Complementary Approaches to Interrogate Mitophagy Flux in Pancreatic β-Cells
Instructor Prep
concepts
Student Protocol
The animal studies presented in this protocol were reviewed and approved by the University of Michigan Institutional Animal Care and Use Committee. Twenty-week-old male C57BL/6J mice, on either a 15-week regular fat diet (RFD) or high-fat diet (HFD), were used for this study.
1. Assessing mitophagy via the dye-based MtPhagy approach (Method 1)
- Mouse islet preparation and treatment
- Perform islet culture and valinomycin exposure following the steps below.
- Isolate islets from either regular fat diet (RFD) or high-fat diet (HFD, 60 kcal% Fat19) mice, following the previously described methods2,10.
- Culture islet samples overnight at 37 °C in RPMI medium supplemented with 100 units/mL antimycotic-antibiotic, 50 units/mL penicillin-streptomycin, 1 mM sodium pyruvate, 10 mM HEPES and 10% fetal bovine serum (FBS) (see Table of Materials). This medium will be referred to as "islet medium".
- Using a pipette, pick 100 islets per condition into 6 cm Petri dishes in 2 mL of islet medium.
NOTE: Conditions used for this protocol: (1) Unstained control: unstained RFD islets; (2) DAPI only control: RFD islets stained with DAPI; (3) Fluozin-3-AM only control: RFD islets stained with Fluozin-3-AM; (4) MtPhagy only control: RFD islets stained with MtPhagy; (5) TMRE only control: RFD islets stained with TMRE; (6) RFD untreated: RFD islets stained with MtPhagy, TMRE, Fluozin-3-AM, and DAPI; (7) Valinomycin-exposed RFD islets: RFD islets exposed to valinomycin and stained with MtPhagy, TMRE, Fluozin-3-AM, and DAPI; (8) HFD untreated: HFD islets stained with MtPhagy, TMRE, Fluozin-3-AM, and DAPI; (9) Valinomycin-exposed HFD islets: HFD islets exposed to valinomycin and stained with MtPhagy, TMRE, Fluozin-3-AM, and DAPI. - For conditions (7) and (9) (see NOTE above) exposed to valinomycin, add 2 µL of 250 nΜ valinomycin stock (see Table of Materials) to islets in the Petri dish for 3 h to induce mitophagy.
- Perform single cell dissociation.
- After 3 h valinomycin exposure, pick 100 islets from each condition into separate microcentrifuge tubes using a pipette.
- Spin islets at 350 x g for 1 min at 10 °C and discard the supernatant using a pipette.
- Wash samples 2x with 1 mL 1x phosphate buffered saline (PBS), with spin steps (350 x g/1 min, 10 °C) between each wash. Discard supernatant with a pipette after each wash.
- To dissociate islets into single cells, add 500 µL of 0.05% trypsin (pre-warmed to 37 °C) to the microcentrifuge tube of one sample to prevent over-digestion of islets. Pipette up and down repeatedly until islets are visibly dispersed.
- Immediately add 1 mL of pre-warmed islet medium to neutralize trypsin. Repeat trypsinization and neutralization for each sample, one at a time.
- Spin samples at 500 x g for 5 min at 10 °C. Remove supernatant with a pipette, careful not to disturb the pellet.
NOTE: Pellet will be delicate for samples exposed to valinomycin. - Wash samples 2x with RPMI medium, no phenol red, supplemented with 100 units/mL antimycotic-antibiotic, 50 units/mL penicillin-streptomycin, 1 mM sodium pyruvate, 10 mM HEPES, and 10% bovine serum albumin (BSA). This medium will be referred to as "islet flow medium". Include spin steps (350 x g/1 min, 10 °C) between each wash. Discard supernatant with a pipette after each wash.
- Stain cells with MtPhagy, TMRE, Fluozin-3-AM, and DAPI to prepare for flow cytometry.
- Resuspend islet samples in 500 µL of islet flow medium.
- Add 0.25 µL of 100 µM MtPhagy stock (see Table of Materials) to tubes receiving MtPhagy dye (conditions 4, 6, 7, 8, and 9, NOTE to step 1.1.1).
- Add 0.25 µL of 100 µM TMRE stock (see Table of Materials) to tubes receiving TMRE dye (conditions 5, 6, 7, 8, and 9).
- Add 0.25 µL of 1 mM Fluozin-3-AM stock (see Table of Materials) to tubes receiving Fluozin-3-AM dye (conditions 3, 6, 7, 8, and 9).
- Vortex tubes at low speed for 5 s to mix.
- Wrap microcentrifuge tubes in aluminum foil to protect from light and incubate at 37 °C for 30 min. Halfway through incubation, vortex tubes at low speed for 5-10 s to mix.
- After incubation, centrifuge samples at 350 x g for 1 min at 10 °C. Discard supernatant using a pipette.
- Resuspend samples in 500 µL islet flow medium. Add 0.2 µg/mL DAPI (see Table of Materials) to conditions 2, 6, 7, 8, and 9.
- Spin samples at 350 x g for 1 min at 10 °C. Discard supernatant using a pipette and resuspend in 500 µL of islet flow medium.
- Place samples on ice.
- Perform islet culture and valinomycin exposure following the steps below.
- Flow cytometry
- Startup the instrument (see Table of Materials).
NOTE: Any flow cytometer with the appropriate filters will work. The following filters were used in this study: (1) VL1 for DAPI: Excitation/Emission – 405 nm/440 nm (50 nm); (2) BL1 for Fluozin-3-AM: Excitation/Emission – 488 nm/530 nm (30 nm); (3) BL2 for MtPhagy dye: Excitation/Emission – 488 nm/590 nm (40 nm); (4) YL1 for TMRE: Excitation/Emission – 561 nm/585 nm (16 nm). - Adjust the voltages for forward (FSC) and side scatter (SSC) to ensure cell populations are at the center of the scatter plot. For this protocol, voltages used were 120 V for FSC and 260 V for SSC to ensure cells fall evenly within the FSC-A vs. SSC-A plot (Figure 1A).
- To exclude non-single cells, add a rectangular gate on FSC-H vs. FSC-W followed by SSC-H vs. SSC-W (Figure 1B,C).
- Adjust voltages and compensation for DAPI to filter for live β-cells. Set fluorescence gates for each fluorophore used based on the unstained RFD sample (Figures 1D-F).
NOTE: Voltages used for each channel: (1) VL1: 320-360 V; (2) BL1: 300-340 V; (3) BL2: 260-300 V; (4) YL1: 300-340 V. - Once gates are established, collect 10,000 events per sample.
NOTE: Utilizing this gating approach, mitophagy levels under basal conditions and upon valinomycin induction were assessed in both RFD and HFD islets (Figure 2). - Save data as FCS files for analysis.
- Startup the instrument (see Table of Materials).
2. Assessing mitophagy using the genetically encoded mt-Keima reporter (Method 2)
- Mouse islet preparation and single-cell dissociation
- Perform islet culture and valinomycin exposure following the steps below.
- Isolate pancreatic islets from wild-type (WT) and mt-Keima/+ (mt-Keima) mice6. In this method, 20-week-old male WT or mt-Keima/+ fed RFD mice were used.
- Culture islet samples overnight at 37 °C in islet medium.
- Using a pipette, pick 100 islets per condition into 6 cm Petri dishes with 2 mL of islet medium.
NOTE: Conditions used for this protocol: (1) Unstained control: Unstained WT islets; (2) DAPI only control: WT islets stained with DAPI; (3) Fluozin-3-AM only control: WT islets stained with Fluozin-3-AM; (4) mt-Keima only control: Unstained mt-Keima/+ islets; (5) Untreated: mt-Keima/+ islets stained with Fluozin-3-AM and DAPI; (6) Valinomycin: mt-Keima/+ islets exposed to valinomycin and stained with Fluozin-3-AM and DAPI. - For condition (6) with valinomycin exposure, add 2 µL of 250 nM valinomycin stock to islets in Petri dish for 3 h to induce mitophagy.
- Perform single-cell dissociation.
- Perform single cell dissociation steps, as described in step 1.1.2.
- Stain cells with Fluozin-3-AM and DAPI.
- Resuspend islet samples in 500 µL of islet flow medium.
- Add 0.25 µL of 1 mM Fluozin-3-AM stock to tubes receiving Fluozin-3-AM dye (conditions 3, 5, and 6).
- Incubate cells at 37 °C and proceed to DAPI treatment, as described in steps 1.1.3.5-1.3.10.
- Perform islet culture and valinomycin exposure following the steps below.
- Flow cytometry
- Startup the instrument. Any flow cytometer with the appropriate filters will work.
NOTE: The following filters were used in this study: (1) VL1 for DAPI: Excitation/Emission – 405 nm/440 nm (50 nm); (2) BL1 for Fluozin-3-AM: Excitation/Emission – 488 nm/530 nm (30 nm); (3) VL3 for mt-Keima neutral: Excitation/Emission – 405 nm/603 nm (48 nm); (4) YL2 for mt-Keima acid: Excitation/Emission – 561 nm/620 nm (15 nm). - Adjust FSC and SSC voltages and exclude non-single cells as described in steps 1.2.2-1.2.3 (Figures 3A-C).
- Adjust voltages and compensation for DAPI, Fluozin-3-AM and mt-Keima using unstained and single-positive controls (conditions 1-4) to ensure that fluorescence-positive cell populations are distinguishable from unstained cells. Once appropriate compensation has been applied to each channel, set up DAPI negative gate and Fluozin-3-AM positive gates to filter for live β-cells (Figures 3D-E).
- Set up a triangle gating scheme using the mt-Keima positive sample (condition 4) to identify acidic and neutral cell populations (Figure 3F).
NOTE: Voltages typically used for each channel: (1) VL1: 320-340 V; (2) BL2: 260-280 V; (3) VL3: 280-300 V; (4) YL2: 280-300 V. - Once gates are established, collect 10,000 events per sample. Gating schemes for conditions 5 and 6 are illustrated in Figure 3A-G.
- Save data as FCS files for analysis.
- Startup the instrument. Any flow cytometer with the appropriate filters will work.
Complementary Approaches to Interrogate Mitophagy Flux in Pancreatic β-Cells
Learning Objectives
Assessing mitophagy via the dye-based MtPhagy approach
This dye-based approach was optimized to analyze mitophagy flux within primary mouse β-cells without the need for a genetic reporter, using Fluozin-3-AM, TMRE, and MtPhagy as well as DAPI to exclude dead cells. By pairing these dyes with valinomycin to induce mitophagy, this protocol outlines a dye-based method to selectively measure mitophagy flux in primary mouse β-cells18. For the data shown using this MtPhagy method, both basal and valinomycin-induced mitophagy were analyzed in islets isolated from either regular fat diet (RFD) or high fat diet (HFD, 60 kcal% Fat) fed mice to assess the effect of metabolic stress on mitophagy flux. To identify the population of interest, cells were gated using untreated RFD islets. FSC and SSC voltages were first adjusted to attain an even distribution of cells on a SSC-A vs. FSC-A plot (Figure 1A). To select for single cells, both FSC-H vs. FSC-W and SSC-H vs. SSC-W plots were used, where multiplets were excluded due to their higher width signal values compared to single cells (Figure 1B,C). Next, DAPI-negative cells were selected to exclude dead cells20 (Figure 1D). After establishing primary gates, single stained controls were utilized to establish fluorescence gates for Fluozin-3-AM, MtPhagy, and TMRE (Figure 1E–G) as well as compensation controls for multi-color fluorescence flow cytometry.
Once these primary and fluorescence gates were established, β-cells with high utilization of mitophagy were defined as the FluozinhighMtPhagyhighTMRElow population in quadrant 3 (Q3) using RFD without valinomycin exposure (Figure 1H). Using this gating strategy, basal and valinomycin-induced mitophagy levels were characterized in both RFD and HFD islets (Figure 2). To quantify mitophagy flux, basal vs. valinomycin-induced mitophagy levels were compared using the following ratio:
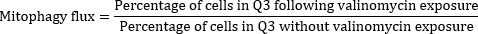
Using this ratio, mitophagy flux was quantified and compared in RFD vs. HFD β-cells to assess differences in mitophagy following the induction of obesity and peripheral insulin resistance. Quantification of mitophagy flux in RFD vs. HFD samples is shown in Figure 2E. This result highlights the feasibility of this assay to quantify mitophagy in β-cells using a straightforward dye-based approach. This method can also be employed in human islets, difficult-to-transfect cells, and islets isolated from complex genetic models where intercrossing to the mt-Keima transgenic model would be cumbersome.
Assessing mitophagy using the genetically encoded mt-Keima reporter
Mt-Keima is a dual excitation fluorescent protein fused with a Cox8-localization sequence that enables its targeting to the inner mitochondrial membrane. The bimodal fluorescent property of mt-Keima allows it to switch its excitation spectra from the neutral (405 nm) to acidic (561 nm) wavelength, depending on the pH of the intracellular compartment6. This enables a robust ratiometric fluorescence analysis of mitophagy, where an increase in acidic-to-neutral ratio indicates mitophagy induction. In this protocol, Fluozin-3-AM was also used to select for β-cells via flow cytometry. In these representative studies, mitophagy flux was assessed using islets isolated from mice fed a RFD diet10,11. FSC and SSC voltages were first adjusted to attain an even distribution of cells on a SSC-A vs. FSC-A plot (Figure 3A). To select for single cells, both FSC-H vs. FSC-W and SSC-H vs. SSC-W plots were used, where multiplets were excluded due to their higher width signal values compared to single cells (Figure 3B,C). The voltage and gating strategy for DAPI and Fluozin-3-AM were determined using single-stained islets (Figure 3D,E). Triangle gates for the acidic and neutral populations were then identified using the mt-Keima positive sample without valinomycin exposure (Figure 3F).
Once these primary and fluorescence gates were established, mitophagy flux was assessed using basal and valinomycin-induced changes in mt-Keima fluorescence (Figure 3F,G). To quantify mitophagy flux, basal mitophagy vs. valinomycin-induced levels were compared using the following ratio:

Using this ratio, mitophagy flux was quantified in RFD cells. Quantitation of this result is shown in Figure 3H. Importantly, these results are comparable to the results in RFD islets generated using the MtPhagy approach (Figure 3H).
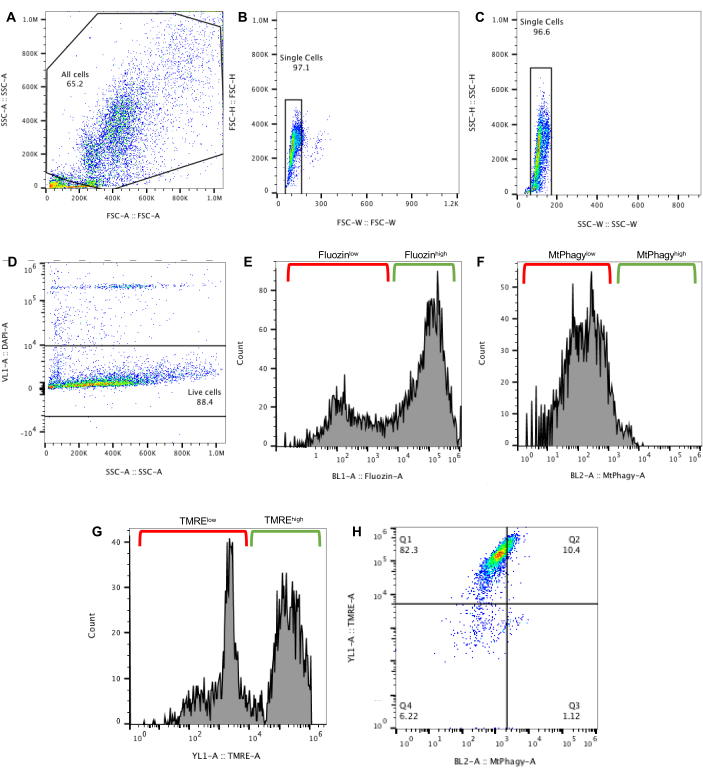
Figure 1: Gating scheme for the MtPhagy method. (A) Flow plot displaying gating scheme to select for all cells. (B) Gating to select for singlets based on FSC-H vs. FSC-W and (C) SSC-H vs. SSC-W. (D) Gating for DAPI–negative cells to exclude dead cells. (E) Gating for Fluozin-3-AMhigh cells to select for β-cells. (F) Gating scheme for MtPhagy dye to identify MtPhagyhigh and MtPhagylow cell populations. (G) Gating scheme for TMRE to identify TMREhigh and TMRElow cell populations. (H) Quadrant gating scheme established with untreated RFD islets to identify FluozinhighMtPhagyhighTMRElow cells in quadrant 3 (Q3) as β-cells undergoing mitophagy. Please click here to view a larger version of this figure.
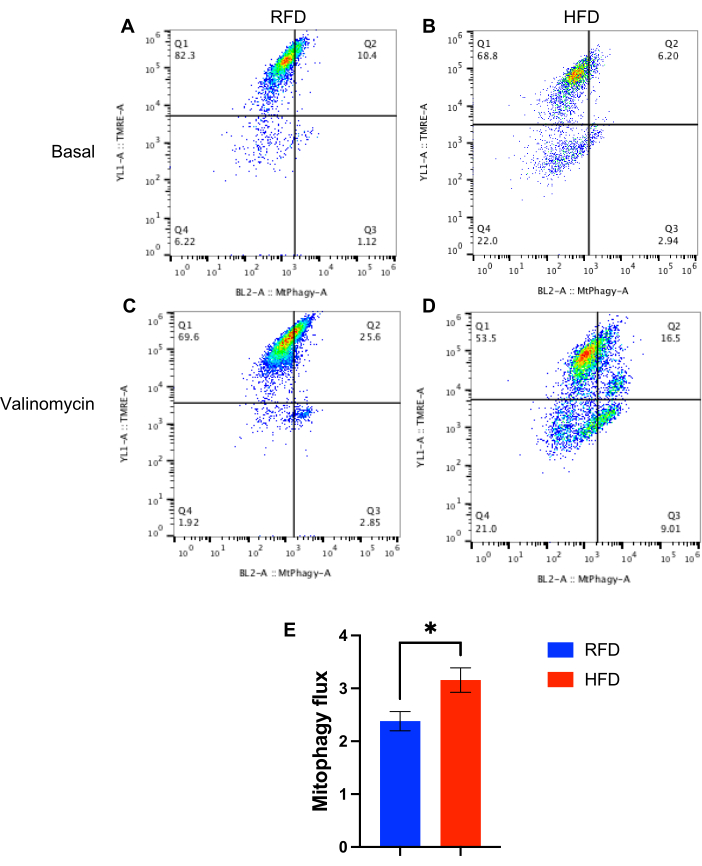
Figure 2: Assessing mitophagy flux differences in mouse β-cells following metabolic stress using the MtPhagy gating scheme. Representative flow cytometry plots of (A) untreated RFD β-cells, (B) untreated HFD β-cells, (C) valinomycin-exposed RFD β-cells, and (D) valinomycin-exposed HFD β-cells. (E) Quantification of mitophagy flux in β-cells, calculated using a ratio of the MtPhagyhighTMRElow cells exposed to valinomycin to the MtPhagyhighTMRElow cells not exposed to valinomycin, for both RFD and HFD samples. *p < 0.05 by Student's unpaired t-test. n = 3/group. Please click here to view a larger version of this figure.
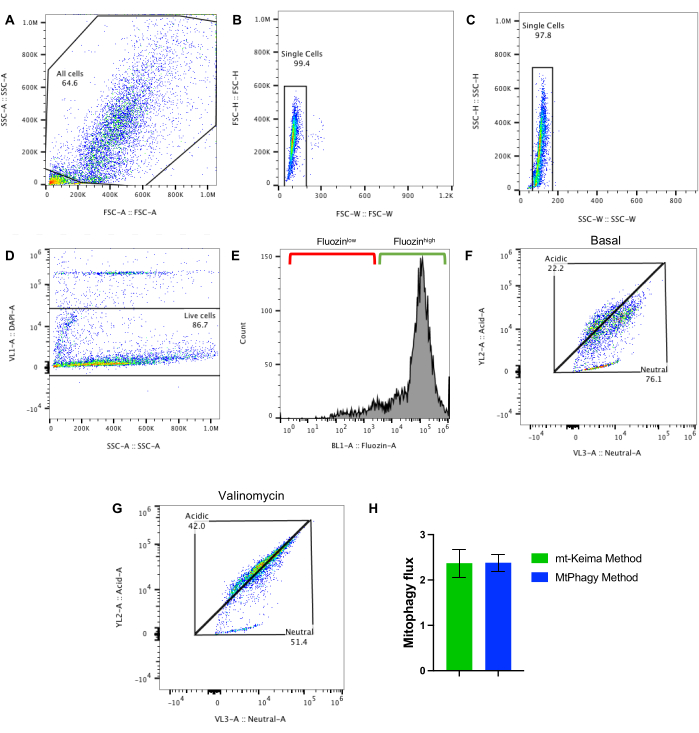
Figure 3: Gating scheme for the mt-Keima method and comparison between both methods. (A) Flow plot displaying gating scheme to select for all cells. (B) Gating to select for singlets based on FSC-H vs. FSC-W and (C) SSC-H vs. SSC-W. (D) Gating for DAPI–negative cells to exclude dead cells. (E) Gating for Fluozin-3-AMhigh cells to select for β-cells. (F) Representative flow cytometry plots of mt-Keima/+ untreated cells and (G) mt-Keima/+ valinomycin-exposed cells. (H) Quantification of mitophagy flux in β-cells from RFD-fed mice, calculated using a ratio of the acidic/neutral cells exposed to valinomycin to ratio of the acidic/neutral cells not exposed to valinomycin using the mt-Keima method and compared to the MtPhagy method (data for MtPhagy protocol originally shown in Figure 2E). n = 3/group. Please click here to view a larger version of this figure.
List of Materials
| Antibiotic-Antimycotic | Life Technologies | 15240-062 | |
| Attune NxT Flow Cytometer | Thermofisher Scientific | A24858 | |
| DAPI (4',6-Diamidino-2-Phenylindole, Dihydrochloride) | Thermofisher Scientific | D1306 | DAPI reconstituted in ddH2O to reach 0.2 µg/mL stock |
| Dimethyl Sulfoxide | Sigma-Aldrich | 317275 | |
| Fatty Acid Free heat shock BSA powder | Equitech | BAH66 | |
| Fetal bovine serum | Gemini Bio | 900-108 | |
| Fluozin-3AM | Thermofisher Scientific | F24195 | 100 μg Fluozin-3AM powder reconstituted in 51 μL DMSO and 51 μL Pluronic F-127 to reach 1 mM stock. |
| Gibco RPMI 1640 Medium | Fisher Scientific | 11-875-093 | |
| HEPES (1M) | Life Technologies | 15630-080 | |
| MtPhagy dye | Dojindo | MT02-10 | 5 μg MtPhagy powder reconstituted with 50 μL DMSO to reach 100 μM stock. |
| MtPhagy dye | Dojindo | MT02-10 | |
| Penicillin-Streptomycin (100x) | Life Technologies | 15140-122 | 1x Solution used in procotol by diluting 1:10 in ddH2O |
| Phosphate buffered saline, 10x | Fisher Scientific | BP399-20 | 1x Solution used in procotol by diluting 1:10 in ddH2O |
| Sodium Pyruvate (100x) | Life Technologies | 11360-070 | 5 μg MtPhagy powder reconstituted with 50 μL DMSO to reach 100 μM stock. |
| TMRE [Tetramethylrhodamine, ethyl ester, perchlorate] | Anaspec | AS-88061 | TMRE powder reconstituted in DMSO to reach 100 μM stock. |
| Trypsin-EDTA (0.05%), phenol red | Thermofisher Scientific | 25300054 | |
| Valinomycin | Sigma | V0627 | Valinomycin powder reconsituted in DMSO to reach 250 nM stock. |
Lab Prep
Mitophagy is a quality control mechanism necessary to maintain optimal mitochondrial function. Dysfunctional β-cell mitophagy results in insufficient insulin release. Advanced quantitative assessments of mitophagy often require the use of genetic reporters. The mt-Keima mouse model, which expresses a mitochondria-targeted pH-sensitive dual-excitation ratiometric probe for quantifying mitophagy via flow cytometry, has been optimized in β-cells. The ratio of acidic-to-neutral mt-Keima wavelength emissions can be used to robustly quantify mitophagy. However, using genetic mitophagy reporters can be challenging when working with complex genetic mouse models or difficult-to-transfect cells, such as primary human islets. This protocol describes a novel complementary dye-based method to quantify β-cell mitophagy in primary islets using MtPhagy. MtPhagy is a pH-sensitive, cell-permeable dye that accumulates in the mitochondria and increases its fluorescence intensity when mitochondria are in low pH environments, such as lysosomes during mitophagy. By combining the MtPhagy dye with Fluozin-3-AM, a Zn2+ indicator that selects for β-cells, and Tetramethylrhodamine, ethyl ester (TMRE) to assess mitochondrial membrane potential, mitophagy flux can be quantified specifically in β-cells via flow cytometry. These two approaches are highly complementary, allowing for flexibility and precision in assessing mitochondrial quality control in numerous β-cell models.
Mitophagy is a quality control mechanism necessary to maintain optimal mitochondrial function. Dysfunctional β-cell mitophagy results in insufficient insulin release. Advanced quantitative assessments of mitophagy often require the use of genetic reporters. The mt-Keima mouse model, which expresses a mitochondria-targeted pH-sensitive dual-excitation ratiometric probe for quantifying mitophagy via flow cytometry, has been optimized in β-cells. The ratio of acidic-to-neutral mt-Keima wavelength emissions can be used to robustly quantify mitophagy. However, using genetic mitophagy reporters can be challenging when working with complex genetic mouse models or difficult-to-transfect cells, such as primary human islets. This protocol describes a novel complementary dye-based method to quantify β-cell mitophagy in primary islets using MtPhagy. MtPhagy is a pH-sensitive, cell-permeable dye that accumulates in the mitochondria and increases its fluorescence intensity when mitochondria are in low pH environments, such as lysosomes during mitophagy. By combining the MtPhagy dye with Fluozin-3-AM, a Zn2+ indicator that selects for β-cells, and Tetramethylrhodamine, ethyl ester (TMRE) to assess mitochondrial membrane potential, mitophagy flux can be quantified specifically in β-cells via flow cytometry. These two approaches are highly complementary, allowing for flexibility and precision in assessing mitochondrial quality control in numerous β-cell models.
Procedure
Mitophagy is a quality control mechanism necessary to maintain optimal mitochondrial function. Dysfunctional β-cell mitophagy results in insufficient insulin release. Advanced quantitative assessments of mitophagy often require the use of genetic reporters. The mt-Keima mouse model, which expresses a mitochondria-targeted pH-sensitive dual-excitation ratiometric probe for quantifying mitophagy via flow cytometry, has been optimized in β-cells. The ratio of acidic-to-neutral mt-Keima wavelength emissions can be used to robustly quantify mitophagy. However, using genetic mitophagy reporters can be challenging when working with complex genetic mouse models or difficult-to-transfect cells, such as primary human islets. This protocol describes a novel complementary dye-based method to quantify β-cell mitophagy in primary islets using MtPhagy. MtPhagy is a pH-sensitive, cell-permeable dye that accumulates in the mitochondria and increases its fluorescence intensity when mitochondria are in low pH environments, such as lysosomes during mitophagy. By combining the MtPhagy dye with Fluozin-3-AM, a Zn2+ indicator that selects for β-cells, and Tetramethylrhodamine, ethyl ester (TMRE) to assess mitochondrial membrane potential, mitophagy flux can be quantified specifically in β-cells via flow cytometry. These two approaches are highly complementary, allowing for flexibility and precision in assessing mitochondrial quality control in numerous β-cell models.
