Computer Vision-Based Biomass Estimation for Invasive Plants
Instructor Prep
concepts
Student Protocol
1. Preparation of datasets
- Selecting the research object
- Select test samples based on the focus of the experimental study, considering options like Mikania micrantha or other invasive plants.
- Collecting UAV images
- Prepare square plastic frames of size 0.5 m*0.5 m and quantity 25-50, depending on the size of the area studied.
- Employ a random sampling approach to determine soil sampling locations in the study area using a sufficient number of biomass samples. Position the sample frame horizontally over the vegetation, fully encompassing the plants with a minimum separation distance of 2 m between each plant.
- Use a drone and a camera to form a UAV remote-sensing filming system, as shown in Figure 1.
- Use the UAV to plot the route within the specified study area. The route planning setup is shown in Figure 2.
- Establish a heading and side overlap rate of 70%, capture photos at uniform time intervals of 2 s, maintain the camera angle perpendicular to the ground at 90°, and position the camera altitude at 30 m. This resulted in continuous visible image data of the study area with a single image resolution measuring 8256 x 5504 pixels, as depicted in Figure 3.
- Store aerial imagery for subsequent processing with Python software for biomass estimation.
- Collecting the aboveground biomass
- Collect the aboveground biomass of Mikania micrantha manually within each sample plot after completing the drone data collection. Bag them and label each bag accordingly.
- When collecting Mikania micrantha, prevent sample plots from moving. First, cut the Mikania micrantha along the inside edge of the sample plot.
- Then, cut the rhizome of the Mikania micrantha from the bottom. Remove any dirt, rocks, or other plants that are mixed in. Finally, bag and label the samples.
- Bring the collected samples of invasive plants from step 1.3.1 to the laboratory. Air-dry all the collected samples to evaporate most of the moisture.
- To further remove moisture from air-dried samples, use an oven. Set the temperature to 55 °C. Dry the samples for 72 h, then weigh each sample on an electronic balance and record the biomass data in grams (g).
- Place the electronic balance in an undisturbed environment, weigh, calibrate, and continue weighing. Place bags of Mikania micrantha on the electronic balance, wait for the readings to stabilize, and record the readings.
- Weigh the Mikania micrantha every hour until the mass no longer changes and record the reading minus the weight of the bag as the measured mass of that sample. Calculate the aboveground biomass of the invasive plant using the formula below:
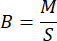
where B represents the biomass of Mikania micrantha in grams per square meter (g/m2), M is the weight of the measured Mikania micrantha, measured in grams (g), S corresponds to the area of the sample plot in square meters (m2).
- Collect the aboveground biomass of Mikania micrantha manually within each sample plot after completing the drone data collection. Bag them and label each bag accordingly.
- Creating a dataset
- Extract the RGB image corresponding to the sample image from the original UAV image. Divide it into a grid of 280 × 280 pixels using Python programming (Supplementray Figure 1).
- Segment the raw image data into smaller images of the same size as the sample images using Python programming. Use the sliding window method for segmentation, setting the horizontal and vertical steps to 280 pixels.
- From the small images segmented in step 1.4.2, randomly select 880 invasive plant images and 1500 background images to create a dataset. Then, split this dataset into training, validation, and test sets in a 6:2:2 ratio (Supplementray Figure 2).
2. Identification of Mikania micrantha
- Preparing the software
- Go to Anaconda's official website (https://www.anaconda.com/) and download and install Anaconda. Then, head to PyCharm's website (https://www.jetbrains.com/pycharm/) and download the PyCharm IDE.
- Creating a Conda environment.
- Open the Anaconda Prompt command line after installing Anaconda, then type conda create -n pytorch python==3.8 to create a new Conda environment. After the environment is created, enter conda info –envs to confirm that the pytorch environment exists.
- Open the Anaconda Prompt and activate the pytorch environment by entering conda activate pytorch. Check the current (Compute Unified Device Architecture)CUDA version by typing nvidia-smi. Then, install PyTorch version 1.8.1 by running the command conda install pytorch==1.8.1 torchvision==0.9.1 torchaudio==0.8.1 cudatoolkit=11.0 -c pytorch.
- Runs for model recognition
NOTE: Use PyTorch to build the Mikania micrantha recognition model used in this paper. The network model employed is ResNet10121, which remains consistent with the original paper in its architecture. Modifications are made to the network's output section to meet the requirements for chamomile recognition.- Preprocess the images to prepare them for model input. Resize the images from 280 x 280 pixels to 224 x 224 pixels and normalize them to ensure they meet the model's size requirements using the following code:
transform = transforms.Compose([
transforms.Resize((224, 224)),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225])]) - Perform image feature extraction and reduce dimensionality using a convolutional neural network.
- First, initialize the convolutional layer for the initial feature extraction through self.conv1. With this convolutional layer, the original image is convolved into a feature map with self.in_channel channels for extracting initial features (Supplementary Figure 3A).
NOTE: Advanced features are extracted in a convolution operation on the residual passes. These layers are produced by invoking the _make_layer function, which comprises a sequence of residual blocks. Each residual block consists of convolution, batch normalization, and activation functions to gradually extract sophisticated features (Supplementary Figure 3B). - Use the layer's function to modify the channel number for dimensionality reduction via 1×1 convolution. This operation decreases the computational load while preserving significant features (Supplementary Figure 3C).
NOTE: Overall, ResNet101 performs feature extraction by using various convolutional layers, and dimensionality reduction is achieved through 1×1 convolutional layers within the residual block. This approach allows the network to learn features more deeply and avoid the issue of gradient vanishing, thus enabling more efficient learning and representation of image features for complex tasks. - After convolutions and pooling operations, input the high-quality features into a fully connected layer.
NOTE: In the ResNet architecture, feature extraction takes place in the convolutional layer. These features are subsequently sent to the fully-connected (FC) layer for classification (Supplementary Figure 4). The operation self.avgpool(x) performs adaptive average pooling to reshape the tensor to a fixed size. The operation torch.flatten(x, 1) spreads the tensor into a one-dimensional vector, and self.fc(x) applies the fully connected layer to the flattened vector, ultimately serving as the final step for classification. This process effectively passes the extracted features through the convolutional layer, transforming them into a format suitable for classification via the fully connected layer. - Use the Softmax function to obtain the final output based on the three classification requirements.
- First, initialize the convolutional layer for the initial feature extraction through self.conv1. With this convolutional layer, the original image is convolved into a feature map with self.in_channel channels for extracting initial features (Supplementary Figure 3A).
- Train a multi-class recognition model with the dataset from step 1.4. Set the number of iterations to 200 and an initial learning rate of 0.0001. Reduce the learning rate by a third every 10 iterations with a batch size of 64. Save the optimal model parameters automatically after each iteration (Supplementary Figure 5).
- Employ a meticulously trained recognition model and systematically traverse the original image from step 1.2.2 for identification purposes.
- Configure horizontal and vertical steps precisely at 280 pixels, resulting in the generation of a comprehensive distribution map highlighting the presence of invasive flora within the boundaries of the study area. Present the selected results visually as shown in Figure 4.
NOTE: The initial image is preprocessed by segmenting it into smaller chunks, classifying each chunk using a trained deep learning model, and combining the results into an output image. If a chunk is classified as an invasive plant, the corresponding location on the output image is set to 255. The resulting output image is saved as a grayscale image file. The specific implementation code is shown in Supplementary Figure 6.
- Configure horizontal and vertical steps precisely at 280 pixels, resulting in the generation of a comprehensive distribution map highlighting the presence of invasive flora within the boundaries of the study area. Present the selected results visually as shown in Figure 4.
- Preprocess the images to prepare them for model input. Resize the images from 280 x 280 pixels to 224 x 224 pixels and normalize them to ensure they meet the model's size requirements using the following code:
3. Estimation of invasive plant biomass
- Perform simple data augmentation with the RandomResizedCrop and RandomHorizontalFlip functions (Supplementary Figure 7) to extend the image set created in step 1.2 and extract the six vegetation indices commonly used for estimating biomass, which are RBRI, GBRI, GRRI, RGRI, NGBDI, and NGRDI. Refer to Table 1 for the calculation formulas for these indices.
- Create a K-nearest neighbor regression (KNNR)22 model using the output of the model to ensure precise estimation of the biomass of invasive plants. Use the extracted vegetation indices as inputs for the estimation model.
- Use the coefficient of determination R2 and Root Mean Square Error (RMSE)23 to assess the model's accuracy, which is calculated as follows:
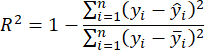
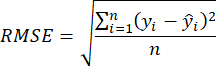
NOTE: The K-Nearest Neighbor Regression (KNNR) algorithm is a nonparametric machine learning technique used to solve regression problems. Its fundamental concept is to predict outcomes by determining the closest K neighbors in feature space based on input sample distances. Key benefits of using KNNR include its simplicity and ease of understanding, and it requires no training phase. Additionally, KNNR does not make excessive assumptions about the data's distribution. KNNR can be applied in regression issues to anticipate continuous objective variables and precisely evaluate the biomass of invasive plants. - Employ the aboveground biomass estimation model chosen in step 3.2 and scan through the invasive plant distribution map from step 2.3.4 with horizontal and vertical strides of 280 pixels.
Computer Vision-Based Biomass Estimation for Invasive Plants
Learning Objectives
We show representative results of a computer vision-based method for the estimation of invasive plants, which is implemented in a programmatic way on a computer. In this experiment, we evaluated the spatial distribution and estimated the biomass of invasive plants in their natural habitats, using Mikania micrantha as a research subject. We utilized a drone camera system to acquire images of the research site, a portion of which is exhibited in Figure 3. We utilized the ResNet101 convolutional neural network to identify the plants present within the study area. Subsequently, we mapped the spatial distribution of invasive plants and illustrated some of our findings in Figure 4. In Figure 3, Mikania micrantha can be observed climbing atop the plant adorned with white flowers. The other plants, as well as the road and accompanying elements, are uniformly depicted in the background. In Figure 4, the model recognizes the red part as Mikania micrantha. Comparing the two sets of images, it is evident that ResNet101 demonstrates robust detection of Mikania micrantha in complex backgrounds. Furthermore, it accurately maps the distribution of Mikania micrantha in the study area with high precision.
The biomass of invasive plants in the study area was estimated by truncating all Mikania micrantha sample plot images from orthophotos at 280 × 280 pixels and extracting the vegetation indices RBRI, GBRI, GRRI, RGRI, NGBDI, and NGBDI. Regression analysis was conducted using the KNNR regression model, with the six indexes as inputs to the estimation model and biomass as the model's output. Figure 5 presents the results: the graph's horizontal coordinates represent the values of the field-measured biomasses, the vertical coordinates represent the values of the model-predicted biomasses, and the gray areas represent the confidence intervals. The results demonstrate strong predictive performance, with an R² value of 0.62 and an RMSE of 10.56 g/m2. The model enhances the accuracy of Mikania micrantha biomass estimation, and the spatial distribution map in Figure 6 effectively captures the distribution of Mikania micrantha biomass.

Figure 1: UAV remote sensing systems. Some examples of RGB image data captured by UAV. Please click here to view a larger version of this figure.
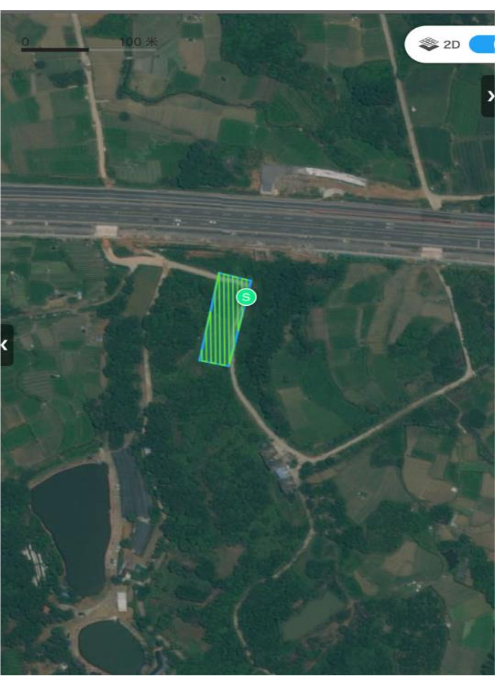
Figure 2: Route planning. Study on regional route planning Please click here to view a larger version of this figure.
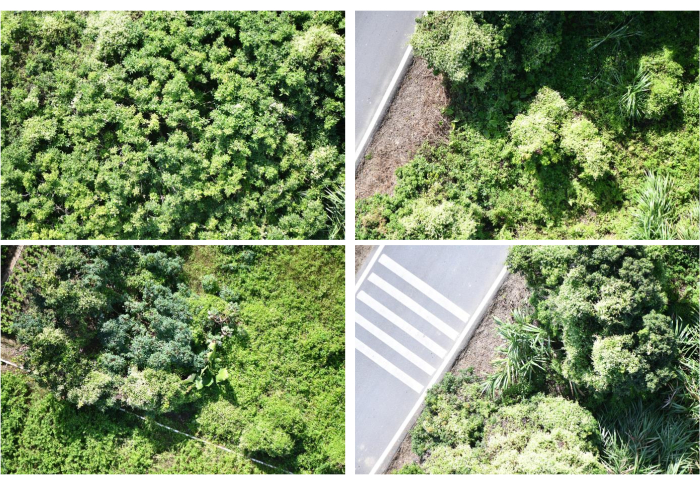

Figure 3: Invasive plant identification results within the study area. The figure displays the findings of identifying Mikania micrantha in the study zone via ResNet101 convolutional neural network. The red region in the picture indicates the area detected by ResNet101 as Mikania micrantha, while the other background signifies the rest of the study area. These outcomes correspond to the sample images portrayed in Figure 1. They represent the recognition results of Figure 1, respectively. Please click here to view a larger version of this figure.
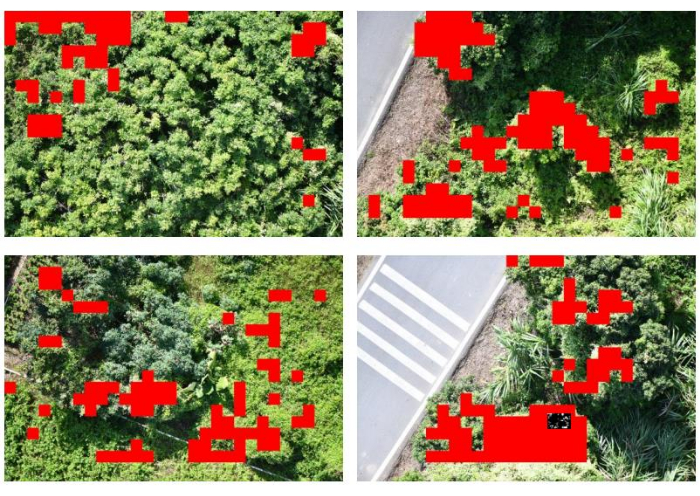

Figure 4: Spatial distribution of invasive plants. The model recognizes the red part as Mikania micrantha. Please click here to view a larger version of this figure.
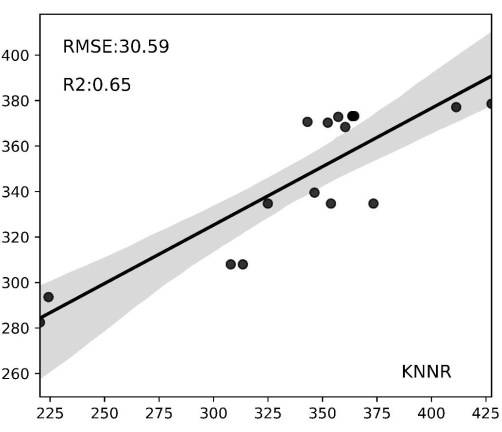
Figure 5: Biomass prediction regression results. The horizontal axis displays biomass values observed in the field, while the vertical axis portrays biomass values estimated by the model. The gray-shaded regions denote confidence intervals. The KNNR model attained an R2 of 0.65 on the test set, while the lowest root mean square error amounted to 30.59 g/m2. In the regression scatter plot of the model, many Mikania micrantha biomass estimations were within the confidence interval, indicating the validity of the biomass prediction. Please click here to view a larger version of this figure.
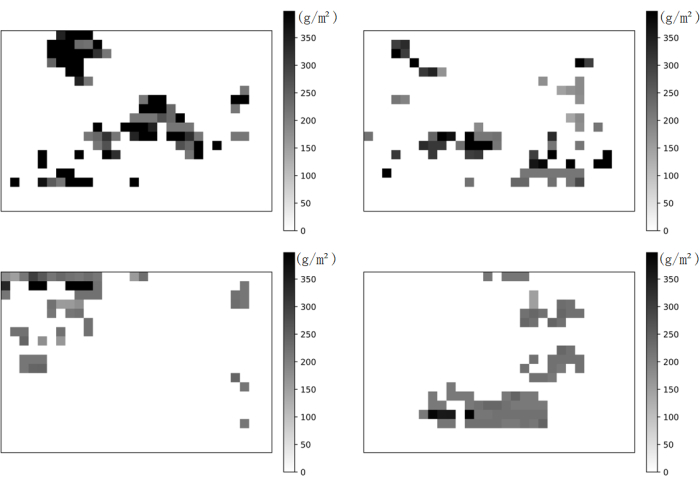
Figure 6: Spatial distribution of Mikania micrantha biomass. The figure illustrates the estimation of Mikania micrantha biomass throughout the research area utilizing KNNR as the predictive model, along with the extracted Mikania micrantha biomass distribution map. Darker shaded regions represent higher quantities of Mikania micrantha biomass. Please click here to view a larger version of this figure.
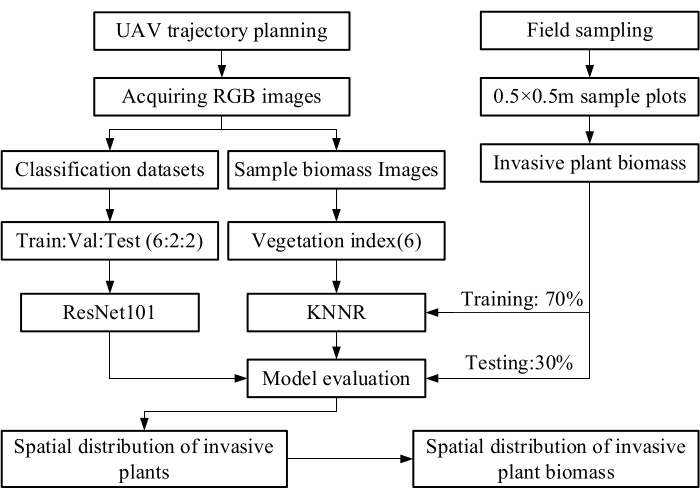
Figure 7: Schematic diagram of the main development of this protocol. The figure illustrates the main steps of the protocol presented. Please click here to view a larger version of this figure.
| Vegetation Index Name | Calculation Formula |
| Green Blue Ratio Index | GBRI = DNG/DNB |
| Green Red Ratio Index | GRRI = DNG/DNR |
| Red Blue Ratio Index | RBRI = DNR/DNB |
| Red Green Ratio Index | RGRI = DNR/DNG |
| Normalized Green Blue Difference Index | NGBDI = (DNG – DNB)/(DNG + DNB) |
| Normalized Green Red Difference Index | NGRDI = (DNG – DNR)/(DNG + DNR) |
Table 1: Vegetation index calculation formula. The vegetation indices used in this protocol and their respective calculation formulas.
Supplementary Figure 1: Cropping an image to 280 x 280 pixels via Python script using the OpenCV library. Please click here to download this File.
Supplementary Figure 2: Partitioning the dataset into a training set, validation set, and test set. Please click here to download this File.
Supplementary Figure 3: Feature extraction and reduction of dimensionality. (A) Initial feature extraction. (B) Convolution operation. (C) Reduction of dimensionality. Please click here to download this File.
Supplementary Figure 4: Conveying the features to the FC layer in ResNet architecture. Please click here to download this File.
Supplementary Figure 5: Setting the parameters. Please click here to download this File.
Supplemnetary Figure 6: The specific implementation code for generating comprehensive distribution map. Please click here to download this File.
Supplementary Figure 7: RandomResizedCrop and RandomHorizontalFlip functions. Please click here to download this File.
List of Materials
| DSLR camera | Nikon | D850 | Sensor type: CMOS; Maximum number of pixels: 46.89 million; Effective number of pixels: 45.75 million; Maximum resolution 8256 x 5504. |
| GPU – Graphics Processing Unit | NVIDIA | RTX3090 | |
| Hexacopter | DJI | M600PRO | Horizontal flight: 65 km/h (no wind environment); Maximum flight load: 6000 g |
| PyCharm | Python IDE | 2023.1 | |
| Python | Python | 3.8.0 | |
| Pytorch | Pytorch | 1.8.1 |
Lab Prep
We report on the detailed steps of a method to estimate the biomass of invasive plants based on UAV remote sensing and computer vision. To collect samples from the study area, we prepared a sample square assembly to randomize the sampling points. An unmanned aerial camera system was constructed using a drone and camera to acquire continuous RGB images of the study area through automated navigation. After completing the shooting, the aboveground biomass in the sample frame was collected, and all correspondences were labeled and packaged. The sample data was processed, and the aerial images were segmented into small images of 280 x 280 pixels to create an image dataset. A deep convolutional neural network was used to map the distribution of Mikania micrantha in the study area, and its vegetation index was obtained. The organisms collected were dried, and the dry weight was recorded as the ground truth biomass. The invasive plant biomass regression model was constructed using the K-nearest neighbor regression (KNNR) by extracting the vegetation index from the sample images as an independent variable and integrating it with the ground truth biomass as a dependent variable. The results showed that it was possible to predict the biomass of invasive plants accurately. An accurate spatial distribution map of invasive plant biomass was generated by image traversal, allowing precise identification of high-risk areas affected by invasive plants. In summary, this study demonstrates the potential of combining unmanned aerial vehicle remote sensing with machine learning techniques to estimate invasive plant biomass. It contributes significantly to the research of new technologies and methods for real-time monitoring of invasive plants and provides technical support for intelligent monitoring and hazard assessment at the regional scale.
We report on the detailed steps of a method to estimate the biomass of invasive plants based on UAV remote sensing and computer vision. To collect samples from the study area, we prepared a sample square assembly to randomize the sampling points. An unmanned aerial camera system was constructed using a drone and camera to acquire continuous RGB images of the study area through automated navigation. After completing the shooting, the aboveground biomass in the sample frame was collected, and all correspondences were labeled and packaged. The sample data was processed, and the aerial images were segmented into small images of 280 x 280 pixels to create an image dataset. A deep convolutional neural network was used to map the distribution of Mikania micrantha in the study area, and its vegetation index was obtained. The organisms collected were dried, and the dry weight was recorded as the ground truth biomass. The invasive plant biomass regression model was constructed using the K-nearest neighbor regression (KNNR) by extracting the vegetation index from the sample images as an independent variable and integrating it with the ground truth biomass as a dependent variable. The results showed that it was possible to predict the biomass of invasive plants accurately. An accurate spatial distribution map of invasive plant biomass was generated by image traversal, allowing precise identification of high-risk areas affected by invasive plants. In summary, this study demonstrates the potential of combining unmanned aerial vehicle remote sensing with machine learning techniques to estimate invasive plant biomass. It contributes significantly to the research of new technologies and methods for real-time monitoring of invasive plants and provides technical support for intelligent monitoring and hazard assessment at the regional scale.
Procedure
We report on the detailed steps of a method to estimate the biomass of invasive plants based on UAV remote sensing and computer vision. To collect samples from the study area, we prepared a sample square assembly to randomize the sampling points. An unmanned aerial camera system was constructed using a drone and camera to acquire continuous RGB images of the study area through automated navigation. After completing the shooting, the aboveground biomass in the sample frame was collected, and all correspondences were labeled and packaged. The sample data was processed, and the aerial images were segmented into small images of 280 x 280 pixels to create an image dataset. A deep convolutional neural network was used to map the distribution of Mikania micrantha in the study area, and its vegetation index was obtained. The organisms collected were dried, and the dry weight was recorded as the ground truth biomass. The invasive plant biomass regression model was constructed using the K-nearest neighbor regression (KNNR) by extracting the vegetation index from the sample images as an independent variable and integrating it with the ground truth biomass as a dependent variable. The results showed that it was possible to predict the biomass of invasive plants accurately. An accurate spatial distribution map of invasive plant biomass was generated by image traversal, allowing precise identification of high-risk areas affected by invasive plants. In summary, this study demonstrates the potential of combining unmanned aerial vehicle remote sensing with machine learning techniques to estimate invasive plant biomass. It contributes significantly to the research of new technologies and methods for real-time monitoring of invasive plants and provides technical support for intelligent monitoring and hazard assessment at the regional scale.
