Differentiation and Imaging of Brown Adipocytes from the Stromal Vascular Fraction of Interscapular Adipose Tissue from Newborn Mice
Summary
Preadipocytes are isolated from the stromal vascular fraction of interscapular brown adipose tissue from newborn mice and differentiated into cells that accumulate lipid droplets, express molecular markers, and show the mitochondrial morphology of mature brown adipocytes. These cells are further analyzed by immunofluorescence and transmission electron microscopy.
Abstract
Brown adipose tissue (BAT) is only present in mammals and has a thermogenic function. Brown adipocytes are characterized by a multilocular cytoplasm with multiple lipid droplets, a central nucleus, a high mitochondrial content, and the expression of uncoupling protein 1 (UCP1). BAT has been proposed as a potential therapeutic target for obesity and its associated metabolic disorders due to its ability to dissipate metabolic energy as heat. To investigate BAT function and regulation, brown adipocyte culturing is indispensable. The present protocol optimizes tissue processing and cell differentiation for culturing brown adipocytes from newborn mice. Additionally, procedures for the imaging of differentiated adipocytes with both confocal immunofluorescence and transmission electron microscopy are shown. In the brown adipocytes differentiated with the techniques described herein, the major defining features of classical BAT are preserved, including high UCP1 levels, increased mitochondrial mass, and very close physical contact between the lipid droplets and mitochondria, making this method a valuable tool for BAT studies.
Introduction
White and brown adipose tissue differ in their anatomical location, cellular origin, function, morphology, and total mass. White adipose tissue (WAT) is the major physiological energy reservoir of the body and stores large amounts of triacylglycerol (TAG) in highly specialized cells that have a single giant lipid droplet occupying most of their cellular volume1. TAG lipolysis releases free fatty acids, which enter the systemic circulation to meet energy demands during fasting or other states of negative energy balance. Additionally, the WAT secretes protein and lipid products, called adipokines and lipokines, respectively, that have metabolic, immune, and reproductive regulatory functions, thus making the WAT the largest endocrine tissue in the body2.
Brown adipose tissue (BAT) is a much smaller organ whose main physiological function is non-shivering thermogenesis to prevent hypothermia. In mice and newborn humans, BAT is a well-defined organ located in the interscapular space. Adult humans lack interscapular BAT (iBAT); nevertheless, they develop clusters of brown adipocyte-like cells integrated in depots that otherwise mostly comprise WAT. These "brown-in-white" (brite) adipocytes share morphological and molecular features with classical iBAT adipocytes, but they have a different cellular origin3,4.
In contrast to white adipocytes, brown adipocytes have multiple small lipid droplets and abundant mitochondria5. Uncoupling protein 1 (UCP1, also known as thermogenin) is uniquely expressed by brown and brite adipocytes and mediates proton leakage in the inner mitochondrial membrane (IMM), thus uncoupling electron transport from ATP synthesis and generating heat. Non-shivering thermogenesis in BAT is activated by norepinephrine (NE), which is released from the sympathetic terminals in the BAT in response to cold stimulation6. NE binds to beta-adrenoceptors (mostly beta 3) on the surface of brown adipocytes and triggers an intracellular cAMP-mediated signaling cascade. This results in TAG lipolysis, the beta-oxidation of mitochondrial fatty acids, and heat generation upon UCP1 activation3. The close functional relationship between lipid droplets and mitochondria in brown adipocytes has structural parallels, such as the interaction between these organelles in areas that are large and have very tight physical contact7,8.
iBAT has abundant blood vessels and sympathetic terminals9. These structures, along with the preadipocytes, immune cells, fibroblasts, and extracellular matrix molecules, compose the adipose stromal vascular fraction (SVF)10. Many protocols have been reported to generate mature adipocytes from preadipocytes11,12,13,14,15 (Supplementary Table 1); nevertheless, they display extreme variations in tissue processing and the composition of the differentiation culture media. The protocol described herein allows the efficient and reproducible differentiation of brown adipocytes that (1) express the key adipogenic transcription factors peroxisome proliferator-activated receptor gamma (PPARγ) and CCAAT/enhancer-binding protein alpha (C/EBPα), (2) express the mature adipocyte markers perilipin1 (PLIN1), and cluster determinant 36 (CD36), (3) accumulate abundant lipid droplets, (4) have high mitochondrial mass and a high abundance of OXPHOS complexes, (5) have thermogenic potential, as determined by high levels of UCP1, and (6) have mitochondrial morphological changes associated with the phenotype of mature brown adipocytes. This methodology is used for studying the molecular mechanisms underlying generalized lipodystrophy15,16,17.
Protocol
The animal procedures were approved by the Institutional Animal Care and Use Committee at Pontificia Universidad Católica de Chile. P0.5 newborn mice of both sexes, derived from a mixed background of C57BL/6J and 129J strains, were used for this study.
1. Tissue extraction
- Clean and disinfect the workbench with a 70% ethanol solution, and use sterile surgical material.
- Euthanize P0.5 newborn pups by decapitation with surgical scissors following institutionally approved protocol.
- Place the mouse in a prone position (facing downward), and make a 1 cm long incision in the skin at the mid-level of the animal's back using sharp scissors. Remove the skin carefully.
NOTE: This step is critical because if the skin is not well removed, it might tear and damage the iBAT and preclude the optimal preparation of the SVF. - Surgically detach the iBAT from the carcass using small scissors.
NOTE: The iBAT is visualized as two fatty lobules that can be removed independently. - Harvest both iBAT lobes by using curved-tip pliers to remove the lobes independently.
- Place the two lobes in a plastic tube with 1.5 mL of sterile PBS at room temperature. Maintain the tissues in their individual tubes until the tissue extraction is completed for all the pups required.
2. Tissue digestion
- Aliquot 300 µL of collagenase type II digestion buffer (0.2% type II collagenase in buffer [25 mM KHCO3, 1.2 mM MgSO4, 120 mM NaCl, 5 mM glucose, 12 mM KH2PO4, 4.8 mM KCl, 1.2 mM CaCl2, 2.5% BSA, and 1% penicillin/streptomycin, at pH 7.4]) (Table 1) in 1.5 mL tubes in a biosafety cabinet.
NOTE: Prepare sufficient tubes for the number of individual tissues to be extracted. In the present study, seven tubes were used, corresponding to a litter of seven pups. - Move to the bench, and place the iBAT lobes in collagenase type II digestion buffer to degrade the extracellular matrix.
- Incubate the iBAT tissues for 45 min at 37 °C with continuous gentle shaking at 800 rpm (Figure 1).
3. Tissue processing
NOTE: After the digestion, perform all the following steps in a class II laminar flow tissue culture hood.
- Prepare and label two sets of 1.5 mL plastic tubes for each individual sample.
- Mechanically disrupt the tissue suspension in collagenase type II digestion buffer by pipetting up and down using a P1,000 micropipette and filtered tips. Use a new tip for each sample.
NOTE: This is a critical step. Ensure to mechanically disaggregate the tissue with the tip and pipette up and down until macroscopic homogenization is complete. - Pass the disaggregated tissues through individual 100 µm cell strainers (see the Table of Materials) into new 1.5 mL tubes.
- Add 500 µL of ACK buffer (see the Table of Materials) to the filtered tissue samples, invert the tubes 4-5 times, and incubate at room temperature for 4 min.
NOTE: Do not exceed the incubation time, as this may damage the cells of interest. - Centrifuge the tissue samples at 300 x g for 5 min at room temperature.
- Discard the supernatant by inverting the tube, and resuspend the pellet in 1 mL of culture medium (DMEM-F12 with 10% FBS, 1% antibiotic antimycotic, pH 7.2) (see the Table of Materials).
- Pass the suspensions through a 40 µm cell strainer into new 1.5 mL tubes using a P1,000 micropipette and filtered tips.
- Centrifuge the samples at 300 x g for 5 min at room temperature.
NOTE: The pellet corresponds to the preadipocyte-containing SVF. - Discard the supernatant by inverting the tube, and resuspend the pellet in 1 mL of culture medium by pipetting.
- Prepare one 15 mL tube for each sample by adding 5 mL of culture medium.
- Add each resuspended pellet to its corresponding 15 mL tube containing 5 mL of culture medium. Invert the tube 4-5 times to mix the cell content.
- Seed each sample into six wells of a 24-well culture plate by adding 1 mL of the suspension to each well (Figure 1).
4. Stromal vascular fraction (SVF) culture
- Change the culture medium every other day until the cells reach 100% confluence.
NOTE: During this period, non-plastic adherent cells are removed, resulting in a primary culture enriched with a preadipocyte-containing SVF. - When the cells become 100% confluent, remove the culture medium, and wash the cells with 800 µL of sterile PBS.
- Add 150 µL of 0.25% trypsin-EDTA (see the Table of Materials) to every well, and place the culture plate for 3 min in the incubator for the detachment of the cells.
NOTE: Verify that the cells are detached using a phase contrast light microscope. - Inactivate the trypsin by adding 850 µL of culture medium to each well.
- Collect the cells from the six wells seeded in step 3.12, and transfer them into a 15 mL tube.
- Make the total volume up to 12 mL by adding 6 mL of culture medium. Invert the tube 4-5 times to mix the content.
- Seed 1 mL of the sample into each well of a 12-well plate. Change the culture medium every other day until the cells reach confluence.
- When the cells become 100% confluent, detach them with 150 µL of 0.25% trypsin-EDTA following the same incubation conditions as in step 4.3, and seed them according to the surface area of the well required for the experiment.
- Perform high-resolution imaging after fixing the cells following step 8.
NOTE: A density of 35,000 cells/cm² is recommended for further experiments (protein and total RNA extraction, 330,000 cells [one well of a 6-well plate]; immunofluorescence [IF], 10,500 cells [one well of a 96-well plate]).
5. Adipogenic differentiation
- Incubate the cells with induction medium (DMEM-F12 with 10% FBS, 1% antibiotic antimycotic, pH 7.2, supplemented with 500 nM dexamethasone, 125 nM indomethacin, 0.5 mM 3-isobutyl-1-methylxanthine [IBMX], 5 µM rosiglitazone, 1 nM T3, and 20 nM insulin) (Table 1) for 72 h (day 0 to day 2) (Figure 2).
NOTE: The culture medium must be changed every other day; thus, the first medium change must occur on day 2 after the initial induction. - Change to maintenance medium (DMEM-F12 with 10% FBS, 1% antibiotic antimycotic, pH 7.2, supplemented with 5 µM rosiglitazone, 1 nM T3, and 20 nM insulin) (Table 1) at day 3, and maintain this maintenance medium until day 7 (Figure 2).
NOTE: The maintenance medium must be changed on day 5 and day 7. Small lipid droplets can be visualized by phase contrast microscopy from day 3 of differentiation. Large lipid droplets are evident on day 7 of differentiation. Differentiation procedures longer than 7 days result in cell loss by detachment, likely due to the high buoyancy of lipid-laden cells.
6. Cellular homogenate preparation
- Remove the culture medium at day 0 and day 7 of differentiation, and wash the cells 3 times with sterile PBS.
- After the last cleaning, add 1.3 mL of sterile PBS to each well.
NOTE: Maintain the cells in ice until the cleaning of the last plate. - Harvest the cells mechanically from one well of a 6-well plate (~330,000 cells) with a cell scraper, and put the cellular suspension in a 1.5 mL tube.
- Centrifuge the samples at 2,800 x g for 5 min at 4 °C.
- Remove the PBS with a micropipette, and maintain the tubes on ice.
- Resuspend the cellular pellet in RIPA buffer with protease and phosphatase inhibitors (see the Table of Materials), and incubate for 30 min on ice.
NOTE: To lysate the cellular pellet, 70-80 µL are sufficient. - Centrifuge the samples at 21,000 x g for 15 min at 4 °C.
- Carefully remove the supernatant to a 1.5 mL tube, and discard the pellet.
NOTE: The supernatant represents the cellular homogenate. - Determine the protein concentration of the cellular homogenate using bicinchoninic acid (BCA) for the colorimetric detection and quantitation of the total protein18,19.
7. Western blotting
- Prepare 30 µg of the cellular homogenate (from step 6.8) and add commercially available SDS-PAGE sample loading buffer (see the Table of Materials).
- Resolve in polyacrylamide gel electrophoresis-SDS20 (PAGE-SDS) according to the molecular weight of every protein.
NOTE: Use 10% PAGE-SDS for PPARγ, C/EBPα, PLIN1, CD36, and UCP1, 12% PAGE-SDS for OXPHOS, and 15% PAGE-SDS for TOM20 (see the Table of Materials). - Electrotransfer the proteins onto the PVDF membrane (see the Table of Materials) at 350 mA for 2 h.
NOTE: The ice container must be changed after 1 h. During the entire process, maintain the chamber on ice. - Block the membranes with 5% BSA or fat-free milk in Tris-buffered saline (TBS) 0.1% Tween 20 (TBS-T) for 2 h following the recommendations from the antibodies' datasheets.
- Incubate with the primary antibodies (the antibodies used in this study are detailed in the Table of Materials) overnight at 4 °C.
NOTE: All the antibodies used in this study were diluted to 1:1,000 in blocking solution. - Wash the membranes 3 times with 0.1% TBS-T, and incubate them with horseradish peroxidase (HRP) conjugated secondary antibody (see the Table of Materials) for 2 h at room temperature.
- Wash the membranes 3 times with 0.1%. TBS-T. Perform one last wash with 1x TBS. Store the membranes in 1x TBS until visualization. Storing for a maximum of 1 h is recommended.
- Visualize the membranes by chemiluminescence20 or any other preferred method.
8. High-resolution imaging
- Perform the immunofluorescence assay following the steps below.
- Seed and differentiate the primary cultures of brown preadipocytes from P0.5 newborn mice (following step 4.8) in 96-well plates with an optical bottom (see the Table of Materials).
- Fix the cells at day 0 and day 7 of differentiation with 4% PFA for 20 min, and wash 3 times with sterile PBS.
- Permeabilize the cells with 0.1% Triton X-100 for 15 min, and block them with 3% fish gelatin (see the Table of Materials) for 1 h at room temperature.
- Wash 3 times with sterile PBS, and incubate with PLIN1 antibody (1:400, see the Table of Materials) in 1% BSA/PBS overnight at 4 °C in a humid chamber.
- Wash 3 times with sterile PBS, and incubate with fluorophore-conjugated secondary antibody at 1:1,000 (1% BSA/PBS) (see the Table of Materials) for 1 h at room temperature.
- Stain the nuclei with 1 µg/mL Hoechst 33342 and the lipid droplets with 1 µg/mL BODIPY for 30 min at room temperature.
NOTE: The fluorescence images were acquired using a cell imaging system (see the Table of Materials). Overall, 9 or 16 photos per well were acquired with two to three channels with a magnification of 20x.
- Perform transmission electron microscopy following the steps below.
- Grow and differentiate the brown preadipocytes in 35 mm x 10 mm cell culture dishes.
- On day 0 and day 7 of differentiation, remove the medium, and fix the cells for 1 h with 2.5% glutaraldehyde in 0.1 M sodium cacodylate buffer, pH 7.0 (see the Table of Materials).
- Wash the cells 3 times with 0.05 M sodium cacodylate buffer for 10 min, and treat them with 1% aqueous osmium tetroxide (see the Table of Materials) for 15 min.
- Wash the cells 3 times with bi-distilled water for 5 min, and treat them with 1% aqueous uranyl acetate (see the Table of Materials) for 15 min.
- Dehydrate the cells with an ethanol gradient of 30%, 50%, 70%, 95%, and 100% for 5 min each time.
- Pre-embed the cells in 1:1 epon/ethanol for 30 min and then in epon for 1 h (see the Table of Materials).
NOTE: The inclusion of cells is performed directly in the dish by adding fresh epon to cover the entire cell culture monolayer. Let the epon polymerize in an oven at 60 °C for 24 h. - Immerse the cell culture dish in liquid nitrogen for 2 min, and then fracture it using pliers to loosen the plastic inclusion.
- Glue the inclusion segments with cells to a block of resin to obtain fine cuts (80 nm) parallel to the surface in an ultramicrotome (see the Table of Materials).
- Stain the cuts with 1% aqueous uranyl acetate for 4 min and lead citrate for 5 min, according to Reynolds et al.21.
- Visualize the grids in an electron microscope (see the Table of Materials).
Representative Results
Adipogenesis is regulated by a network of transcription factors that are responsible for both the expression of key proteins that induce brown adipocyte formation and functioning22, including classical adipogenic regulators such as PPARγ and C/EBPα23,24,25, as well as markers of mature adipocytes26,27. Through testing the different concentrations of rosiglitazone that allow the acquisition of the thermogenic brown adipose phenotype, the method described herein allows the differentiation of preadipocytes present in the SVF of iBAT of newborn P0.5 mice to mature brown adipocytes. The addition of 5 µM rosiglitazone in the induction and maintenance medium markedly increased the protein levels of the adipogenic regulators and UCP1 at day 7 of differentiation compared with a lower concentration of this PPARγ agonist (1 nM) (Figure 3). Undifferentiated preadipocytes (day 0) present very low or undetectable levels of PPARγ, C/EBPα, PLIN1, and CD36. In contrast, these markers were significantly higher on day 7 of differentiation (Figure 4A–F). Consistent with the previously published results15, brown preadipocytes accumulated multiple lipid droplets across the differentiation protocol (Figure 5).
The mitochondrial mass increases during brown adipogenesis and is strongly associated with lipid droplet accumulation28. The present protocol markedly increased the mitochondrial mass marker TOM20 (Figure 6) and the OXPHOS complex (Figure 7) protein levels at day 7 of differentiation. UCP1 is highly expressed in mature brown adipocytes and is a bona fide marker of this cell type29. As shown in Figure 6, UCP1 was not detectable in undifferentiated preadipocytes (day 0) but was markedly increased at day 7 of differentiation.
Brown adipogenic differentiation is associated with changes in mitochondrial morphology and their physical association with other organelles15,30. As shown in Figure 8, the mitochondria evolved from an elongated tubular shape at day 0 toward a "rounded" or "bean-like" shape at day 7 (Figure 8A,B). The mitochondrial inner structure was also modified by adipogenesis, resulting in higher density of the parallel-packed cristae. Importantly, the mitochondria were intimately associated with lipid droplets in the differentiated brown adipocytes, so no discernible distance between the outer membrane of the mitochondria and the surfaces of the lipid droplets could be detected, even with high-resolution transmission electron microscopy (Figure 8C,D).
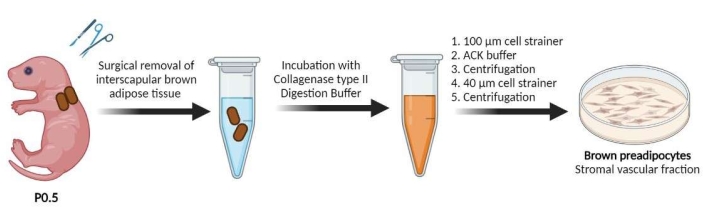
Figure 1: Generation of primary cultured preadipocytes from the SVF of the iBAT. Preadipocyte-containing SVFs were obtained from the interscapular adipose tissue of newborn (P0.5) mice. The interscapular brown adipose tissue was digested with collagenase type II digestion buffer. The blood cells were lysed with ACK buffer. The preadipocytes contained in the SVF were seeded in 24-well plates. Please click here to view a larger version of this figure.
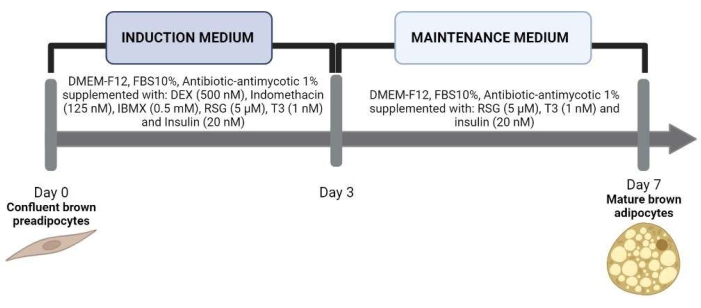
Figure 2: Induction of adipogenesis in primary cultured preadipocytes. Once the SVF cells reached confluency, adipogenesis was induced with the induction medium. On the third day of differentiation, this medium was replaced by a maintenance medium until day 7. Abbreviations: DMEM-F12 = Dulbecco's Modified Eagle Medium/Nutrient Mixture F-12; FBS = fetal bovine serum; DEX = dexamethasone; IBMX = isobutylmethylxanthine; RSG = rosiglitazone; T3 = triiodothyronine. Please click here to view a larger version of this figure.
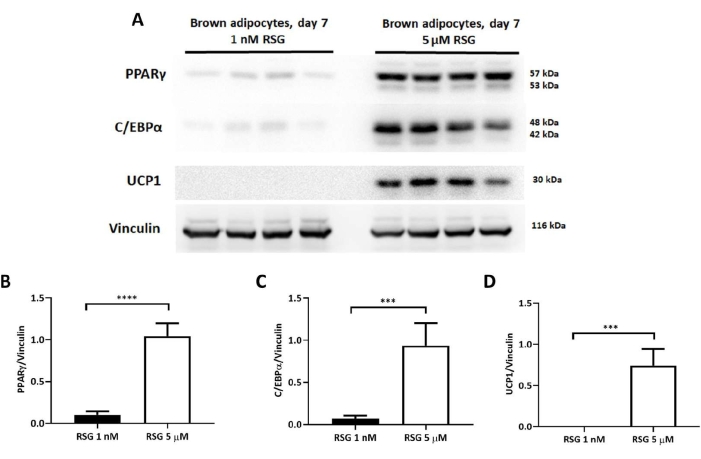
Figure 3: Protein levels of PPARγ, C/EBPα, and UCP1 in differentiated brown adipocytes differentiated with 1 nM or 5 µM rosiglitazone. (A) Representative immunoblot images of adipogenic regulators and UCP1 on day 7 of differentiation. Immunoblot quantification of (B) PPARγ, (C) C/EBPα, and (D) UCP1 protein levels; normalized to vinculin levels (n = 4 per experimental condition). Results expressed as mean ± SD. ***p < 0.001, and ****p < 0.0001 between adipocytes treated with 1 nM versus 5 µM rosiglitazone. The p-values were calculated using a Student's t-test. Please click here to view a larger version of this figure.
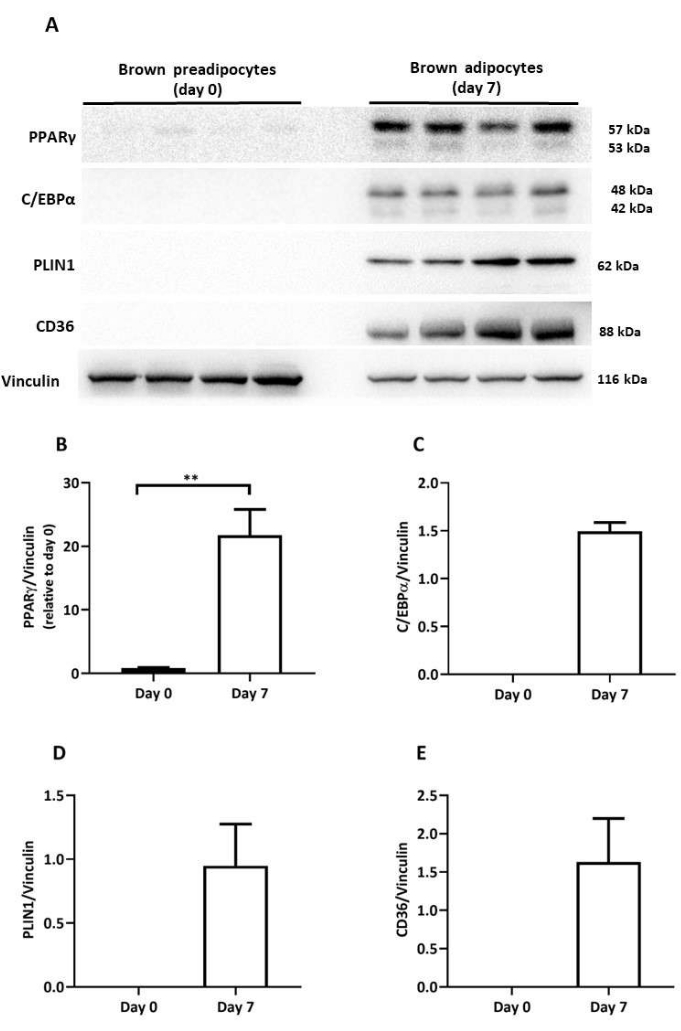
Figure 4: Protein levels of brown adipogenic regulators and mature adipocyte markers in differentiated brown adipocytes. (A) Representative immunoblot images of PPARγ, C/EBPα, PLIN1, and CD36of differentiated brown adipocytes on day 0 and day 7 of differentiation. Immunoblot quantification of (B) PPARγ, (C) C/EBPα, (D) PLIN1, and (E) CD36 protein levels; normalized to vinculin levels (n = 4 per differentiation day). Results expressed as mean ± SD. **p < 0.01 between undifferentiated (day 0) and differentiated brown adipocytes (day 7). The p-values were calculated using a Student's t-test. Please click here to view a larger version of this figure.
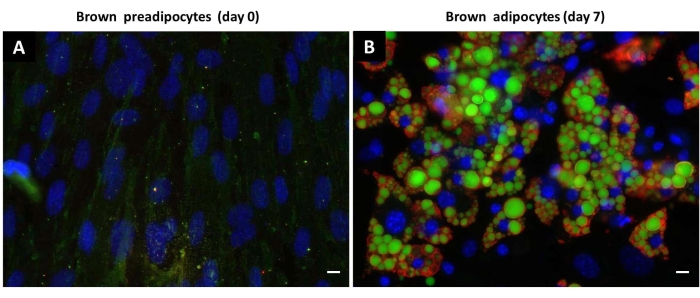
Figure 5: Accumulation of lipid droplets by differentiated brown adipocytes. Representative immunofluorescence images showing the staining of neutral lipids with BODIPY (green) and PLIN1 (red) and the staining of nuclei with Hoechst 33342 (blue) in brown (pre)adipocytes on day 0 and day 7 of differentiation. (A) Brown preadipocytes, day 0 of differentiation. (B) Brown adipocytes, day 7 of differentiation. Scale bar: 20 µm. Please click here to view a larger version of this figure.
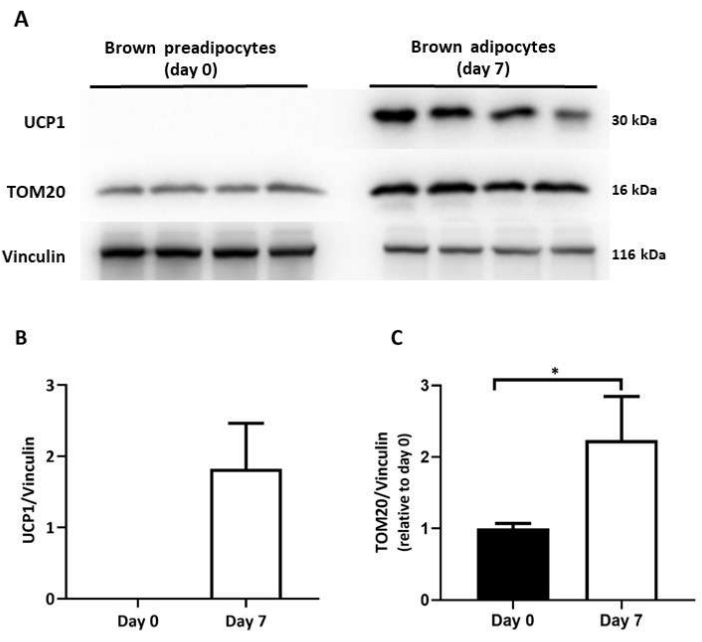
Figure 6: Protein levels of mitochondrial mass and thermogenic markers in differentiated brown adipocytes. (A) Representative immunoblot images of TOM20 and UCP1 on day 0 and day 7 of differentiation. Immunoblot quantification of (B) UCP1 and (C) TOM20 protein levels; normalized to vinculin levels (n = 4 per differentiaion day). Results expressed as mean ± SD. *p < 0.05 between undifferentiated (day 0) and differentiated brown adipocytes (day 7). The p-values were calculated using a Student's t-test. Please click here to view a larger version of this figure.
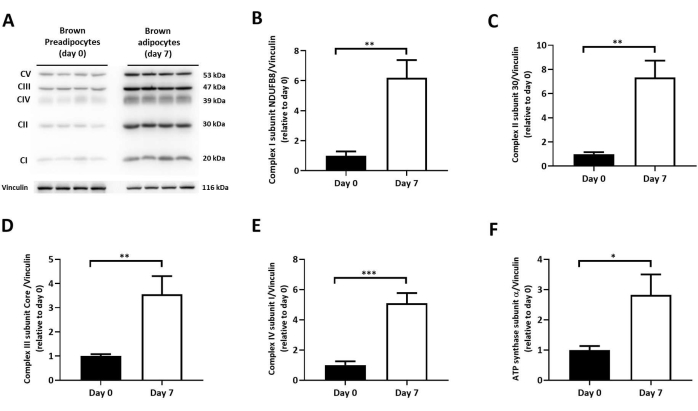
Figure 7: Protein levels of OXPHOS subunits in differentiated brown adipocytes. (A) Representative OXPHOS immunoblotting on day 0 and day 7 of differentiation. Immunoblot quantification of (B) complex I, subunit NDUFB8, (C) complex II, subunit 30, (D) complex III, subunit core, (E) complex IV, subunit I, (F) ATP synthase, subunit α; normalized to vinculin levels (n = 4 per differentiation day). Results expressed as mean ± SD. *p < 0.05, **p < 0.01, and ***p < 0.001 between undifferentiated (day 0) and differentiated brown adipocytes (day 7). The p-values were calculated using a Student's t-test. Please click here to view a larger version of this figure.
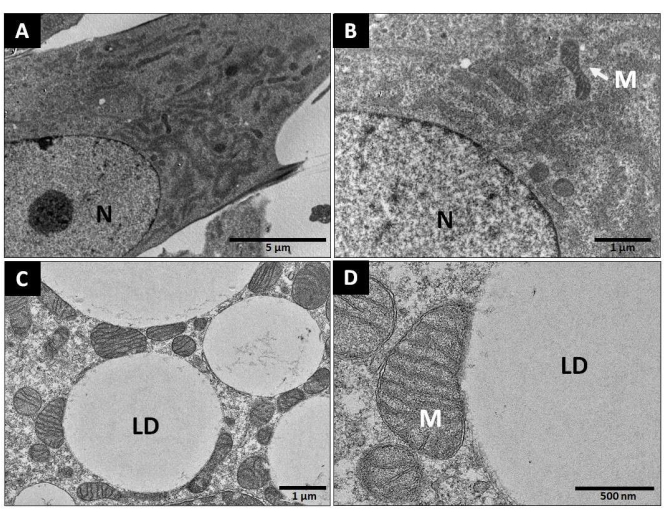
Figure 8: Mitochondrial morphology in differentiated brown adipocytes. Representative images of transmission electron microscopy on day 0 and day 7 of differentiation. (A,B) Brown preadipocytes on day 0 of differentiation; magnification: 4,200x and 16,500x, respectively. (C,D) Brown adipocytes on day 7 of differentiation; magnification: 8,500x and 28,000x, respectively. Abbreviations: LD = lipid droplet; M = mitochondria. Scale bar: (A) = 5 µm; (B,C) = 1 µm; (D) = 500 nm. Please click here to view a larger version of this figure.
Table 1: Buffer and medium preparation. Collagenase type II digestion buffer: Prepared in sterile deionized water, filtered, and aliquoted in a biosafety cabinet. For long-term storage, it is recommended to maintain it at −20 °C and thaw right before use. Culture medium: Adjusted to pH 7.2. For storage, it is recommended to maintain it at 4 °C. Induction and maintenance medium: Prepared in DMEM-F12, 10% FBS, 1% antibiotic-antimycotic, pH 7.2. Both these media must be freshly prepared just before use. Please click here to download this Table.
Supplementary Table 1: Comparison of the differentiation media used in various reported studies. Please click here to download this Table.
Discussion
The present protocol is a simple and replicable two-phase differentiation procedure (Figure 2) for generating cells with the molecular and morphological characteristics of mature brown adipocytes. The surgical harvesting of the iBAT is the first critical step because tissue tearing severely limits the viability of the starting material. Tissue processing is also key because a homogeneous cell suspension that is free of debris greatly increases the amount of SVF that can be cultured. In the current study, filtration was performed twice using 100 µm and 40 µm cell strainers in contrast to other protocols that only used a 100 µm cell strainer11,12,13,14.
For the stromal vascular fraction culture, two rounds of passages were performed to multiply the number of cells available for the experiments. This standardization allowed for increasing the reproducibility of our SVF differentiations. In contrast, another published protocol12 suggested a variable number of passages in the case of a limited number of newborn mice. This practice is not recommended because the differentiation capacity changes with the number of passages. Additional critical considerations are (1) the preadipocyte culture confluence level, which must be 100% at the time of adipogenic induction, and (2) the chemical stability of each compound in the differentiation cocktail once they have been reconstituted.
The use of P0.5 newborn mice is not strictly essential for obtaining primary cultures of brown preadipocytes. However, compared with recently published protocols that used P2.5 pups13,14 or adult mice12, the surgical removal of iBAT from newborn mice is a quick and easy procedure since newborns mostly lack the layer of subcutaneous white adipose tissue that covers the iBAT in older mice. This is important because the inadequate separation of the two types of adipose tissue may lead to contamination of the iBAT SVF preparation by white adipocyte precursors. Additionally, the use of older animals increases the costs of maintaining bigger animal colonies. Finally, the efficiency of in vitro adipocyte differentiation is higher in newborns compared to adult mice, likely because of a higher abundance of preadipocytes in the SVF at early ages (data not shown).
This protocol has delivered consistent results in our laboratory over several years15. It is important to consider that many published protocols have similar qualitative compositions of the induction and maintenance media but differ greatly in the concentrations of these components and the duration of the differentiation11,12,13,14,15 (Supplementary Table 1). Indomethacin, a non-steroidal anti-inflammatory compound, promotes the differentiation of murine brown adipocytes and the expression of UCP1 and PGC1α in a dose-dependent manner31 by binding and activating PPARγ32. In other protocols, indomethacin is present in a higher concentration11,13 than in the current method, but it has also been reported absent in other differentiation media12,14. The PPARγ agonist rosiglitazone increases the degree of differentiation of adipogenic progenitor cells33. Rosiglitazone was used in both the induction and maintenance media because it results in higher mitochondrial biogenesis34 and UCP1 levels35 and promotes adipocyte browning in vitro and in vivo36,37. The addition of rosiglitazone during the induction and maintenance phase increases the lipid content and PPARγ, C/EBPα, and UCP1 expression compared to the standard adipogenic protocol38. Consistent with this previous report38, at the beginning of this research, a lower concentration of rosiglitazone (1 nM) was tested, and this was included only in the induction medium. This concentration was associated with a lower abundance of PPARγ and C/EBPα and undetectable levels of UCP1 compared with the 5 µM rosiglitazone recommended in this protocol.
Immortalized cell lines derived from human and mouse brown adipocytes have been generated and used to study brown adipocyte differentiation and function39,40,41. These offer advantages compared to work with in vitro differentiated brown adipocytes, including in terms of bioethical and practical considerations. Nonetheless, these cell lines must be strictly validated to ensure they retain the cellular phenotype of mature and functional brown adipocytes. Additionally, the use of an SVF derived from the iBAT of genetically modified mice allows for the direct assessment of the effect of these modifications on brown adipogenesis and comparison with the phenotype in living mice. The main limitations of using the SVF from mice compared with immortalized brown preadipocyte cell lines are the time required for generating the animals for processing and the higher overall costs.
In conclusion, the procedures described herein provide tools for a simple and effective protocol for in vitro brown adipocyte differentiation, which can be easily performed on a regular laboratory setup with reliable and reproducible outcomes.
Disclosures
The authors have nothing to disclose.
Acknowledgements
Funding was provided by FONDECYT (1181214 and 1221146) and Anillos (ACT210039) to VC and doctoral scholarships ANID 21171743 to AMF and ANID 21150665 to FS. We thank Alejandro Munizaga for help in the processing of the samples and technical advice for the transmission electron microscopy. The illustrations were produced using BioRender.
Materials
| 10x Tris/Glycine Buffer | BioRad | 1610734 | |
| 16% Paraformaldehyde Aqueous Solution | Electron Microscopy Sciences | 15710 | |
| 35 mm TC-treated Easy-Grip Style Cell Culture Dish | Falcon | 353001 | |
| 3-Isobutyl-1-methylxanthine (IBMX) | Calbiochem | 410957 | |
| 40% Acrylamide/Bis Solution, 37.5:1 | BioRad | 1610148 | |
| 6-well plate | SPL Life Science | – | |
| 96 well optical black w/lid cell culture sterile | Thermo Scientific | 165305 | |
| AccuRuler RGB Plus Pre-stained Protein Ladder | Maestrogen | 02102-250 | |
| ACK lysing buffer | Gibco | A10492-01 | |
| Ammonium Persulfate | BioRad | 1610700 | |
| Antibiotic-antimycotic | Gibco | 15240062 | |
| Anti-mouse IgG, HRP-linked Antibody | Cell Signaling | 7076 | |
| Anti-rabbit IgG, HRP-linked Antibody | Cell Signaling | 7074 | |
| Blotting-grade blocker | BioRad | 170-6404 | |
| BODIPY 493/503 | Invitrogen | D3922 | |
| BSA | Sigma | A1470 | |
| C/EBPα antibody | Cell Signaling | 2295 | |
| CaCl2 | Calbiochem | 208291 | |
| CD36 antibody | Invitrogen | PA1-16813 | |
| Cell Strainer 100 µm, nylon | Falcon | 352360 | |
| Cell Strainer 40 µm, nylon | Falcon | 352340 | |
| Collagenase type II | Gibco | 17101-015 | |
| Cytation 5 Cell Imaging Multimode Reader | Biotek | ||
| Dexamethasone | Sigma | D4902 | |
| DMEM/F-12, powder | Gibco | 12500062 | |
| EMBed-812 EMBEDDING KIT (Epon) | Electron Microscopy Sciences | 14120 | |
| Ethanol absolute | Merck | 100983 | |
| Fetal bovine serum | Gibco | 16000-044 | |
| Gelatin from cold water fish skin | Sigma | G7041 | |
| Glucose | Gibco | 15023-021 | |
| Glutaraldehyde 25% Aqueous Solution | Electron Microscopy Sciences | 16210 | |
| Goat anti-Rabbit IgG (H+L) Cross-Adsorbed Secondary Antibody, Alexa Fluor 594 | Life Technologies | A11012 | |
| Halt Phosphatase Inhibitor Cocktail 100X | Thermo Scientific | 78427 | |
| Halt Protease Inhibitor Cocktail 100X | Thermo Scientific | 78429 | |
| Hoechst 33258 | Invitrogen | H1398 | |
| Immun-Blot PVDF Membrane | BioRad | 1620177 | |
| Indomethacin | Sigma | I7378 | |
| Insulin | Sigma | I3536 | |
| KCl | Calbiochem | 529552 | |
| KH2PO4 | Calbiochem | 529568 | |
| KHCO3 | Sigma | 60339 | |
| Lane Marker Reducing Sample Buffer | Thermo Scientific | 39000 | |
| MgSO4 | Sigma | M2643 | |
| MilliQ water sterile | – | – | |
| Mitoprofile Total OXPHOS Rodent WB antibody Cocktail | Abcam | MS604 | |
| NaCl | Merck | 1064041000 | |
| OmniPur 10x PBS Liquid Concentrate | Calbiochem | 6505-OP | |
| Osmium Tetroxide | Electron Microscopy Sciences | 19100 | |
| Perilipin 1 antibody | Cell Signaling | 9349 | |
| PPARγ (81B8) antibody | Cell Signaling | 2443 | |
| RIPA buffer lysis | Thermo Scientific | 89901 | |
| Rosiglitazone | Merck | 557366 | |
| Sodium bicarbonate | Sigma | S5761 | |
| Sodium Cacpdylate | Electron Microscopy Sciences | 12300 | |
| Sodium Dodecyl Sulfate | BioRad | 1610301 | |
| T3 | Sigma | T6397 | |
| Talos F200C G2 | Thermo Scientific | ||
| TEMED | BioRad | 1610800 | |
| TOM20 antibody | Cell Signaling | 42406 | |
| Tris Buffered Saline (TBS-10X) | Cell Signaling | 12498 | |
| Triton X-100 | Sigma | 93443 | |
| Trypsin-EDTA (0,25%) | Gibco | 25200056 | |
| Tween 20, Molecular Biology Grade | Promega | H5152 | |
| UCP1 antibody | Cell Signaling | 14670 | |
| Ultracut R | Leica | ||
| Uranyl Acetate | Electron Microscopy Sciences | 22400 | |
| Westar Sun | Cyanagen | XLS063 | |
| Westar Supernova | Cyanagen | XLS3 |
References
- Luo, L., Liu, M. Adipose tissue in control of metabolism. Journal of Endocrinology. 231 (3), 77-99 (2016).
- Gaspar, R. C., Pauli, J. R., Shulman, G. I., Muñoz, V. R. An update on brown adipose tissue biology: A discussion of recent findings. American Journal of Physiology-Endocrinology and Metabolism. 320 (3), 488-495 (2021).
- Richard, D., Picard, F. Brown fat biology and thermogenesis. Frontiers in Bioscience. 16, 1233-1260 (2011).
- Rosenwald, M., Wolfrum, C. The origin and definition of brite versus white and classical brown adipocytes. Adipocyte. 3 (1), 4-9 (2014).
- Smorlesi, A., Frontini, A., Giordano, A., Cinti, S. The adipose organ: White-brown adipocyte plasticity and metabolic inflammation. Obesity Reviews. 13, 83-96 (2012).
- Townsend, K., Tseng, Y. -. H. Brown adipose tissue: Recent insights into development, metabolic function and therapeutic potential. Adipocyte. 1 (1), 13-24 (2012).
- Benador, I. Y., et al. Mitochondria bound to lipid droplets have unique bioenergetics, composition, and dynamics that support lipid droplet expansion. Cell Metabolism. 27 (4), 869-885 (2018).
- Benador, I. Y., Veliova, M., Liesa, M., Shirihai, O. S. Mitochondria bound to lipid droplets: Where mitochondrial dynamics regulate lipid storage and utilization. Cell Metabolism. 29 (4), 827-835 (2019).
- Lee, P., Swarbrick, M. M., Ho, K. K. Brown adipose tissue in adult humans: A metabolic renaissance. Endocrine Reviews. 34 (3), 413-438 (2013).
- Han, S., Sun, H. M., Hwang, K. C., Kim, S. W. Adipose-derived stromal vascular fraction cells: Update on clinical utility and efficacy. Critical Reviews in Eukaryotic Gene Expression. 25 (2), 145-152 (2015).
- Klein, J., et al. Beta(3)-adrenergic stimulation differentially inhibits insulin signaling and decreases insulin-induced glucose uptake in brown adipocytes. Journal of Biological Chemistry. 274 (3), 34795-34802 (1999).
- Gaom, W., Kong, X., Yang, Q. Isolation, primary culture, and differentiation of preadipocytes from mouse brown adipose tissue. Methods in Molecular Biology. 1566, 3-8 (2017).
- Rocha, A. L., Guerra, B. A., Boucher, J., Mori, M. A. A Method to induce brown/beige adipocyte differentiation from murine preadipocytes. Bio-protocol. 11 (24), 4265 (2021).
- Galmozzi, A., Kok, B. P., Saez, E. Isolation and differentiation of primary white and brown preadipocytes from newborn mice. Journal of Visualized Experiments. (167), e62005 (2021).
- Tapia, P. J., et al. Absence of AGPAT2 impairs brown adipogenesis, increases IFN stimulated gene expression and alters mitochondrial morphology. Metabolism. 111, 154341 (2020).
- González-Hódar, L., et al. Decreased caveolae in AGPAT2 lacking adipocytes is independent of changes in cholesterol or sphingolipid levels: A whole cell and plasma membrane lipidomic analysis of adipogenesis. Biochimica et Biophysica Acta – Molecular Basis of Disease. 1867 (9), 166167 (2021).
- Fernández-Galilea, M., Tapia, P., Cautivo, K., Morselli, E., Cortés, V. A. AGPAT2 deficiency impairs adipogenic differentiation in primary cultured preadipocytes in a non-autophagy or apoptosis dependent mechanism. Biochemical and Biophysical Research Communications. 467 (1), 39-45 (2015).
- Walker, J. M. The bicinchoninic acid (BCA) assay for protein quantitation. The Protein Protocols Handbook. , 11-15 (2009).
- Smith, P. K., et al. Measurement of protein using bicinchoninic acid. Analytical Biochemistry. 150 (1), 76-85 (1985).
- Kurien, B. T., Scofield, R. H. Western blotting. Methods. 38 (4), 283-293 (2006).
- Reynolds, E. S. The use of lead citrate at high pH as an electron-opaque stain in electron microscopy. Journal of Cell Biology. 17 (1), 208-212 (1963).
- Ghaben, A. L., Scherer, P. E. Adipogenesis and metabolic health. Nature Reviews Molecular Cell Biology. 20 (4), 242-258 (2019).
- Ambele, M. A., Dhanraj, P., Giles, R., Pepper, M. S. Adipogenesis: A complex interplay of multiple molecular determinants and pathways. International Journal of Molecular Sciences. 21 (12), 4283 (2020).
- Lefterova, M. I., et al. PPARγ and C/EBP factors orchestrate adipocyte biology via adjacent binding on a genome-wide scale. Genes & Development. 22 (21), 2941-2952 (2008).
- Ntambi, J. M., Young-Cheul, K. Adipocyte differentiation and gene expression. The Journal of Nutrition. 130 (12), (2000).
- Itabe, H., Yamaguchi, T., Nimura, S., Sasabe, N. Perilipins: A diversity of intracellular lipid droplet proteins. Lipids in Health and Disease. 16, 83 (2017).
- Christiaens, V., Van Hul, M., Lijnen, H. R., Scroyen, I. CD36 promotes adipocyte differentiation and adipogenesis. Biochimica et Biophysica Acta – General Subjects. 1820 (7), 949-956 (2012).
- Carobbio, S., Rosen, B., Vidal-Puig, A. Adipogenesis: New insights into brown adipose tissue differentiation. Journal of Molecular Endocrinology. 51 (3), 75-85 (2013).
- Ricquier, D. Uncoupling protein 1 of brown adipocytes, the only uncoupler: A historical perspective. Frontiers in Endocrinology. 2, 85-91 (2011).
- Cui, L., Mirza, A. H., Zhang, S., Liang, B., Liu, P. Lipid droplets and mitochondria are anchored during brown adipocyte differentiation. Protein & Cell. 10 (12), 921-926 (2019).
- Hao, L., et al. Indomethacin enhances brown fat activity. Journal of Pharmacology and Experimental Therapeutics. 365 (3), 467-475 (2018).
- Lehmann, J. M., Lenhard, J. M., Oliver, B. B., Ringold, G. M., Kliewer, S. A. Peroxisome proliferator-activated receptors alpha and gamma are activated by indomethacin and other non-steroidal anti-inflammatory drugs. Journal of Biological Chemistry. 272 (6), 3406-3410 (1997).
- Scott, M. A., Nguyen, V. T., Levi, B., James, A. W. Current methods of adipogenic differentiation of mesenchymal stem cells. Stem Cells and Development. 20 (10), 1793-1804 (2011).
- Wilson-Fritch, L., et al. Mitochondrial remodeling in adipose tissue associated with obesity and treatment with rosiglitazone. The Journal of Clinical Investigation. 114 (9), 1281-1289 (2004).
- Cannon, B., Nedergaard, J. Brown adipose tissue: Function and physiological significance. Physiological Reviews. 84 (1), 277-359 (2004).
- Ohno, H., Shinoda, K., Spiegelman, B. M., Kajimura, S. PPARgamma agonists induce a white-to-brown fat conversion through stabilization of PRDM16 protein. Cell Metabolism. 15 (3), 395-404 (2012).
- Merlin, J., et al. Rosiglitazone and a β3-adrenoceptor agonist are both required for functional browning of white adipocytes in culture. Frontiers in Endocrinology. 9, 249 (2018).
- Fayyad, A. M., et al. Rosiglitazone enhances browning adipocytes in association with MAPK and PI3-K pathways during the differentiation of telomerase-transformed mesenchymal stromal cells into adipocytes. International Journal of Molecular Sciences. 20 (7), 1618-1633 (2019).
- Zennaro, M. -. C., et al. Hibernoma development in transgenic mice identifies brown adipose tissue as a novel target of aldosterone action. The Journal of Clinical Investigation. 101 (6), 1254-1260 (1998).
- Klaus, S., et al. Characterization of the novel brown adipocyte cell line HIB 1B. Adrenergic pathways involved in regulation of uncoupling protein gene expression. Journal of Cell Science. 107 (1), 313-319 (1994).
- Guennoun, A., et al. Comprehensive molecular characterization of human adipocytes reveals a transient brown phenotype. Journal of Translational Medicine. 13, 135 (2015).

.
