Native Chromatin Immunoprecipitation Using Murine Brain Tumor Neurospheres
Summary
Epigenetic mechanisms are frequently altered in glioma. Chromatin immunoprecipitation could be used to study the consequences of genetic alterations in glioma that result from changes in histone modifications which regulate chromatin structure and gene transcription. This protocol describes native chromatin immunoprecipitation on murine brain tumor neurospheres.
Abstract
Epigenetic modifications may be involved in the development and progression of glioma. Changes in methylation and acetylation of promoters and regulatory regions of oncogenes and tumor suppressors can lead to changes in gene expression and play an important role in the pathogenesis of brain tumors. Native chromatin immunoprecipitation (ChIP) is a popular technique that allows the detection of modifications or other proteins tightly bound to DNA. In contrast to cross-linked ChIP, in native ChIP, cells are not treated with formaldehyde to covalently link protein to DNA. This is advantageous because sometimes crosslinking may fix proteins that only transiently interact with DNA and do not have functional significance in gene regulation. In addition, antibodies are generally raised against unfixed peptides. Therefore, antibody specificity is increased in native ChIP. However, it is important to keep in mind that native ChIP is only applicable to study histones or other proteins that bind tightly to DNA. This protocol describes the native chromatin immunoprecipitation on murine brain tumor neurospheres.
Introduction
Epigenetic events are frequent in gliomas and likely play an important role in tumor pathogenesis. Indeed, in pediatric high-grade glioma, mutations in genes encoding histone variants H3.3 and H3.1 occur frequently1. The mutations affect histone modifications and have major epigenetic consequences2,3. In the adolescent to adult spectrum, recurrent mutations in isocitrate dehydrogenase gene 1/2 (IDH1/2), a mutation that inhibits α-KG dependent histone and DNA de-methylasaes, and genetic alterations in other chromatin regulators such as ATRX and DAXX occur4. Therefore, it is of critical importance to study how mutations that affect epigenetic regulators alter chromatin structure and regulatory histone modifications, which, in turn, have a dramatic impact of the tumor cells' transcriptome.
Chromatin immunoprecipitation (ChIP) is a powerful tool used to evaluate the impact of epigenetic modifications in the genome5,6,7. In native ChIP, chromatin is digested with micrococcal nuclease (MNase), immunoprecipitated using an antibody raised against the protein of interest, and then DNA is purified from the immunoprecipitated chromatin complex6. Cells are not fixed during the procedure so this technique is only applicable for the study of proteins that interact tightly with DNA6. The absence of cross-linking aids antibody specificity since antibodies are usually raised against unfixed peptides or proteins7. In addition, since there is no cross-linking step, this reduces the chances of fixing transient protein-DNA interactions that are non-specific and not regulatory7,8. ChIP can be used to identify the enrichment of histone modifications in a specific genomic region. Here, we detail a protocol for performing native ChIP in neurospheres (NS) generated from genetically engineered mouse models of glioma.
Protocol
All methods described here have been approved by the Institutional Animal Care and Use Committee (IACUC) of the University of Michigan.
1. Generation of brain tumor NS and culturing conditions.
- Prepare Neural Stem Cell Medium (NSC) with Dulbecco's Modified Eagle Medium/F12B27 supplement (1x), N2 supplement (1x), and an antimicrobial reagent. Supplement NSC medium on the day of use with human recombinant endothelial growth factor (EGF) and basic-fibroblast growth factor (FGF) at a final concentration of 20 ng/mL each.
- Induce the brain tumor in mouse using the Sleeping Beauty transposon system in which plasmids encoding oncogenes or shRNA targeting tumor suppressors is injected into the lateral ventricles of neonates to generate spontaneous brain tumors9. Select a mouse with a large tumor, which is confirmed with bioluminescence imaging.
NOTE: More detailed information about the Sleeping Beauty Transposon System and plasmid constructs used can be found in Calinescu et al.9.- To image the mice, inject 100 µL of 30 mg/mL luciferin solution intraperitoneally using a 26 gauge needle.
- 5 min after injection of luciferin, anesthetize the animal using an anesthesia chamber with oxygen/isoflurane flow set to 2% isoflurane.
- When the animals are anesthetized (usually 2-3 min), place the animals in the bioluminescence chamber with oxygen isoflurane set to 2% isoflurane. If the animals will be under anesthesia for longer than 5 min, use an ophthalmic ointment to prevent dryness while under anesthesia.
- Image the mice using an in vivo optical imaging system with the following parameters: automatic exposure, large binning, and aperture f=1. The parameter to consider it a large tumor is a bioluminescence >106 photons/s/cm2/sr.
- Euthanize the mouse with an overdose of isoflurane inhalation anesthetic in a closed chamber, check by toe pinch that the animal is anesthetized, and decapitate the mouse.
- Extract the brain and dissect it as described previously in Calinescu et al.9.
- Place the brain in a 10 cm Petri dish. If the tumor induced expresses a fluorescent protein, use a fluorescent dissecting microscope set to the appropriate fluorescent channel and 0.3X magnification, otherwise identify tumor mass by the differences in tissue color and texture. Use scalpel and forceps to separate the tumor tissue from the normal brain.
- Mince the tumor into small pieces in the Petri dish. Place the dissected tumor in a 1.5 mL tube with 300 µL of the NSC medium.
- Use a disposable plastic pellet pestle that fits to the walls of a conical microcentrifuge tube to gently homogenize the tumor. Add 1 mL of cell detachment medium and incubate the sample for 5 min at 37 °C.
NOTE: The use of a pestle may kill some cells. If the reader wants to maximize the cell viability, the use of a pestle may be omitted. - Pass the cell suspension from step 1.7 through a 70 µm cell strainer and wash the strainer with 25 mL of NSC medium.
- Centrifuge the suspension at 300 x g for 5 min at room temperature, decant the supernatant and re-suspend the pellet into 6 mL of NSC medium.
- Plate the tumor cell suspension from step 1.9 into a T-25 culture flask and culture it at 37 °C in a tissue culture incubator with an atmosphere of 95% air and 5% CO2. Usually after 3 days, there will be a mixture of some dead cells, adhered cells and some cells forming NS which float in the medium.
- Transfer the cell suspension into a 15 mL conical tube, collect only the cells that sink to the bottom using a micropipette and re-plate them onto a T-75 tissue culture flask.
- Maintain the cells at relatively high density (1-3 X 106 cells in a T-25). Passage the cells when they are confluent.
NOTE: Confluency is determined by the spheres size. Black centers of large spheres mean that there are dead cells in the center and indicate that the cells should be passaged immediately. - Add the growth factors (EGF and basic-FGF; 20 ng/mL each) every three days. Change the medium when the cells are not confluent and the medium turns light orange.
- To change the medium, collect the old medium and centrifuge it at 300 x g at room temperature for 5 min and re-suspend the cell pellet in NSC medium supplemented with EGF and FGF at the concentration specified in step 1.1.
NOTE: If the spheres are at mid-to high confluency, but not ready to be passaged (sphere size is small with a diameter < 100 µm), then the pellet can be split into two flasks. - To passage the cells, centrifuge the cells at 300 x g for 5 min at room temperature, decant the supernatant and incubate the cells in cell detachment medium for 5 min at 37 °C.
- Add 5 mL of balanced salt solution to dilute detachment medium and centrifuge it at 300 x g for 5 min at room temperature.
- Decant the supernatant, resuspend the cell pellet in 18 mL of NSC medium and divide the suspension equally into three T-25 flasks.
- To freeze the aliquots of cells, dissociate the cells as in step 1.15, and freeze the cells in 1 mL aliquots of 90% fetal bovine serum with 10% dimethyl sulfoxide. Slowly cool down the cells by placing the cells on ice for 10 min, then keeping the cells -20 °C for 30 min, and -80 °C for 1 day. The next day, transfer the cells to a liquid nitrogen storage container.
2. Native ChIP
- Chromatin preparation
- After making stock of derived NS, culture NS in a T-75 in 15 mL of NSC medium at 37 °C in a tissue culture incubator with an atmosphere of 95% air and 5% CO2 until NS become confluent. One confluent T-75 plate will yield 3-5 X 106 cells.
- When the cells become confluent, centrifuge NS at 300 x g for 5 min, decant the supernatant and add 1 mL of cell dissociation medium. Incubate the cells at 37 °C for 5 min. Add 5 mL of medium and count the cells using a hemocytometer10.
NOTE: 1 x 106 cells are needed per immunoprecipitation (IP) reaction. Note that a pre-immune serum or IgG control must also be included11. - Centrifuge the NS in one 1.5 mL microcentrifuge tube at 300 x g for 5 min, decant the supernatant and re-suspend NS pellet with 1 mL of balanced salt solution and transfer the suspension to low protein binding 1.5 mL microcentrifuge tubes.
- Centrifuge NS at 300 x g for 5 min, decant the supernatant and re-suspend the cell pellet in 95 µL of digestion buffer (50 mM Tris-HCl, pH 8.0, 1 mM CaCl2, 0.2% polyethylene glycol octylphenyl ether) per 1 x 106 cells supplemented with protease inhibitor cocktail at a high concentration (1:100). Immediately pipet the cells up and down to prevent clumping and avoid creating any bubbles.
- Resuspend MNase in 50% glycerol to make 1 mg/mL solution (1.15 units/µL, stored at -20 °C) that will be referred to as "1x" stock. Make a "0.1x" stock by doing a second 1:9 dilution with 50% glycerol (e.g., 900 µL of "1x" MNase and 100 µL of 50% glycerol) and store it at -20 °C.
- Mix 5 µL of "0.1x" MNase and 145 µL of digestion buffer. Keep the enzyme on ice all the times. Add 5 µL of diluted MNase in digestion buffer per 1 x 106 cells. Flick the tube to mix and place the tube in 37 °C block for exactly 12 min. Process the samples one at a time to ensure an accurate incubation time of 12 min.
- Add 10 µL of 10x MNase Stop Buffer (110 mM Tris-HCl, pH 8.0, 55 mM EDTA) per 1 x 106 cells. Samples must be kept on ice from this point onwards.
- Add 110 µL "2x RIPA buffer" (280 mM NaCl, 1.8% polyethylene glycol octylphenyl ether, 0.2% sodium dodecyl sulfate (SDS), 0.2% Na-deoxycholate, 5 mM ethylene glycol-bis(β-aminoethyl ether)-N,N,N',N'-tetraacetic acid (EGTA); stored at 4 °C) supplemented with protease inhibitor cocktail at high concentration (1:100) per 1 x 106 cells. Flick the tube to mix.
- Centrifuge the tubes for 15 min in a microfuge at 4 °C and 1700 x g. Transfer the supernatant to a new regular 1.5 mL microcentrifuge tubes (low protein binding microcentrifuge tubes are no longer necessary) and store them on ice. Discard the pellet.
- Immunoprecipitation (IP)
- Mix Protein A and Protein G magnetic beads in a 1:1 ratio. For every IP, prepare 25 µL of beads.
- Wash the beads by adding 1 mL of RIPA buffer (10 mM Tris pH 8.0, 1 mM EDTA, 140 mM NaCl, 1% polyethylene glycol octylphenyl ether, 0.1% SDS, 0.1% Na-deoxycholate; stored at 4 °C) supplemented with protease inhibitors at low concentration (1:1000).
- Use a magnet to allow beads to separate from the washing solution and decant the washing solution (approximately 1 min). Perform the wash step twice.
- Re-suspend the beads to the original volume using RIPA buffer supplemented with protease inhibitors (1:1000).
- If necessary, dilute the chromatin using RIPA buffer supplemented with protease inhibitors (1:1000) so that each IP has a volume of 100-200 µL. Reserve 10% of the volume used per IP of chromatin for the input.
NOTE: For example, if 200 µL are used per IP, 20 µL of diluted chromatin should be reserved for the input. For a total of four IPs, total chromatin volume should be diluted to 820 µL. - Prepare the input. Add 100 µL of TE buffer (10 mM Tris-HCl pH 8.0, 1 mM EDTA) supplemented with proteinase K (0.5 mg/mL) to the input and incubate the input at 55 °C for 1 h.
NOTE: Steps 2.2.6-2.2.7 can be performed along with steps 2.3.4 and 2.4.1. - Purify the input using a PCR purification kit following the manufacturer's instructions and elute in 50 µL of elution buffer provided in kit.
- Measure the concentration of the input using a microvolume spectrophotometer and record the results in a notebook. Blank with elution buffer from kit.
NOTE: In mouse neurospheres, we used approximately 6 µg of DNA for each IP. - Run input on a 1% gel or load 1 µL on a DNA Bioanalyzer using standard settings. Store the remainder for use as input in downstream applications.
- Add 10 µL of washed protein A & G magnetic beads for every IP and incubate IP for 1 h at 4 °C.
NOTE: For example, add 30 µL of protein A & G magnetic beads for three IPs. - Place the samples on a magnet and allow beads to separate. Divide chromatin by transferring the previously determined amount into a new 1.5 mL tube.
- Add the antibody and wrap the caps with plastic paraffin film to avoid evaporation. Incubate the samples overnight at 4 °C with rotation at 20 rpm.
NOTE: Concentration should be determined empirically following manufacturer recommendations. - The next day, spin the tubes using a mini centrifuge at room temperature, 2000 x g, for 10 s. Then add 10 µL of beads to each IP and incubate IP for 3 h at 4 °C with rotation at 20 rpm.
- IP Washes and Protein Digestion
- Add 150 µL of RIPA buffer supplemented with protease inhibitors at low concentration (1:1000) to each IP and incubate IP at 4 °C with rotation at 20 rpm for 5 min. Place the sample on a magnetic stand and allow the magnetic beads to separate. Use a pipet to remove the supernatant. Repeat this tep five washes.
- Add 150 µL of LiCl buffer (250 mM LiCl, 10 mM Tris pH 8.0, 1 mM EDTA, 0.5% NP-40, 0.5 % Na-deoxycholate; store at 4 °C) supplemented with protease inhibitors at low concentration (1:1000) to each IP and incubate IP at 4 °C with rotation at 20 rpm for 5 min. Place the sample on a magnetic stand and allow magnetic beads to separate. Use a pipet to remove supernatant.
- Add 150 µL of cold TE (no protease inhibitors) and incubate the sample at 4 °C with rotation at 20 rpm for 5 min. Place the sample on a magnetic stand and allow magnetic beads to separate. Use a pipet to remove the supernatant.
- After the final washing step, re-suspend the sample in 100 µL of TE buffer supplemented with proteinase K (0.5 mg/mL) and incubate the sample at 55 °C for 1 h.
- DNA Purification and ChIP Quantitative Real-Time PCR (qPCR)
- Purify immunoprecipitated DNA using a PCR purification kit. After addition of DNA binding buffer from the kit to each sample, place the tube on a magnet and wait until magnetic beads aggregate, then transfer the supernatant to the cleanup column. Follow the manufacturer's instructions and elute in 50 µL of elution buffer provided in kit.
- To check if the IP is successful, perform a qPCR. Ideally, primers should be designed for positive and negative control regions.
NOTE: For example, for an IP performed with H3K4me3 and H3K27me3, the following samples should be run on qPCR: H3K4me3, H3K27me3, and IgG ChIP DNA, input DNA, and a no template control (NTC) using primers for regions to be enriched with H3K4me3 (positive control) and regions to be enriched with H3K4me3 (negative control); likewise, for H3K27me3. - Follow standard protocols to perform qPCR12. Briefly, perform a qPCR using SYBR green master mix, 0.5 µL of 10 µM working concentration of each forward and reverse primer, and 2 µL of ChIP or input DNA. Read plates using a Real Time PCR system. Primer sequences are provided in Table 2. The Gapdh primer sequences from Hwang et al. were used13.
- Analyze ChIP data relative to the input, i.e., percent input method (% IP)14. Calculate the percent input using the following formulas:
Adjusted input = mean ct(input)-log2(DF)
where DF is the dilution factor of the starting input and
DF = total volume of IP/ volume of input
% IP = 100×2^(Adjusted input – Ct (IP))
where Ct is the threshold factor.
Representative Results
A schematic representation of tumor NS generated from a brain tumor where brain tumor cells are Katushka positive is presented in Figure 1. Figure 2 is a schematic representation of the entire ChIP technique. Figure 3 shows the representative results of chromatin from brain tumor NS digested with MNase for 12 min, yielding a majority of mono, di-, and tri- nucleosomes. Following ChIP, a qPCR may be performed on the ChIP and input DNA samples. Figure 4 shows representative ChIP qPCR data from a qPCR for glyceraldehyde-3-phosphate dehydrogenase (Gapdh). Gapdh is a housekeeping gene that is enriched with H3K4me3, a modification that is associated with active transcription. The results show that Gapdh is enriched with H3k4me3 and are not enriched with H3K27me3 which is a modification associated with regions of repressed chromatin.
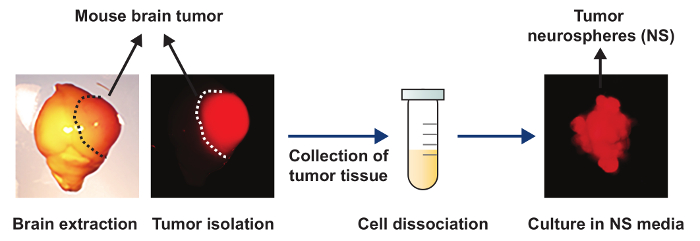
Figure 1: Schematic of tumor NS generated from a Katushka positive brain tumor. A bright field and fluorescent image of a brain harvested from a mouse that reached the end point stage. The tumor area is delineated by the dotted line and is positive for the fluorescent reporter, Katushka. The tumor is then isolated and the tumor tissue is collected in a 1.5 mL tube. The cells are then dissociated and cultured in NSC medium and tumor NS form. Please click here to view a larger version of this figure.
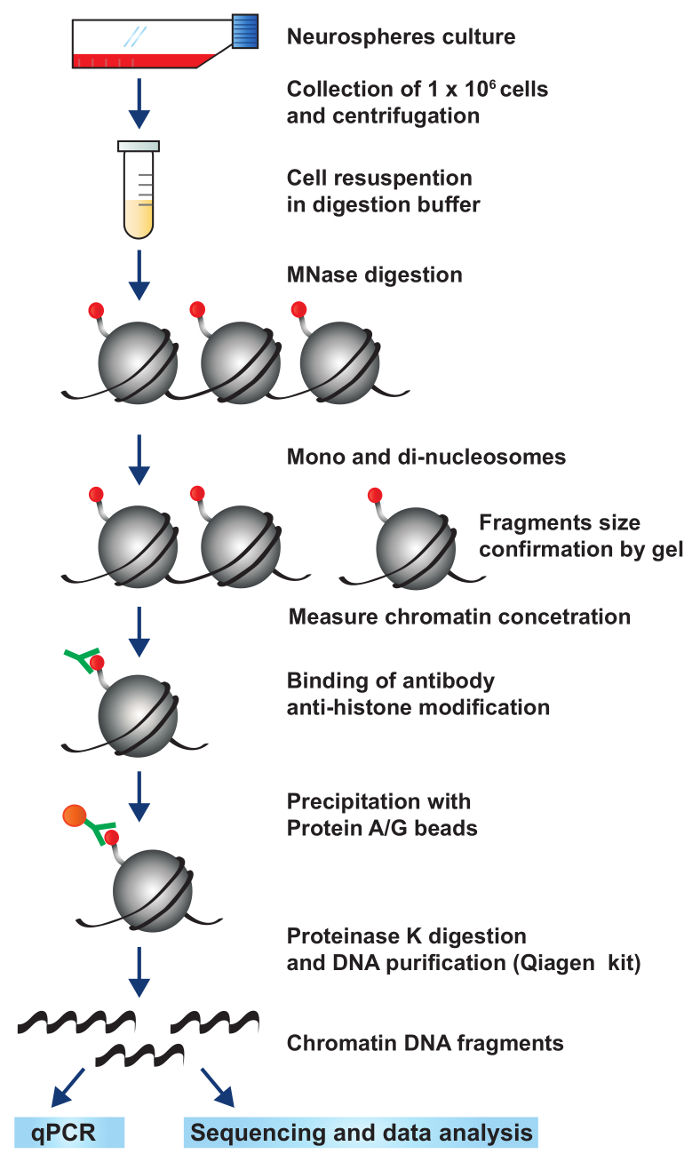
Figure 2: A schematic representation of the workflow for native ChIP. Tumor NS are cultured and expanded. Each IP is performed with 1 X 106 cells. Chromatin is fragmented using by MNase digestion to obtain mono, di-, and tri- nucleosomes. The chromatin is incubated with an antibody specific for a histone modification or DNA-associated protein of interest. The antibody-DNA complex is immunoprecipitated with magnetic protein A/G beads. Finally, protein is digested and DNA is purified to obtain only DNA enriched with histone modification or DNA-associated protein of interest. Please click here to view a larger version of this figure.
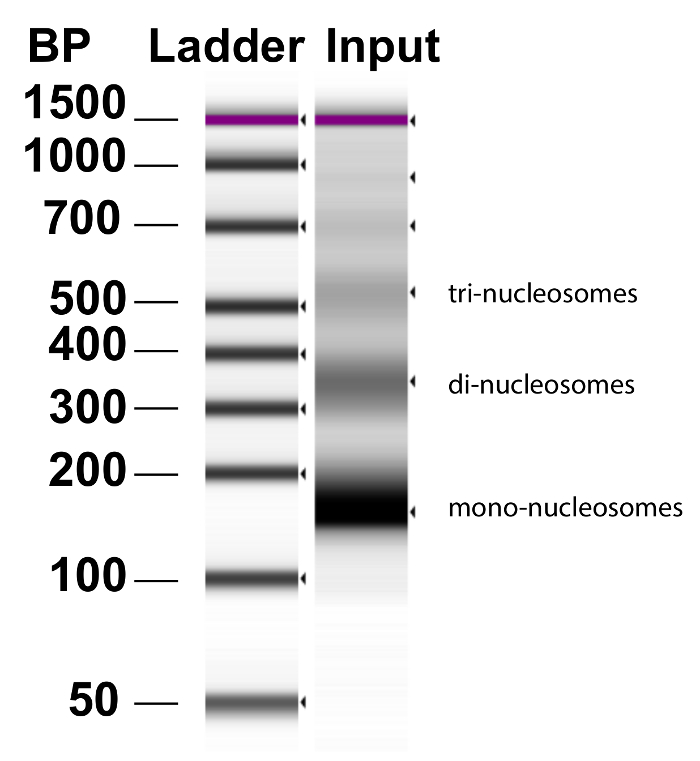
Figure 3: Chromatin fragmentation by MNase. Chromatin was prepared from brain tumor NS by the addition MNase and incubation at 37 °C for exactly 12 min. Representative DNA results from Bioanalyzer analysis of an input sample demonstrate that the majority of the DNA has been fragmented into mono-, di-, and tri- nucleosomes. Lane 1 is the ladder in base pairs (BP). Please click here to view a larger version of this figure.
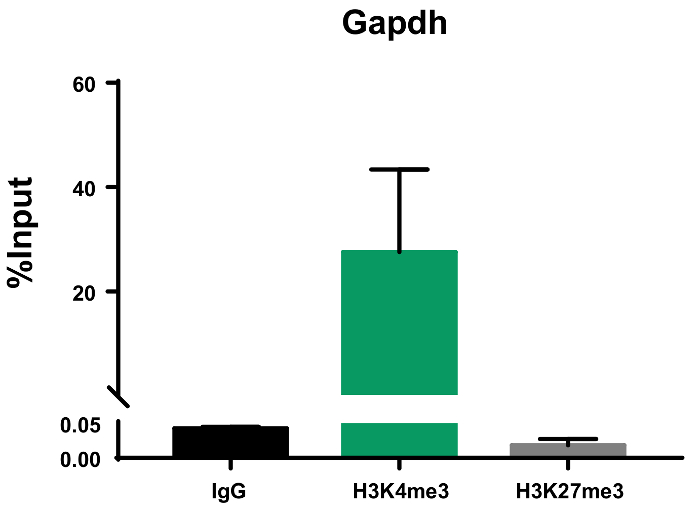
Figure 4: Representative ChIP qPCR data presented as percent input. qPCR was performed with IgG, H3K4me3, and H3K27me3 ChIP DNA using primers for glyceraldehyde-3-phosphate dehydrogenase (Gapdh). Representative results demonstrate that Gapdh, a housekeeping gene, is only enriched with the H3K4me3 modification, associated with active transcription, and not the H3K27me3 modification, associated with repressed chromatin. This graph represents the results of two biological experiments run with three replicate wells each. Error bars represent standard error of the mean (SEM). Please click here to view a larger version of this figure.
| Gene | Forward primer | Reverse primer |
| Gapdh | TCCCCTCCCCCTATCAGTTC | GACCCGCCTCATTTTTGAAA |
Table 1: Primers used for ChIP qPCR experiments. The sequences for primers used for Gapdh are provided in this table.
| mean Ct (input) | Standard Deviation | Adjusted Input | |
| 24.265 | 0.071 | 20.943 | |
| Percent input | |||
| Sample | Raw Mean Ct | Standard Deviation | Percent input = 100*2^ (adjusted input-Ct(IP)) |
| IgG | 32.23 | 0.112 | 0.04 |
| H3K4me3 | 22.148 | 0.128 | 43.37 |
| H3k27me3 | 32.79 | 0.519 | 0.027 |
| NTC | undetermined | undetermined | undetermined |
Table 2: Sample calculation of the percent input method for ChIP qPCR analysis. Sample calculation of percent input for ChIP performed with 10% starting input; DF = 10. The numbers in this table illustrate the raw values of one biological experiment with 3 replicate wells run for each sample.
Discussion
The protocol presented here will enable the user to perform native ChIP on NS derived from genetically engineered brain tumors. In contrast to cross-linking ChIP, this protocol is limited for the study of proteins that associate tightly with DNA6. The number of cells used can be modified as necessary and the protocol can be scaled up. We used 1 X 106 cells per IP, however, native ChIP may also be performed with as little as 4 x 104 cells5. In this protocol, we analyzed ChIP DNA via qPCR; however, this protocol may also be combined with next generation sequencing to study protein-DNA interactions on a genome wide level. We have used this ChIP protocol successfully to analyze the effect of glioma oncogenes on the deposition of histone modifications within specific regions of the tumor cells genome.
There are a few critical steps when performing native ChIP. First, the digestion conditions for each cell type must be empirically determined. The majority of the chromatin should be mono-nucleosomes (150 bp) with some di- and tri- nucleosome peaks (Figure 3). Our protocol results in more than 70% mono-, di-, or tri- nucleosome peaks, however, some undigested DNA remains. If the protocol will be used for ChIP-seq, additional steps will be required to remove undigested DNA. In addition, it is important to keep in mind that each batch of MNase purchased may differ in activity and thus require optimization of the digestion conditions. Second, the quality of a ChIP depends highly on the specificity of the antibody used11. A high-quality ChIP antibody should have high reactivity against the intended target and low cross reactivity with other DNA-associated proteins or off target histone modifications11. For this reason, we recommend testing antibody specificity using a peptide binding test. ENCODE guidelines suggest that a ten-fold enrichment should be observed for the modification of interest relative to other modifications11. It is also critical to perform necessary control immunoprecipations, i.e., pre-immune or IgG control. When these considerations are met, native ChIP data is a strong reliable method to study epigenetic mechanisms regulated by protein-DNA interactions.
Disclosures
The authors have nothing to disclose.
Acknowledgements
This work was supported by National Institutes of Health/National Institute of Neurological Disorders & Stroke (NIH/NINDS) Grants R37-NS094804, R01-NS074387, R21-NS091555 to M.G.C.; NIH/NINDS Grants R01-NS076991, R01-NS082311, and R01-NS096756 to P.R.L.; NIH/NINDS R01-EB022563; the Department of Neurosurgery; Leah's Happy Hearts, and Chad Though Foundation to M.G.C. and P.R.L. RNA Biomedicine Grant F046166 to M.G.C. F.M.M. is supported by an F31 NIH/NINDS-F31NS103500. R.I.Z.-V. is supported by the NIH/ NIGMS grant 5T34GM007821-37.
Materials
| Agilent 2100 Bioanalyzer | Agilent | G2946-90004 | bioanalyzer |
| AccuSpin Micro 17R, refrigerated | Fisher Scientific | 13-100-676 | |
| C57BL/6 | Taconic | B6-f | C57BL/6 mouse |
| Calcium Chloride | Aldrich | 22350-6 | buffer reagent |
| D-Luciferin, Potassium Salt | Goldbio | LuckK-1g | |
| DiaMag1.5 magnetic rack | Diagenode | B04000003 | for magnetic bead washes |
| DiaMag Rotator EU | Diagenode | B05000001 | rotator |
| DMEM/F-12 | Gibco | 11330-057 | NSC component |
| Dynabeads Protein A | Thermo Fisher Scientific | 10001D | protein A magnetic beads |
| Dynabeads Protein G | Thermo Fisher Scientific | 10003D | protein G magnetic beads |
| EGF | PeproTech | AF-100-15 | prepare 20 μg/mL stock in 0.1% BSA and aliquot. |
| Ethylenediaminetetraacetic acid (EDTA) | Sigma | E-4884 | buffer reagent |
| ethylene glycol-bis(β-aminoethyl ether)-N,N,N',N'-tetraacetic acid) (EGTA) | Sigma | E-4378 | buffer reagent |
| Fast SYBR Green Master Mix | Applied Biosystems | 4385612 | qPCR reagent |
| Fetal Bovine Serum | Gibco | 10437028 | for freezing cells |
| FGF | PeproTech | 100-18b | Prepare 20 μg/mL stock in 0.1% BSA and aliquot. |
| Forceps | Fine Science Tools | 11008-13 | for dissection of tumor |
| Glycerol, MB Grade | EMD- Millipore | 356352 | |
| H3K4me3 | Abcam | Ab8580 | |
| H3K27me3 | Millipore | 07-449 | |
| HBSS | Gibco | 14175-103 | balanced salt solution |
| Fluriso | VETone | 501017 | inhalation anesthetic |
| Ivis Spectrum | Perkin-Elmer | 124262 | in vivo optical imaging system |
| Hyqtase | HyClon | SV3003001 | cell detachment media |
| Lithium Chloride | Sigma | L8895 | buffer reagent |
| Low binding microtubes | Corning Costar | CLS3207 | low protein binding microcentrifuge tube |
| Microcentrifuge tube | Fisher | 21-402-903 | regular microcentrifuge tube |
| Micrococcal Nuclease | Thermo Fisher Scientific, Affymetrix | 70196Y | each batch may differ; purchase sufficient amount for experiments and aliquot. |
| N2 | Gibco | 17502-048 | NSC component |
| Normal Rabbit IgG | Millipore | 12-372 | |
| Normocin | Invivogen | NOL-36-063 | anti-microbial agent, use at 0.1 mg/mL. |
| NP-40 (Igepal CA-630) | Sigma | 18896-50ML | buffer reagent |
| Kimble Kontes Pellet Pestle | Fisher Scientific | K749515-0000 | |
| Protease Inhibitor Cocktail | Sigma-Aldrich | P8340 | aliquot and store at -20 °C. |
| Protinase K | Sigma-Aldrich | P2308 | make 10 mg/mL stock in water; aliquot and store at -20 °C. |
| QIAquick PCR Purification Kit | Qiagen | 28104 | DNA purification kit |
| Scalpel | Fine Science Tools | 10007-16 | for dissection of tumor |
| Sodium Chloride | VWR | 0241-5KG | buffer reagent |
| Sodium Deoxycholate | Sigma-Aldrich | D670-25G | buffer reagent |
| Sodium Dodecyl sulfate (SDS) | Sigma | L-4390 | buffer reagent |
| Tris Base | Thermo Fisher Scientific | Bp152-1 | buffer reagent |
| Triton X-100 | Thermo Fisher Scientific | BP 151-500 | polyethylene glycol octylphenyl ether |
| Standard Mini Centrifuge | Fisherbrand | 12-006-901 | standard mini centrifuge |
| SZX16 microscope | Olympus | SZX16 | flourescent dissecting microscope |
| ViiA 7 Real-Time PCR System with Fast 96-Well Block | Applied Biosystems | 4453535 | |
| Nanodrop One | Thermo-Fisher Scientific | ND-ONEC-W |
References
- Schwartzentruber, J., et al. Driver mutations in histone H3.3 and chromatin remodelling genes in paediatric glioblastoma. Nature. 482 (7384), 226-231 (2012).
- Bjerke, L., et al. Histone H3.3. mutations drive pediatric glioblastoma through upregulation of MYCN. Cancer Discov. 3 (5), 512-519 (2013).
- Bender, S., et al. Reduced H3K27me3 and DNA hypomethylation are major drivers of gene expression in K27M mutant pediatric high-grade gliomas. Cancer Cell. 24 (5), 660-672 (2013).
- Maleszewska, M., Kaminska, B. Is glioblastoma an epigenetic malignancy?. Cancers (Basel). 5 (3), 1120-1139 (2013).
- Gilfillan, G. D., et al. Limitations and possibilities of low cell number ChIP-seq. BMC Genomics. 13, 645 (2012).
- Thorne, A. W., Myers, F. A., Hebbes, T. R. Native chromatin immunoprecipitation. Methods Mol Biol. 287, 21-44 (2004).
- Turner, B. . Mapping Protein/DNA Interactions by Cross-Linking. , (2001).
- Tseng, Z., Wu, T., Liu, Y., Zhong, M., Xiao, A. Using native chromatin immunoprecipitation to interrogate histone variant protein deposition in embryonic stem cells. Methods Mol Biol. 1176, 11-22 (2014).
- Calinescu, A. A., et al. Transposon mediumted integration of plasmid DNA into the subventricular zone of neonatal mice to generate novel models of glioblastoma. J Vis Exp. (96), (2015).
- Sambrook, J., Russell, D. W. Estimation of cell number by hemocytometry counting. CSH Protoc. 2006 (1), (2006).
- Landt, S. G., et al. ChIP-seq guidelines and practices of the ENCODE and modENCODE consortia. Genome Res. 22 (9), 1813-1831 (2012).
- Kubista, M., et al. The real-time polymerase chain reaction. Mol Aspects Med. 27 (2-3), 95-125 (2006).
- Hwang, W. W., et al. Distinct and separable roles for EZH2 in neurogenic astroglia. Elife. 3, e02439 (2014).
- Haring, M., et al. Chromatin immunoprecipitation: optimization, quantitative analysis and data normalization. Plant Methods. 3, 11 (2007).

