Single Molecule Fluorescence Energy Transfer Study of Ribosome Protein Synthesis
Instructor Prep
concepts
Student Protocol
1. Preparation of labeled ribosome and tRNA for FRET detection
- Isolate ribosome without L27 from E. coli strain IW312 according to standard protocols20,21. Extract the regular ribosome from E. coli strain MRE600.
- Clone the L27's rpmA gene with C-terminal His-tag into pET-21b (+) plasmid, which is transformed and expressed in BL21(DE3)pLysS cells15. Purify the protein via a prepacked sepharose column.
- Labeling of L27
- Incubate 20-100 μM of purified L27 in 100 μL with 2-10-fold excess of TCEP (Tris-2-carboxyethyl-phosphine) at room temperature (RT) for 30 min. Buffer exchange the solution into PBS buffer (10 mM Na2HPO4, 1.8 mM KH2PO4, 2.7 mM KCl; 0.137 M NaCl), using a nap-5 column. Add 10-fold excess Cy5-maleimide mono-reactive dye in DMSO into the solution and incubate at RT in the dark for 2 h.
- Load the labeled protein solution (500 μL) mentioned above on a Nap-5 column to remove the excess dye. Ensure that the protein elutes in the first colored band, followed by the free dye elute in the second band, respectively. Collect the eluted protein in 1 mL of TAM10 buffer (20 mM tris-HCl (pH 7.5), 10 mM Mg (OAc)2, 30 mM NH4Cl, 70 mM KCl, 0.5 mM EDTA, and 7 mM BME (2-mercaptoethanol)).
- Reconstitute the ribosome with L27. Mix the labeled L27 protein with an equal amount of IW312 ribosome in TAM10 buffer and incubate at 37 °C for 1 h. Remove the free protein by sucrose cushion ultracentrifugation (100,000 x g, overnight) to allow the ribosome to form a blue-colored pellet at the bottom of the tube.
- Label the tRNAPhe according to the previously published report22. First, incubate the tRNA (10 A260 in 1 mL of water) with 100 μL of NaBH4 (100 mM stock, adding dropwise) at 0 °C for 1 h. Free the tRNAs from unreacted NaBH4 via three-time ethanol precipitation by adding 1/10 volume of 3 M KAc (pH 5.0) and 2.5x volume of ethanol. Incubate the reduced tRNA with 1 μL of Cy3-NHS dye solution (1 mg in 10 μL of DMSO) in a minimum volume (~20 μL) of 0.1 M sodium formate (pH 3.7). After 2 h of incubation at 37 °C, separate the tRNA from the unreacted dye via a small G25 Sephadex column, and then remove the free dye via two-time ethanol precipitation.
2. Preparation of the ribosome complexes
- Express and purify the His-tagged proteins of IF1, IF2, IF3, EF-G, EF-Tu, EF-Ts, and tRNA synthetases using standard methods15.
- Prepare the initiation mix by adding the following ingredients in TAM10 buffer at 37 °C for 15 min: 1 µM of the labeled ribosome; 1.5 µM each of IF1, IF2, and IF3; 2 µM biotinylated mRNA (coding MF for the first two amino acids); 4 µM of charged fMet-tRNAfMet; and 1 mM GTP.
- Prepare the EF-Tu_G mix by mixing the following ingredients in TAM10 buffer at 37 °C for 15 min: 4 µM EF-Tu, 0.4 µM EF-Ts, 2 µM EF-G, 4 mM GTP, 4 mM PEP, and 0.02 mg/mL of Pyruvate Kinase.
- Prepare the EF-Tu_NoG mix similar to step 2.3 without EF-G.
- Prepare the Phe mix by mixing the following ingredients in nuclease-free water at 37 °C for 15 min: 100 mM Tris (pH 7.5), 20 mM MgAc2, 1 mM EDTA, 4 mM ATP, 7 mM BME, 2 mM Phenylalanine-specific synthetase, 2 A260/mL of labeled tRNAPhe, and 50 μM phenylalanine.
- Prepare the pre-translocation complex (PRE) by mixing the initiation, EF-Tu_NoG, and Phe mixes in the ratio of 1:2:2, at 37 °C for 2 min. After incubation, purify the PRE by 1.1 M sucrose cushion ultracentrifugation overnight at 100,000 x g.
- Prepare the post-translocation complex (POST) by mixing the initiation, EF-Tu_WG, and Phe mixes in the ratio of 1:2:2, at 37 °C for 10 min. After incubation, purify the POST by 1.1 M sucrose cushion ultracentrifugation overnight at 100,000 x g.
3. Preparation of sample slides
- Clean the microscope glass slides (75 mm x 25 mm) containing six pairs of holes (1 mm in diameter). Each pair forms an inlet-outlet of the sample chamber. Ensure that the distance between each inlet-outlet hole is 13 mm and the center-to-center distance between two channels is 6 mm. Place six slides in a glass crucible.
- Fill the crucible with acetone and sonicate them for 5 min. Decant the acetone and rinse the slides three times with ultrapure water. Fill the crucible again with water and 1 mL of 10 M KOH. Sonicate for 20 min.
- Rinse the slides three times with ultrapure water. Fill the crucible again with ethanol and sonicate for 5 min. Completely decant the solvent. Let the slides dry in the fume hood.
- Clean the microscope glass coverslips (#1.5 thickness, 24 x 40 mm). Perform the cleaning steps as described in steps 3.1-3.3.
- Bake the cleaned slides and coverslips at 300 °C for 3 h. Keep them in the furnace overnight to cool.
- Coat the coverslip with aminosilane. In the crucible containing coverslips, pour in methanol mix (100 mL of methanol, 5 mL of water, 0.5 mL of HAc, and 1 mL of aminosilane). Warm the crucible in a water bath for 10 min, sonicate it for 10 min, and then warm the crucible in the water bath for another 10 min. Decant the methanol mixture, rinse the coverslip well in three crucibles of clean water. Purge the surfaces with a dry nitrogen stream through a needle.
- Coat the coverslips with biotin-PEG in a laminar clean hood.
- To make the PEG solution, use 98 mg of PEG and 2-3 mg of biotin-PEG in 200 μL of 100 mM NaHCO3 solution.
- Lay one coverslip flat on a surface. Carefully drop 60 μL of PEG solution on the top edge. Then, lay another coverslip on top of it, letting the capillary effect spread the solution between the two coverslips. Ensure no bubbles are formed.
- Repeat step 3.7.2 for two more pairs of coverslips. Coat six pieces of coverslips with biotin. If more coverslips are needed, adjust the amount of PEG and Biotin PEG accordingly to make more PEG solution.
- Store these coverslips in an empty tip-box filled with water and incubate for 3 h in the dark.
- After 3 h, separate the coverslips, rinse with water in three consecutive crucibles and purge with a dry nitrogen stream. Place the coated coverslips on a ceramic rack. Mark the surface with coating.
- Assemble the sample chambers19.
- Pull sharp pipette tips through the holes on the glass slides until they fit tight. Scrape the sharp ends off until it is completely flat to the glass surface. Cut the other end of the tip to be approximately 5 mm for sample loading.
- Cut a double-faced tape with the channel pattern on it (25 x 40 mm).
- Stick the double-faced tape onto the glass slides on the flat side.
- Stick the coated coverslip onto the double-faced tape. Press on it to make a tight seal of the sample chamber. Make sure the coated side is facing inward.
4. Single molecule FRET imaging
- Turn on the computer.
- Turn on the microscope switch.
- Start the laser control interface and turn on the laser. Push the enable button on the laser control box to warm up the laser.
- Turn on the cameras.
- Start the microscope-associated software program. Click the upper shutter Off and click the lamp On.
- Prepare the TAM10_WT buffer by adding 10 μL of 5% Tween-20 into 1 mL of TAM10 buffer.
- Prepare the deoxy solution by weighing 3 mg of glucose oxidase in a small microcentrifuge tube. Add 45 μL of catalase. Vortex gently to dissolve the solid. Spin at 20,000 x g in a microcentrifuge for 1 min. Take the supernatant.
- Prepare 30% glucose solution in water and 200 mM Trolox solution in DMSO.
- Prepare 0.5 mg/mL of streptavidin solution in nuclease-free water.
- Add one drop of imaging oil to the TIRF objective.
- Put the sample chambers on the objective.
- Fill the chamber with 10 μL of streptavidin solution. Wait for 1 min.
- Wash the chamber with 30 mL of TAM10_WT buffer and collect the run-through solution with a folded filter paper. Wait for 1 min.
- Dilute the ribosome complexes (PRE or POST) to 10-50 nM concentration with TAM10_WT buffer.
- Load 20 μL of the ribosome sample into the channel. Wait for 2 min.
- Make the imaging buffer (50 μL of TAM10 buffer plus 0.5 μL each of deoxy, glucose, and Trolox solutions). Mix the buffer well.
- Flush the chamber with 30 μL of the imaging buffer.
- Set the camera to acquire 100 ms/frame, 16 bits, 4 x 4 binning.
- Click the upper shutter On and the lamp Off.
- Start camera acquisition. Spread the images into three windows representing two individual cameras and the overlay.
- Click on Auto-focus. Fine-tune to find the best focus.
- Click on a method. Set the field points, file storage, and the loop number.
- Click on Run.
NOTE: A new window appears with real-time imaging. The progress of the time series and field of view series are shown at the top of the window. - Once the image acquisition is complete, close the popup window.
- Open the ND file (acquired file).
- Click on ROI/ simple ROI editor. Choose the option Circle.
- Select the ROI on the image. Click on Finish when it is complete.
- Click on Measurement/ Time Measurement to show the data either as a plot or as a spreadsheet.
- Open the Export tab. Set the export parameters.
- Click on Export to save the intensities.
Single Molecule Fluorescence Energy Transfer Study of Ribosome Protein Synthesis
Learning Objectives
The smFRET had the ribosome labeled at the middle position of tRNA traffic, to distinguish the tRNA translocation from the A- to the P-site (Figure 1)15. The distance from the L27 labeling residue to the A- or P-site tRNA is 52 or 61 Å, respectively, corresponding to FRET efficiency of 0.47 and 0.65. After the image collection, fluorescence intensities from the donor and acceptor channels were retrieved and plotted as time lapses (Figure 1). FRET efficiency was calculated by the formula Iacceptor/(Iacceptor+Idonor). A homemade program detected the single-step bleaching of the donor/acceptor and fitted the data before the bleaching points to avoid calculation of FRET from noises. After donor bleaching, both traces approached the baseline because no excitation can occur directly on the acceptor (Figure 2A). After acceptor bleaching, the donor intensity increased because fewer reaction pathways dissipate the excitation energy (Figures 2B,C).
It was found that the individual ribosomes exhibit different fluctuation, such as in the examples shown in Figure 2. These fluctuations are due to the wobbling motion of the tRNAs, which causes distance fluctuations to the L2717,18. In Figure 2, the dotted lines are the original data, and solid lines are the fitted data from the program. The fitted traces do not change the raw data reading but truncate the time-lapse traces after bleaching. The blue traces are the calculated FRET efficiencies. By tracking the exact same ribosomes after 5 min incubation at RT, it was found that one type of dynamics can switch to another23. Because these signals report on tRNA motions dictated by the surrounding ribosome, similar FRET efficiencies were grouped into various ribosome subpopulations.
Figure 3 shows very diverse subpopulations in the PRE complex. There are approximately 70% (640/951) of fluctuating species and 30% (311/951) of non-fluctuating species. These two categories can be further grouped into finer subpopulations. On the other hand, Figure 4 shows the opposite dynamics distribution. In POST, 65% of subpopulations are non-fluctuating, and the majority of them exhibited high FRET efficiency. These results indicate that the ribosome is more flexible in the PRE state than the POST state, which corroborates a cryo-EM study that concluded that the peptidyl chain at the P-site locks the ribosome dynamics in the POST state. However, in the PRE state, the peptidyl chain is transferred to the A-site, unlocking the ribosome and promoting5. The results supported the structure study conclusion under more physiological conditions. The unlocked state is correlated with the ratcheting motion between the 30S and 50S, and the locked state is correlated with the un-ratcheted conformation.
The subpopulation sorting method reveals the ribosome conformation upon inhibition by the antibiotic viomycin, as shown in Figure 514. The overall FRET efficiency histogram shows a major peak at 0.47, the classic state, without sorting. In this state, the ribosome is locked and un-ratcheted. However, sorting the subpopulations has revealed that 60% of the population is fluctuating as the unlocked PRE complex. Therefore, viomycin has trapped the ribosome at the ratcheted state. The results of this study contradicted an x-ray structure result at the time of publishing (2010) but were supported recently by another structure study (2020).
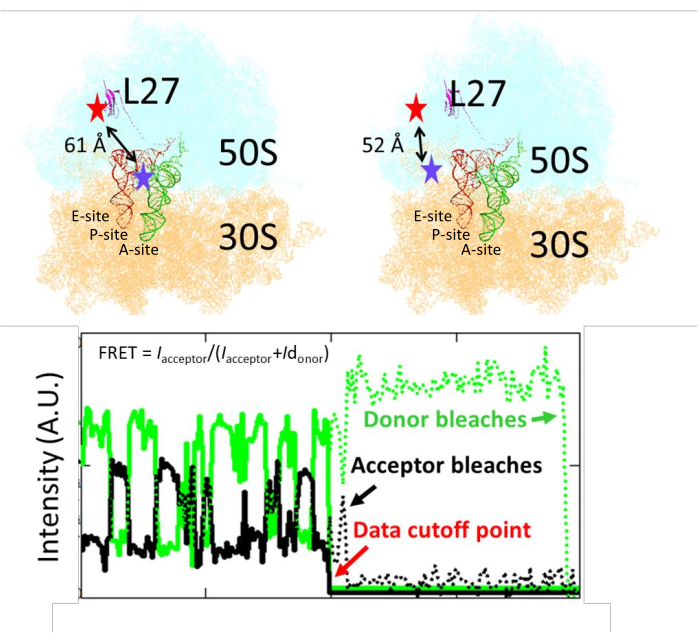
Figure 1: The labeling position of the Cy3/Cy5 on the ribosome and the tRNA. The distances between the labeling residue of L27 to the A- and P-site tRNA are shown (the red and blue stars show the approximate labeling locations of the Cy5 and Cy3, respectively). One representative time-lapse trace of fluorescence intensities from the CY3/Cy5 FRET pair is shown. A program detects bleaching points on donor/acceptor traces and truncates the trace before that point. This figure has been modified from Altuntop, M. E. et al.15. Please click here to view a larger version of this figure.
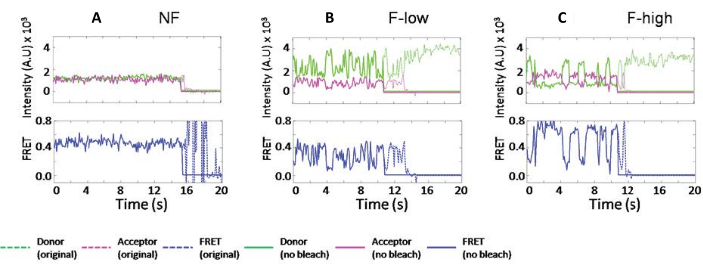
Figure 2: The typical ribosome traces were classified by their different dynamics. (A) A non-fluctuating ribosome trace. (B) A fluctuating ribosome trace that only samples FRET values lower than 0.6. (C) A fluctuating ribosome trace that samples FRET value higher than 0.6. The legends are shown in the plot. The fluorescence intensities of donor and acceptor are green and magenta, respectively. FRET values are calculated as Iacceptor/(Iacceptor+Idonor) and plotted separately in blue. The original data are displayed in dotted lines, and data truncated before the bleaching points are displayed in solid lines. This figure has been modified from Altuntop, M. E. et al.15. Please click here to view a larger version of this figure.
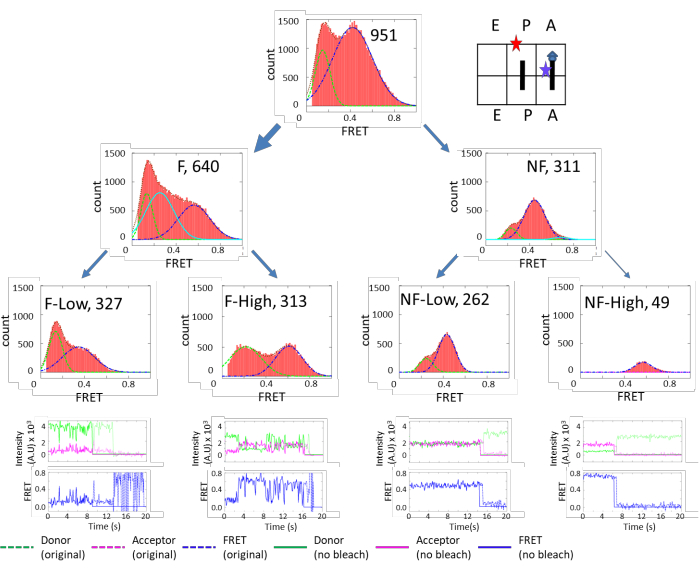
Figure 3: FRET efficiency histograms of the Pre-complex. The top-tier plots show the FRET histogram of the total ribosomes. The second-tier plots show the FRET histograms of the ribosomes separated into fluctuating (F) and non-fluctuating (NF) groups. The third-tier plots show the FRET histograms of the ribosomes further separated into groups of fluctuations that were above or below a FRET value of 0.6. Similar criteria were applied to the NF ribosomes to separate them into stable FRET states below or above 0.6 (NF-low, NF-high). The fourth-tier plots display the representative traces for each subpopulation. This figure has been modified from Altuntop, M. E. et al.15. Please click here to view a larger version of this figure.
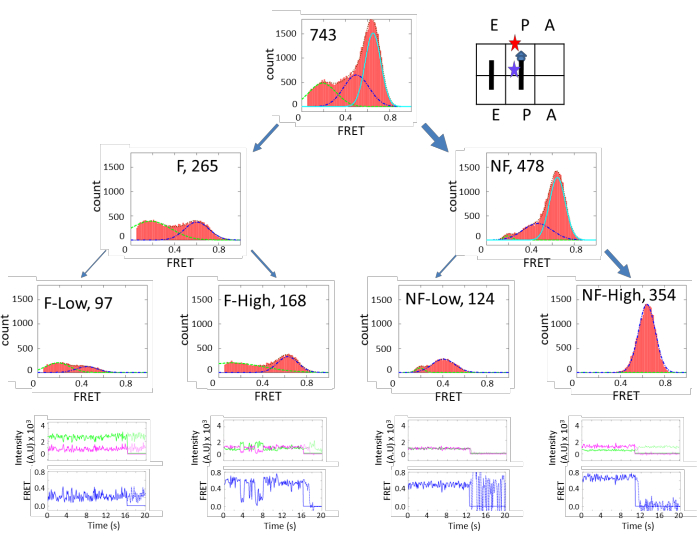
Figure 4: FRET efficiency histograms of POST-Complex. The arrangement and grouping are similar to Figure 3. Contrary to Figure 3, the NF-High population is the majority. This figure has been modified from Altuntop, M. E. et al.15. Please click here to view a larger version of this figure.
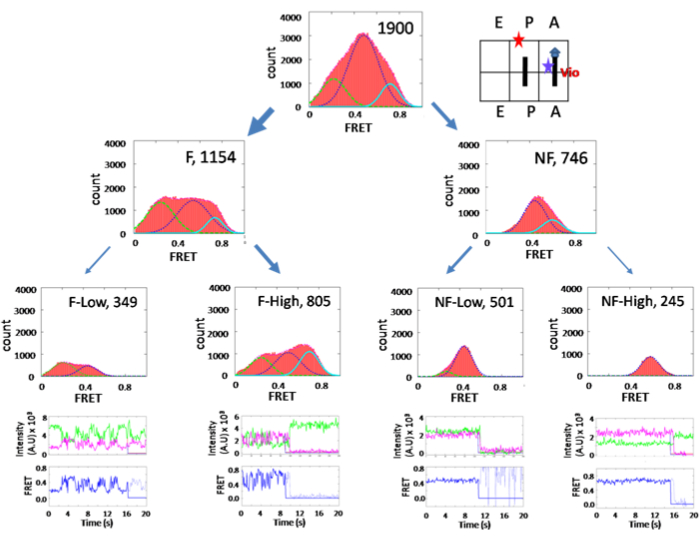
Figure 5: Histograms of the PRE complex in the presence of 100 µM viomycin. The arrangement and grouping are similar to Figure 3. This figure has been modified from Ly, C. T. et al.14. Please click here to view a larger version of this figure.
List of Materials
| Aminosilane | Laysanbio | MPEG-SIL-5000 | |
| Biotin-PEG | Laysanbio | Biotin-PEG-SVA-5000 | |
| BL21(DE3)pLysS cells | Novagen | 71403 | |
| Catalase | millipore sigma | C3515 | |
| CS150FNX Micro Ultracentrifuge | nuaire | ||
| Cy3/C5-maleimide | ApexBio | A8138/A8140 | |
| ECLIPSE Ti2 inverted microscope | Nikon | ||
| EdgeGARD Laminar Flow Hood | Baker | ||
| Glucose oxidase | millipore sigma | G2133 | |
| Histrap HP column (Prepacked sepharose column) | Cytiva | 17524701 | |
| Microscope cover slip | VWR | 48393-230 | |
| Microscope glass slides | VWR | 470235-792 | |
| ORCA-Flash4.0 V3 camera | Hamamatsu | ||
| PEG (5,000) | Laysanbio | MPEG-SVA-5000 | |
| pET-21b (+) plasmid | Novagen | 69741 | |
| Sonicator | VWR | CPX-952-518R | |
| TCEP | Apexbio | B6055 | |
| Trolox | millipore sigma | 238813 |
Lab Prep
The ribosome is a large ribonucleoprotein complex that assembles proteins processively along mRNA templates. The diameter of the ribosome is approximately 20 nm to accommodate large tRNA substrates at the A-, P- and E-sites. Consequently, the ribosome dynamics are naturally de-phased quickly. Single molecule method can detect each ribosome separately and distinguish inhomogeneous populations, which is essential to reveal the complicated mechanisms of multi-component systems. We report the details of a smFRET method based on the Nikon Ti2 inverted microscope to probe the ribosome dynamics between the ribosomal protein L27 and tRNAs. The L27 is labeled at its unique Cys 53 position and reconstituted into a ribosome that is engineered to lack L27. The tRNA is labeled at its elbow region. As the tRNA moves to different locations inside the ribosome during the elongation cycle, such as pre- and post- translocation, the FRET efficiencies and dynamics exhibit differences, which have suggested multiple subpopulations. These subpopulations are not detectable by ensemble methods. The TIRF-based smFRET microscope is built on a manual or motorized inverted microscope, with home-built laser illumination. The ribosome samples are purified by ultracentrifugation, loaded into a home-built multi-channel sample cell and then illuminated via an evanescent laser field. The reflection laser spot can be used to achieve feedback control of perfect focus. The fluorescence signals are separated by a motorized filter-turret and collected by two digital CMOS cameras. The intensities are retrieved via the NIS-Elements software.
The ribosome is a large ribonucleoprotein complex that assembles proteins processively along mRNA templates. The diameter of the ribosome is approximately 20 nm to accommodate large tRNA substrates at the A-, P- and E-sites. Consequently, the ribosome dynamics are naturally de-phased quickly. Single molecule method can detect each ribosome separately and distinguish inhomogeneous populations, which is essential to reveal the complicated mechanisms of multi-component systems. We report the details of a smFRET method based on the Nikon Ti2 inverted microscope to probe the ribosome dynamics between the ribosomal protein L27 and tRNAs. The L27 is labeled at its unique Cys 53 position and reconstituted into a ribosome that is engineered to lack L27. The tRNA is labeled at its elbow region. As the tRNA moves to different locations inside the ribosome during the elongation cycle, such as pre- and post- translocation, the FRET efficiencies and dynamics exhibit differences, which have suggested multiple subpopulations. These subpopulations are not detectable by ensemble methods. The TIRF-based smFRET microscope is built on a manual or motorized inverted microscope, with home-built laser illumination. The ribosome samples are purified by ultracentrifugation, loaded into a home-built multi-channel sample cell and then illuminated via an evanescent laser field. The reflection laser spot can be used to achieve feedback control of perfect focus. The fluorescence signals are separated by a motorized filter-turret and collected by two digital CMOS cameras. The intensities are retrieved via the NIS-Elements software.
Procedure
The ribosome is a large ribonucleoprotein complex that assembles proteins processively along mRNA templates. The diameter of the ribosome is approximately 20 nm to accommodate large tRNA substrates at the A-, P- and E-sites. Consequently, the ribosome dynamics are naturally de-phased quickly. Single molecule method can detect each ribosome separately and distinguish inhomogeneous populations, which is essential to reveal the complicated mechanisms of multi-component systems. We report the details of a smFRET method based on the Nikon Ti2 inverted microscope to probe the ribosome dynamics between the ribosomal protein L27 and tRNAs. The L27 is labeled at its unique Cys 53 position and reconstituted into a ribosome that is engineered to lack L27. The tRNA is labeled at its elbow region. As the tRNA moves to different locations inside the ribosome during the elongation cycle, such as pre- and post- translocation, the FRET efficiencies and dynamics exhibit differences, which have suggested multiple subpopulations. These subpopulations are not detectable by ensemble methods. The TIRF-based smFRET microscope is built on a manual or motorized inverted microscope, with home-built laser illumination. The ribosome samples are purified by ultracentrifugation, loaded into a home-built multi-channel sample cell and then illuminated via an evanescent laser field. The reflection laser spot can be used to achieve feedback control of perfect focus. The fluorescence signals are separated by a motorized filter-turret and collected by two digital CMOS cameras. The intensities are retrieved via the NIS-Elements software.
