2D and 3D Matrices to Study Linear Invadosome Formation and Activity
Summary
This protocol describes how to prepare a 2D mixed matrix, consisting of gelatin and collagen I, and a 3D collagen I plug to study linear invadosomes. These protocols allow for the study of linear invadosome formation, matrix degradation activity, and the invasion capabilities of primary cells and cancer cell lines.
Abstract
Cell adhesion, migration, and invasion are involved in many physiological and pathological processes. For example, during metastasis formation, tumor cells have to cross anatomical barriers to invade and migrate through the surrounding tissue in order to reach blood or lymphatic vessels. This requires the interaction between cells and the extracellular matrix (ECM). At the cellular level, many cells, including the majority of cancer cells, are able to form invadosomes, which are F-actin-based structures capable of degrading ECM. Invadosomes are protrusive actin structures that recruit and activate matrix metalloproteinases (MMPs). The molecular composition, density, organization, and stiffness of the ECM are crucial in regulating invadosome formation and activation. In vitro, a gelatin assay is the standard assay used to observe and quantify invadosome degradation activity. However, gelatin, which is denatured collagen I, is not a physiological matrix element. A novel assay using type I collagen fibrils was developed and used to demonstrate that this physiological matrix is a potent inducer of invadosomes. Invadosomes that form along the collagen fibrils are known as linear invadosomes due to their linear organization on the fibers. Moreover, molecular analysis of linear invadosomes showed that the discoidin domain receptor 1 (DDR1) is the receptor involved in their formation. These data clearly demonstrate the importance of using a physiologically relevant matrix in order to understand the complex interactions between cells and the ECM.
Introduction
Extracellular matrix (ECM) remodeling occurs during physiological and pathological processes, such as angiogenesis and tumor cell invasion. In physiological conditions, many cell types, mostly from the hematopoietic lineage, are able to degrade ECM elements. For example, macrophages are able to cross anatomical barriers to reach tissues, and osteoclasts degrade bone matrix to ensure calcium homeostasis. More globally, all matrices in the body renew to maintain their physical and chemical properties. In cancer tissues, the tumor microenvironment composition is altered. For example, in breast and lung cancer, type I collagen is overexpressed and accumulates around the tumor. Moreover, this accumulation is associated with an increased risk of developing metastasis1,2. Cancer metastasis is dependent on the ability of cancer cells to degrade the ECM and invade adjacent tissues.
The invasive activity of cells is attributed to specialized actin-rich structures known as invadosomes. This term includes podosomes and invadopodia, which are present, respectively, in normal and cancer cells (e.g., macrophages, endothelial cells, and cancer cells such as the MDA-MB-231 breast cancer cell line). In vitro, invadosomes can organize into different shapes: dots, aggregates, or rosettes3. Classically, invadosomes are composed of an F-actin core containing several proteins, such as the scaffold protein Tks5 and cortactin, surrounded by adhesion plaque molecules, such as integrins and vinculin4. Recently, it was shown that Tks5 and the RhoGTPase Cdc42 can be used as a minimum molecular signature for functional invadosomes5. These structures are able to degrade the ECM via the recruitment and activation of specific metalloprotease proteins, such as MT1-MMP and MMP-26. In a previous study, we reported that the interaction between type I collagen fibrils and the discoidin domain receptor 1 (DDR1), a specific receptor of type I collagen fibrils, leads to the formation of a new class of invadosomes, named linear invadosomes. Linear invadosomes are formed along type I collagen fibrils7. These structures are able to degrade type I collagen fibrils via the recruitment and activation of MT1-MMP/MMP2. Moreover, their formation is dependent on Tks5 and Cdc425. In vitro, we demonstrated the existence of linear invadosomes and the involvement of DDR1 in their formation and activity8 using different strategies consisting of a combination of various matrix elements, including: i) fluorescent gelatin-coated coverslips, ii) fluorescent type I collagen fibril-coated coverslips, and iii) 3D type I collagen plugs. Due to the use of different matrices, we were able to study and characterize linear invadosome formation and their degradation activity7,8.
Finally, to better understand the biology of invasive cells, a combination of ECM components in both 2D and 3D culture systems were used, mimicking the complexity of the tumor microenvironment composition. Below are protocols for coating coverslips with gelatin/type I collagen in 2D and 3D culture systems.
Protocol
NOTE: Prior to fixation, all steps are completed in a sterile laminar flow hood.
1. 2D matrix
NOTE: 2D matrices are used to determine the percentage of cells forming invadosomes and the matrix degradation area per cell.
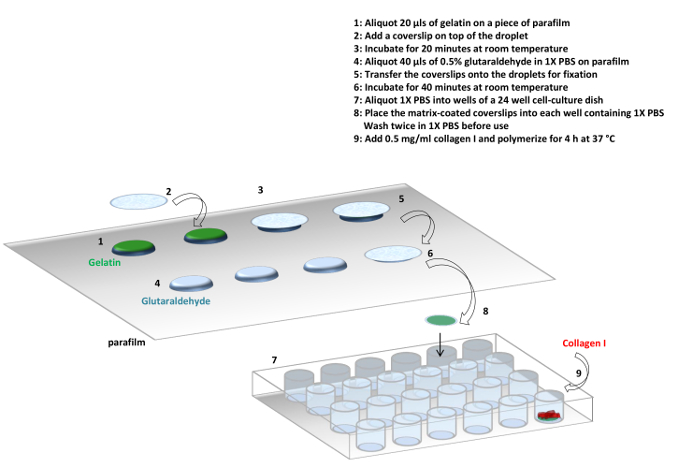
Figure 1: Protocol. Summary of the protocol used to prepare gelatin and collagen I matrices. It is possible to make the gelatin matrix only or a gelatin and collagen I mixed matrix. Please click here to view a larger version of this figure.
- Gelatin matrix
NOTE: Fluorescent gelatin matrices are used to quantify matrix degradation, and non-fluorescent gelatin is used to promote the adhesion of collagen I fibers. Fluorescent gelatin matrices are also used for podosome and invadopodia studies (see Figure 1).- Under a sterile laminar flow hood, place a piece of parafilm (20 cm x 15 cm) on the work area and disinfect it with 70% ethanol.
- Aliquot 20 µL droplets of 1 mg/mL gelatin in a line, spaced approximately 1 cm apart; refer to Figure 1 for droplet organization.
- Cover each 20 µL gelatin droplet with an autoclaved glass coverslip (12 mm in diameter).
- Incubate for 20 min at room temperature. If the gelatin is fluorescent, protect it from light.
- Aliquot 40 µL droplets of 0.5% glutaraldehyde in 1x phosphate-buffered saline (PBS) in a line, each spaced approximately 1 cm below the line of coverslips from step 1.1.3.
CAUTION: Glutaraldehyde is a toxic fixative agent. Use gloves and do not inhale. - Using jeweled forceps, transfer each gelatin-coated coverslip to a glutaraldehyde droplet.
NOTE: It is normal that a small gelatin droplet remains on the parafilm. Do not reuse this for another experiment. - Incubate for 40 min at room temperature. For fluorescent coverslips, protect them from light.
- Take a 24-well cell culture plate and fill each well with 500 µL of sterile 1x PBS. Place the coverslips, gelatin side up, into each well.
- Wash twice in 1x PBS for 5 min each.
NOTE: At this step, the coverslips can be used for experimentation or stored in 1x PBS at 4 °C. Non-fluorescent coverslips can be stored for one week, and fluorescent coverslips can be stored for 48 h.
- Fibrillar collagen type I matrix
NOTE: Fibrillar collagen type I matrix is used to study linear invadosome formation and the associated matrix degradation. A combination of fluorescent collagen I and non-fluorescent gelatin coverslips is used to colocalize invadosome markers with collagen fibers (e.g., F-actin in the far-red channel, collagen I in the red channel, Tks5 in the green channel, and Hoechst dye for a nuclear stain). On the other hand, a combination of non-fluorescent collagen I with non-fluorescent gelatin coverslips is used to maximize the number of invadosome markers stained (e.g., F-actin in the far-red channel, cortactin in the red channel, Tks5 in the green channel, and Hoechst dye for a nuclear stain). Finally, a combination of non-fluorescent collagen I with fluorescent gelatin coverslips is used to quantify the gelatin-associated degradation (e.g., F-actin in the far-red channel, Tks5 in the red channel, in the green channel, and Hoechst dye for a nuclear stain) (see Figure 1). Hoechst dye is used to stain the nucleus, and it also checks for mycoplasma contamination, in addition to a PCR test done regularly on the cell lines used.- Fluorescent collagen I
- Use commercial collagen I extracted from rat-tail tendon in 0.02 N acetic acid. Withdraw the quantity of collagen I necessary to prepare 500 µL of solution per coverslip at a final concentration of 0.4 mg/mL of collagen I.
- Add 5-carboxy x rhodamine succinimidyl ester at a final concentration of 10 µg/mL, calculated for the final volume of collagen I solution (n x 500 µL).
- Mix the solution by pipetting up and down and incubate for 5 min at room temperature, protected from light.
- Dilute to the final volume with 1X Dulbecco's Phosphate-Buffered Saline (DPBS) containing calcium and magnesium, and use directly.
- Fibrillar collagen I coverslip
- Use the non-fluorescent or fluorescent gelatin coverslips previously prepared in step 1.1.
- Prepare a solution of fluorescent or non-fluorescent collagen I at a final concentration of 0.4 mg/mL in 1X DPBS containing calcium and magnesium.
- Add 500 µL of fluorescent or non-fluorescent collagen I solution per coverslip in a 24-well cell culture plate.
- Incubate for 4 h at 37 °C for collagen I polymerization.
- After incubation, remove the excess collagen I by tilting the plate and pipetting on the side of the well (not directly on the coverslip) to avoid damaging the matrix.
NOTE: This matrix cannot be stored. Seed the cells immediately after removing excess collagen I.
- Fluorescent collagen I
2. Cell seeding, time of culture, and fixation
- Depending on the cell type, prepare the cells in the appropriate culture medium and add cells for a final volume of 500 µL per well.
- To characterize the ability of a cell line to form linear invadosomes and to degrade the matrix, plate the cells for 4 h, 12 h, and 24 h on collagen I matrix to determine the kinetics of linear invadosome formation and the associated degradation activity.
NOTE: Table 1 contains examples of cell lines tested in the laboratory.
Table 1: Cell seeding and incubation time reference. This table presents a non-exhaustive list of cell lines used to study linear invadosomes. It indicates the number of cells to seed on the matrix, the time necessary to observe linear invadosomes, and the time necessary to observe the associated matrix degradation. Linear invadosomes have a short lifetime. To be able to observe linear invadosomes in the maximum number of cells, the cells have to be in contact with the collagen I matrix for a short period of time. On the other hand, to be able to visualize the reported gelatin degradation, the cells must be in contact with the collagen I matrix for a longer period of time than those listed above.Cell line Number of cells per coverslip Time of contact with matrix for linear invadosome quantification Time of contact with matrix for quantification of linear invadosome degradation activity MDA MB 231 40,000 4 h 12 h HUH7 40,000 12 h 24 h HEP 3B 40,000 12 h 24 h SNU 398 40,000 12 h 24 h NIH 3T3 src 30,000 4 h 12 h A431 50,000 6 h 24 h A549 40,000 12 h 24 h RAW 40,000 12 h 12 h PAE 40,000 12 h 12 h - After the removal of the cell culture medium, fix the coverslips for 10 min in 500 µL of 4% paraformaldehyde in 1X PBS.
- Wash twice with 1x PBS.
- Store at 4 °C, protected from light.
3. Immunofluorescence
- Permeabilize the cells for 10 min with a freshly prepared solution of 0.2% Triton X-100 in 1x PBS. Do not reuse this solution.
- Wash the coverslips twice with 1x PBS.
- Place a piece of parafilm (20 cm x 15 cm) on the bench.
- Dilute primary antibody in 4% bovine serum albumin (BSA) in 1x PBS, as described in the Materials Table.
- Aliquot 50 µL droplets of primary antibody solution in a line, each spaced approximately 1 cm apart.
- Place each coverslip on a droplet.
- Incubate for 40 min at room temperature, protected from light.
- After the incubation period, wash twice with 1x PBS.
- Dilute the secondary antibody, phalloidin, and Hoechst dye in 4% BSA in 1x PBS.
- Aliquot 50 µL droplets of the mix, each spaced approximately 1 cm below the first line of coverslips, as done previously.
- Place each coverslip on a droplet.
- Incubate for 30 min at room temperature, protected from light.
- After incubation, wash twice with 1x PBS.
- Wash once with distilled water to remove the salts.
- Mount the coverslips on a glass slide with polymerizing mounting medium.
- Let the coverslips dry overnight at room temperature, protected from light, before microscopy.
4. Quantifications
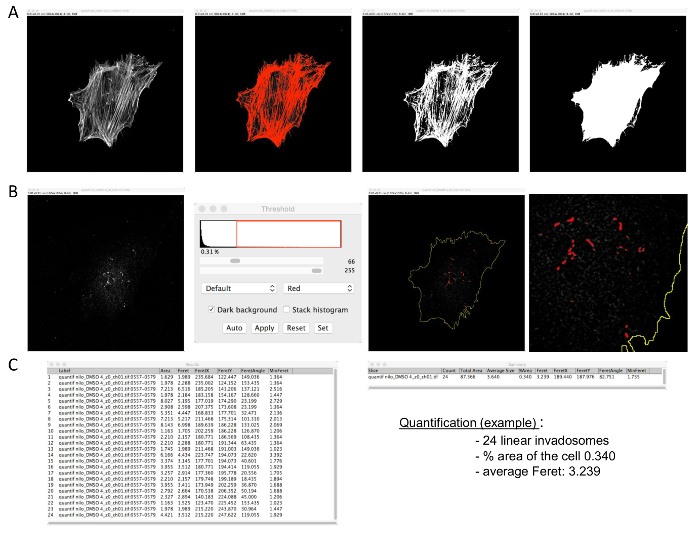
Figure 2: Linear invadosome quantification. The macro requires confocal images of F-actin and Tks5 staining (format: 1,024 x 1,024). (A) First, open the F-actin picture and make a mask. (B) Then, open the Tks5 picture and threshold it. Apply the macro. (C) Analyzed results will appear in two tables. The number of linear invadosomes per cell and other information, such as the percentage area used by linear invadosomes in the cell, will be in the Summary table. The size of each linear invadosome will be in the Results table. Please click here to view a larger version of this figure.
- Percentage of cells forming linear invadosomes
- Count the cells forming linear invadosomes, approximately 100 cells per coverslip (use technical triplicates). Quantify at least 300 cells per n, with n = 3.
- Calculate the percentage of cells forming linear invadosomes by taking the average of three independent experiments, resulting in 900 quantified cells.
- Linear invadosome size and number per cell
NOTE: Use a confocal microscope to take sequential images of the F-actin and Tks5 stains. The macro is adapted to a 1,024 x 1,024 image format.- Use ImageJ with the supplementary macro.
- Degradation per cell
- Use three coverslips per condition.
- Take 30 images per coverslip of fluorescent gelatin and Hoechst-stained nuclei.
- Use the ImageJ software as described in Martin KH et al.9.
5. 3D Collagen Invasion Assay
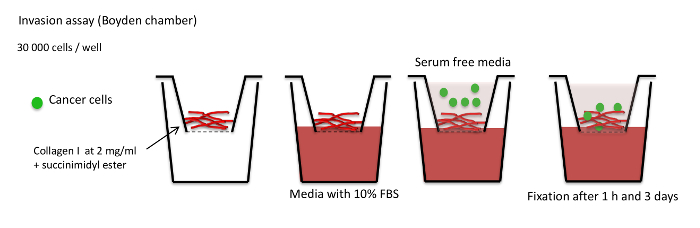
Figure 3: Invasion assay schema. Summary of the invasion assay protocol. Collagen I is cross-linked with succinimidyl ester dye and polymerized for 1 h at 37 °C. Cells are added in serum-free medium on top of the collagen I plug. The insert is placed in medium with 10% FBS. Collagen I plugs are fixed 1 h after cell seeding in the control and 3 days after cell seeding in the experimental condition to determine the invasion capacity of the cells. Please click here to view a larger version of this figure.
- Collagen I plug preparation
- Make a time course of 4 h, 12 h, 24 h, 48 h, and 72 h for the first experiment to determine the kinetics of invasion into the gel according to the cell type. For each experiment, add a 1 h time point as a reference time point when no invasion occurs.
- Prepare the collagen I solution on ice under a sterile laminar flow hood. Prepare 500 µL of solution for 3 Boyden chamber inserts.
- Aspirate 1 mg of commercial rat-tail collagen I to prepare a solution with a final concentration of 2 mg/mL collagen I.
- Add 5-carboxy x Rhodamine succinimidyl ester at a final concentration of 1 µg/mL, calculated for the final volume of collagen I solution (in this case, 500 µL).
- Mix well and incubate for 5 min on ice under the sterile laminar flow hood, protected from light.
- Add 50 µL of ice-cold, filtered 10X PBS and 6.25 µL of ice-cold 1 M sodium hydroxide (NaOH).
- Add sterile water according to the formula 443.75 – x µL of collagen I.
- Add 100 µL of solution per Boyden chamber insert.
- Incubate for 1 h at 37 °C to allow for polymerization.
- Add cell culture medium enriched with fetal bovine serum (FBS) into new wells; place inserts into these wells.
- Seed 30,000 cells on top of the collagen I plug in FBS-free medium.
- After culturing the cells at 37 °C, fix the collagen I plug for 30 min in 4% paraformaldehyde in 1x PBS.
- Complete immunofluorescence as in step 3, with the exception of the following steps: permeabilize with 0.2% Triton X-100 in 1x PBS for 30 min, not 10 min; incubate with primary and secondary antibody solutions for 1.5 h each, rather than 40 min and 30 min, respectively. Stain for F-actin or nuclei to localize the cells.
- After immunofluorescence, use a scalpel to cut around the membrane of the insert and carefully transfer the collagen I plug to a glass-bottom dish. Make sure that the top of the collagen I plug is in contact with the glass. Keep the insert membrane attached to the bottom of the collagen I plug in order to maintain the integrity of the plug.
NOTE: The collagen I plug remains intact in the glass-bottom dish. The depth between the glass bottom and the plastic surrounding the glass bottom is equivalent to the thickness of the plug. - Place a coverslip on the collagen I plug and mount with polymerizing mounting medium to prevent the plug from drying out.
- Use an inverted confocal microscope to image the collagen I plug; the top of the plug is in contact with the glass bottom of the dish.
- Acquire a z-stack from the top to the bottom of the collagen I plug to visualize cell invasion. Determine the thickness of the z-stack by examining how deeply the cells invade the imaging field. Perform acquisitions with a 40X oil objective in a 512 x 512 format with a z-step size of 0.25 μm.
NOTE: For reference, the average z-stack for MDA-MB-231 cells 3 days after seeding is 30 µm.
- Acquire a z-stack from the top to the bottom of the collagen I plug to visualize cell invasion. Determine the thickness of the z-stack by examining how deeply the cells invade the imaging field. Perform acquisitions with a 40X oil objective in a 512 x 512 format with a z-step size of 0.25 μm.
Representative Results
Using a combination of two types of matrices, gelatin and type I collagen (Figures 1 and 4), we have highlighted a new type of invadosome, known as linear invadosomes. The labelling of these matrices allows for the observation of linear invadosome formation along collagen I fibers and of their degradation capabilities (Figure 5). The number of invasive structures can then be quantified by the macro described previously (step 4.2), and the degradation activity can also be determined using the ImageJ software, as describe by Martin et al. and Diaz et al.9,10. The mixed matrix allows for the characterization of linear invadosomes by defining DDR1 as the receptor necessary for linear invadosome formation and functionality (Figure 6).
In order to understand the invasive capacity of cells, we have also developed a 3D type I collagen plug assay (Figures 3 and 7). This assay allows us to determine the number of cells that have invaded into the collagen I plug, as well as the distance traveled, by using z-stack reconstructions.
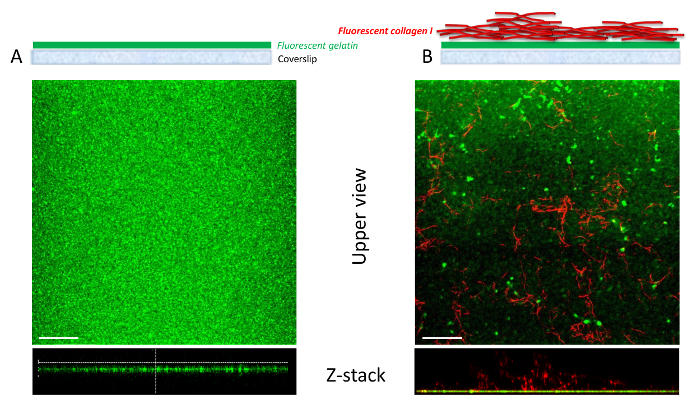
Figure 4: Comparison of gelatin and gelatin-collagen I mixed matrices using confocal microscopy. Schematic representation showing the organization of fluorescent gelatin coverslips on the top panel. (A) Confocal z-stack reconstruction demonstrates that the fluorescent gelatin matrix forms a thin, uniform layer that covers the entire surface of the coverslip. (B) In the mixed matrix, after the deposition of fluorescent gelatin on the coverslips, type I collagen fibrils polymerize on top. The collagen I fibrils are stained in red. The collagen I matrix is thick and heterogeneous on the coverslip. The distribution of the collagen I fibers is dependent on the polymerization of collagen I α chains. Scale bar = 10 µm. Please click here to view a larger version of this figure.
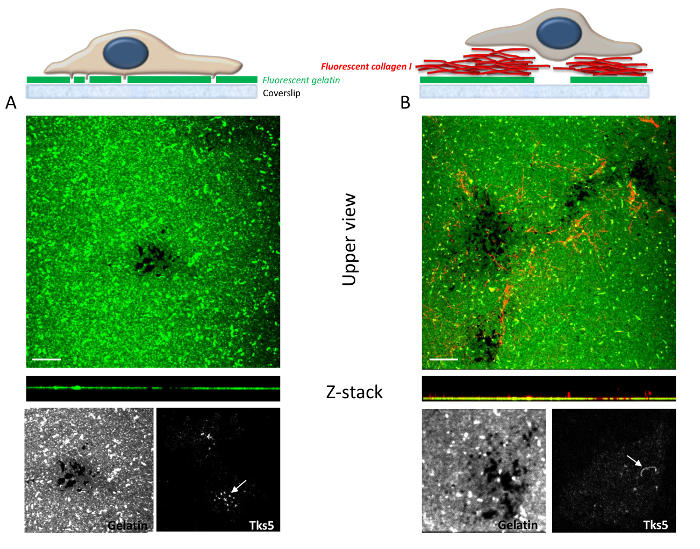
Figure 5: Impact of gelatin and gelatin-collagen I matrices on invadosome formation and activity. The cells used for these assays are MDA-MB-231 breast cancer cells. (A) A schematic representation shows cells seeded on a fluorescent gelatin coverslip on the top panel. Degradation areas are visualized in black due to the reduction of fluorescence. Tks5 staining was used as an invadosome marker. On gelatin, the invadosomes that form are organized as dots. (B) A schematic representation shows cells seeded on a mixed matrix of gelatin and collagen I. Collagen I is labeled in red; the addition of type I collagen fibrils increases the ability of the cell to degrade gelatin. Interestingly, the invadosome marker Tks5 is reorganized, and the dots are replaced by linear structures representing linear invadosomes. Degradation areas are visualized in black due to the reduction of fluorescence. Scale bar = 5 µm. Please click here to view a larger version of this figure.
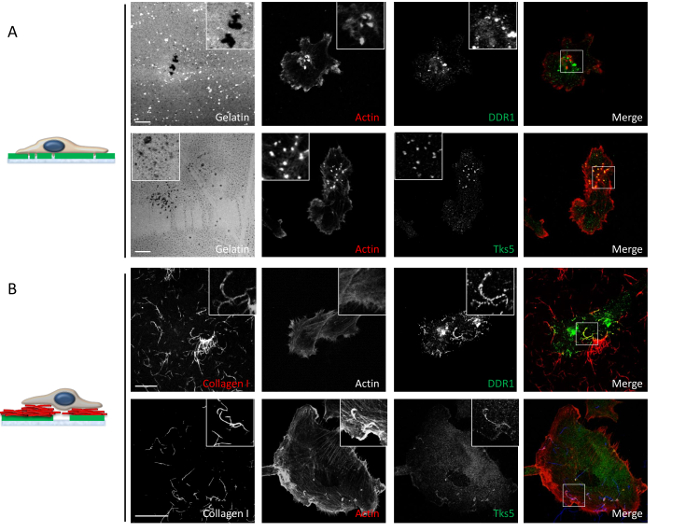
Figure 6: Confocal microscopy analysis of the molecular composition and organization of invadosomes in both gelatin and mixed-matrix conditions. (A) In the gelatin condition, invadosomes are organized in dots, and F-actin (red) co-localizes with Tks5 (green, bottom panel), but not with DDR1 (green, top panel). These are classical invadosomes. Scale bars = 5 µm. (B) In the gelatin-collagen I mixed-matrix condition, DDR1 (green, top panel) colocalizes with the collagen I fibrils (red, top panel), Tks5 (green, bottom panel), and F-actin (red, bottom panel). The type I collagen fibrils induce the formation of linear invadosomes. Scale bars = 10 µm. Please click here to view a larger version of this figure.
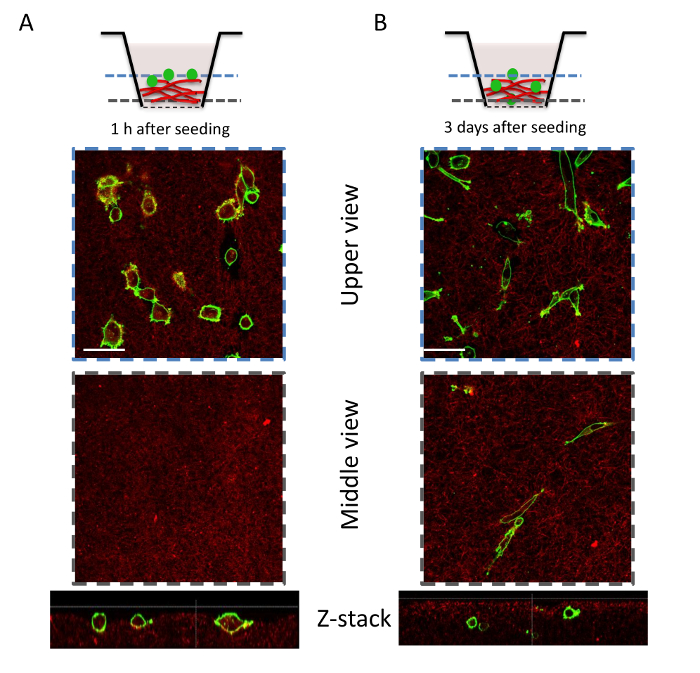
Figure 7: Cell invasion assay of MDA-MB-231 cells in a collagen I plug. 1 h after seeding the cells, control inserts are fixed and stained. (A) z-stack acquisition of the collagen I plug is performed using confocal microscopy. The cells do not invade the collagen I plug at this time point; rather, they remain on top of the collagen I plug. Thus, this is used as the control time point when no invasion occurs. MMP activation is necessary for the cells to invade a collagen I plug at this density. (B) Three days after seeding, some cells invade the collagen I plug. This assay allows for the observation and quantification of cell invasion. In addition, it can be used to study the impact of various drugs or siRNAs, for example. Scale bar = 20 µm. Please click here to view a larger version of this figure.
Discussion
Classically, invadosomes are studied in vitro without regard to the microenvironment and the matrix on which the cells are plated. Several types of matrices are currently used, including gelatin, fibronectin, vitronectin, or high-density fibrillar collagen (HDFC)7,11; however, these are often not representative of the microenvironment in which cells reside and are not physiologically relevant. Here, a novel type of matrix, which consists of an association between gelatin and fibrillar type I collagen, was used. The use of type I collagen fibrils allows us to highlight a novel class of invadosomes, known as linear invadosomes, which specifically form on collagen I in its physiological architecture7. Labeling the collagen I allows us to observe linear invadosomes along the fibers and to quantify their formation using cellular markers such as Tks5 and cortactin. Using the mixed matrix, we have identified a new receptor, the discoidin domain receptor 1 (DDR1), involved in the formation of linear invadosomes8.
The 2D mixed matrix allows us to quantify the matrix degradation activity of linear invadosomes via the zymography in situ assay. This assay reports the proteolysis activity of MMPs by analyzing the presence of black holes in the fluorescent gelatin layer. Interestingly, the organization of F-actin into linear invadosomes increases the matrix degradation activity compared to gelatin-only7. One limitation of this assay is that the gelatin is a non-physiological matrix, and it has been demonstrated that a more physiological matrix changes the cellular response to the microenvironment. An additional limitation exists in the way in which MMP activity is quantified on the gelatin-collagen I mixed matrices. Only the gelatin degradation that localizes under the collagen I fibril layer can be quantified; therefore, this is an indirect method to quantify collagen I matrix degradation. An alternative method to visualize the degradation activity of linear invadosomes is to label the collagen I fibers with a specific antibody against collagen cleavage sites (Col1-3/4C, immunoglobulin). This allows for the visualization of cleaved fibers by immunofluorescence. However, due to cell migration, this antibody is not very specific in 2D for degradation due to linear invadosome formation. Another way to visualize and quantify the degradation activity of the collagen I fibers is to use multiphoton microscopy and second harmonic generation8. This method allows for the imaging of collagen I fibers without any staining.
Our method of polymerizing type I collagen is different compared to other methods used in the literature12,13. For example, Artym et al. centrifuge coverslips coated with an alpha helix collagen I solution and allow for a brief polymerization of 30 min. This high-density fibrillar collagen (HDFC) is much stiffer and denser than our collagen I fibers but is less polymerized. This difference in matrix structure shifts the invadosome shape from linear to dot-like. The time, pH, concentration, and temperature are parameters that have to be controlled for ECM polymerization. For example, the lower the temperature of polymerization, the larger the fibers will be.
To study the involvement of linear invadosomes in cell invasion, the 3D collagen I plug assay was used. The 3D collagen I plug is already used by the scientific community in order to study invasive structures or the involvement of metalloproteinases in the invasion process14,15,16. These types of 3D matrices have limits regarding their rigidity-for example, the 3D collagen I plug is less rigid than the 2D matrix. Moreover, the type and origin of the collagen I is also important regarding the different organization of collagen fibers in vivo17. Finally, in regards to the microenvironment composition in vivo, only one element of the extracellular matrix, the type I collagen, was focused on. Further study is necessary to determine the relevance of linear invadosomes in vivo.
In addition to the study of linear invadosomes, the mixed matrix, described herein, could be used with other types of matrix components, such as fibronectin, vitronectin, and other types of collagens (e.g., type IV collagen). This protocol could be adapted, depending on the types of matrix elements that are of interest and the processes to be studied. However, it is clear that generating complex and physiological matrices will allow for the identification of new pathways involved in cell adhesion, migration, and invasion.
Disclosures
The authors have nothing to disclose.
Acknowledgements
J.D.M. was supported by a PhD fellowship from INSERM/Région Aquitaine and is now supported by a post-doctoral ARC fellowship and the Tisch Cancer Institute at the Mount Sinai School of Medicine. E.H. is supported by a PhD from the Ministère de l’Enseignement Supérieur et de la Recherche. Z.E. is supported by a post-doctoral fellowship from Agence Nationale de la Recherche (ANR). C.M. is supported by the Tisch Cancer Institute at the Mount Sinai School of Medicine and J.J.B.C. is supported by the NIH/NCI grant K22CA196750, the TCI Young Scientist Cancer Research Award JJR Fund, and the Tisch Cancer Institute at Mount Sinai School of Medicine. This work was supported by a grant from ANR-13-JJC-JSV1-0005. F.S. is supported by the “Ligue nationale contre le cancer.” V.M. and F.S. are supported by funding from “Equipe Labellisée Ligue Nationale contre le Cancer 2016” and Institut National du Cancer, INCA_8036, and PLBio2012.
Materials
| Gelatin solution | Sigma | G1393 | Stock 20 mg/ml used at 1 mg/ml |
| Gelatin from pig skin Oregon green 488 conjugate | Molecular probe life technologies | G13186 | 5mg dilute at 1 mg/ml in sterile water |
| Microscope cover glasses | Marlenfeld GmbH & Co | 111520 | Ø12mm No.1 |
| 2.5% Glutaraldehyd in 0.1M sodium cacodylate buffer pH 7.4 | Electron microscopy science | 15960 | Dilute at 0.5 % in sterile water |
| 1X PBS pH 7.4 | Gibco by life technologies | 10010-015 | Use this PBS in all steps before fixation |
| Collagen I Rat tail | Corning | 354236 | |
| 5 carboxy X rhodamin siccinimidyl ester | Life technologies | C-6125 | |
| DPBS 1X + calcium + magnesium | Gibco by life technologies | 14040-091 | |
| Paraformaldehyde 16% solution | Electron microscopy science | 15710 | Dilute at 4% in 1X PBS |
| Triton X 100 | Sigma | T9284 | |
| 10X PBS buffer pH 7.4 | Ambion | AM9625 | Dilute at 1X in water Use in steps after fixation |
| Tks5 antibody | Santa Cruz | sc-30122 | Invadosome markers Dilution : 1/100 |
| Cortactin 4F11 antibody | Millipore | 5180 | Invadosome markers Dilution :1/100 |
| DDR1 antibody | Cell signaling | 5583 | Linear invadosome receptor |
| Dilution :1/100 | |||
| Phalloidin FluoProbes | Interchim | FT-AZ0330 | Fibrillary actin marker Dilution :1/200 |
| Hoechst | Sigma | 33258 | Nucleus marker Dilution :1/200 |
| Secondary antibodies FluoProbes | Interchim | FP-488 FP-547H or FP-647H | |
| Albumin from bovine serum | Sigma | A2153 | Dilute at 4% in 1X PBS |
| Fluoromount G mounting medium | Interchim | FP 483331 | |
| ImageJ software | Public domain | http://www.macbiophotonics.ca/imagej/ | |
| Cell culture insert | Corning | 353097 | 8.0µm pore size / 24 wells |
| Sodium hydroxide (NaOH) | Sigma | 221465 |
References
- Ramaswamy, S., Ross, K. N., Lander, E. S., Golub, T. R. A molecular signature of metastasis in primary solid tumors. Nat. Genet. 33, 49-54 (2003).
- Gilkes, D. M., et al. Collagen prolyl hydroxylases are essential for breast cancer metastasis. Cancer Res. 73, 3285-3296 (2013).
- Linder, S., Wiesner, C., Himmel, M. Degrading Devices: Invadosomes in Proteolytic Cell Invasion. Annu. Rev. Cell Dev. Biol. 27, 185-211 (2011).
- Murphy, D. A., Courtneidge, S. A. The ‘ins’ and ‘outs’ of podosomes and invadopodia: characteristics, formation and function. Nat. Rev. Mol. Cell Biol. 12, 413-426 (2011).
- Di Martino, J., et al. Cdc42 and Tks5: a minimal and universal molecular signature for functional invadosomes. Cell Adh. Migr. 8, 280-292 (2014).
- Poincloux, R., Lizarraga, F., Chavrier, P. Matrix invasion by tumour cells: a focus on MT1-MMP trafficking to invadopodia. J. Cell Sci. 122, 3015-3024 (2009).
- Juin, A., et al. Physiological type I collagen organization induces the formation of a novel class of linear invadosomes. Mol. Biol. Cell. 23, 297-309 (2012).
- Juin, A., et al. Discoidin domain receptor 1 controls linear invadosome formation via a Cdc42-Tuba pathway. J. Cell Biol. 207, 517-533 (2014).
- Martin, K. H., et al. Quantitative measurement of invadopodia-mediated extracellular matrix proteolysis in single and multicellular contexts. J. Vis. Exp. , e4119 (2012).
- Diaz, B. Invadopodia detection and gelatin degradation assay. Bio-protocol. 3, 1-8 (2013).
- Artym, V. V., et al. Dense fibrillar collagen is a potent inducer of invadopodia via a specific signaling network. J. Cell Biol. 208, 331-350 (2015).
- Schachtner, H., et al. Megakaryocytes assemble podosomes that degrade matrix and protrude through basement membrane. Blood. 121, 2542-2552 (2013).
- Schoumacher, M., Glentis, A., Gurchenkov, V. V., Vignjevic, D. M. Basement membrane invasion assays: native basement membrane and chemoinvasion assay. Methods Mol. Biol. 1046, 133-144 (2013).
- Wolf, K., Muller, R., Borgmann, S., Brocker, E. B., Friedl, P. Amoeboid shape change and contact guidance: T-lymphocyte crawling through fibrillar collagen is independent of matrix remodeling by MMPs and other proteases. Blood. 102, 3262-3269 (2003).
- Geraldo, S., et al. Do cancer cells have distinct adhesions in 3D collagen matrices and in vivo?. Eur. J. Cell Biol. 91, 930-937 (2012).
- Monteiro, P., et al. Endosomal WASH and exocyst complexes control exocytosis of MT1-MMP at invadopodia. J. Cell Biol. 203, 1063-1079 (2013).
- Wolf, K., et al. Collagen-based cell migration models in vitro and in vivo. Semin. Cell Dev. Biol. 20, 931-941 (2009).

