Adeno-associated Virus-mediated Transgene Expression in Genetically Defined Neurons of the Spinal Cord
Summary
Intraspinal injection of recombinase dependent recombinant adeno-associated virus (rAAV) can be used to manipulate any genetically labelled cell type in the spinal cord. Here we describe how to transduce neurons in the dorsal horn of the lumbar spinal cord. This technique enables functional interrogation of the manipulated neuron subtype.
Abstract
Selective manipulation of spinal neuronal subpopulations has mainly been achieved by two different methods: 1) Intersectional genetics, whereby double or triple transgenic mice are generated in order to achieve selective expression of a reporter or effector gene (e.g., from the Rosa26 locus) in the desired spinal population. 2) Intraspinal injection of Cre-dependent recombinant adeno-associated virus (rAAV); here Cre-dependent AAV vectors coding for the reporter or effector gene of choice are injected into the spinal cord of mice expressing Cre recombinase in the desired neuronal subpopulation. This protocol describes how to generate Cre-dependent rAAV vectors and how to transduce neurons in the dorsal horn of the lumbar spinal cord segments L3-L5 with rAAVs. As the lumbar spinal segments L3-L5 are innervated by those peripheral sensory neurons that transmit sensory information from the hindlimbs, spontaneous behavior and responses to sensory tests applied to the hindlimb ipsilateral to the injection side can be analyzed in order to interrogate the function of the manipulated neurons in sensory processing. We provide examples of how this technique can be used to analyze genetically defined subsets of spinal neurons. The main advantages of virus-mediated transgene expression in Cre transgenic mice compared to classical reporter mouse-induced transgene expression are the following: 1) Different Cre-dependent rAAVs encoding various reporter or effector proteins can be injected into a single Cre transgenic line, thus overcoming the need to create several multiple transgenic mouse lines. 2) Intraspinal injection limits manipulation of Cre-expressing cells to the injection site and to the time after injection. The main disadvantages are: 1) Reporter gene expression from rAAVs is more variable. 2) Surgery is required to transduce the spinal neurons of interest. Which of the two methods is more appropriate depends on the neuron population and research question to be addressed.
Introduction
The dorsal spinal cord is essential for information exchange between the periphery of the body and the brain. Sensory stimuli such as heat, cold, touch, or noxious stimuli are detected by specialized peripheral neurons, which convey this information to neurons of the spinal cord dorsal horn. Here, a complex network of inhibitory and excitatory interneurons modulates and eventually relays sensory information via spinal projection neurons to supraspinal sites1,2. The computations carried out by spinal inter- and projection neurons gate sensory information, thus determining which information is suppressed or relayed at which intensity. Changes in the integration of sensory stimuli, such as an altered balance between inhibition and excitation, can cause sensory dysfunctions such as hypersensitivity or allodynia (painful sensations after normally non-painful stimulation). These changes are thought to be the underlying cause of various chronic pain states3,4. Thus, spinal circuits are of high importance in sensory processing and consequently in the perception of an organism's environment and self. With the recent advent and combination of molecular, genetic, and surgical techniques that allow the precise manipulation of genetically identified spinal neuron subpopulations, scientists are now beginning to understand the underlying spinal circuits responsible for the processing of distinct sensory modalities.
Intraspinal injection of rAAV into wild-type or transgenic mice has greatly contributed to the manipulation, analysis, and understanding of the function of specific subsets of spinal neurons5,6,7,8,9,10,11. This technique allows the delivery of marker proteins (such as GFP/ GFP fusion proteins), reporter proteins (such as GCaMP), or effector proteins (such as bacterial toxins, channelrhodopsin, or pharmacogenetic receptors) in a spatially restricted manner to spinal neurons. Local injection of Cre-dependent rAAVs into transgenic mice expressing Cre recombinase in a specific subset of spinal neurons allows the specific analysis of the respective neuronal population. We have employed this technique to label, ablate, inhibit or activate spinal glycinergic neurons demonstrating that they are an essential part of the spinal gate controlling pain and itch transmission7. In these experiments, intraspinal injection of Cre-dependent rAAV into GlyT2::Cre mice enabled the selective manipulation of glycinergic neurons in the lumbar spinal cord. Thereby, simultaneous manipulation of supraspinal circuits that contain glycinergic neurons critical for the survival of the animal can be avoided.
While an intraspinal injection of rAAVs limits infection to the site of injection, viral transduction can occur not only in local neurons but also in neurons that connect to the injection site via axonal projections. The latter is often used to trace CNS areas providing neuronal input to a particular nucleus in the brain. The infection of axonal projections can, however, also be a confounding factor when a defined population of neurons shall be studied at a particular site. To address these issues, we have recently conducted a comprehensive analysis of AAV serotypes and expression cassettes to identify serotypes and promoters that can be used to either minimize or maximize retrograde transduction. In the context of this specific research in spinal circuits, we analyzed the ability of different serotypes and promoters to retrogradely transduce neurons in the dorsal root ganglia (DRG), the rostral ventromedial medulla (RVM), and the somatosensory cortex12. The technique outlined in this protocol can therefore be used either to analyze spinal neurons at the injection site or to analyze projection neurons that provide input to the injected site of the spinal cord. In the protocol described here, three injections of rAAV into the left side of the lumbar spinal cord are performed to enable transduction of neurons in the three lumbar segments (L3-L5). The L3-L5 segments receive the majority of the sensory input from the hindlimb ipsilateral to the injection site. We demonstrate that functional manipulation of genetically labeled neurons in L3-L5 is sufficient to evoke robust behavioral changes, thus providing functional evidence for the circuit function of such a genetically labeled neuron subtype.
Protocol
All animal experiments were approved by the Swiss cantonal veterinary office (Zurich) and are in accordance and compliance with all relevant regulatory and institutional guidelines.
NOTE: All materials along with respective manufacturers and/or vendors are listed in the Table of Materials.
1. Generation of Cre-dependent AAV Vectors
NOTE: A variety of Cre-dependent vectors with different promoters can be purchased (see Table of Materials) or, if the desired expression construct is not available, it can be generated by modifying existing AAV constructs. Note, the promoter and serotype can have an impact on the spread of viral transduction (see 12). The first part of this protocol briefly describes the generation of two different Cre-dependent AAV vectors suitable for gain and loss of function experiments, respectively.
- Pharmacogenetic Activation (Gain of Function)
- Order pAAV.hSyn.flex.hM3D(Gq)-mCherry (see Table of Materials) for designer receptors exclusively activated by designer drugs (DREADD)-mediated activation.
- Upon receiving the bacterial stab culture, streak out bacteria on LB plates (bacteriological-grade agar dissolved in Luria-Bertani liquid medium) supplemented with the appropriate antibiotic. Incubate over night at 37 °C and pick a single colony on the next day.
NOTE: Plasmids containing AAV vector genomes can be unstable and recombination of the viral genome, especially within the viral inverted terminal repeats (ITR), can occur during amplification in E. coli. We suggest using recA-deficient/-suppressed strains such as MDS42 or Stbl3 to avoid recombination within the plasmid containing the AAV genome. - Amplify bacteria following the instructions given by the vendor of the chosen DNA-Maxi-Kit and prepare high quality plasmid DNA with a commercially available DNA-Maxi-Kit (e.g., see Table of Materials). Perform a quality control of the plasmid DNA preparation by verifying the integrity and identity of the plasmid DNA through restriction digest of an aliquot of the plasmid DNA.
NOTE: There are recognition sites for the restriction endonuclease SmaI within the ITRs. Include a SmaI digest in the quality control to verify the integrity of the ITRs. - Send the DNA to a viral vector core facility in order to produce high quality rAAV.
NOTE: High titer, high quality viral preps are crucial for avoiding confounding phenotypes caused by contaminations in the viral prep. We therefore recommend having the rAAV of choice produced in an established vector core facility, unless the AAV production is well established in the laboratory.
- Ablation (Loss of Function)
NOTE: Bacterial toxins or toxin receptors can be used to mediate cell ablation or neuronal silencing. We have generated pAAV.EF1α.flex.DTA to express diphtheria toxin fragment A (DTA) in a Cre-dependent manner.- To generate a Cre-dependent viral vector, choose a Cre-dependent vector with a suitable promoter (e.g., pAAV.EF1α.flex.hChR2(H134R)-eYFP). Amplify the coding sequence of choice (e.g., DTA) by polymerase chain reaction with primers that include AscI and NheI or compatible restriction endonuclease recognition sites in their overhangs (see Foster et al.7)
- Using standard molecular biology techniques, perform restriction digests of the vector DNA (pAAV.EF1α.FLEX.hChR2(H134R)-eYFP) and of the PCR-amplified insert (NheI-DTA-AscI) with the restriction endonucleases AscI and NheI. Ligate the purified fragments.
- Transform the ligation reaction into suitable bacteria, according to the protocol given by the supplier of the competent bacteria of choice, and plate the bacteria onto LB plates. Use recA-deficient/-suppressed strains, such as MDS42 or Stbl3, for cloning and subsequent amplification of the plasmid DNA.
- On the day after transformation, pick clones and inoculate them in LB medium supplemented with the appropriate antibiotic overnight at 37 °C. On the next day, extract plasmid DNA from cultured bacteria. Verify successful cloning by restriction digests and sequencing of the extracted plasmid DNA.
- Amplify high quality, bacterial plasmid DNA (see step 1.1.3). Send amplified DNA to a viral vector core facility for virus production.
2. Transduction of Spinal Cells
- Preparation of Virus Solution
Caution: Viruses are infectious reagents and should be handled according to the relevant guidelines. In most cases, rAAVs can be handled at biosafety level 1 (BSL1).- On the day of injection, defrost a stock aliquot of the desired purified virus on ice and keep it on ice until directly before the injection. Avoid repeated freeze-thaw-cycles, as these will reduce the effective titer of the virus.
NOTE: If necessary, the defrosted virus aliquot can be kept at 4 °C for up to 3 days. - Dilute the virus particles in sterile 0.9% NaCl or phosphate buffered saline (PBS).
NOTE: The appropriate titer depends on the experimental aims and should be determined experimentally. A good total viral load to start with is 3×109 genome copies (3.33×1012 GC/mL, 3×300 nL injected). Taking excess and some loss while loading the capillary into account, about 2.5 µL of virus solution will be needed for each mouse.
- On the day of injection, defrost a stock aliquot of the desired purified virus on ice and keep it on ice until directly before the injection. Avoid repeated freeze-thaw-cycles, as these will reduce the effective titer of the virus.
- Preparation of Micropipettes for Intraspinal Injections
- 'Pull' thin-wall glass capillaries (outer diameter 1 mm) on a micropipette puller to create a ~5.5 mm long, shallow shank. Use a 3.0 mm heater filament and settings of program 00 with P(A) adapted as in Table 1.
NOTE: Pulling can be done in advance. The pulled capillaries should be stored in a closed container on modeling clay to avoid dust, which could later clog the micropipette. Autoclaving of micropipettes is not necessary. - Clip the shank of the pulled capillary with laminectomy forceps to a length of about 4.5 to 5 mm to create a tip opening with an inner diameter of 25-35 μm. Using a microscope with a micrometer scale, measure the inner diameter of several micropipettes to get a feeling for how large the opening should be, or measure for every micropipette.
NOTE: A smaller tip can lead to clogging; a larger tip might cause tissue damage and difficulties to penetrate the spinal cord.
- 'Pull' thin-wall glass capillaries (outer diameter 1 mm) on a micropipette puller to create a ~5.5 mm long, shallow shank. Use a 3.0 mm heater filament and settings of program 00 with P(A) adapted as in Table 1.
- Preparation of Surgical Setup and Tools
- Ensure the equipment is clean and ready to be used. Disinfect the work area by wiping with 70% ethanol and sterilize tools by autoclaving or disinfecting them. Do not leave any disinfectant on the microliter syringe that would harm the viral vector to be used.
- Anesthesia and Preparation of the Animal for Surgery
NOTE: The total duration of the surgery is 60-90 min.- Choose 6- to 10-week old mice to facilitate the surgical procedure.
NOTE: If the experimental design requires it, the injection of younger or older animals is possible. Any strain and both sexes can be used in principle, but use the same background strain (e.g., C57BL/6J) for comparison of behaviors between different transgenic mouse lines. If sex differences are expected or to be investigated, analyze males and females in separate groups. - Induce anesthesia using ~5% isoflurane. Place the animal in the stereotaxic frame on a heat mat and maintain anesthesia at 1-2.5% isoflurane (air flow rate 900 mL/min). Monitor the respiration rate throughout the surgery.
- Apply lubricant eye ointment to prevent corneal drying during the surgery.
- Shave the animal's back and remove hair carefully with wet tissues. Disinfect the shaved skin with iodine solution and allow the skin to dry.
- Give analgesic treatment (e.g., 0.1-0.2 mg/kg buprenorphine subcutaneously).
- Choose 6- to 10-week old mice to facilitate the surgical procedure.
- Exposure of the Vertebral Column at the Lumbar Spinal Cord Level
NOTE: Due to the differential development of the spinal cord and the vertebral column, the respective levels are not aligned, but the lumbar spinal cord segment L4 is at the same level as the thoracic vertebra T13.- Locate the most caudal rib pair by palpating along the vertebral column. Using a scalpel, make a 1.5-2.5 cm longitudinal cut into the skin starting from just rostral to the most caudal rib pair.
- Lift the skin with forceps and detach it from the underlying muscle with scissors. Throughout the surgery, keep the exposed tissues moist with sterile 0.9% NaCl.
- Using fine forceps and small scissors, make an incision into the next, thin membranous layer right next to the midline and cut it off from the spinous processes. It is separate from the underlying paraspinous muscle, but attached to the spinous processes.
- Identification of the Vertebrae at the Lumbar Spinal Cord Level
NOTE: Once the vertebral column is exposed, the intertransverse ligaments and the dorsal spinous processes should be visible. Several anatomical landmarks can aid in identifying the correct vertebra to target (Figure 1B).- Pull the skin back towards the tail to expose the iliac crest. At the same level, the most caudal pair of visible intertransverse ligaments joins the L6 spinous process. Count backwards in a caudal to rostral direction to identify the vertebra of interest.
- Palpate along the vertebral column. The most caudal rib pair is located just rostral to the T13 vertebra and the pelvic bone at hip level is at the level of the L6 vertebra.
- Locate the spot where the tendon along the side of the vertebral column is whitest and most medial. The T13 vertebra is located just rostral. The lumbar spinal cord segment L4 is located within this vertebra.
- Fixation of the Vertebral Column and Exposure of the Lumbar Spinal Cord
- Place the animal onto a cushion of rolled up tissues to elevate it to the spinal clamps of the stereotaxic frame.
- Fix the target vertebra to avoid movements of the vertebral column due to breathing. For this aim, align the clamps adjacent to the target vertebra, fix one clamp in position and hold the vertebral column with Adson forceps while fixing the second clamp.
NOTE: The column should be perpendicular to the injection plane and firmly fixed so that it does not move upon pressure from above. If necessary, a second pair of clamps can be fixed to the vertebral column. In this case, it is useful to fix the two pairs of clamps to the two vertebrae adjacent to the target vertebra. - Remove the paraspinous muscle above the vertebrae of interest. Using a scalpel, make a parallel incision just medial to the tendons that are parallel to the vertebral column, as well as perpendicular incisions rostral and caudal to the target vertebra. Be careful not to cut too deep. Tear/cut away the muscle using rongeurs. Use forceps as needed to remove tissue remaining on the vertebra or above the dura in the intervertebral space.
- In the intervertebral space, the dorsal blood vessel should be visible marking the midline of the spinal cord. For unilateral injections, perform a partial laminectomy by drilling a hole into the middle of the target side of the vertebra. Use a fine dentistry drilling apparatus with a 0.5 mm spherical cutter and drill especially carefully when approaching the spinal cord. Remove any remaining bone fragments with a 26G beveled needle to expose the spinal cord.
- Using a 26G beveled needle, perforate the dura in the drilled hole, and in the intervertebral space rostral and caudal to the target vertebra, all approximately 200 µm lateral to the dorsal blood vessel. Cerebrospinal fluid should be escaping from the holes and the spinal cord should be slightly bulging out.
- Preparation of the Injection Syringe
NOTE: The injection syringe should be prepared immediately before the start of the injection to reduce the risk of dust particles clogging the micropipette.- Mount the glass capillary onto a microliter syringe using the removable needle compression fitting kit. Fasten the nut tightly to ensure a tight and secure fit.
- Fill the microliter syringe with sterile distilled water and start pressing it out with the plunger.
NOTE: If there is too much resistance so that the water does not easily come out of the tip, the micropipette is likely blocked and should be exchanged. - Mount the syringe onto a micromanipulator connected to an electronically controlled microinjector. Be careful not to touch anything with the micropipette.
- Using the microinjector, draw up about 1 µL of air to create a bubble between the water and the virus.
- Place a 2.5 µL droplet of virus solution onto a piece of paraffin film, carefully move the tip of the micropipette into the droplet using the stereotaxic frame and draw it up. Then carefully press dispense until a bit of virus solution emerges at the tip.
- Retract the syringe and, using a pen, mark the micropipette with a scale and note the level of the virus solution; this will facilitate the monitoring of whether the infusion is progressing as programmed.
- Intraspinal Injections
- Move the tip of the micropipette above one of the holes in the dura and then down until a slight dent is noted in the dura. To target the injection to the spinal dorsal horn, move down 500 µm in 100 µm consecutive increments (speed 1 mm/s) and then 200 µm up to allow for mechanical stabilization of the tissue.
NOTE: The final depth of the tip is 300 µm in this case, but can be adapted depending on the target area.- If the tissue is resistant to penetration, the micropipette might be catching on the dura. In this case, retract and move slightly to overlie the hole, or use the needle again to perforate the dura properly before repeating the penetration attempt.
- Program the pump to a target injection volume of 300 nL at an injection speed of 50 nL/min and press the start button to begin the infusion.
- After the injection is completed, check the scale to see whether the virus level has dropped and leave the micropipette in place for an additional 3 min to allow the pressure to equilibrate before slowly retracting the syringe.
- Repeat step 2.9.1.-2.9.3. for the other two injection sites.
- Move the tip of the micropipette above one of the holes in the dura and then down until a slight dent is noted in the dura. To target the injection to the spinal dorsal horn, move down 500 µm in 100 µm consecutive increments (speed 1 mm/s) and then 200 µm up to allow for mechanical stabilization of the tissue.
- Suturing and Recovery
- Remove the spinal clamps and the cushion below the mouse.
- Close the wound in layers with interrupted stiches, suturing the superficial tissue layers with absorbable sutures and the skin with non-absorbable sutures. Apply iodine disinfectant on the sutured wound.
- Terminate the anesthesia and leave the animal on the heat mat until it recovers before returning it to its home cage.
- Postoperative Care
- Monitor the health of the animal on the day of surgery, the next day, and then every 2-3 days. Ensure that the animal has a normal gait, a healthy appearance (weight, eyes, fur, and behavior), and that the wound is healing.
- Continue analgesic treatment (e.g., 0.1-0.2 mg/kg buprenorphine subcutaneously) as necessary, three times per day.
3. Behavioral and Morphological Analyses
- Behavioral Analysis
NOTE: Allow animals to accommodate to the respective setup for at least 30 min before commencing measurements to let them adapt to the new experimental environment.- von Frey Test
- Place animals individually into individual compartments of about 10 cm x 10 cm on a metal grid floor.
- Stimulate the plantar surface of the hindpaw ipsilateral to the injection site with a dynamic von Frey filament. Note the force at which the animal withdraws its paw from the stimulus.
- Repeat five times and calculate the average withdrawal threshold from the six measurements for each animal.
- Pin Prick Test
- Place animals into individual compartments of about 10 cm x 10 cm on a metal grid floor.
- Stimulate the plantar surface of the hindpaw ipsilateral to the injection site with a blunted 26-G needle without penetration of skin. Score behavior as zero (no reaction) or one (reaction).
- Repeat 9 times with intervals of 3 min between stimulations and calculate percentage response from the 10 measurements for each animal.
- Injection of DREADD-ligand clozapine-N-oxide (CNO)
- Dissolve CNO in dimethyl sulfoxide (DMSO) to create a stock solution of 0.2 mg/µL, which can be stored at room temperature.
- On the day of the experiment, dilute stock solution in sterile 0.9% NaCl to 0.2 µg/µL and inject 10 µL/g body weight intraperitoneally. Inject control animals with vehicle (0.2% DMSO in 0.9% NaCl).
NOTE: According to experience, peak effects on behaviors can be expected 1-3 h after injection, and effects should completely subside after 24 h. In the case of activation of glycinergic neurons, observed behaviors include reduced pain and itch responses. Behaviors evoked by DREADD-mediated activation of neurons will depend on the targeted subset of spinal neurons.
- Injection of Pruritogens to Induce Itch
- Dissolve pruritogens in 0.9% NaCl to solutions of 8 mg/mL (chloroquine) or 10 mg/mL (histamine). 2 h after CNO administration, inject 10 µL of pruritogen or vehicle solution subcutaneously into the plantar surface of the hindpaw ipsilateral to the spinal injection site. Alternatively, inject the pruritogen intradermally into the calf, which has been shaved one day before to reduce aversive behavior caused by the shaving itself.
- Videotaping to Analyze Spontaneous Nocifensive or Pruritogen-induced Itch Behaviors
- Place animals individually into transparent compartments (about 10 cm diameter) with a bit of bedding from their respective home-cage on the floor.
- Videotape the animals for 5 min to record spontaneous and for 30 min to record pruritogen-induced behaviors. Analyze the videos off-line in bins of 5 min.
- Pain Versus Itch Behaviors
- Observe and note flinching, licking, and biting behaviors.
NOTE: Flinching and licking of the affected site are considered responses to pain, whereas biting is associated with itch. - Analyze videos at normal speed, measure the time that the mouse spent licking or biting the affected site, or measure in slow motion for a more precise distinction between licking and biting reflexes. Alternatively, count the number of licking/biting/flinching bouts.
- Observe and note flinching, licking, and biting behaviors.
- von Frey Test
- Morphological Analysis
- Perform morphological analyses from one to three weeks after injection of the virus or at the end of a behavioral experiment.
NOTE: We have detected eGFP expression as soon as 48 h after injection, but also found that expression increases significantly within the next days and even weeks. For cells with strong reporter expression, one week of incubation (from intraspinal injection until perfusion) may be enough. For cells with weak transgene expression, viruses with weak promoters or weaker fluorophores, a longer expression may be required to ensure detectable levels. - Tissue Fixation
- Perfuse each mouse transcardially, first with of 20 mL of ice-cold artificial cerebrospinal fluid (ACSF) solution (NaCl 125 mM, NaHCO3 25 mM, NaH2PO4 1.25 mM, D-glucose 20 mM, KCl 2.5 mM, and CaCl2 2 mM MgSO4 1 mM), then with 100 mL of 4% ice-cold paraformaldehyde (in 0.1 M sodium phosphate buffer, pH 7.4).
Caution: Paraformaldehyde is toxic and must be handled with care. - Immediately dissect the lumbar spinal cord, and post-fix the tissue for 2 h with 4% paraformaldehyde on ice. Briefly wash the post-fixed tissue with 0.1 M sodium phosphate buffer (pH 7.4) and then incubate it in 25% sucrose solution (in 0.1 M sodium phosphate buffer, pH 7.4) overnight at 4 °C.
- Cut the cryoprotected tissue at 30 μm on a cryostat and mount the sections on microscope slides.
- Perfuse each mouse transcardially, first with of 20 mL of ice-cold artificial cerebrospinal fluid (ACSF) solution (NaCl 125 mM, NaHCO3 25 mM, NaH2PO4 1.25 mM, D-glucose 20 mM, KCl 2.5 mM, and CaCl2 2 mM MgSO4 1 mM), then with 100 mL of 4% ice-cold paraformaldehyde (in 0.1 M sodium phosphate buffer, pH 7.4).
- Immunofluorescence Staining
- After brief washes in PBS, apply 300 μL of blocking solution (10% Normal donkey serum in 0.3% Triton-PBS) on the sections for 1 h at room temperature.
- Incubate the slides with 300 μL of the respective combinations of primary antibodies in blocking solution (see Table of Materials) over night at 4 °C. Wash the sections 3 times for 5 min each in PBS.
- Incubate the slides with the respective combinations of secondary antibodies in blocking solution for 1 h at room temperature. Wash the sections 3 times for 5 min each in PBS.
- Briefly rinse the slides in ddH2O, then mount the coverslips using fluorescent mounting medium.
- Acquire fluorescent images using a confocal microscope equipped with a 20x and a 40x objective.
- Perform morphological analyses from one to three weeks after injection of the virus or at the end of a behavioral experiment.
Representative Results
In order to illustrate the expression levels that can be obtained by the intraspinal injection of rAAV encoding a marker protein, we first injected AAV1.CAG.eGFP into the lumbar spinal cord of wild-type mice. Three injections spaced approximately 1 mm apart produced a nearly continuous infection of lumbar spinal segments L3 to L5 (Figure 1A-C). Virus injection at a depth of 300 µm from the spinal surface leads to predominant infection of cells in the spinal cord dorsal horn. However, infected cells could also be found in the ventral horn (Figure 1D). Next, the goal was to illustrate the difference in rAAV-mediated expression when injecting a Cre-dependent rAAV into a Cre transgenic mouse. We therefore injected the Cre-dependent AAV1.CAG.flex.eGFP vector into the spinal cord of GlyT2::Cre transgenic mice. As before, eGFP expression was observed in the dorsal and ventral horn. However, as expected, the expression of eGFP became more restricted, reflecting the distribution of GlyT2+ neurons, i.e. the relatively sparse expression in the superficial dorsal horn and dense expression in the deep dorsal horn (Figure 1E).
Next, to demonstrate the feasibility of addressing circuit function of spinal neuronal subpopulations, two different Cre-dependent AAVs coding for different effector proteins (DTA and hM3Dq) were injected into GlyT2::Cre transgenic mice. Analogous to the viral expression observed after three injections of AAV1.CAG.eGFP into wild-type mice, there was robust ablation of inhibitory neurons (Pax2+) in lumbar segments L3-L5 after the injection of AAV1.EF1a.flex.DTA into Glyt2::Cre mice (Figure 2A,C). Loss of glycinergic inhibitory neurons in these segments evoked a marked mechanical hypersensitivity (Figure 2D) and spontaneous aversive behavior directed towards the ipsilateral hindlimb (Figure 2E). The aversive behavior led to self-inflicted lesions, which could be observed on the paws, calf, and thigh (for data, see Foster et al.7) Opposite effects were observed when activating glycinergic neurons through injection of AAV1.hSyn.flex.hM3Dq and subsequent intraperitoneal injection of CNO (Figure 2B). Mice became desensitized to noxious mechanical stimulation (Figure 2F) and other noxious stimuli (see Foster et al.7) In addition, when treated with the pruritogens histamine or chloroquine, hM3Dq-mediated activation of glycinergic neurons efficiently suppressed prurifensive responses (Figure 2G). These results demonstrate that three injections of rAAV into the lumbar spinal cord are able to transduce an area of the lumbar spinal cord sufficient to observe robust behavioral changes evoked by stimulation of the corresponding hindlimb.
In a final set of experiments, we wanted to demonstrate the potential differences in using Cre reporter mice compared to Cre reporter viruses. Therefore, a Cre driver gene was chosen that has previously been described as displaying a restricted expression pattern in the mouse spinal cord. The gene RORβ has been suggested to be expressed predominantly in inhibitory interneurons of the deep dorsal horn13,14. This study used RORβCre knock-in mice and crossed them to Rosa26lox-STOP-lox-tdTomato (R26Tom) Cre reporter mice, which lead to the expression of tdTomato in all cells displaying Cre expression at any time point before analysis (Figure 3A). Characterization of tdTomato+ cells in RORβCre; R26Tom mice revealed expression in neurons and astrocytes of the dorsal horn (Figure 3B,C). In fact, quantification of the tdTomato+ cells suggested that the majority of cells undergoing Cre-mediated recombination (58%) were non-neuronal. We then injected a Cre-dependent tdTomato reporter rAAV (AAV1.CAG.flex.tdTomato) into the spinal cord of P40 RORβCre mice (Figure 3D). In contrast to R26Tom-mediated reporter expression, we found all tdTomato+ cells labeled by the reporter virus to be neurons (Figure 3E,F). Finally, the identity of the tdTomato+ neurons was analyzed in both sets of mice (RORβCre; R26Tom and RORβCre injected with AAV1.CAG.flex.tdTomato). In both cases, the majority of neurons were inhibitory (>85%) and the minority of neurons excitatory (<20%) (Figure 3 G-I), which is in agreement with previous assessments of the identity of RORβ+ neurons13,14.
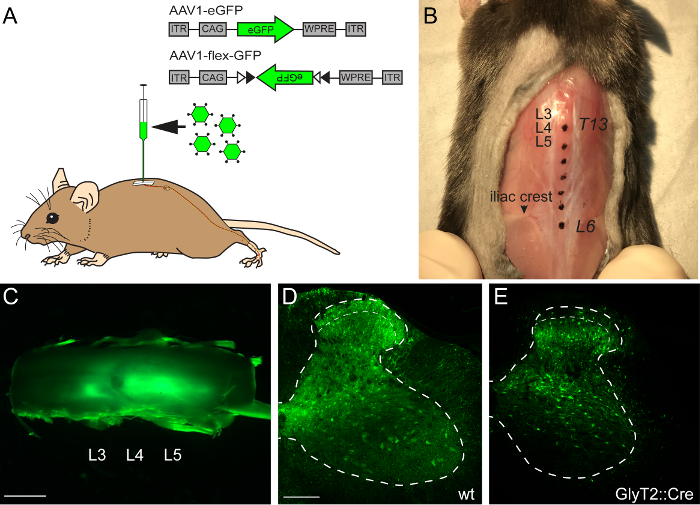
Figure 1: Intraspinal Injection of AAV1-eGFP/AAV1-flex-eGFP
(A) Schematic illustration of an intraspinal injection of an AAV1.CAG.eGFP (AAV1-eGFP) into the lumbar spinal cord, which is innervated from the hindlimb. (B) Anatomical location of the lumbar spinal cord segments L3-L4 can be seen in a top down view of the back of a mouse. The skin was opened to expose the vertebral column. Spinal processes of vertebrae T13-L6 are colored as anatomical references and an arrow indicates the iliac crest. (C) Representative image of a whole mount lumbar spinal cord. Green fluorescence indicates virus-transduced areas of the spinal cord. (D) Representative image of a cross section through an AAV1-eGFP-transduced lumbar spinal cord from a wild-type mouse. (E) Representative image of a cross section of an AAV1.CAG.flex.eGFP-transduced spinal cord of a GlyT2::Cre mouse. Dashed lines represent outline of the gray matter and superficial dorsal horn of the spinal cord. Scale bars C = 1 mm, D = 100 µm. Please click here to view a larger version of this figure.
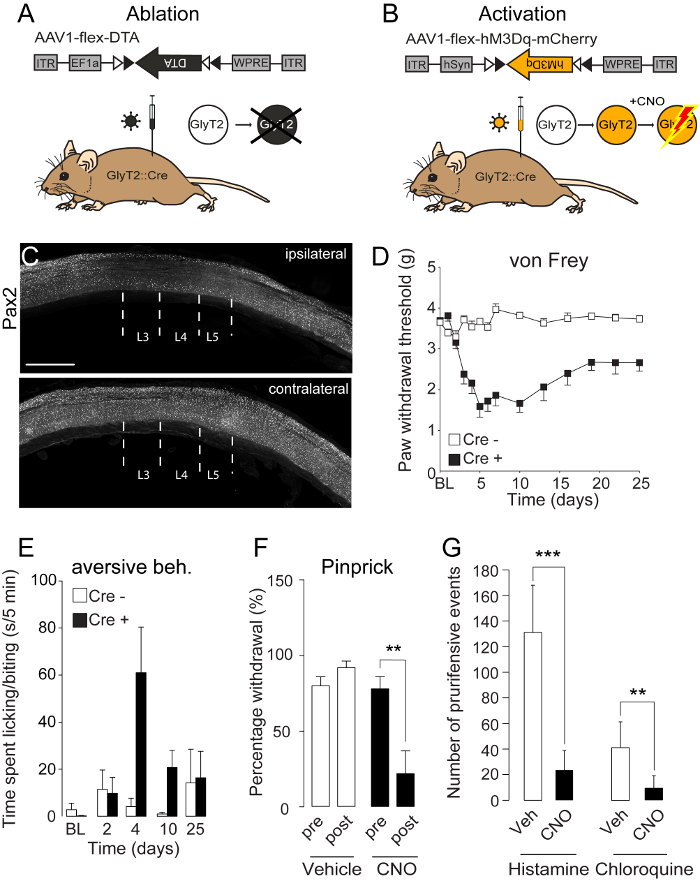
Figure 2: Functional Manipulation of Spinal Cre-expressing Glycinergic Neurons
(A+B) Schematic illustration of an intraspinal injection of an AAV1.EF1a.flex.DTA (A) or an AAV1.EF1a.flex.hM3Dq (B) into the lumbar spinal cord. Cre-dependent expression of diphtheria toxin fragment A (DTA) will lead to ablation of the glycinergic neurons (GlyT2+) (A), while Cre-dependent expression of the pharmacogenetic designer receptor hM3Dq will render GlyT2 neurons activatable by clozapine-N-oxide (CNO) (B). (C) Three injections of AAV1.EF1a.flex.DTA into the L3-L5 segments of GlyT2::Cre mice led to a marked loss of inhibitory neurons in the respective segments of the ipsilateral but not of the contralateral side. (D) Loss of GlyT2 neurons evoked a long-lasting hypersensitivity to mechanical von Frey stimulation in the ipsilateral hind paw of GlyT2::Cre mice injected with AAV1.EF1a.flex.DTA, but no change was observed if Cre-negative mice were injected. (E) Loss of GlyT2 neurons evoked spontaneous aversive behavior reminiscent of chronic itch. (F) hM3Dq-mediated activation of GlyT2 neurons alleviated noxious mechanical pain evoked by pinprick stimulation. (G) hM3Dq-mediated activation of GlyT2 neurons reduced pruritogen (chloroquine or histamine)-evoked aversive behavior. Data are represented as mean ± SEM. *** p < 0.001; ** p < 0.01. Scale bar C = 1 mm. g = grams. Images are re-used and modified from Foster et al.7 Please click here to view a larger version of this figure.
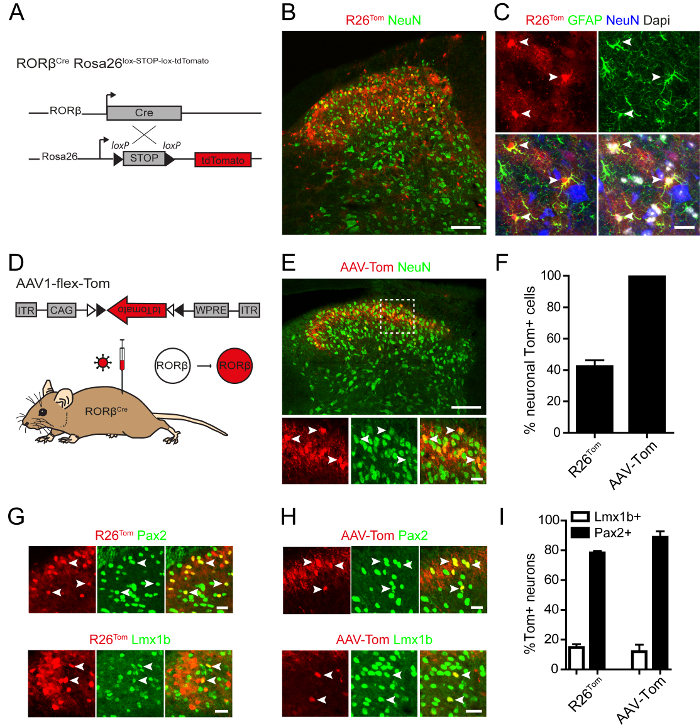
Figure 3: Genetic and Virus-mediated Labeling of RORβ-expressing Cells. (A) Diagram showing the strategy for Cre-dependent expression of fluorescent tdTomato reporter in RORβCre; Rosa26lox-STOP-lox-tdTomato (R26Tom) mice. (B) Immunostaining on spinal cord sections of RORβCre; R26Tom mice revealed that NeuN+ RORβ-Tom neurons can be found in laminae I-IV. (C) Cre-dependent expression of tdTomato was also observed in GFAP+ cells of RORβCre; R26Tom mice, suggesting that RORβ is expressed in astrocytes during development. Arrowheads indicate double-labeled astrocytes in lamina III. (D) Diagram showing intraspinal injection of AAV-flex-Tom (rAAV1.CAG.flex.tdTomato) virus into RORβCre mice to drive local Cre-dependent expression of tdTomato. (E) Immunostaining on spinal cord sections of RORβCre mice injected with AAV-flex-Tom virus. RORβ-Tom neurons were localized to the superficial laminae of the spinal dorsal horn and expression of tdTomato was absent from astrocytes. (F) Percentage of RORβ-Tom cells expressing the neuronal marker NeuN in spinal sections of RORβCre; R26Tom mice and RORβCre mice injected with AAV-flex-Tom virus. (G-H) Immunostaining on spinal cord sections of (G) RORβCre; R26Tom mice and (H) RORβCre mice injected with AAV-flex-Tom virus showing colocalization between RORβ-Tom neurons and the inhibitory marker Pax2 or the excitatory marker Lmx1b. (I) Percentage of RORβ-Tom neurons expressing Lmx1b and Pax2 in RORβCre; R26Tom mice and RORβCre mice injected with AAV-Tom virus. Data are represented as mean ± SEM. Data are from 2-3 mice and 1-3 sections per mouse. Scale bars represent 100 µm (B,E) and 20 µm (C,E in high magnification images, G and H). Please click here to view a larger version of this figure.
| Variable | Set value | Unit |
| heat (H) | 450 | – (value proportional to power of the radiated heat) |
| force preliminary pull (F(TH)) | 20 | – (value proportional to voltage applied to force coil) |
| distance threshold (s(TH)) | 25 | 0.12 mm |
| delay heatstop (t(H)) | 30 | 0.5 ms |
| distance heatstop (s(H)) | 0 | 0.12 mm |
| delay pull 1 (t(F1)) | 200 | 0.5 ms |
| force pull 1 (F1) | 300 | – (value proportional to voltage applied to force coil) |
| distance pull 2 (s(F2)) | 30 | 0.12 mm |
| force pull 2 (F2) | 600 | – (value proportional to voltage applied to force coil) |
| adjust (AD) | 0 | – |
Table 1: Puller Settings
Discussion
Intraspinal injection of AAVs may become a powerful technique in a research laboratory, enabling the analysis of spinal cells with a high temporal and spatial solution. This protocol enables the transduction of the three main spinal segments innervated by sensory neurons extending their peripheral axons to the hindlimb. Transducing three segments produces robust and reproducible behavioral data. It also enables testing of a larger sensory area than possible after a single intraspinal injection. For example, the same injection regime allows for testing the paw (von Frey, Hargreaves, etc.) and the thigh (intradermal injections, mechano-receptor stimulation), thus expanding the sensory modalities that can be addressed.
Critical Steps:
There are several steps critical for obtaining robust morphological and behavioral data and for avoiding artefacts. The design of the viral vector and the choice of the promoter and serotype can have a strong impact on transduction efficiency and extent of the transduced area, and therefore on behavioral results (for details on viral spread and transduction efficiencies, see 12). High quality viral preparations are required to minimize side effects of the virus injection, which can occur if contaminants or cell debris are present in the virus solution. The surgery that is required to inject the spinal cord has to be undertaken with great care in order to avoid morphological and behavioral artefacts introduced by the surgery. Particular attention is required in handling the spinal cord, especially during the laminectomy and during the perforation of the dura with the needle. Damage to the spinal cord could impair somatic sensations and motor coordination of the mouse, thus compromising any planned behavioral experiments. In addition, it is important to assemble the syringe carefully. Improper assembly or clogging of the micropipette can hamper the experiment. Spinal tissue at the injection site has to be examined at the end of the experiment in order to verify successful injection and transduction of the injected area and to exclude excessive tissue damage as a result of the surgery. This protocol has used viral titers up to a concentration of 1×1013 GC/mL without apparent toxicity, yet higher titers of virus might become toxic at some time point. In addition, toxicity will most likely also depend on the protein encoded by the injected AAV.
Modifications:
Several parameters in the protocol can be adapted to the neuronal population and research question to be studied. Injection of 300 nL virus solution at a depth of 300 µm leads to transduction of neurons throughout, and predominantly in the dorsal horn. If the aim is to target neurons further ventral, the depth can be adjusted. Injection of larger volumes of virus (we have used up to 500 nL per injection) will lead to a larger spread in dorsoventral and rostrocaudal directions. This may be beneficial to achieve targeting of additional cells in the rostrocaudal extent, but can increase the spillover to the contralateral side, which may hamper the use of the contralateral side as an internal control. In addition, the dorsoventral extent of transduced cells will increase, which may be the desired effect or should be avoided. Injection of 300 nL at a depth of 300 µm avoided side effects on motor control in GlyT2::Cre mice, while injection of 500 nL or injection at a greater depth occasionally produced motor phenotypes, such as spastic extension of the hindlimb (data not shown). Injection of smaller volumes can be used to restrict targeting to even more confined areas5. The viral titer is another parameter that can be modified and should in fact be determined for each virus. For bacterial toxins, such as the highly efficient DTA, low expression levels may be sufficient to induce the desired effect, whereas for fluorescent reporters and DREADDs, higher titers may be necessary for detectable or effective expression.
Significance of the Method with Respect to Existing/Alternative Methods:
To date, mainly two techniques have been used to functionally interrogate the neuronal circuits required for transmission of sensory signals. Many researchers have used intraspinal injections of rAAV coding for reporter and effector proteins into recombinase-expressing mice in order to label and manipulate spinal neuronal subpopulations. Intraspinal injection as a technique to manipulate spinal cells has previously been described15,16. The protocol outlined here was specifically designed to transduce genetically labeled neurons in three consecutive segments of the lumbar spinal cord while minimizing surgical manipulation. This is achieved by placing two of the three injections in the intervertebral space rostral and caudal of the T13 vertebrae and second, by drilling a hole into T13 for the third injection instead of removing the vertebrae. The targeted L3-L5 segments are the main termination area of sensory neurons innervating the hindlimb. We demonstrate that this type of transduction is sufficient to produce robust behavioral changes after the manipulation of glycinergic neurons and suggest that it is suitable for the analysis of a variety of different genetically labeled spinal neurons.
The second technique that has been used in order to manipulate the function of spinal neuron subsets is based on crossing transgenic animals. Here, mice expressing a recombinase in a specific subset of spinal neurons have to be crossed to reporter mice in order to achieve expression of the desired marker or effector protein in the respective neuronal subset17,18,19. To illustrate some of the differences that can occur when comparing results obtained by using reporter mice or intraspinal virus injections we either crossed RORβCre knock-in animals to a Cre-dependent reporter mouse or injected RORβCre mice with a Cre-dependent reporter rAAV. We compared reporter gene expression obtained from the rAAV to reporter gene expression obtained in RORβCre; R26Tom mice. Reporter gene expression obtained from the intraspinal injection of a Cre-dependent rAAV was restricted to neurons of the spinal cord, while R26Tom reporter mouse-evoked tdTomato expression is also found in astrocytes. This suggests that RORβ is transiently expressed in astrocytes. Similarly, Gutierrez-Mecinas et al. found a more restricted reporter expression (spatially and cell-type specific) after viral injection compared to mouse-evoked reporter expression in Tac1Cre mice20. By using intraspinal injections of a rAAV reporter into the spinal cord of adult mice, reporter and/or effector gene expression can efficiently be restricted to the population of cells that express the gene in the adult and thus avoid recombination/expression in those populations with earlier transient gene activity.
A second main advantage of intraspinal rAAV injections is the ability to restrict the expression of the rAAV-encoded genes to the injection site. In transgenic mice, Cre driver-mediated expression is often not only found in the neuronal subpopulation of interest, but also in other populations within the nervous system. Dymecki and colleagues, as well as the groups of Ma and Goulding, have developed elegant ways of using intersectional reporter mouse-based approaches that avoid recombination in parts of the nervous system that shall remain unaltered17,18,19,21,22,23. Using knock-in mice to obtain expression of marker or effector proteins can also be assumed to be less variable, as there is only one genomic copy of the respective expression cassette per cell and the expression cassette is present in all cells in the region of interest. In contrast, the presence of a respective expression cassette and also the copy number of the expression cassette after viral transduction is dependent on the transduction efficiency, which may vary from cell to cell, from injection to injection, and depends on the virus lot. Finally, the main disadvantage of virus-mediated gene expression over traditional genetic methods is the necessity of surgery. Surgery-mediated injury/inflammation may affect neurons and circuits directly. Inclusion of control mice that have undergone the same surgery is therefore mandatory.
Future Applications:
Intraspinal injection can be used to analyze and manipulate any genetically identified spinal subpopulation through overexpression of marker and effector proteins. It is therefore likely in the future that many will adopt this technique to study neuronal populations in their specific focus. Moreover, besides using intraspinal injection of rAAVs to achieve overexpression of exogenous effector proteins, this technique can also be used to silence or overexpress endogenous proteins and therefore to study the function of any given gene with high spatial and temporal resolution.
Disclosures
The authors have nothing to disclose.
Acknowledgements
We thank Hanns Ulrich Zeilhofer for generously supporting this work. Hendrik Wildner was supported by the Olga Mayenfisch foundation. We thank Carmen Birchmeier for the Lmx1b antibody.
Materials
| Equipment | |||
| micropipette puller: DMZ-Universal-Electrode-Puller | Zeitz | NA | |
| anesthesia unit: Oxymat3 oxygen concentrator | Weinmann | NA | |
| anesthesia unit: VIP 3000 Veterinary Vaporizer | Midmark | NA | |
| Heat mat: Mio Star Thermocare 100 | Migros | 717614700000 | |
| Electric shaver | Philips | BT9290 | |
| surgical microscope (OPMI pico) | Zeiss | NA | |
| Small animal stereotaxic apparatus | Kopf | NA | |
| Neurostar StereoDrive (optional) | Neurostar | NA | |
| Model 51690 Cunningham mouse spinal adaptor | Harvard Apparatus | 72-4811 | |
| PHD Ultra syringe pump with nanomite | Harvard Apparatus | 70-3601 | |
| Hamilton 701 RN 10 μl glass microliter syringe | Hamilton | 7635-01 | |
| Hamilton Removable needle (RN) compression fitting 1 mm | Hamilton | 55750-01 | |
| fine dentistry drilling apparatus: Osada success 40 | Osada | OS-40 | |
| spherical cutter, 0.5mm | Busch | 12001005B | |
| electronic von Frey anesthesiometer | IITC | 23905 | |
| flexible von Frey hairs | IITC | #7 | |
| LSM710 Pascal confocal microscope | Zeiss | NA | |
| 0.8 NA × 20 Plan-apochromat objective | Zeiss | NA | |
| 1.3 NA × 40 EC Plan-Neofluar oil-immersion objective | Zeiss | NA | |
| Name | Company | Catalog Number | Comments |
| Surgical Tools | |||
| Scalpel Handle #4, 13cm | Fine Science Tools | 10004-13 | |
| Extra Fine Bonn Scissors | Fine Science Tools | 14084-08 | |
| Adson forceps, 1 x 2 teeth, 12 cm | Fine Science Tools | 11027-12 | |
| Friedman-Pearson rongeurs, curved, 0.7 mm cup | Fine Science Tools | 16121-14 | |
| Dumont #2 laminectomy forceps | Fine Science Tools | 11223-20 | |
| Olsen-Hegar needle holders, serrated, 8.5 mm clamp length | Fine Science Tools | 12002-12 | |
| Fine forceps #5 | Fine Science Tools | 11254-20 | |
| Name | Company | Catalog Number | Comments |
| Consumables and Chemicals | |||
| Thin-wall glass capillary, 1mm outside diameter | World Precision Instruments | TW 100-3 | |
| Syringes (1, 5 and 20 ml) | B. Braun | (9166917V, 4606051V, 4606205V) | |
| 26G beveled needle | B. Braun | 4665457 | |
| Sterile scalpel blades | B. Braun | BB523 | |
| Surgical sutures Safil Quick+ 4/0, absorbable | B. Braun | C1046220 | |
| Surgical sutures Premilene 5/0, non-absorbable | B. Braun | C0932191 | |
| Sterile PBS or saline (0.9%) | NA | ||
| Ethanol, 70% (disinfectant) | NA | ||
| Iodine solution (e.g. Braunol) | B. Braun | 18380 | |
| Anaesthetics (e.g. Attane isoflurane) | Provet | 2222 | |
| Aldasorber | Provet | 333526 | |
| analgesics (e.g. buprenorphine: temgesic) | Indivior | GTIN: 7680419310018 | |
| Ophthalmic ointment (e.g. vita-pos) | Pharma medica | GTIN: 4031626710635 | |
| Cotton swabs (e.g. from) | IVF Hartmann | 1628100 | |
| Facial tissues (e.g. from) | Uehlinger AG | 2015.10018 | |
| Superfrost plus microscope slides | ThermoScientific | J1800AMNZ | |
| Name | Company | Catalog Number | Comments |
| Mice | |||
| C57BL/6J mice (wildtype) | The Jackson Laboratory | RRID:IMSR_JAX:000664 | |
| Rorbtm1.1(cre)Hze/J mice (RORβCre) | The Jackson Laboratory | RRID:IMSR_JAX:023526 | |
| Gt(ROSA)26Sortm14(CAG-tdTomato)Hze/J mice (R26Tom) | The Jackson Laboratory | RRID: IMSR_JAX:007914 | |
| Name | Company | Catalog Number | Comments |
| Viral vectors | |||
| AAV1.CB7.CI.eGFP.WPRE.rBG (AAV1.CAG.eGFP) | Penn Vector Core | AV-1-PV1963 | |
| AAV1.CAG.flex.eGFP.WPRE.bGH (AAV1.CAG.flex.eGFP) | Penn Vector Core | AV-1-ALL854 | |
| AAV1.CAG.flex.tdTomato.WPRE.bGH (AAV1.CAG.flex.tdTomato) | Penn Vector Core | AV-1-ALL864 | |
| AAV1.EF1a.flex.DTA.hGH (AAV1.EF1a.flex.DTA) | Penn Vector Core | Custom production | |
| AAV1.hSyn.DIO.hM3D(Gq)-mCherry.hGH (AAV.flex.hM3D(Gi)) | Penn Vector Core | Custom production | |
| Name | Company | Catalog Number | Comments |
| Plasmids | |||
| pAAV.hSyn.flex.hM3D(Gq)-mCherry | Addgene | 44361 | |
| pAAV.EF1α.flex.hChR2(H134R)-eYFP | Addgene | 20298 | |
| Name | Company | Catalog Number | Comments |
| Bacteria | |||
| MDS42 | ScarabGenomics | ||
| Stbl3 | ThermoScientific | C737303 | |
| Name | Company | Catalog Number | Comments |
| Reagents | |||
| EndoFree Plasmid Maxi Kit | Quiagen | 12362 | |
| NucleoBond PC 500 | Machery & Nagel | 740574 | |
| clozapine-N-oxide (CNO) | Enzo Life Sciences | BBL-NS105-0025 | |
| chloroquine diphosphate salt | Sigma | C6628 | |
| histamine | Sigma | H7125 | |
| Dapi | Invitrogen | D3571 | |
| Name | Company | Catalog Number | Comments |
| Antibodies (dilution) | |||
| Rabbit anti-GFP (1:1000) | Molecular Probes | RRID:AB_221570 | |
| Rabbit anti-NeuN (1:3000) | Abcam | RRID:AB_10711153 | |
| Goat anti-Pax2 (1 : 200) | R & D Systems | RRID:AB_10889828 | |
| Guinea pig anti-Lmx1b (1 : 10 000) | Dr Carmen Birchmeier | Muller et al. 2002 | |
| Rabbit anti-GFAP (1 : 1000) | DakoCytomation | RRID:AB_10013382 | |
| Secondary antibodies raised in donkey (1:800) | Jackson ImmunoResearch Laboratories | NA |
References
- Goulding, M., Bourane, S., Garcia-Campmany, L., Dalet, A., Koch, S. Inhibition downunder: an update from the spinal cord. Curr Opin Neurobiol. 26, 161-166 (2014).
- Todd, A. J. Neuronal circuitry for pain processing in the dorsal horn. Nat Rev Neurosci. 11 (12), 823-836 (2010).
- Sandkuhler, J. Models and mechanisms of hyperalgesia and allodynia. Physiol Rev. 89 (2), 707-758 (2009).
- Zeilhofer, H. U., Wildner, H., Yevenes, G. E. Fast synaptic inhibition in spinal sensory processing and pain control. Physiol Rev. 92 (1), 193-235 (2012).
- Azim, E., Jiang, J., Alstermark, B., Jessell, T. M. Skilled reaching relies on a V2a propriospinal internal copy circuit. Nature. 508 (7496), 357-363 (2014).
- Cui, L., et al. Identification of Early RET+ Deep Dorsal Spinal Cord Interneurons in Gating Pain. Neuron. 91 (6), 1413 (2016).
- Foster, E., et al. Targeted ablation, silencing, and activation establish glycinergic dorsal horn neurons as key components of a spinal gate for pain and itch. Neuron. 85 (6), 1289-1304 (2015).
- Francois, A., et al. A Brainstem-Spinal Cord Inhibitory Circuit for Mechanical Pain Modulation by GABA and Enkephalins. Neuron. 93 (4), 822-839 (2017).
- Peirs, C., et al. Dorsal Horn Circuits for Persistent Mechanical Pain. Neuron. 87 (4), 797-812 (2015).
- Petitjean, H., et al. Dorsal Horn Parvalbumin Neurons Are Gate-Keepers of Touch-Evoked Pain after Nerve Injury. Cell Rep. 13 (6), 1246-1257 (2015).
- Zhang, Y., et al. Identifying local and descending inputs for primary sensory neurons. J Clin Invest. 125 (10), 3782-3794 (2015).
- Haenraets, K., et al. Spinal nociceptive circuit analysis with recombinant adeno-associated viruses: the impact of serotypes and promoters. J Neurochem. , (2017).
- Abraira, V. E., et al. The Cellular and Synaptic Architecture of the Mechanosensory Dorsal Horn. Cell. 168 (1-2), 295-310 (2017).
- Wildner, H., et al. Genome-wide expression analysis of Ptf1a- and Ascl1-deficient mice reveals new markers for distinct dorsal horn interneuron populations contributing to nociceptive reflex plasticity. J Neurosci. 33 (17), 7299-7307 (2013).
- Inquimbert, P., Moll, M., Kohno, T., Scholz, J. Stereotaxic injection of a viral vector for conditional gene manipulation in the mouse spinal cord. J Vis Exp. (73), e50313 (2013).
- Kohro, Y., et al. A new minimally-invasive method for microinjection into the mouse spinal dorsal horn. Sci Rep. 5, 14306 (2015).
- Bourane, S., et al. Gate control of mechanical itch by a subpopulation of spinal cord interneurons. Science. 350 (6260), 550-554 (2015).
- Bourane, S., et al. Identification of a spinal circuit for light touch and fine motor control. Cell. 160 (3), 503-515 (2015).
- Duan, B., et al. Identification of spinal circuits transmitting and gating mechanical pain. Cell. 159 (6), 1417-1432 (2014).
- Gutierrez-Mecinas, M., et al. Preprotachykinin A is expressed by a distinct population of excitatory neurons in the mouse superficial spinal dorsal horn including cells that respond to noxious and pruritic stimuli. Pain. 158 (3), 440-456 (2017).
- Awatramani, R., Soriano, P., Rodriguez, C., Mai, J. J., Dymecki, S. M. Cryptic boundaries in roof plate and choroid plexus identified by intersectional gene activation. Nat Genet. 35 (1), 70-75 (2003).
- Kim, J. C., et al. Linking genetically defined neurons to behavior through a broadly applicable silencing allele. Neuron. 63 (3), 305-315 (2009).
- Kim, J. C., Dymecki, S. M. Genetic fate-mapping approaches: new means to explore the embryonic origins of the cochlear nucleus. Methods Mol Biol. 493, 65-85 (2009).

