Cell-Free Production of Proteoliposomes for Functional Analysis and Antibody Development Targeting Membrane Proteins
Instructor Prep
concepts
Student Protocol
1. Preparation of pEU expression plasmid
NOTE: pEU expression plasmid should include start codon, open reading frame of target membrane protein, and stop codon in the fragment (see Figure 1). Add detection/purification tag sequence(s) at the appropriate position when required. Either restriction enzyme digestion or seamless cloning is applicable for subcloning. Here we describe a protocol using a seamless cloning method.
- Prepare insert DNA fragment.
- Amplify the gene of interest by PCR using the cDNA template, primer 1 and primer 2. Primer 1 and primer 2 contain 15 bp overlaps for seamless cloning, respectively (see the Table of Materials).
NOTE: Do not include sequences to be processed and removed from mature protein in the cells (e.g., signal sequence). Processing of synthesized protein is not performed in wheat cell-free system. Do not add Kozak sequence. pEU-E01-MCS vector has E01 translation enhancer. - Add 1/25 volume of DpnI restriction enzyme to the PCR product to remove template plasmid DNA. Incubate for 30 min at 37 °C.
- Use a PCR purification kit to purify the PCR product and adjust concentration at 20–50 ng/µL.
- Amplify the gene of interest by PCR using the cDNA template, primer 1 and primer 2. Primer 1 and primer 2 contain 15 bp overlaps for seamless cloning, respectively (see the Table of Materials).
- Linearize pEU-E01-MCS vector.
- Conduct inverse PCR using pEU-E01-MCS, primer 3, and primer 4.
- Add 1/25 volume of DpnI restriction enzyme to the inverse PCR product. Incubate for 30 min at 37 °C.
- Use a PCR purification kit to purify the PCR product as per the manufacturer’s recommendation. Adjust the concentration at 20–50 ng/µL.
- Mix 2 µL of insert DNA fragment, 2 µL of linearized vector, and 4 µL of 2x seamless cloning enzyme mixture.
- Transform Escherichia coli strain JM109 with the seamless cloning product. Using a spreader, spread the bacterial suspension on a LB-ampicillin agar plate.
NOTE: pEU vector has an ampicillin resistance marker. - Confirm the sequence of the expression plasmid constructed using primer 5 and primer 6 from 5’ and 3’ side of MCS in pEU plasmid, respectively.
- Amplify and purify the expression plasmids.
- Culture the plasmid-transformed E. coli strain JM109 in 150 mL of LB-ampicillin medium at 37 ˚C and 125 strokes per minute shaking overnight.
- Extract and purify the plasmids using commercially available plasmid prep midi kit. Dissolve plasmids in 500 µL of TE buffer.
CAUTION: Do not use a mini prep kit for plasmid extraction. It does not provide sufficient quality and quantity of plasmid. - Add 500 µL of phenol/chloroform/isoamyl alcohol (25:24:1). Mix vigorously for 5 min, and centrifuge for 5 min at 17,800 x g and room temperature. Transfer the upper plasmid solution to a new tube.
CAUTION: Wear disposable gloves to protect the skin from phenol and chloroform.
NOTE: In order to remove contamination of RNase from plasmid extraction kit, purify the plasmids using phenol-chloroform purification. - Add 500 µL of chloroform to remove phenol completely. Mix vigorously for 5 min, and centrifuge for 5 min at 17,800 x g and room temperature. Transfer the upper plasmid solution to a new tube.
- Add 2.5 volume of ethanol and 1/8 volume of 7.5 M ammonium acetate, and store at -30 °C for 1 h.
- Centrifuge at 17,800 x g at 4 °C for 10 min. Wash the pellet with 500 µL of 70% ethanol. Remove the supernatant carefully and leave the pellet for 5 min to dry up.
- Dissolve the expression plasmids in 100 µL of ultrapure water completely. Measure the concentration of plasmids with absorbance at 260 nm. Adjust the concentration to 1 mg/mL.
2. In vitro transcription
CAUTION: Use DNase and nuclease-free plastic tubes and tips in steps of transcription and translation. Avoid autoclaving plastic wares to prevent contamination.
- Harvest ultrapure water in a new plastic tube.
CAUTION: Do not use DEPC-treated water because residual DEPC strongly inhibits the reaction. Use freshly purified ultrapure water for transcription and translation. - Prepare transcription reaction mix by mixing 115.2 µL of ultrapure water, 40 µL of Transcription Buffer LM, 20 µL of NTP mix, 2.4 µL of 80 U/µL RNase inhibitor, 2.4 µL of 80 U/µL SP6 polymerase, and 20 µL of 1 mg/mL pEU expression plasmids. Mix the reagents gently by inverting. Perform a quick spin.
- Incubate the transcription reaction at 37 °C for 6 h.
- Mix the reaction gently by inverting and quickly spin down. Use it immediately for translation, otherwise freeze and store at -80 °C.
- Confirm the transcription product by electrophoresis.
- Mix 100 mL of 1x TAE buffer and 1 g of agarose. Heat the suspension in a microwave oven to prepare 1% agarose TAE gel.
- Take 1 µL of transcription reaction and mix with 3 µL of water and 4 µL of 2x loading dye.
NOTE: Denaturing of RNA is not required. - Load 4 µL of the mixture and 2 µL of DNA ladder marker to the agarose TAE gel.
- Electrophorese at 100 V for 20 min.
- Stain the gel in ethidium bromide for 30 min. Check the ladder band pattern of mRNA using UV transilluminator and gel imager.
NOTE: When a smeared band of less than 500 bp is observed, mRNA degradation is suspected.
3. Preparation of materials for translation
- Prepare the translation buffer.
- Mix 27 mL of freshly prepared ultrapure water and 0.75 mL of each 40x stock solution for S1, S2, S3, and S4, respectively in a 50 mL tube.
NOTE: Modulate the amounts of materials according to the final required amount of 1x translation buffer.
CAUTION: Do not store or refreeze the excess 1x translation buffer after use.
- Mix 27 mL of freshly prepared ultrapure water and 0.75 mL of each 40x stock solution for S1, S2, S3, and S4, respectively in a 50 mL tube.
- Prepare creatine kinase stock solution. Dissolve lyophilized creatine kinase in ultrapure water to a final concentration of 20 mg/mL. Dispense the solution in small amounts (10 to 50 µL each) in 0.2 mL 8-strip PCR tubes. Freeze the tubes in liquid nitrogen, and store at -80 °C.
CAUTION: Do not re-freeze creatine kinase solution after thawing. - Wash dialysis cups (0.1 mL size) to remove glycerol from the dialysis membrane.
NOTE: There are several different-sized dialysis cups. Small-sized cups (0.1 mL) are used for small-scale test (section 5.4), and large-sized cups (2 mL) for large-scale production (section 5.5), respectively. The washing step of dialysis membrane of large-size cups is avoidable.- Put 1 mL of ultrapure water into a new 1.5 mL tube. Insert small-sized dialysis cup (0.1 mL) to the tube. Add 0.5 mL of ultrapure water into the cup.
- Incubate for more than 30 min at room temperature.
4. Preparation of liposomes
NOTE: Here we describe two protocols for preparation of liposomes. One uses ready-to-use lyophilized liposomes (section 4.1), while the other produces liposomes by hydrating a thin lipid film (section 4.2).
- Prepare liposomes using lyophilized liposomes.
NOTE: An easier way to produce proteoliposome is to use commercially available Asolectin liposome. Asolectin is a kind of natural lipid extracted from soybeans.- Open the vial containing 10 mg of lyophilized Asolectin liposomes (see the Table of Materials) and add 200 µL of translation buffer (section 3.1) to the bottom of the vial. Seal the vial and incubate for 10 min.
- Mix vigorously by putting the vial on the vortex-mixer for 1 min.
- Insert the vial into a 50 mL tube. Spin down the tube by centrifuging at 500 x g for 1 min.
- Using a pipette, transfer the Asolectin liposome suspension (50 mg lipid/mL) to a new 1.5 mL tube. Use liposome immediately for translation, otherwise freeze in liquid nitrogen and store at -80 °C.
- Prepare liposomes by hydrating a thin lipid film.
- If a lipid is sold in powder form, dissolve in chloroform or appropriate organic solvent to 10-100 mg/mL concentration.
NOTE: A thin lipid film can be prepared using purified and/or synthesized amphiphilic lipids. The purification method of asolectin is previously described38. Functionally modified lipids, such as biotinylated lipids, fluorescent lipids, and adjuvant lipids, can be added to the basal lipids to produce functional liposomes. - Transfer the lipid solution containing 50 mg of lipid(s) to an evaporation flask.
- Using a rotary evaporator, evaporate solvent and evenly spread the lipid on the wall of the flask bottom to form a thin film of lipid.
- Put the flask in a vacuum desiccator and leave under negative pressure overnight to remove the solvent completely.
- Add 1 mL of translation buffer to the evaporation flask. Rotate the flask to spread the buffer over the thin lipid film. Incubate for 5 min to hydrate the film.
- Sonicate the flask with an ultrasonic homogenizer or ultrasonic cleaner. Change the angle of the flask occasionally to allow the solution to touch the film thoroughly. Ensure that the thin lipid film is peeled from the bottom and emulsified completely and homogenously.
NOTE: Electron micrograph of biotinylated lipids containing liposomes is shown in Figure 1. - Transfer the liposome suspension (50 mg lipids/mL) to a new 1.5 mL tube. If it is not to be used immediately, freeze the liposomes in liquid nitrogen, and store at -80 °C.
- If a lipid is sold in powder form, dissolve in chloroform or appropriate organic solvent to 10-100 mg/mL concentration.
5. In vitro translation
- Thaw the wheat germ extract quickly by floating the tubes on water at room temperature for a few minutes. After thawing, immediately mix gently by inverting the tubes, spin down, and chill on ice until use.
NOTE: Freeze wheat germ extract in liquid nitrogen after use, and store at -80 °C. It withstands several freeze/thaw cycles. - Thaw 20 mg/mL creatine kinase stock solution. Mix 5 µL of stock solution and 45 µL of translation buffer to prepare 2 mg/mL creatine kinase working solution.
CAUTION: Refreezing creatine kinase is not recommended. - Thaw liposomes or mRNAs when required.
- Conduct a small-scale protein translation.
- Remove water from both the tube and the dialysis cup (0.1 mL), as prepared in step 3.3.2.
- Inject 1 mL and 300 µL of translation buffer in the tube and the dialysis cup, respectively.
CAUTION: In case the bottom of the dialysis cup does not reach the surface of translation buffer in the 1.5 mL tube, inject an additional 50–100 µL of buffer to the tube. - Prepare translation reaction mixture by mixing 15.6 µL of translation buffer, 2.4 µL of 2 mg/mL creatine kinase, 12 µL of 50 mg/mL liposomes, 15 µL of wheat germ extract, and 15 µL of mRNA. Mix gently by inverting the tubes, and spin down.
- Aspirate 60 µL of the translation reaction mixture using a 200 µL pipette.
- Insert the pipette tip into the undersurface of translation buffer in the dialysis cup. Pipette out the reaction mixture slowly and gently. Cover the dialysis cup with a lid to prevent evaporation.
NOTE: The reaction mixture sinks naturally to the bottom of the cup and forms a bilayer. Do not disturb the bilayer by mixing or shaking the cup.
- Conduct a large-scale translation (Figure 1).
- Pour 22 mL of translation buffer into a 25 mL tube. Insert a large-sized dialysis cup (2 mL) into the tube and add 2 mL of translation buffer in the cup.
- Prepare a translation reaction mixture by mixing 130 µL of translation buffer, 20 µL of 2 mg/mL creatine kinase, 100 µL of 50 mg/mL liposome, 125 µL of wheat germ extract, and 125 µL of mRNA. Mix gently by pipetting.
- Aspirate all the translation reaction mixture (500 µL) using a 1,000 µL pipette. Inject translation reaction mixture into the dialysis cup in the same way as described in step 5.4.5. Cover dialysis cup with a lid to prevent evaporation.
- Incubate the reactions at 15 °C for 24 h.
- Mix the reaction well in the dialysis cup by pipetting. Transfer the crude proteoliposome suspension to a new tube.
NOTE: A flat-bottomed 1.5 mL tube is recommended to collect the proteoliposome from the small-scale translation. After centrifugation, liposomes form compact and easily visible pellet on the bottom corner of the tube.
6. Purification of proteoliposomes
- Centrifuge the tube containing crude proteoliposome suspension at 17,800 x g at 4 °C for 10 min.
- Remove the supernatant. Suspend the proteoliposome pellet in PBS (small scale: 1 mL, large scale: 10 mL) by pipetting.
- Repeat the centrifugation and washing of proteoliposomes for another two circles.
- After washing, add a small amount of PBS and re-suspend proteoliposome pellet well by pipetting. Measure the volume of suspension using a micro pipette. Add PBS to adjust the volume to 60 µL (small scale) or 500 µL (large scale). Transfer the suspension to a new 1.5 mL tube.
- Transfer 10 µL of proteoliposome suspension to a new PCR tube for SDS-PAGE. Divide the rest of the samples into smaller portions for use when necessary. Freeze in liquid nitrogen and store at -80 °C.
7. SDS-PAGE and CBB staining
- Add 70 µL of water and 40 µL of 3x SDS-PAGE sample buffer to 10 µL of proteoliposome suspension.
CAUTION: Do not boil the SDS-PAGE sample, as membrane proteins aggregate, and hardly penetrate into acrylamide gel in electrophoresis. Also, add enough reducing agent to SDS-PAGE sample buffer (e.g., 2-mercaptoethanol at 3% final concentration) to prevent oxidation. - Set a 5%–20% gradient SDS-PAGE gel in an electrophoresis chamber. Load 3 µL, 6 µL, 12 µL of proteoliposome samples, 2 µL of protein size marker, and BSA standard series as well.
- Electrophorese at 52 mA, 400 V for 30 min.
- Stain the gel with CBB dye for 1 h. Decolorize in hot water and scan the gel image.
- Using NIH Image J software (https://imagej.nih.gov/ij/), quantify the band intensity of membrane protein in each lane. Estimate the amount of synthesized membrane proteins with BSA standard series.
Cell-Free Production of Proteoliposomes for Functional Analysis and Antibody Development Targeting Membrane Proteins
Learning Objectives
Using this protocol, partially purified proteoliposomes can be obtained in a short time. Representative results are shown in Figure 2A. Twenty five GPCRs of Class A, B, and C were successfully synthesized using the bilayer-dialysis method (small scale) and partially purified by centrifugation and buffer wash. Although the amount of synthesized proteins varies according to the type of protein, 50 to 400 µg of membrane proteins usually can be synthesized per reaction when large dialysis cups are used. Several milligrams of membrane proteins can be easily produced by increasing the number of reactions, due to the high scalability of wheat cell-free system. A pre-test using a small dialysis cup is sufficient to determine the production efficacy of the target protein in bilayer-dialysis method. According to the obtained productivity, the amount of the target protein to be produced using large dialysis cups can be estimated.
This protocol is suitable for expression of membrane proteins, particularly for those with multiple transmembrane helices. In most cases, membrane proteins with three or more transmembrane helices are easily incorporated into proteoliposomes after synthesis (Figure 2B), which makes a good productivity of proteoliposomes. Single-transmembrane-helix proteins are usually synthesized smoothly; however, they hardly integrate into liposomes due to the small hydrophobic region. Regarding proteins with two transmembrane helices, whether or not they are anchored to liposomes is dependent on the way their transmembrane helices are exposed.
Synthesized proteoliposomes are collected by simple centrifugation, and partially purified with a washing buffer, which greatly shortens the purification process of membrane proteins. Although both biological membranes and membrane proteins have been removed from wheat germ extracts beforehand, small amounts of wheat proteins are sometimes co-precipitated by binding to liposomes or membrane proteins synthesized (Figure 2A). Such protein contaminants are difficult to remove by simple centrifugation and buffer wash. When a highly purified membrane protein is required, it is necessary to solubilize the partially purified proteoliposomes with a surfactant and purify them by column chromatography.
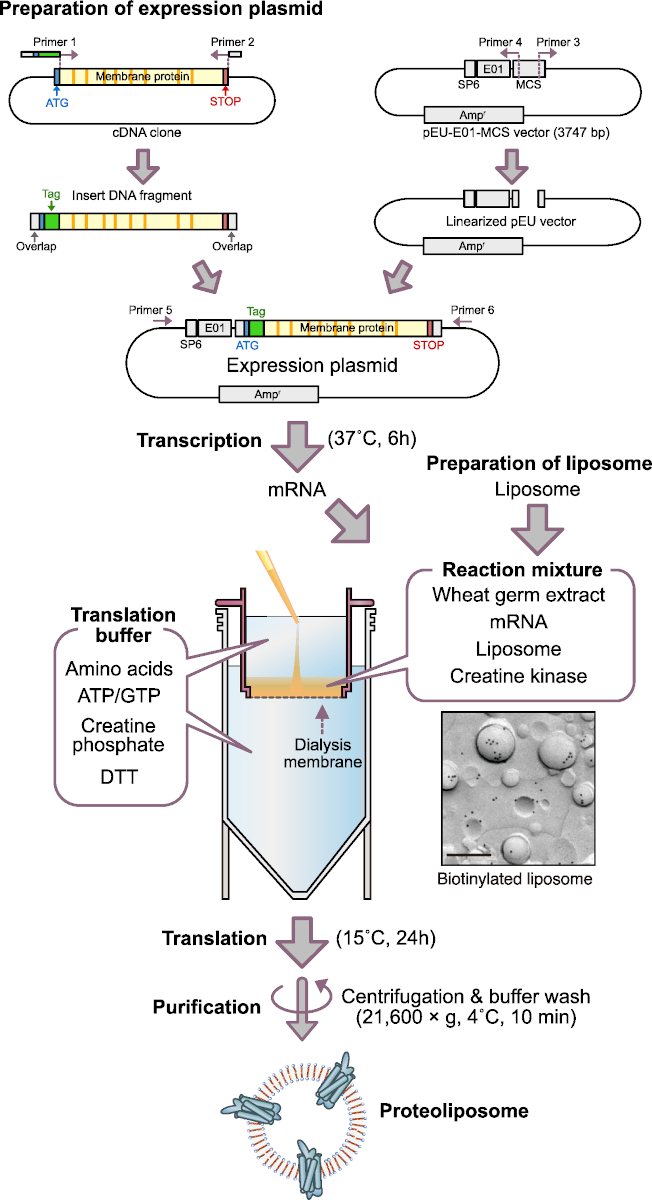
Figure 1: Scheme of cell-free proteoliposome production. SP6, SP6 promoter sequence; E01, E01 translation enhancer sequence; Ampr, ampicillin resistance gene; DTT, dithiothreitol. Electron micrograph shows immunogold labeling of biotinylated lipid containing liposome. Bar, 0.2 μm. This electron micrograph image was from Figure 1D in Takeda et al., 201545. Please click here to view a larger version of this figure.
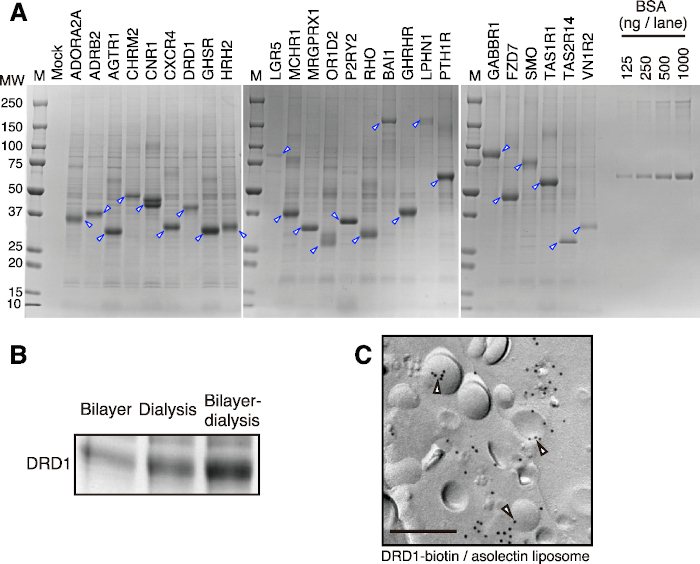
Figure 2: Representative results of proteoliposome production by bilayer-dialysis method. (A) SDS-PAGE image of cell-free synthesized GPCRs. Twenty-five selected GPCRs were synthesized by the bilayer-dialysis method. Proteoliposomes were partially purified and applied to SDS-PAGE and CBB staining. Arrowheads indicate target GPCRs. (B) Comparison of membrane protein productions between different translation methods. Dopamine receptor D1 (DRD1) protein was synthesized by each method in the same ratio of wheat germ extract, liposomes, and mRNA, respectively. DRD1 proteoliposome was partially purified by centrifugation, and subjected to SDS-PAGE and CBB staining. (C) Immunogold labeling of DRD1-biotin/liposome complex. DRD1 was enzymatically biotinylated by BirA biotin ligase. Bar, 0.2 μm. Blank arrowheads indicate DRD1-biotin on liposomes. This figure was modified from Figure 1 in Takeda et al., 201545. Please click here to view a larger version of this figure.
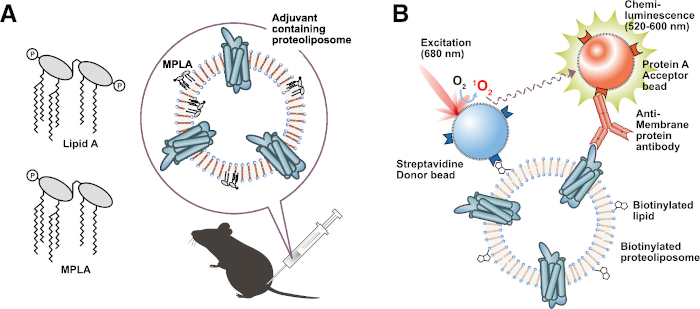
Figure 3: Application of functional proteoliposomes. (A) Immunization of adjuvant lipid-containing proteoliposome. (B) Biotinylated liposome-based interaction assay (BiLIA). Interaction between membrane protein and anti-membrane protein antibody was detected by AlphaScreen. Please click here to view a larger version of this figure.
List of Materials
| ×3 SDS-PAGE sample buffer | Containing 10% 2-mercaptoethanol | ||
| 5-20% gradient SDS-PAGE gel | ATTO | E-D520L | |
| 70% ethanol | Diluted ethanol by ultrapure water. | ||
| Agarose | Takara Bio | ||
| Ammonium acetate | Nakalai tesque | 02406-95 | As this reagent is deliquescent, dissolve all of it in water once opened and store it at -30°C. |
| Ampicillin Sodium | Nakalai tesque | 02739-74 | |
| Asolectin Liposome, lyophilized | CellFree Sciences | CFS-PLE-ASL | A vial contains 10 mg of lyophilized liposomes. |
| BSA standard | 1000 ng, 500 ng, 250 ng, 125 ng BSA / 10 µL ×1 SDS-PAGE sample buffer | ||
| CBB gel stain | |||
| cDNA clone of interest | Plasmid harboring cDNA clone or synthetic DNA fragment | ||
| Chloroform | Nakalai tesque | 08402-84 | |
| Cooled incubator | Temperature ranging from 0 to 40 °C or wider. | ||
| Creatine kinase | Roche Diagnostics | 04524977190 | |
| Dialysis cup (0.1 mL) | Thermo Fisher Scientific | 69570 | Slide-A-Lyzer MINI Dialysis Device, 10K MWCO, 0.1 mL |
| Dialysis cup (2 mL) | Thermo Fisher Scientific | 88404 | Slide-A-Lyzer MINI Dialysis Device, 10K MWCO, 2 mL |
| DNA ladder marker | Thermo Fisher Scientific | SM0311 | GeneRuler 1 kb DNA Ladder |
| DpnI | Thermo Fisher Scientific | FD1703 | FastDigest DpnI |
| E. coli strain JM109 | |||
| Electrophoresis chamber | ATTO | ||
| Ethanol (99.5%) | Nakalai tesque | 14713-95 | |
| Ethidium bromide | |||
| Evaporation flask, 100 mL | |||
| Gel imager | |||
| Gel scanner | We use document scanner and LED immuninator as a substitute. | ||
| LB broth | |||
| Lipids of interest | Avanti Polar Lipids | ||
| Micro centrifuge | TOMY | MX-307 | |
| NTP mix | CellFree Sciences | CFS-TSC-NTP | Mixture of ATP, GTP, CTP, UTP, at 25 mM each |
| Nuclease-free 25 mL tube | IWAKI | 362-025-MYP | |
| Nucrease-free plastic tubes | Watson bio labs | Do not autoclave. Use them separately from other experiments. | |
| Nucrease-free tips | Watson bio labs | Do not autoclave. Use them separately from other experiments. | |
| PBS buffer | |||
| PCR purification kit | MACHEREY-NAGEL | 740609 | NucleoSpin Gel and PCR Clean-up |
| pEU-E01-MCS vector | CellFree Sciences | CFS-11 | |
| Phenol/chloroform/isoamyl alcohol (25:24:1) | Nippon Gene | 311-90151 | |
| Plasmid prep Midi kit | MACHEREY-NAGEL | 740410 | NucleoBond Xtra Midi |
| Primer 1 | Thermo Fisher Scientific | Custom oligo synthesis | 5’-CCAAGATATCACTAGnnnnnnnnnnnnnnnnnnnnnnnn-3’ Gene specific primer, forward. Upper case shows overlap sequence to be added for seamless cloning. Lower case nnnn…. (20-30 bp) shows gene specific sequence. |
| Primer 2 | Thermo Fisher Scientific | Custom oligo synthesis | 5'-CCATGGGACGTCGACnnnnnnnnnnnnnnnnnnnnnnnn-3’ Gene specific primer, reverse. Upper case shows overlap sequence to be added for seamless cloning. Lower case nnnn…. (20-30 bp) shows gene specific sequence. |
| Primer 3 | Thermo Fisher Scientific | Custom oligo synthesis | 5'-GTCGACGTCCCATGGTTTTGTATAGAAT-3' Forward primer for vector linearization. Underline works as overlap in seamless cloning. |
| Primer 4 | Thermo Fisher Scientific | Custom oligo synthesis | 5'-CTAGTGATATCTTGGTGATGTAGATAGGTG-3' Reverse primer for vector linearization. Underline works as overlap in seamless cloning. |
| Primer 5 | Thermo Fisher Scientific | Custom oligo synthesis | 5’-CAGTAAGCCAGATGCTACAC-3’ Sequencing primer, forward |
| Primer 6 | Thermo Fisher Scientific | Custom oligo synthesis | 5’- CCTGCGCTGGGAAGATAAAC-3’ Sequencing primer, reverse |
| Protein size marker | Bio-Rad | 1610394 | Precision Plus Protein Standard |
| Rotary evaporator | |||
| seamless cloning enzyme mixture | New England BioLabs | E2611L | Gibson Assembly Master Mix Other seamless cloning reagents are also avairable. |
| SP6 RNA Polymerase & RNase Inhibitor | CellFree Sciences | CFS-TSC-ENZ | |
| Submarine Electrophoresis system | |||
| TAE buffer | |||
| Transcription Buffer LM | CellFree Sciences | CFS-TSC-5TB-LM | |
| Translation buffer | CellFree Sciences | CFS-SUB-SGC | SUB-AMIX SGC (×40) stock solution (S1, S2, S3, S4). Prepare ×1 translation buffer before use by mixing stock S1, S2, S3, S4 stock and ultrapure water. |
| Ultrapure water | We recommend to prepare ultrapure water by using ultrapure water production system every time you do experiment. Do not autoclave. We preparaed ultrapure water by using Milli-Q Reference and Elix10 system. Commercially available nuclease-free water (not DEPC-treated water) can be used as a substitute. Take care of contamination after open the bottle. |
||
| Ultrasonic homogenizer | Branson | SONIFIER model 450D-Advanced | Ultrasonic cleaner can be used as a substitute. |
| UV transilluminator | |||
| Vacuum desiccator | |||
| Wheat germ extract | CellFree Sciences | CFS-WGE-7240 | WEPRO7240 |
Lab Prep
Membrane proteins play essential roles in a variety of cellular processes and perform vital functions. Membrane proteins are medically important in drug discovery because they are the targets of more than half of all drugs. An obstacle to conducting biochemical, biophysical, and structural studies of membrane proteins as well as antibody development has been the difficulty in producing large amounts of high-quality membrane protein with correct conformation and activity. Here we describe a “bilayer-dialysis method” using a wheat germ cell-free system, liposomes, and dialysis cups to efficiently synthesize membrane proteins and prepare purified proteoliposomes in a short time with a high success rate. Membrane proteins can be produced as much as in several milligrams, such as GPCRs, ion channels, transporters, and tetraspanins. This cell-free method contributes to reducing the time, cost and effort for preparing high-quality proteoliposomes, and provides suitable means for functional analysis of membrane proteins, drug targets screening, and antibody development.
Membrane proteins play essential roles in a variety of cellular processes and perform vital functions. Membrane proteins are medically important in drug discovery because they are the targets of more than half of all drugs. An obstacle to conducting biochemical, biophysical, and structural studies of membrane proteins as well as antibody development has been the difficulty in producing large amounts of high-quality membrane protein with correct conformation and activity. Here we describe a “bilayer-dialysis method” using a wheat germ cell-free system, liposomes, and dialysis cups to efficiently synthesize membrane proteins and prepare purified proteoliposomes in a short time with a high success rate. Membrane proteins can be produced as much as in several milligrams, such as GPCRs, ion channels, transporters, and tetraspanins. This cell-free method contributes to reducing the time, cost and effort for preparing high-quality proteoliposomes, and provides suitable means for functional analysis of membrane proteins, drug targets screening, and antibody development.
Procedure
Membrane proteins play essential roles in a variety of cellular processes and perform vital functions. Membrane proteins are medically important in drug discovery because they are the targets of more than half of all drugs. An obstacle to conducting biochemical, biophysical, and structural studies of membrane proteins as well as antibody development has been the difficulty in producing large amounts of high-quality membrane protein with correct conformation and activity. Here we describe a “bilayer-dialysis method” using a wheat germ cell-free system, liposomes, and dialysis cups to efficiently synthesize membrane proteins and prepare purified proteoliposomes in a short time with a high success rate. Membrane proteins can be produced as much as in several milligrams, such as GPCRs, ion channels, transporters, and tetraspanins. This cell-free method contributes to reducing the time, cost and effort for preparing high-quality proteoliposomes, and provides suitable means for functional analysis of membrane proteins, drug targets screening, and antibody development.
