CB-myc6-Rb1 animals for further studies. - Breed adult TetO-DN-CB-myc6-Rb1 mice to the ROSA-CAG-rtTA tetracycline inducer line (Stock #006965) (alternatively, breed to a tTA inducer line) to generate experimental DN-CBRb mice. Genotype the offspring of this cross using the following primer set: for rtTA-WT R, use GGAGCGGGAGAAATGGATATG; for rtTA Mutant R, use GCGAGGAGTTTGTCCTCAACC; and for rtTA Common, use AAAGTCGCTCTGAGTTGTTAT. The PCR product sizes are 340 bp (mutant), 340 bp and 650 bp (heterozygote), and 650 bp (wild-type).
- Generate a dose-response curve of the RB1 protein following the Dox treatment to determine the time and length of the Dox treatment necessary to elicit a transgene expression.
NOTE: In this experiment, western blot and qRT-PCR were used to assess tissue-specific transgene activation and RB1 expression. - Perform fluorescence in situ hybridization (FISH) to confirm the genomic insertion of the transgene.
NOTE: This was carried out at the Centre for Applied Genomics of the Hospital for Sick Children (Toronto, Canada). ELISA or western blotting (using protein-specific antibody) can be used to assess transgene activation, tissue-specificity, and changes in the levels of the endogenous POI. For the DN-CB-myc6-Rb1 construct, we used an anti-RB1 antibody.
Inducible and Reversible Dominant-negative (DN) Protein Inhibition
Learning Objectives
Generally, designing a DN mutation requires a considerable amount of information on the structure and function of the POI. In contrast, the DN strategy presented here is particularly useful when the structural and functional information for the POI is limited. If the POI is a multimeric protein, a fusion of one subunit to a lysosomal protease can dominantly inhibit the assembled multimer and, potentially, other ligands through a combination of proteolysis of the endogenous subunits and subcellular diversion of the multimer to the lysosome. To confirm successful cloning and test the effectiveness of Dox-regulated transgene activation, purified pTet-Splice plasmid containing the TetO-DN-CB-myc6-Rb1 construct was transfected into mouse NIH3T3 cells that stably expressed the rtTA protein, but not Rb113. In the absence of Dox, cells expressing rtTA displayed no RB1 expression, whereas a robust RB1 expression was observed in the presence of Dox (Figure 2A). Next, to test the reversibility of the system, we cotransfected HEK293 cells (which endogenously express Rb114) with purified pTet-Splice/TetO-DN-CB-myc6-Rb1 and the pCMV-Tet3G vectors. As expected, the expression of the endogenous RB1 protein was significantly inhibited on transgene activation by the addition of Dox in the cell culture media (Figure 2B). To test the reversibility of the system, following a 24 h period, in a subset of HEK293 cells, we replaced the Dox-containing cell culture media with fresh Dox-free media and incubated the cells for an additional 24 h. Supporting the Dox-regulated reversibility, RB1 expression was restored upon Dox removal from the culture media (Figure 2B).Transfected HEK293 cells that were not treated with Dox or Dox-treated HEK293 cells transfected with the pCMV-Tet3G vector only (Figure 2B) showed basal levels of endogenous RB1 expression. Because NIH3T3 cells do not express RB1 protein naturally, the positive RB1 reactivity is due to the accumulation of the transgenic RB1 protein. Given our interest in the potential proliferative effect of the TetO-DN-CB-myc6-Rb1 construct in the auditory system, we measured the total DNA content in pTet-Splice/TetO-DN-CB-myc6-Rb1– and pCMV-Tet3G-transfected HEI-OC1 cells (inner ear organ of a Corti-derived cell line) in the presence and absence of Dox. Consistent with the working hypothesis presented here, a significant increase in the DNA content (corresponding to an increase in cell number) was observed in Dox-treated but not untreated (control) transfected HEI-OC1 cells (Figure 3A – 3C).
Following the in vitro confirmation of the transgene activity and function, a transgenic mouse model was generated. FISH, using a TetO-DN-CB-myc6-Rb1 digoxigenin (DIG)-horse radish peroxidase (HRP)/tyramine-biotin/avidin-Cy5-labeled probe, and G-banding of male mice splenocytes and ear fibroblasts were performed to confirm the transgene insertion in the transgenic mice genome. Among the five germline transmitting founders, line 14 showed a consistent transgene insertion within segment 10C3~D2 (Figure 4A – 4C). Mice from this line were used to establish the breeding colonies and generate experimental animals for further studies. To determine the optimum time for the Dox induction and TetO-DN-CB-myc6-Rb1 transgene activation, we divided the transgenic mice into three different groups, wherein Dox was administered in drinking water for variable lengths of time: Group 1 (3 days of treatment), Group 2 (7 days of treatment), and Group 3 (10 days of treatment). Upon completion of the treatment, changes in the RB1 protein expression were assessed by western blotting (Figure 5A – 5D). Because the DN and native RB1 proteins have similar molecular weights, they cannot be resolved from each other on a western blot. However, consistent with an initial increase in RB1 protein (due to the accumulation of DN-RB1 and endogenous proteins), there was an initial increase in total RB1 protein expression in the transgenic mouse cochleae compared to the control group (Figure 5A). Moreover, consistent with the inhibitory and proteolytic activity of the TetO-DN-CB-myc6-Rb1 transgene product, a quantifiable reduction in RB1 expression was observed in groups 2 and 3 compared to group 1 and control groups (Figure 5A – 5D). Of note, treatment longer than 10 days did not result in any further change in RB1 protein expression. Thus, 10 days of Dox treatment was considered optimal and used for further studies.
Because the rtTA and, consequently, TetO-DN-CB-myc6-Rb1 transgene activation are under the ubiquitous ROSA-CAG promoter, RB1 could be potentially inhibited in any tissue or cell where RB1is endogenously expressed (e.g., lungs, heart, retina). To test this premise and evaluate the transgene's tissue specificity, cochleae, lungs, heart, and eye biopsies were dissected from TetO-DN-CB-myc6-Rb1/ROSA-CAG-rtTA, and control mice (not carrying the ROSA-CAG-rtTA transgene) were assessed for RB1 expression (Figure 6A – 6C). Regardless of the tissue analyzed, a significant reduction in RB1 protein was observed in TetO-DN-CB-myc6-Rb1/ROSA-CAG-rtTA but not in the control group (Figure 6A). In sharp contrast, qRT-PCR analyses in the eye, heart, and cochleae of Dox-treated TetO-DN-CB-myc6-Rb1/ROSA-CAG-rtTA and age-matched control mice showed a significant upregulation of the DN-CBRb transcript in the tissues of the former but not the latter mice (Figure 6C). Altogether, these results confirm the efficiency of the construct. An increase in transcript expression reflects the effective Dox-regulated transgene induction. Likewise, a reduction in protein expression is consistent with the routing and proteolytic degradation of the endogenous RB1.
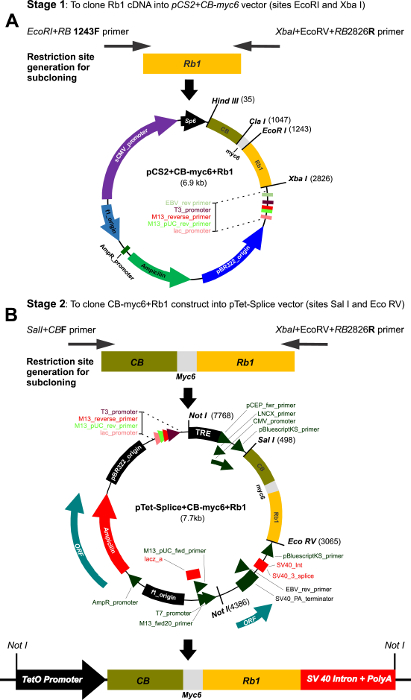
Figure 1: Multi-step cloning of the CBRb and generation of the inducible TetO-DN-CB-myc6-Rb1/ROSA-CAG-rtTA. (A) This panel shows the cloning of the Rb1 fragment into the pCS2-CB-Myc6 vector. A 1583-bp Rb1 gene product was PCR amplified using the EcoRI+RB1243 forward and Xba+EcoRV+RB2826 reverse primers. The resulting Rb1 fragment was cloned between the EcoRI and Xbal restriction sites of the pCS2 vector containing the CB-Myc6 construct to generate the pCS2+CB-myc6+Rb1 vector. (B) The CB-myc6+Rb1 fragment was cloned into the pTetSplice vector between the SalI and EcoRV restriction sites. Please click here to view a larger version of this figure.
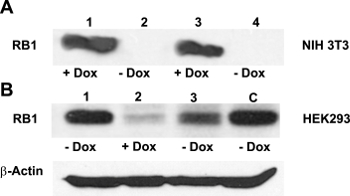
Figure 2: Effectiveness of the Dox-regulated transgene activation. (A) NIH3T3 cells do not usually express RB113. Confirming the efficient activation of the TetO-DN-CB-myc6-Rb1 construct, a western blot analysis with the anti-RB1 antibody revealed a robust RB1 expression in the presence of Dox (lanes 1 and 3), but not in its absence (lanes 2 and 4). (B) HEK293 cells, which endogenously express Rb114, were cotransfected with TetO-DN-CB-myc6-Rb1 and the pCMV-Tet3G transactivator vector. In the absence of Dox (lane 1), a positive RB1 expression was observed. The addition of Dox to the system caused the TetO-DN-CB-myc6-Rb1 activation and RB1 downregulation (lane 2). The RB1 expression was resumed 24 h after Dox was removed from the media (lane 3). C = control, untransfected HEK293 cells not treated with Dox. This is an adaptation from a figure originally published in the journal Frontiers in Cellular Neuroscience2. Its reproduction agrees with Frontiers' policy on authors' copyright retention. Please click here to view a larger version of this figure.
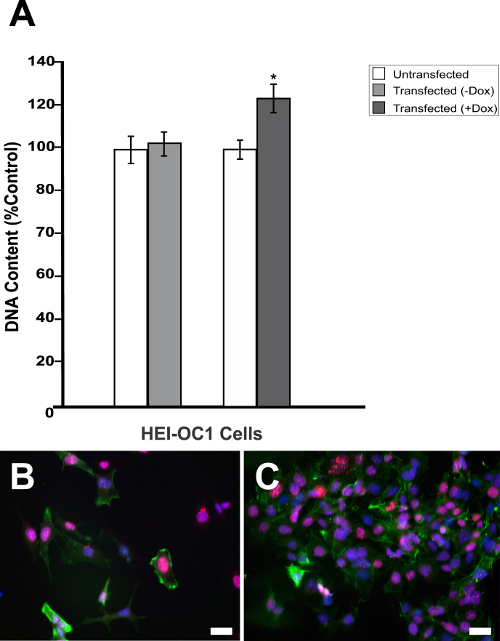
Figure 3: Dox-mediated DN-CBRb transgene activation increases cell proliferation. (A) HEI-OC1 cells cotransfected with purified TetO-DN-CB-myc6-Rb1, and the pCMV-Tet3G vectors were cultured in the absence (−) or presence (+) of Dox. Cell proliferation was determined 48 h after the transfection using a proliferation assay. The percentage change in proliferation was calculated using a change in fluorescence values with untransfected cells as a control. A modest but significant increase in cell number was observed in the transfected cells, following Dox treatment. In contrast, no significant changes were observed between transfected HEI-OC1 cells not treated with (−) Dox and the untransfected control. (B) HEI-OC1 cells untransfected or (C) cotransfected with purified TetO-DN-CB-myc6-Rb1 and the pCMV-Tet3G vectors cultured in the presence (+) of Dox were labeled with Ki-67 (red), Phalloidin (green), and DAPI (blue). * P < 0.05. Scale bar = 10 µm. This is an adaptation from a figure originally published in the journal Frontiers in Cellular Neuroscience2. Its reproduction agrees with Frontiers' policy on authors' copyright retention. Please click here to view a larger version of this figure.
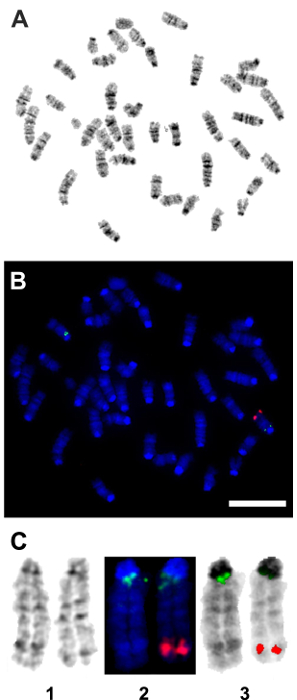
Figure 4: Fluorescent in situ hybridization (FISH) confirmation of the genomic insertion of the TetO-DN-CB-myc6-Rb1 transgene. A pCS2-CBRb (digoxigenin probe/anti-DIG-HRP/tyramide-biotin/avidin-Cy5, in red) test probe was hybridized to splenocytes and ear fibroblast cell metaphase to see if the probe signal could be detected or not with a one-copy target insert size of 2.5 kb. The pCS2-CBRb test probe hybridized to one chromosome 10 with bright, doublet probe signals that showed the insertion of the TetO-DN-CB-myc6-Rb1 transgene within the segment 10C3-D2. The RP23-267G24 (Spectrum Green) control probe hybridized to chromosome 10 at band 10A1 as expected and confirmed that the CBRb DNA of interest inserted into chromosome 10. (A) This panel shows the G-band metaphase. (B) This panel shows the tyramide signal amplification (TSA) FISH image from the same metaphase as shown in panel A. (C) This panel shows composite images of chromosome 10 showing (1) G-banding, (2) TSA FISH, and (3) TSA FISH with inverted DAPI banding. Red = pCS2-CBRb probe; blue = DAPI banding; green = RP23-267G24 control probe. Scale bar = 10 µm. Please click here to view a larger version of this figure.
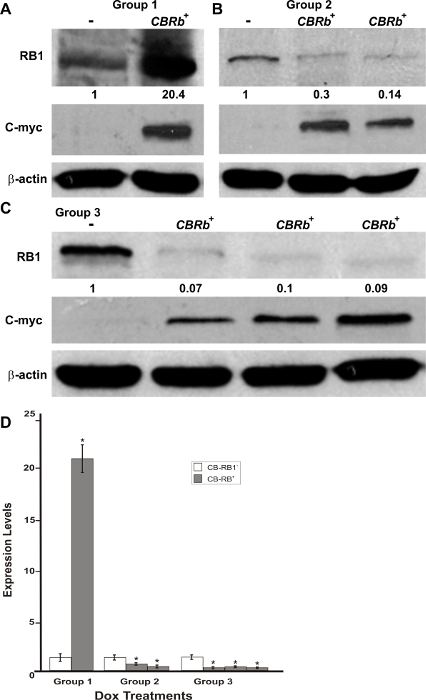
Figure 5: DN-CBRb transgene activation in the inner ear of postnatal (P) 36 TetO-DN-CB-myc6-Rb1/ROSA-CAG-rtTA+ (CBRb+) and ROSA-CAG-rtTA+ (CBRb−) mice after 3 (group 1), 7 (group 2), and 10 (group 3) days of Dox treatment. (A–C) Anti-RB1, which reacts with both hyperphosphorylated and hypophosphorylated forms of RB1, as well as anti-c-myc antibodies were used for detection. Densitometric analysis was done using ImageJ software. The corresponding values obtained were normalized to the negative control CBRb− (-) and then to β-actin. The relative RB1 expression levels are shown underneath each blot. (D) The graphic representation of normalized RB1 expression levels plotted against the different treatments highlights a consistently lower RB1 detection in CBRb+ (+) samples. Each lane on the western blot gel and each column on the graphic correspond to a different individual. * P < 0.05.This figure was originally published in the journal Frontiers in Cellular Neuroscience2. Its reproduction agrees with Frontiers' policy on authors' copyright retention. Please click here to view a larger version of this figure.
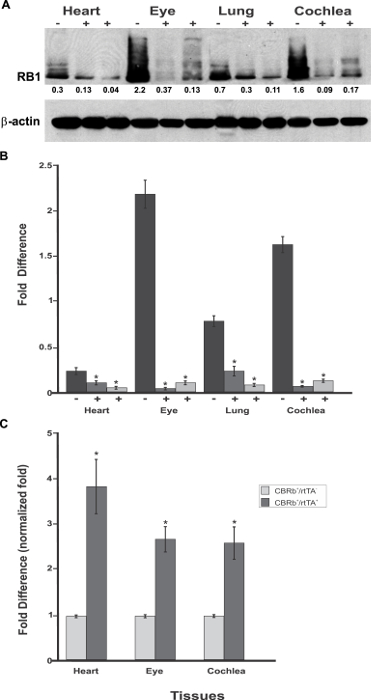
Figure 6: Spatial analyses of the TetO-DN-CB-myc6-Rb1/ROSA-CAG-rtTA transgene activation in P18 TetO-DN-CB-myc6-Rb1/ROSA-CAG-rtTA+ (+) and DN-CBRb+/ROSA-CAG-rtTA− negative control (-) mice heart, eye, lungs, and cochlea. (A)Confirming the efficiency of Dox-mediated transgene activation, the expression of the endogenous RB1 was downregulated in the CBRb+/ROSA-CAG-rtTA+ tissues but not in the negative control mice tissues. Densitometric analysis was performed as described in Figure 5. The relative RB1 expression levels are shown underneath each blot. (B) This panel shows a graphic representation of normalized RB1 expression levels plotted against the different tissues. An interindividual variation on endogenous RB1 levels may explain differences in individual responses to transgene activation and RB1 downregulation. (C) RT-PCR analyses in eye, heart, and cochleae revealed a significant upregulation of the TetO-DN-CB-myc6-Rb1/ROSA-CAG-rtTA transcript in Dox-treated TetO-DN-CB-myc6-Rb1/ROSA-CAG-rtTA+ tissues, but not in the DN-CBRb+/ROSA-CAG-rtTA− negative control group. CBRb+/rtTA+ = TetO-DN-CB-myc6-Rb1/ROSA-CAG-rtTA+; CBRb−/rtTA+ = ROSA-CAG-rtTA+; * P < 0.05. This figure was originally published in the journal Frontiers in Cellular Neuroscience2. Its reproduction agrees with Frontiers' policy on authors' copyright retention. Please click here to view a larger version of this figure.
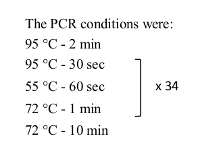

Table 1: PCR condition used for checking the CB-Rb1 fusion region.
List of Materials
| Dulbecco's Modified Eagle's Medium, DMEM | GIBCO-BRL | 11965-084 | |
| Minimum Essential Medium Eagle, MEME | Sigma | M8042 | |
| Fetal bovine serum | Sigma | F2442 | |
| Lipofectamine | DharmaFECT | T-2010-03 | |
| Sal I | Roche | 11745622 | |
| EcoRV-HF | New England BioLabs | R3195S | |
| NotI | Roche | 13090730 | |
| CaspaTag | Millipore | APT523 | |
| DAPI | Sigma | D9542 | |
| Staurosporine | Sigma | S4400 | |
| CyQuant NF cell proliferation kit | Invitrogen | C35007 | |
| Retinoblastoma 1 antibody | Abcam | Ab6075 | |
| c-Myc antibody | Sigma | M5546 | |
| b-actin | Sigma | A5316 | |
| Ki-67 | Thermofisher scientific | MA5-14520 | |
| Phallodin | Thermofisher scientific | A12379 | |
| Fluorescence microplate reader | FLUOstar OPTIMA, BMG Labtech | ||
| Epifluorescence microscope | NikonEclipse80i | ||
| The TetO-DN-CB-myc6-Rb1 (DN-CBRb) mouse line is available from the Jackson Laboratory as JAX#032011. |
Lab Prep
Dominant-negative (DN) protein inhibition is a powerful method to manipulate protein function and offers several advantages over other genome-based approaches. For example, although chimeric and Cre-LoxP targeting strategies have been widely used, the intrinsic limitations of these strategies (i.e., leaky promoter activity, mosaic Cre expression, etc.) have significantly restricted their application. Moreover, a complete deletion of many endogenous genes is embryonically lethal, making it impossible to study gene function in postnatal life. To address these challenges, we have made significant changes to an early genetic engineering protocol and combined a short (transgenic) version of the Rb1 gene with a lysosomal protease procathepsin B (CB), to generate a DN mouse model of Rb1 (CBRb). Due to the presence of a lysosomal protease, the entire CB-RB1 fusion protein and its interacting complex are routed for proteasome-mediated degradation. Moreover, the presence of a tetracycline inducer (rtTA) element in the transgenic construct enables an inducible and reversible regulation of the RB1 protein. The presence of a ubiquitous ROSA-CAG promoter in the CBRb mouse model makes it a useful tool to carry out transient and reversible Rb1 gene ablation and provide researchers a resource for understanding its activity in virtually any cell type where RB1 is expressed.
Dominant-negative (DN) protein inhibition is a powerful method to manipulate protein function and offers several advantages over other genome-based approaches. For example, although chimeric and Cre-LoxP targeting strategies have been widely used, the intrinsic limitations of these strategies (i.e., leaky promoter activity, mosaic Cre expression, etc.) have significantly restricted their application. Moreover, a complete deletion of many endogenous genes is embryonically lethal, making it impossible to study gene function in postnatal life. To address these challenges, we have made significant changes to an early genetic engineering protocol and combined a short (transgenic) version of the Rb1 gene with a lysosomal protease procathepsin B (CB), to generate a DN mouse model of Rb1 (CBRb). Due to the presence of a lysosomal protease, the entire CB-RB1 fusion protein and its interacting complex are routed for proteasome-mediated degradation. Moreover, the presence of a tetracycline inducer (rtTA) element in the transgenic construct enables an inducible and reversible regulation of the RB1 protein. The presence of a ubiquitous ROSA-CAG promoter in the CBRb mouse model makes it a useful tool to carry out transient and reversible Rb1 gene ablation and provide researchers a resource for understanding its activity in virtually any cell type where RB1 is expressed.
Procedure
Dominant-negative (DN) protein inhibition is a powerful method to manipulate protein function and offers several advantages over other genome-based approaches. For example, although chimeric and Cre-LoxP targeting strategies have been widely used, the intrinsic limitations of these strategies (i.e., leaky promoter activity, mosaic Cre expression, etc.) have significantly restricted their application. Moreover, a complete deletion of many endogenous genes is embryonically lethal, making it impossible to study gene function in postnatal life. To address these challenges, we have made significant changes to an early genetic engineering protocol and combined a short (transgenic) version of the Rb1 gene with a lysosomal protease procathepsin B (CB), to generate a DN mouse model of Rb1 (CBRb). Due to the presence of a lysosomal protease, the entire CB-RB1 fusion protein and its interacting complex are routed for proteasome-mediated degradation. Moreover, the presence of a tetracycline inducer (rtTA) element in the transgenic construct enables an inducible and reversible regulation of the RB1 protein. The presence of a ubiquitous ROSA-CAG promoter in the CBRb mouse model makes it a useful tool to carry out transient and reversible Rb1 gene ablation and provide researchers a resource for understanding its activity in virtually any cell type where RB1 is expressed.
