Inducible, Cell Type-Specific Expression in Arabidopsis thaliana Through LhGR-Mediated Trans-Activation
Instructor Prep
concepts
Student Protocol
Vectors and modules can be obtained from the non-profit repository, Addgene (https://www.addgene.org).
1. Cloning using GreenGate
- Design primers using the overhangs listed in Table 1, replacing ‘NNNN’ with module type-specific adapter sequences listed below: Forward: 5´AACA GGTCTC A NNNN (n) CA* + specific sequence 3´, Reverse: 5´AACA GGTCTC A NNNN (n) + reverse complement of specific sequence 3´. Add the underlined bases to ensure maintenance of the reading frame.
Module overhangs:
NOTE: In the default GreenGate framework, six modules, A-F, are assembled with a plant transformation vector backbone into one expression plasmid (Figure 1C). Typically, the A module will harbor promoter sequences, a B-module an N-terminal tag or “dummy” sequence6, a C-module a CDS, a D-module a C-terminal tag or dummy, an E-module a terminator, and an F-module a resistance cassette for selection of transgenic plants. Adapter for coupling with other preassembled units are available for use instead of the F module2. - PCR amplification
- Amplify the sequence of interest with the designed primers by polymerase chain reaction (PCR) according to standard protocols.
- Separate the PCR product on an agarose gel. Excise and column purify the correct fragment using a commercial kit (see Table of Materials) as per the manufacturer’s instructions.
- Entry module creation
- Separately digest the module vector and the PCR fragment (step 1.2.) (Figure 1 A, B) with Eco31I/BsaI using 100-500 ng of DNA (Vector or Fragment), 3 µL 10x digestion buffer, and 5-10 U Eco31I in a tube and bring the volume to 30 µL with ddH2O.
- Mix gently by pipetting up and down and spin down briefly at 1,000 x g. Incubate at 37 °C on a heat block for 15 min (or following the time recommendation of the purchased restriction enzyme).
- After the digestion, column purify each sample using a commercial kit (see Table of Materials) and quantify the obtained DNA by determining the optical density at 260 nm (OD260) using a spectrophotometer.
- Mix the 30 – 100 ng digested insert and the 10 – 50 ng digested vector by pipetting up and down several times and incubate with 1 µL T4 Ligase (5 U/µL, and 3 µL 10 x T4 ligation buffer in a total volume of 30 µL at room temperature for 1 h (following the recommendation of the T4 ligase supplier) (see Table of Materials).
NOTE: Calculate the desired molar ratio entry module:destination vector (e.g., 3:1) for the ligation depending on the concentration and length of the entry module inserts. - To increase the efficiency of transformation, heat inactivate the T4 ligase by Incubating the reaction at 65 °C for 20 min.
- Use the ligation product to transform competent E. coli cells according to the standard lab protocols and spread the bacteria on agar plates supplemented with ampicillin (100 µg/mL)
- Incubate the plate with bacteria at 37 °C for overnight.
- Screen for colonies with the desired entry module by colony PCR using the primers 86A1 (5′-GTTGTGTGGAATTGTGAGC-3′) and 86A2 (5′-GTTTTCCCAGTCACGACG-3′) and standard PCR conditions.
NOTE: This primer combination is valid for all entry modules as the primers bind to the entry vector backbone. - Select single colonies for the plasmid isolation. Use 2 mL LB liquid culture supplemented with ampicillin (100 µg/mL) and incubate overnight in a 37 °C shaker.
- Isolate plasmids from the overnight liquid culture with a Mini-Prep Kit or the desired extraction protocol (see Table of Materials).
- To identify plasmids with the correct insert, perform restriction enzyme analysis, for example by selecting two enzymes that cut uniquely in your insert by mixing 200 ng of plasmid, 2 µL 10 x digestion buffer, 5-10 U of selected restriction enzymes and ddH2O to 20 µL.
- Sequence the selected plasmids using the primers 86A1 and 86A2 (see step 1.3.9.).
- Destination module creation:
NOTE: For the destination plasmid use the desired destination module pGGZ001, pGGZ002 or pGGZ0032 (Figure 1C).- In a tube add and mix 50 – 150 ng empty destination vector (pGGZ003, for example), 50-300 ng of each filled entry module, 2 µL 10 x digestion buffer buffer, 1.5 µL 10 mM ATP, 1 µL T4 Ligase (30 U/µL), 5-10 U µL Eco31I and dH2O to a total volume of 20 µl.
NOTE: Calculate the desired molar ratio entry module:destination vector (e.g. 3:1) for the ligation depending on the concentration and length of the entry module inserts. - Mix and perform the GreenGate reaction2 using a PCR thermocycler alternating 30 times between 37 °C for 5 min and 16 °C for 2 min, followed by single steps of 5 min at 50 °C and 5 min at 80 °C.
- To increase the efficiency, add 1 µL T4 Ligase (30 U/µL) and 1.5 µL 10 mM ATP to the reaction and incubate for 1 h at room temperature, followed by heat-inactivation of T4 DNA Ligase at 65 °C for 20 min.
- Use the ligation reaction to transform competent E. coli and spread the bacteria on agar plates supplemented with the appropriate selective agent spectinomycin (50 µg/mL).
- Incubate the transformed E. coli on the plate at 37 °C for overnight.
- To confirm transformation, re-streak each of the selected single colonies on spectinomycin (50 µg/mL) and ampicillin (100 µg/mL) containing plates, respectively. Use only colonies that grow on spectinomycin but not on ampicillin plates.
- Incubate the E. coli colonies overnight at 37 °C.
- Select single colonies for plasmid isolation. Use 2 mL LB liquid culture supplemented with spectinomycin (50 µg/mL) and incubate overnight at a 37 °C shaker.
- Isolate the corresponding plasmids with a Mini-Prep Kit (see Table of Materials).
- Check by restriction enzyme analysis (see 1.3.12.) and sequence selected positive constructs using primers 88C3 (5′-ACCTCTCGGGCTTCTGG-3′), 88C4 (5′-CCTTTTTACGGTTCCTG-3′). If the insert cannot be fully sequenced with these primers, design internal primers for sequencing.
NOTE: To identify clones with the correct number of pOp repeat sequences (see Discussion), the primers pOp6 (pOp6_F, 5′-TGCATATGTCGAGCTCAAGAA-3′; and pOp6_R, 5′-CTTATATAGAGGAAGGGTCTT-3′) that bind in the short flanking sequences can be used for PCR amplification and subsequent gel electrophoresis to discriminate by size.
- In a tube add and mix 50 – 150 ng empty destination vector (pGGZ003, for example), 50-300 ng of each filled entry module, 2 µL 10 x digestion buffer buffer, 1.5 µL 10 mM ATP, 1 µL T4 Ligase (30 U/µL), 5-10 U µL Eco31I and dH2O to a total volume of 20 µl.
- Intermediate supermodule creation
- To combine two independent sets of entry modules, as required for generating the GR-LhG4 driver lines with integrated reporter-effector cassette, first build two intermediate plasmids, called supermodules2, before assembling the final expression plasmid in pGGZ001 or pGGZ003.
- Be sure to add F-H adaptor at the end of the first supermodule and H-A adaptor at the beginning of the second supermodule (those serve as connections between two constructs) as described2.
NOTE: Only the pGGN000 intermediate module carries the resistance cassette. To generate the expression plasmid, perform the GreenGate reaction with the destination vector and the pGGN000 and pGGM000 intermediate supermodules. Alternatively, mix destination vector, pGGN000 intermediate supermodule, and the remaining single modules to perform the GreenGate reaction2. The latter method is less efficient than the former but can be faster. - To create a pGGM000/pGGN000 supermodule, mix 1.5 µL (100-300 ng/µL) of each of the entry modules with 1 µL (30 ng/µL) empty intermediate vector (pGGM000 or pGGN000), 2 µL 10 x digestion buffer, 1.5 µL 10 mM ATP, 1 µL T4 Ligase (30 U/µL), and 5-10 U Eco31I in a total volume of 20 µL.
NOTE: Calculate the desired molar ratio entry module:destination vector (e.g., 3:1) for the ligation depending on the concentration and length of the entry module inserts - Mix and perform the GreenGate reaction as in step 1.4.2
- To increase efficiency, add 1 µL T4 Ligase and 1.5 µL ATP (10 mM), and incubate for 1 h at room temperature, followed by heat-inactivation at 65 °C for 20 min.
- Use 10 µL of the ligation reaction to transform competent E. coli and streak on plates containing kanamycin (50 µg/mL).
- Perform overnight incubation of the transformed E. coli on the plate at 37 °C.
- Re-streak each of the selected single colonies on kanamycin (50 µg/mL) and ampicillin (100 µg/mL) containing plates, respectively.
- Perform overnight incubation of the E. coli colonies on the plate at 37 °C.
- Select single colonies that grow only on kanamycin but not on ampicillin for plasmid isolation. Use 2 mL LB liquid culture supplemented with kanamycin (50 µg/mL) and incubate overnight in a 37 °C shaker.
- Isolate the corresponding plasmids using a mini-prep kit (see Table of Materials).
- Check the plasmids by restriction enzyme analysis (see 1.3.12.)and confirm by sequencing using primers 87E2 (5´-AGGCATCAAACTAAGCAGAAG-3´) and 87E3 (5´-CGTTTCCCGTTGAATATGGC-3´) annealing to the pGGM000/pGGN000 backbone. If the insert cannot be fully sequenced with these primers, design internal primers for sequencing.
NOTE: If cloning into the pGGM000 or pGGN000 vectors is unsuccessful, one possible solution is to digest the pGGM000 or pGGN000 with Eco31I, run on a gel and purify from the gel the pGGM000 or pGGN000 backbone (approximately 2000 bp) fragment without the ccdB cassette (approximately 1400 bp). Then proceed normally with the ligation reaction.
- Transform an A. tumefaciens pSOUP+ strain (e.g. ASE), as the pSa origin of replication (ori A. tum.) in the destination vectors require the presence of the helper plasmid pSOUP12. Streak out bacteria on LB plates containing chloramphenicol (34 µg/mL), kanamycin (50 µg/mL), spectinomycin (50 µg/mL) and tetracycline (12.5 µg/mL). Incubate the transformed A. tumefaciens on the plate in a 28 °C incubator for two to three days.
2. Generation of Arabidopsis transgenic plants
- Transformation of Arabidopsis thaliana.
NOTE: Transform A. thaliana plants according to Zhang et al., 200613. - Selection of transgenic lines.
- To select the transformed plants, use the selection scheme used at GABI-Kat14 for resistance to sulfadiazine15.
- To select stable lines with a possible single integration event, choose those in generation T2, that show the 3:1 segregation ratio in the resistance marker. Propagate these lines to T3 generation and select the plants homozygous for the resistance gene. Alternatively, perform a Southern Blot analysis or a standard quantitative real-time PCR (SA-qPCR)16 to select single insertion lines.
3. Induction of trans-activation in Arabidopsis driver lines
- Root
- To test reporter expression in A. thaliana driver lines generated through steps 1 and 2, sterilize seeds as described below.
- Add 0.5-1.0 mL of 70 % Ethanol + 0.01 % non-ionic detergent to approximately 100 seeds (20 mg) in a 1.5 ml reaction tube and invert the tube a few times. Spin down at 1000 x g for 15 s and aspirate the supernatant.
- Add 0.5-1.0 mL of absolute ethanol. Invert the tube several times, spin down for 1,000 x g for 15 s and discard the supernatant.
- Add 0.5-1.0 mL of absolute ethanol and invert the tube t a few times, then spin down at 1,000 x g for 15 s and discard the supernatant.
- Allow seeds to dry inside the hood. If seeds are going to be stratified in tubes continue with step 3.1.1.6.
- Add 0.5-1.0 mL of sterile water and invert the tube a few times. Spin down at 1,000 x g for 15 s and discard the supernatant.
- Add 0.5-1.0 mL of sterile water and invert the tube several times. Spin down at 1,000 x g for 15 s and discard the supernatant.
- Add 0.5-1.0 mL of sterile water.
- Prepare half-strength Murashige and Skoog medium, pH 5.8 and add 0.9% plant agar and 1% sucrose. After autoclaving add Dexamethasone (Dex), dissolved in DMSO, to a final concentration of 10 – 30 µM to induction plates and an equal amount of DMSO to control plates, respectively.
- Put seeds for root imaging on plates and stratify them for 48 h in darkness and cold (4 °C).
- Put plates in a vertical position in a plant incubator (long day (16/8 h), 22 °C, humidity, 65 %) and grow them for five days.
- Five days after germination (dag), image the seedlings using confocal laser scanning microscopy.
- To test reporter expression in A. thaliana driver lines generated through steps 1 and 2, sterilize seeds as described below.
- Stem
- Sterilize seeds as described in 2.1.1. and put seeds on ½ MS, 0.9% plant agar and 1 % sucrose plates. Stratify for 48 h in darkness and cold (4 °C).
- Six to seven days after germination, transfer the seedlings to soil, each plant in a single pot (long day (16/8 h), 22 °C, humidity, 65%).
- Induce trans-activation in stems by watering with Dex or mock solution, or by dipping the plants in Dex or mock solutions, respectively.
- For watering, use a 25 µM Dex solution in water prepared from a stock of 25 mM Dex dissolved in ethanol. Water every 2-3 days until the desired time of induction.
- For dipping, prepare a 1 L beaker, 750 mL of water containing 0.02% silwet L-77 with 25 µM of Dex or equivalent amount of DMSO for the Dex and mock treatment, respectively. Dip single plants for 30 s in the induction or in the mock solution. Repeat every 2-3 days until the desired time of imaging.
NOTE: Following the dipping, the plants should be maintained in high humidity for one hour.
- Shoot Apical Meristem (SAM)
- Sterilize seeds as described in 2.1.1. Prepare plates with ½ MS, 0.9 % plant agar and 1 % sucrose medium and put seeds on plates. Store the plates for two days in darkness and cold for stratification.
- Put plates in a vertical position in a plant incubator. Six to seven days after germination, transfer the seedlings to the soil, each plant in a single pot (long day (16/8 h), 22 °C, humidity, 65 %).
- When the stem is around 1 cm long (25-30 dag), spray the inflorescence SAM with 10-50 µM Dex in H20 wearing a face mask. For induction and imaging at later stages of development, induce SAMs of longer stems or side shoots.
NOTE: SAMs can also be induced by paintbrushing the Dex solution to prevent spray droplets. Do use a mask when Dex is applied by spraying. - 24-48 h after induction, dissect the SAMs and proceed to imaging (see 4.3).
NOTE: Only un-induced plants were used for propagation to reduce the probability of gene silencing. Plants in T2/T3 were used for imaging.
4. Imaging of reporter expression in Arabidopsis driver lines.
- Root
- Transfer seedlings from plate to a 10 µg/mL propidium iodide (PI) solution and counter-stain them for 5 min.
- Place the roots into a microscope imaging chamber (1-well tissue culture chamber on cover glass II, see Table of Materials) and image them using a confocal laser scanning microscope with a 63x water immersion objective (see Table of Materials).
- To visualize PI fluorescence, use an excitation wavelength of 488 nm and collect emission between 590 and 660 nm. For mTurquoise2 fluorescence use 458 nm excitation and collect emission between 460 and 615 nm.
- Stem
- Perform horizontal hand cut sections of plant stems with a razor blade, excising a segment of approximately 3 cm. Fix the stem with the finger on the opposite side of the desired section to cut. Perform several fine cuts sequentially but note that only sections from a similar position in the stem should be compared.
- After each cut, immerse the razorblade in a petri dish containing tap water and collect the stem sections. Stain sections at this point or directly mount on microscope slides.
- For staining, prepare a 1 mL of 250 µg/mL of PI solution of water in a small petri dish (see Table of Materials). Immerse the sections for 5 min and rinse them in water. Transfer them to microscope slides with fine forceps or a fine paint brush. Take care to not squeeze the samples with the coverslip.
- Image samples using a confocal laser scanning microscope with a 25x dipping water immersion lens (see Table of Materials). Use 561 nm laser light to excite PI fluorescence and collect from 570 to 620 nm. Use 405 nm to excite the mTurquoise2 fluorophore effector encoded by the driver line constructs and collect emission from 425 to 475 nm.
- Shoot Apical Meristem
- Cut the stem 2 cm below the shoot tip with forceps. Hold the stem in one hand and use fine forceps to remove the flower buds and large primordia. To remove young primordia, fix the SAM in an upright position into a petri dish containing 3 % agarose. Use a binocular and forceps to remove young primordia close to the SAM. Alternatively, use an injection cannula (see Table of Materials) to cut off these primordia.
- Stain the dissected SAM in a 250 µg/mL PI solution for 5-10 min. Perform staining either directly by pipetting a few drops of staining solution on top of the prepared SAM or transfer the sample to a tube filled with staining solution. Keep the SAM fully submerged during the staining procedure.
- Place the stained SAM into a small petri dish (see Table of Materials) with 3 % agarose medium and cover the SAM with ddH2O.
- Image dissected SAMs using a confocal laser scanning microscope equipped with a 25 x dipping water immersion lens (see Table of Materials).
- Use 561 nm laser light to excite PI fluorescence and collect from 570 to 620 nm. Use 405 nm to excite the mTurquoise2 fluorophore and collect emission from 425 to 475 nm.
Record image stacks spanning 50 µm in z-direction with a step size of 0.5 µm.
NOTE: SAM imaging as described here requires an upright confocal laser scanning microscope and a lens (here, a water dipping lens) with long working distance.
Inducible, Cell Type-Specific Expression in Arabidopsis thaliana Through LhGR-Mediated Trans-Activation
Learning Objectives
Generation of driver and effector lines through GreenGate cloning
The GreenGate cloning system is based on GoldenGate cloning and use the type IIS restriction endonuclease BsaI or its isoschizomer Eco31I. As the enzyme produces overhangs distant from its asymmetric recognition site, the base composition of the overhangs can be freely chosen, which is the basis form the modularity of the system. Each PCR-generated element, for example, a promoter sequence, CDS, or terminator, is first inserted into a designated entry vector with matching overhangs produced by restriction digest to generate a module. After subcloning, a number of matching modules, usually six, are used for the GreenGate ligation reaction resulting in the assembly of the construct in a binary plant destination vector.
For driver lines, modules containing the DNA sequences of tissue-specific promoter (pTS), the GR-LHG4 transcription factor, the pOp6 promoter, and the mTurquoise2 reporter fused to an N-terminal signal peptide and a C-terminal ER retention signal were fused including terminators and various adapter modules and a module for transgenic selection as described previously 2,8 (Figure 2). Effector lines were constructed with pOp6 promoter, and an effector cassette, for example consisting of a gene of interest and terminator as well as a module encoding a resistance gene for transgenic plant selection (Figure 2).
Induction of driver lines and visualization of reporter fluorescence
Induction with Dex leads to cell type specific mTurquoise2 expression in the root endodermis (pSCR>>SP-mTurquoise2-HDEL, Figure 3A), phloem precursors and cambium (pSMXL5>>SP-mTurquoise2-HDEL 17, Figure 3B), and the stem cells in the shoot apical meristem (pCLV3>>SP-mTurquoise2-HDEL 18, Figure 3C).
Trans-activation of VND7 in starch sheath cells of the cambium
As a test case for trans-activation, we generated an effector line encoding the secondary cell wall master transcription factor VND7 fused to the VP16 activation domain, which has been shown to induce secondary cell wall formation cell autonomously when misexpressed 8,9,19,20.
After 5 days of a single treatment with either 15 µM of Dex or DMSO for the induced or the mock plants respectively, stem sections were prepared for the visualization of ectopic lignification in the starch sheath. Propidium Iodide shows in the stem strong affinity to lignified tissue. The starch sheath cells in induced samples by Dex, but not in the mock showed a strong signal for the PI channel and some cells show the typical reticulate thickening of the cell wall in xylem cells (Figure 4).
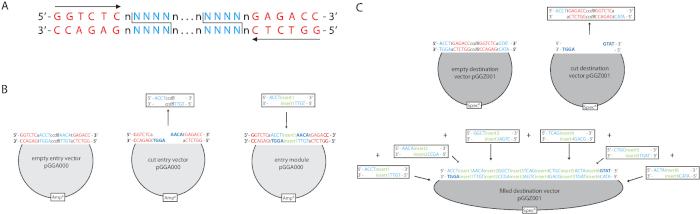
Figure 1. The GreenGate cloning principle. A) Type IIS restriction endonucleases, like BsaI/Eco31I, recognize non-palindromic sequences (red) and cut asymmetrically in a defined distance independently of the sequence (blue). Eco31I recognizes ‘GGTCTC’, cuts from the second nucleotide downstream of the recognition site and creates a four base 5’ overhang. B) The GreenGate cloning system is based on a modular system with six different entry vectors pGGA000-pGGF000. These vectors contain an Ampicillin resistance cassette (AmpR) and a ccdB cassette flanked by the specific adaptors for each entry vector (e.g pGGA000 entry vector) and the ‘GGTCTC’ Eco31I recognition sites. Eco31I digestion of pGGA000 releases ccdB and creates the pGGA000-specific four nucleotide overhangs (dark blue). Insert1 is amplified by primers harboring ‘GGTCTC’ and pGGA000 specific adaptors and after digestion ligated into pGGA000. The same procedure is followed with the other modules. C) The final GreenGate reaction combines the Eco31I digestion of the destination vector pGGZ001 and the six entry vectors pGGA000-pGGF000 and the simultaneous ligation of all modules into the destination vector. Please click here to view a larger version of this figure.
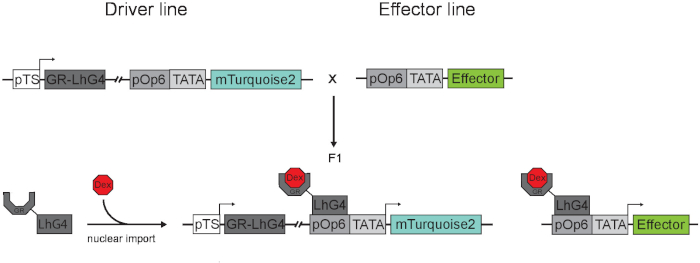
Figure 2. Overview of the Dex-inducible LhGR/pOp system with driver and effector lines. In driver lines, tissue-specific promoters (pTS) control the expression of the synthetic transcription factor LhG4, which is translationally fused to the ligand binding domain of rat glucocorticoid receptor (GR) and thereby prevents, in the absence of Dex, the nuclear translocation. The effector line harbors a transcriptional cassette driven by a pOp element and a TATA box with a minimal 35S promoter. Crossed with a driver line and upon Dex induction, GR-LhG4 binds to the pOp-type elements in the reporter cassette and to those in the effector module, inducing the transcription of mTurquoise2 and the effector. Please click here to view a larger version of this figure.
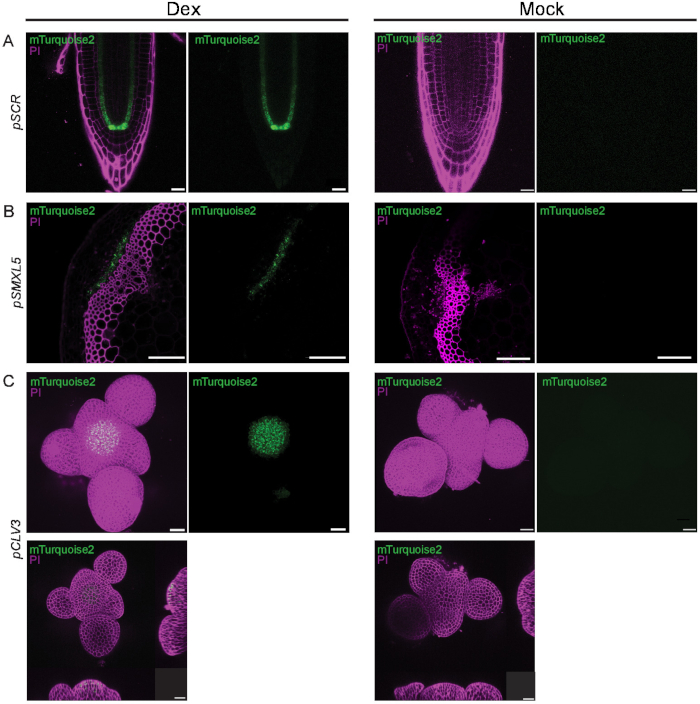
Figure 3. Induced driver lines in root, stem, and SAM. A) Dex-induced driver line pSCR>>SP-mTurquoise2-HDEL (germinated in 50 µM Dex) in the root and mock treatment. The SCARECROW promoter (pSCR) mediates expression in the endodermis, cortex/endodermis initial (CEI) and quiescent center (QC). Cells are counter-stained with propidium iodide (PI). Scale bars = 20 µm. B) Dex-induced driver line pSMXL5>>SP-mTurquoise2-HDEL (dipped in a 50 µM Dex solution and visualized after 3 days) in in the stem and mock treatment. SMXL5 promoter (pSMXL5) mediates expression in the cambium stem cell domain and phloem precursors. Cells are counter-stained with propidium iodide (PI). Scale bars = 100 µm. C) Dex-induced driver line pCLV3>>SP-mTurquoise2-HDEL (10 µM, 48 h) in the SAM. The CLAVATA3 promoter (pCLV3) mediates expression in the stem cell domain. Pictures in the bottom show XZ and YZ cross sections, Dex-induced and mock treated, respectively. Cells are counter-stained with propidium iodide (PI). Scale bars = 20 µm. Please click here to view a larger version of this figure.
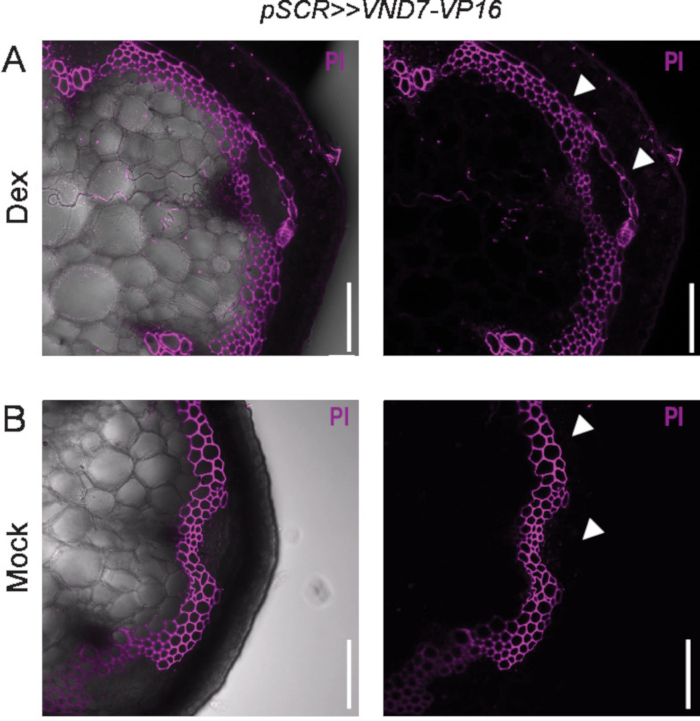
Figure 4. Ectopic lignification in the starch sheath of the stem. A) Ectopic lignification is seen five days after Dex induction of the driver line pSCR>>VND7-VP16 in the stem. The SCARECROW promoter (pSCR) mediates expression in the starch sheath cells in the stem. Left image shows merge of PI channel and bright field, right image only shows PI channel. B) The mock control shows no signal in the starch sheath cells. Cells are counter-stained with propidium iodide (PI). Scale bars = 100 µm. PI fluorescence is false-colored in green while chloroplast auto fluorescence is red in all images. White arrowheads point to starch sheath cells. Left image shows merge of PI channel and bright field, right image only shows PI channel. Please click here to view a larger version of this figure.
| Module | 5'overhang | typically used for | 3' overhang |
| A | ACCT | promoter | AACA |
| B | AACA | N-terminal tag | GGCT |
| C | GGCT | Coding sequence | TCAG |
| D | TCAG | C-terminal tag | CTGC |
| E | CTGC | terminator | ACTA |
| F | ACTA | resistance cassette | GTAT |
Table 1: Overhangs used for primer design
List of Materials
| Ampicillin | Carl Roth GmbH + Co. KG | K029.1 | |
| ATP | Sigma-Aldrich | A9187 | |
| Chloramphenicol | Sigma-Aldrich | C1919 | |
| Column purification | Qiagen | QIAquick PCR Purification Kit (250) | |
| Culture chamber for imaging | Sarstedt AG & Co. KG | 1-well tissue culture chamber, on cover glass II | |
| Dexamethasone | Sigma-Aldrich | D4903 | |
| DMSO | Fisher Scientific, UK | D139-1 | |
| Eco31I | Thermo Fisher Scientific | FD0294 | |
| injection cannula (0.30 x 12 mm, 30 G x 1/2) | Sterican, Braun | ||
| Kanamycin | Carl Roth GmbH + Co. KG | T832.2 | |
| Leica TCS SP5 CLSM, HCX PL APO lambda blue 63x water immersion objectiv | Leica, Wetzlar, Germany | ||
| MS medium | Duchefa, Haarlem, Netherlands | M0221.0050 | |
| Nikon A1 CLSM, Apo LWD 25x 1.1 NA water immersion objective | Nikon, Minato, Tokyo, Japan | ||
| Petri dish 35/10 mm | Greiner Bio-One GmbH, Germany | 627102 | |
| Petri dish 60/150 mm | Greiner Bio-One GmbH, Germany | 628102 | |
| Petri dish 120/120/17 | Greiner Bio-One GmbH, Germany | 688102 | |
| Plant agar | Duchefa, Haarlem, Netherlands | P1001 | |
| Plasmid extraction | Qiagen | QIAprep Spin Miniprep Kit | |
| Propidium iodide (PI) | Sigma-Aldrich | P4170 | |
| Razorblade | Classic, Wilkinson Sword GmbH | 7005115E | |
| Reaction tubes | Sarstedt AG & Co. KG | 72.690.001 | |
| Silwet L-77 | Kurt Obermeier GmbH & Co. KG, Bad Berleburg, Germany | ||
| Spectinomycin | AppliChem GmbH | 3834.001 | |
| Spectrophotometer | Thermo Fisher Scientific | NanoDrop 2000c | |
| Sucrose | Carl Roth GmbH + Co. KG | 4621.1 | |
| Sulfadiazine | Sigma-Aldrich | S6387 | |
| Tetracycline | AppliChem GmbH | 2228.0025 | |
| T4 Ligase 5 U/µl | Thermo Fisher Scientific | EL0011 | |
| T4 Ligase 30 U/µl | Thermo Fisher Scientific | EL0013 |
Lab Prep
Inducible, tissue-specific expression is an important and powerful tool to study the spatio-temporal dynamics of genetic perturbation. Combining the flexible and efficient GreenGate cloning system with the proven and benchmarked LhGR system (here termed GR-LhG4) for the inducible expression, we have generated a set of transgenic Arabidopsis lines that can drive the expression of an effector cassette in a range of specific cell types in the three main plant meristems. To this end, we chose the previously developed GR-LhG4 system based on a chimeric transcription factor and a cognate pOp-type promoter ensuring tight control over a wide range of expression levels. In addition, to visualize the expression domain where the synthetic transcription factor is active, an ER-localized mTurquoise2 fluorescent reporter under control of the pOp4 or pOp6 promoter is encoded in driver lines. Here, we describe the steps necessary to generate a driver or effector line and demonstrate how cell type specific expression can be induced and followed in the shoot apical meristem, the root apical meristem and the cambium of Arabidopsis. By using several or all driver lines, the context specific effect of expressing one or multiple factors (effectors) under control of the synthetic pOp promoter can be assessed rapidly, for example in F1 plants of a cross between one effector and multiple driver lines. This approach is exemplified by the ectopic expression of VND7, a NAC transcription factor capable of inducing ectopic secondary cell wall deposition in a cell autonomous manner.
Inducible, tissue-specific expression is an important and powerful tool to study the spatio-temporal dynamics of genetic perturbation. Combining the flexible and efficient GreenGate cloning system with the proven and benchmarked LhGR system (here termed GR-LhG4) for the inducible expression, we have generated a set of transgenic Arabidopsis lines that can drive the expression of an effector cassette in a range of specific cell types in the three main plant meristems. To this end, we chose the previously developed GR-LhG4 system based on a chimeric transcription factor and a cognate pOp-type promoter ensuring tight control over a wide range of expression levels. In addition, to visualize the expression domain where the synthetic transcription factor is active, an ER-localized mTurquoise2 fluorescent reporter under control of the pOp4 or pOp6 promoter is encoded in driver lines. Here, we describe the steps necessary to generate a driver or effector line and demonstrate how cell type specific expression can be induced and followed in the shoot apical meristem, the root apical meristem and the cambium of Arabidopsis. By using several or all driver lines, the context specific effect of expressing one or multiple factors (effectors) under control of the synthetic pOp promoter can be assessed rapidly, for example in F1 plants of a cross between one effector and multiple driver lines. This approach is exemplified by the ectopic expression of VND7, a NAC transcription factor capable of inducing ectopic secondary cell wall deposition in a cell autonomous manner.
Procedure
Inducible, tissue-specific expression is an important and powerful tool to study the spatio-temporal dynamics of genetic perturbation. Combining the flexible and efficient GreenGate cloning system with the proven and benchmarked LhGR system (here termed GR-LhG4) for the inducible expression, we have generated a set of transgenic Arabidopsis lines that can drive the expression of an effector cassette in a range of specific cell types in the three main plant meristems. To this end, we chose the previously developed GR-LhG4 system based on a chimeric transcription factor and a cognate pOp-type promoter ensuring tight control over a wide range of expression levels. In addition, to visualize the expression domain where the synthetic transcription factor is active, an ER-localized mTurquoise2 fluorescent reporter under control of the pOp4 or pOp6 promoter is encoded in driver lines. Here, we describe the steps necessary to generate a driver or effector line and demonstrate how cell type specific expression can be induced and followed in the shoot apical meristem, the root apical meristem and the cambium of Arabidopsis. By using several or all driver lines, the context specific effect of expressing one or multiple factors (effectors) under control of the synthetic pOp promoter can be assessed rapidly, for example in F1 plants of a cross between one effector and multiple driver lines. This approach is exemplified by the ectopic expression of VND7, a NAC transcription factor capable of inducing ectopic secondary cell wall deposition in a cell autonomous manner.
