Electroporation-Mediated Delivery of Cas9 Ribonucleoproteins and mRNA into Freshly Isolated Primary Mouse Hepatocytes
Summary
This protocol describes techniques for isolating primary mouse hepatocytes from the liver and electroporating CRISPR-Cas9 as ribonucleoproteins and mRNA to disrupt a therapeutic target gene associated with an inherited metabolic disease of the liver. The methods described result in high viability and high levels of gene modification after electroporation.
Abstract
This protocol describes a fast and effective method for isolating primary mouse hepatocytes followed by electroporation-mediated delivery of CRISPR-Cas9 as ribonucleoproteins (RNPs) and mRNA. Primary mouse hepatocytes were isolated using a three-step retrograde perfusion method resulting in high yields of up to 50 × 106 cells per liver and cell viability of >85%. This protocol provides detailed instructions for plating, staining, and culturing hepatocytes. The results indicate that electroporation provides a high transfection efficiency of 89%, as measured by the percentage of green fluorescent protein (GFP)-positive cells and modest cell viability of >35% in mouse hepatocytes.
To demonstrate the utility of this approach, CRISPR-Cas9 targeting the hydroxyphenylpyruvate dioxygenase gene was electroporated into primary mouse hepatocytes as proof-of-principle gene editing to disrupt a therapeutic gene related to an inherited metabolic disease (IMD) of the liver. A higher on-target edit of 78% was observed for RNPs compared to 47% editing efficiency with mRNA. The functionality of hepatocytes was evaluated in vitro using an albumin assay that indicated that delivering CRISPR-Cas9 as RNPs and mRNA results in comparable cell viability in primary mouse hepatocytes. A promising application for this protocol is the generation of mouse models for human genetic diseases affecting the liver.
Introduction
IMDs of the liver are genetic disorders characterized by the deficiency of a crucial hepatic enzyme involved in metabolism that leads to the accumulation of toxic metabolites. Without treatment, IMDs of the liver result in organ failure or premature death1,2. The only curative option for patients with IMDs of the liver is orthotopic liver transplantation, which is limited due to the low availability of donor organs and complications from immunosuppressive therapy following the procedure3,4. According to recent data collected by the Organ Procurement and Transplantation Network, only 40%-46% of adult patients on the liver transplant waiting list receive an organ, while 12.3% of these patients die while on the waiting list5. Moreover, only 5% of all the rare liver diseases have an FDA-approved treatment6. It is clear that there is a critical need for novel treatments for IMDs of the liver. However, appropriate disease models are required to develop new therapeutic options.
Modeling human diseases using in vitro and in vivo systems remains an obstacle for developing effective therapies and studying the pathology of IMDs of the liver. Hepatocytes from patients with rare liver diseases are challenging to obtain7. Animal models are crucial for developing an understanding of disease pathology and for testing therapeutic strategies. However, one obstacle is generating models from embryos carrying lethal mutations. For example, attempts to create mouse models of Alagille Syndrome (ALGS) with embryos containing homozygous deletions of a 5 kb sequence near the 5′-end of the Jag1 gene resulted in the early death of the embryos8. In addition, generating mouse models by gene editing in embryonic stem cells can be time- and resource-intensive9. Lastly, mutations will appear outside the targeted tissue, leading to confounding variables that may impede study of the disease9. Somatic gene editing would allow for easier editing in liver tissue and bypasses the challenges associated with generating models using embryonic stem cells.
Electroporation enables the delivery of CRISPR-Cas9 directly into the nucleus by applying high-voltage currents to permeabilize the cell membrane and is compatible with many cell types, including those that are intransigent to transfection techniques, such as human embryonic stem cells, pluripotent stem cells, and neurons10,11,12. However, low viability is a potential drawback of electroporation; optimizing the procedure can yield high levels of delivery while limiting toxicity13. A recent study demonstrates the feasibility of electroporating CRISPR-Cas9 components into primary mouse and human hepatocytes as a highly efficient approach14. Ex vivo electroporation in hepatocytes has the potential to be applied to generate new mouse models for human IMDs of the liver.
This protocol provides a detailed step-by-step procedure for isolating mouse hepatocytes from the liver and subsequently electroporating CRISPR-Cas9 as RNP complexes, consisting of Cas9 protein and synthetic single-guide RNA (sgRNA), or Cas9 mRNA combined with sgRNA to obtain high levels of on-target gene editing. In addition, the protocol provides methods for quantifying gene editing efficiency, viability, and functionality following electroporation of CRISPR-Cas9 into freshly isolated mouse hepatocytes.
Protocol
The animal experiments were all performed in compliance with the Institutional Animal Care and Use Committee guidelines and approved protocols at Clemson University. Surgical procedures were performed in anesthetized wild-type C57BL/6J mice between 8 and 10 weeks old.
1. Animal surgery
- Preparation of solutions and instruments
NOTE: Perfusion solutions and anesthesia cocktail recipes are shown in the Table of Materials.- Prepare Perfusion Solution 1 (HEPES, EGTA, and EBSS), Perfusion Solution 2 (HEPES, EGTA, EBSS, CaCl2, and MgSO4), and Perfusion Solution 3 (Solution 2 and Liberase). Make sure the solutions are well mixed.
- Set the water bath to 42 °C and prewarm Perfusion Solutions 1, 2, and 3 for a minimum of 30 min before starting the procedure and keep them in the water bath.
NOTE: Perfusion Solutions 1 and 2 will chelate the calcium and flush out the blood in the liver. Perfusion Solution 3 contains the digestive enzyme liberase to dissociate the extracellular matrix. - Place 150 mL of DMEM + 10% fetal bovine serum (FBS) in conical tubes and keep them on ice.
NOTE: This medium will be used to release the cells from the liver capsule and wash the isolated hepatocytes in steps 2.1 and 2.2, respectively. - Set the centrifuge to 4 °C.
- Sterilize the pump tubing by placing both ends of the tubing in a flask filled with 70% ethanol and cycling the ethanol three times.
- Wash the tubing with distilled water and empty it.
- Fill the tubing with prewarmed 30 mL of Perfusion Solution 1, followed by 8 mL of Perfusion Solution 2. If the tubing still has space for more fluids, fill the rest of the tubing with Perfusion Solution 3. Stop the pump when switching to a different perfusion solution to avoid introducing air bubbles into the tubing.
- Place the excess tubing into the 42 °C water bath to keep the perfusion solutions warm.
- Place the end of the pump tubing in the conical tube containing Perfusion Solution 3, while kept in the water bath.
NOTE: This step is necessary to allow the pump to continuously fill with Perfusion Solution 3 during perfusion.
- Animal preparation
- Anesthetize the mouse by injecting the anesthetic cocktail intraperitoneally at a dose of 10-11.7 µL/g body weight. The anesthetic cocktail has a final concentration of 7.5 mg/mL ketamine, 0.25 mg/mL acepromazine and 1.5 mg/mL xylazine.
- Monitor the mouse breathing and check that the mouse is not reacting to pain. Wait until the mouse has a breathing rate of ~55-65 breaths per min, is unresponsive to having its toes pinched, and has a flaccid tail. Additionally, check the palpebral (blinking) reflex by lightly touching the skin on the medial side of the closed eye. If the mouse blinks or the eye muscles twitch, increase the dose of the entire anesthetic cocktail by 5-10 µL.
- Place the mouse on its back in a supine position and secure the limbs to the surface using pins or tape (Supplemental Figure S1).
- Spray the abdomen with 70% ethanol and pat it dry using a cotton ball.
- Liver perfusion
- Make a U-shaped lateral incision in the abdomen skin using scissors and expand the hole through the sides.
- Continue opening the skin to the rib cage.
- Using scissors, cut through the peritoneum to expose the abdominal cavity up to the rib cage, being careful not to nick any organs. Move the intestines to the right side using the back of forceps or the blunt surface of scissors. Identify the inferior vena cava and the portal vein (Supplemental Figure S1).
- Start the pump to flush prewarmed perfusion solution through the catheter.
- Stop the pump and remove the catheter once all air has been flushed through the system. Insert the tip of the needle connected to the catheter into the inferior vena cava at a 10°-20° angle to the vein. Remove the needle from the catheter and gently push the catheter into the vein in a movement parallel to the vein.
NOTE: The user should observe flashback of blood in the catheter to verify proper cannulation in the vein. - Connect the tubing to the catheter without moving the catheter further into the vein.
- Start the pump with a 2 mL/min flow rate. Look for the liver immediately turning pale as a sign that the perfusion is successful.
- Cut the portal vein using scissors.
- Gradually increase the flow rate to 5 mL/min.
- Apply pressure periodically to the portal vein at the approximate cut site for 3 s using forceps or a cotton swab to block the drainage and facilitate good perfusion.
- Once Perfusion Solution 3 is flowing through, carefully monitor the elasticity of the liver. To determine if the liver is softened, gently press the liver with a cotton swab or forceps and check whether indentations form on the liver. Be careful not to overperfuse to avoid loss of cell viability.
- Once the liver is softened (occurring after 30-50 mL of Perfusion Solution 3 has flowed through), stop the pump, remove the catheter, and carefully dissect the liver. Place the liver into a 100 mm Petri dish containing ice-cold DMEM + 10% FBS medium. Excise the gallbladder, being careful not to spill its contents. Swirl the Petri dish to gently remove possible blood clots.
- Transfer the liver to a new Petri dish containing ice-cold DMEM + 10% FBS medium.
2. Hepatocyte isolation
- Release cells from the liver capsule.
- Place the Petri dish on ice and using two pairs of sterile forceps, gently tear apart the liver capsule on all the lobes. Gently swirl the liver around in the medium using a cell lifter or a forceps to release the hepatocytes. Continue this motion until the liver becomes very small and the suspension turns brown and opaque.
- Remove the remains of the liver tissue from the plate and using a 25 mL serological pipette set to low speed, carefully transfer the cell suspension to a 50 mL conical tube (prechilled on ice) fitted with a 100 μm cell strainer.
- Add fresh, ice-cold DMEM + 10% FBS medium to the Petri dish to wash out and collect the remaining cells from the surface and transfer to the 50 mL conical tube. Repeat this step until all the cells are collected.
- Wash and purify hepatocytes from nonparenchymal cells.
- Centrifuge the cells collected in the 50 mL conical tube at 50 × g at 4 °C for 5 min at low deceleration or without any brakes.
- Discard the supernatant containing debris and nonparenchymal cells and resuspend the pellet in the leftover supernatant by gently swirling the tube. After the pellet is resuspended, add 30 mL of fresh, ice-cold DMEM + 10% FBS medium.
- Repeat the centrifugation (step 2.2.1) and washing (step 2.2.3) three times.
- Resuspend the final cell pellet in 10 mL of ice-cold DMEM + 10% FBS medium.
NOTE: The volume of medium used to resuspend the pellet depends on the pellet size. If the pellet is considerably smaller than 15 mm, use 1-5 mL of ice-cold DMEM + 10% FBS medium.
- Cell viability quantification
- In a microfuge tube, add 50 µL of 0.4% Trypan Blue solution to 350 µL of Hepatocyte Plating Medium (PM). Then, add 100 µL of cell suspension to make a final 1:5 dilution and pipette up and down multiple times to mix.
- Count cell density and viability using a hemocytometer. While counting, place the hepatocyte suspension on ice, and then place the ice bucket on an orbital shaker set to 30 rpm to keep the hepatocytes from settling.
- Follow the Percoll treatment and centrifugation steps below if the cell viability is <70%.
- Dilute Percoll with 10x phosphate-buffered saline (PBS) in a 9:1 ratio.
- Mix the hepatocyte suspension with the diluted Percoll in a 1:1 ratio.
- Centrifuge at 200 × g at 4 °C for 10 min.
- Discard the supernatant containing dead cells. Resuspend the pellet in ice-cold DMEM + 10% FBS medium.
- Count the cells again.
- Testing hepatocytes for plateability
- Plate some of the cells on collagen I-coated 6-well plates at a density of 0.5 × 106 cells per mL using 1.5 mL of prewarmed PM.
- Keep the remaining cells on ice while on an orbital shaker until ready to perform electroporation.
- Place the plate of cells in a water-jacketed CO2 incubator at 37 C. Move the plate horizontally and vertically gently in a north-to-south and east-to-west motion to ensure homogeneous plating. Repeat the movement two more times every 1.5 h.
- Verify cell attachment at 3 h after plating.
NOTE: At least 60% of the cells should adhere to the plate as an indicator of good-quality hepatocytes. - Change the medium to prewarmed Hepatocyte Maintenance Medium (MM) at 24 h after plating to maintain the cells in culture.
- Glycogen staining
- After the cells have attached to the 6-well collagen I-coated plates, remove the spent medium from the cultured cells.
- Fix the cells for 10-15 min in ice-cold ethanol.
- Wash thoroughly with sterile water.
- Incubate in 1% aqueous periodic acid for 5 min and then wash with sterile water.
- Incubate in 100% Schiff Reagent for 30 min.
NOTE: The Schiff reagent must not be diluted before adding to the cells. - Wash with water three times over 10 min.
- Mount in mounting medium.
- Image using a microscope on brightfield setting.
3. Design sgRNAs for CRISPR-Cas9 gene editing
NOTE: This section describes the design of an sgRNA targeting the mouse hydroxyphenylpyruvate dioxygenase (Hpd) gene as proof-of-principle gene editing to disrupt a therapeutic target gene related to an IMD of the liver.
- Design the guide sequence targeting the Hpd using the referenced software15.
- Verify the sequence for Hpd with the Ensembl Genome Browser.
- Use the following parameters for designing the gRNA: single guide length of 20 nucleotides and an NGG (N can be any nucleotide) protospacer-adjacent motif (PAM) sequence.
- Select a target region from exon 3 in the Hpd.
- Generate a list of guide sequences from the target region with On- and Off-Target Scores.
NOTE: The On-Target Score indicates predicted on-target editing efficiency for the given design: a higher score is correlated with higher editing efficiency. The Off-Target Score correlates with the specificity of the guide sequence: a higher score indicates lower probability of off-target editing. Ideally both scores should be high: the On-Target Score >60, and the Off-Target Score >50. - Select the guide sequence with the highest On-Target and Off-Target Scores. Have the sgRNA containing the guide sequence chemically synthesized.
4. Electroporation and cell culture
- Preparation of CRISPR-Cas9 substrates, media, cells, and the electroporation instrument
- Add the entire accompanying supplement to the PM and warm in the 37 °C water bath for 10 min.
- Turn on the electroporation device by pressing the power button and set the electroporation program by pressing the x button and then the down arrow button until the program T-028 appears on the screen.
- Prepare the destination wells for the electroporated hepatocytes by adding 1.5 mL of PM to six-well collagen I-coated plates. Incubate the plates in a 37 °C incubator until ready to use.
- Add the entire accompanying supplement to the electroporation buffer solution and incubate at room temperature for 10 min.
NOTE: Label the electroporation buffer expiration date on the vial (3 months after the date of supplement addition). - Remove the electroporation vessels from the packaging and label them for each sample.
- In PCR split-strip tubes, prepare the Cas9 RNP complexes by combining 30 µg of sgRNA with 300 pmol of Cas9 protein and incubate the complexes at room temperature for 10 min. Prepare the Cas9 mRNA by combining 30 µg of sgRNA with 4 µL of Cas9 mRNA (1 µg/µL). Store the mRNA-sgRNA mix on ice until electroporation. Separately prepare a tube containing 1.0 µg of eGFP mRNA to verify successful electroporation using fluorescence microscopy.
- Centrifuge 1.2 × 106 cells per electroporation reaction at 100 × g for 2 min at 4 °C.
- Remove the supernatant from the cell pellet and add 100 µL of electroporation solution per reaction along the side of the tube wall. Resuspend the hepatocytes in the electroporation solution by rocking the tube gently by hand.
- Electroporation of CRISPR-Cas9 RNPs and mRNA into hepatocytes
- Once the hepatocytes appear evenly dispersed in the electroporation solution, transfer 100 µL of the hepatocyte suspension to the strip tubes containing the Cas9 complexes.
NOTE: Use wide-bore pipette tips when transferring hepatocytes to preserve cell viability. - Transfer the contents of the strip well to a electroporation vessel.
- Place the nucleovette vessel into the slot on the electroporation device. Electroporate the hepatocytes by pressing the x button. After electroporation, wait for a screen to pop up on the device that says OK. Press the x button again and remove the vessel.
- Incubate the electroporated vessel on ice for 15 min.
- Add 500 µL of prewarmed PM to the electroporation vessel.
- Transfer 300 µL of the electroporation reaction to each destination well on the prewarmed plate.
NOTE: There should be two destination wells per electroporation reaction. One well will be used to quantify gene editing efficiency; the other well will be used for the MTT and albumin assays. - To prevent hepatocytes from accumulating in the center of the well, disperse the cells by gently moving plates horizontally and vertically in a north-to-south and east-to-west motion. Ensure that the plate maintains contact with the incubator shelf while it is being moved. Repeat this motion at 15, 30, 45, 60, and 90 min after plating the electroporated hepatocytes.
- Incubate the plates overnight at 37 °C.
- Remove the medium from the wells designated for gene editing analysis 24 h after plating the cells, and replace with 0.25 mg/mL basement membrane matrix overlay.
NOTE: If stored in the freezer, transfer the membrane matrix to 4 °C and allow it to thaw. Because the basement membrane matrix is solid above 10 °C, it is crucial for the basement membrane matrix to remain on ice or at 4 °C until ready to use.
- Once the hepatocytes appear evenly dispersed in the electroporation solution, transfer 100 µL of the hepatocyte suspension to the strip tubes containing the Cas9 complexes.
5. Calculate the volume of the membrane matrix to mix with MM to give a final concentration of 0.25 mg/mL
NOTE: For a 6-well plate, 2 mL of overlay is needed per well.
6. Add the calculated volume of the membrane matrix to ice cold MM and mix by pipetting up and down 10 times
- Pipette the overlay mixture slowly on top of the cells, and place the plate back in the 37 °C incubator.
- Replace the medium with fresh maintenance medium every 24 h.
- At 24 h after plating, transfer the conditioned medium from the well designated for MTT and albumin assays. Store the conditioned medium in a microfuge tube until ready to perform the albumin assay. Proceed to step 7.1 for the MTT assay.
NOTE: Store the conditioned medium at 4 °C for short-term storage (less than a day). For longer storage, store the medium at -80 °C.
7. Analysis of delivery efficiency, viability, and on-target editing in electroporated mouse hepatocytes
- Viability measurement using MTT assay
- Prepare 12 mM MTT stock solution by adding 1 mL of PBS to one 5 mg vial of MTT.
- Add 150 µL of the MTT stock solution to the wells and incubate for 24 h.
- After incubation, add 1.5 mL of sodium dodecyl sulfate-hydrochloride (SDS-HCl) solution to each well and then pipette to mix.
- Incubate the plate for 4 h at 37 °C and look for a change in the color of the medium from clear to purple to indicate viable cells.
- Mix each sample again using a pipette and read the absorbance at 570 nm using a microplate reader.
- Calculate the normalized viability for Cas9-treated samples using Eq (1):
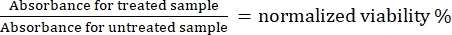 (1)
(1)
- Albumin assay to evaluate hepatocyte functionality
- Transfer 50 µL of the conditioned medium to the wells in the plate provided by the albumin kit. Cover the wells and incubate for 2 h at room temperature.
- Wash the wells 5x with 200 µL of wash buffer.
- Invert the plate to empty each well on tissue paper and tap a few times.
- After washing, add 50 µL of the biotinylated antibody provided in the albumin assay kit to each well and incubate at room temperature for 1 h.
- Repeat steps 5.2.2-5.2.3 to wash the wells.
- Add 50 µL of streptavidin-peroxidase provided in the albumin assay kit to each well and incubate at room temperature for 30 min.
- Repeat step 5.2.2.
- Add 50 µL of the chromogen substrate provided in the albumin assay kit per well and incubate for 30 min at room temperature. Gently tap the plate to ensure even mixing. Remove any air bubbles with a pipette tip.
- Lastly, add 50 µL of stop solution to each well and look for a color change from yellow to blue to indicate functional cells.
- Proceed immediately to read the absorbance at 450 nm using a microplate reader.
- Estimate the delivery efficiency in wells electroporated with eGFP mRNA.
- Capture phase contrast and green fluorescence images of hepatocytes 24 h after electroporation. Take three images at different locations within the well.
- Upload images of the cells to ImageJ. Under the Image tab, select Type | 16-bit to convert image to greyscale.
- To highlight cell structures in the image, under the Image tab, select Adjust | Threshold. Adjust the slider to highlight cells.
- Under the Process tab, select Subtract Background to remove background noise.
- Count the cells by clicking the Analyze tab and then select Analyze particles. Click Ok to generate a list of counted particles.
- Calculate the percentage of GFP-positive cells by dividing the number of counted particles in the green fluorescence images by the number of counted particles in the corresponding phase contrast image.
- Analysis of on-target Cas9 activity
- Extract genomic DNA from electroporated cells.
- Remove medium from the plate 3 days after electroporation, and add 500 μL of 0.25% trypsin. Incubate at 37 °C for 5 min. Pipette several times after incubation to ensure hepatocytes have detached from the plate.
- Transfer the trypsinized cell suspension to microcentrifuge tubes and spin in a tabletop centrifuge at 800 × g for 10 min at room temperature. After spinning, examine the tubes for pellets.
- Remove the supernatant containing trypsin by pipetting. Make sure not to disturb the pellet. Resuspend the pellet in 80 µL of DNA extraction solution. If the pellet is difficult to see, resuspend the content of the tube in 50 µL of DNA extraction solution.
- Vortex for 30 s to ensure the pellet is resuspended. Transfer the suspension to PCR split-strip tubes and place them in the thermal cycler. Run for 15 min at 95 °C and 8 min at 68 °C.
- PCR-amplify the on-target locus.
- Follow the procedure described below for amplifying the Hpd on-target region using DNA polymerase.
- Prepare a reaction containing 0.5 µL of 10 mM dNTPs, 0.5 µL of 10 µM Forward primer, 0.5 µL of 10 µM Reverse primer, 0.125 µL of Taq polymerase, 5 µL of genomic DNA extracted from the hepatocytes, and 18.375 µL of nuclease free water. See Supplemental Table S1 for primer sequences.
- Briefly vortex and quickly centrifuge the PCR mix to ensure that the solution is at the bottom of the tube.
- Place the PCR mix in a thermal cycler and amplify under these conditions: 94 °C for 30 s for denaturation, 30 cycles of 94 °C for 30 s, 57 °C for 45 s for annealing, 68 °C for 1 min, and a final extension of 68 °C for 5 min.
NOTE: The annealing temperature for the PCR reaction will vary depending on the composition of the primers. Consider using a melting temperature calculator before performing PCR. - To confirm the correct product size, mix 3 µL of the PCR products with 2 µL of water and 1 µL of 6x dye. Run on a 1.5% agarose gel and confirm the correct product size.
- Purify the DNA using magnetic beads.
NOTE: Spin columns could also be used for PCR clean-up. Below are procedures for purifying PCR products using magnetic beads.- Remove magnetic beads from 4 °C and allow them to reach room temperature.
- Briefly vortex and quickly centrifuge the PCR mix.
- Add a volume of beads equal to 1.8× the volume of the PCR mix. Pipette the PCR-bead mix 10x to ensure the solution is equally mixed.
- Incubate at room temperature for 10 min.
- Transfer the PCR-bead mix to a PCR plate and place it on a magnetic plate.
- Incubate the plate on the magnet for 5 min. After the 5 min, check that the solution is clear, and the beads are bound to the magnet before moving to the next step.
- Discard the supernatant.
- Add 200 µL of 70% ethanol to the well and incubate for 30 s.
NOTE: The 70% ethanol should be prepared shortly before performing the clean-up to prevent DNA loss. - Discard the supernatant and repeat step 7.4.4.8.
- Remove the supernatant and allow to dry for 3 min at room temperature to remove traces of ethanol.
- Remove the plate from the magnet and add 22 µL of water to the sample. Pipette up and down 10x to mix water and beads.
- Incubate the samples for 3 min at room temperature.
- Transfer the plate back to the magnet and incubate for 3 min or until the solution is clear.
- Transfer 20 µL of the sample to a PCR tube. Take care that no beads are carried over.
- Sequence purified DNA amplicon by Sanger sequencing.
- Upload .ab1 sequence files for treated and control (untreated) samples to Tracking of indels by Decomposition (TIDE)16 to quantify indels at the target site.
- Extract genomic DNA from electroporated cells.
Representative Results
Isolation of plateable primary hepatocytes from the liver
The overall process of liver perfusion and hepatocyte isolation is illustrated in Figure 1. In this experiment, wild-type, 8-10-week-old C57BL6/6J mice were used. The procedure is expected to yield 20-50 × 106 cells per mouse with a viability between 85% and 95%. If the viability is <70%, percoll treatment should be followed to remove dead cells. Freshly isolated hepatocytes should be plated on collagen I-coated or nitrogen-containing tissue culture plates. Within 3-12 h of plating, the hepatocytes are expected to adhere to the plate, and the cell morphology assumes a typical polygonal or hexagonal appearance within 24 h (Figure 2). Hepatocytes are the only cells in the liver that store glucose in the form of glycogen. To verify the purity of the isolated cells, glycogen staining is performed using Periodic acid-Schiff reagent at 24 h after plating. The cytoplasm of the stained hepatocytes appears magenta (Figure 3).
Electroporation of CRISPR-Cas9 RNPs and mRNA into isolated mouse hepatocytes
Freshly isolated mouse hepatocytes were electroporated with eGFP mRNA, Cas9 mRNA along with Hpd-targeting sgRNA, or Cas9 protein complexed with Hpd-sgRNA (RNP). The Hpd-sgRNA were chemically modified with 2′-O-methyl phosphorothioate linkages to the first and last three consecutive nucleotides on the 5′ and 3′ ends14. Streptococcus pyogenes Cas9 was used for experiments. The hepatocytes were imaged 24 h after electroporation using a fluorescence microscope, and the percentage of GFP-positive cells was counted in the images (Figure 4A). On average, 89.8% [range 87.1%-92.4%] of the hepatocytes were GFP-positive. At 3 days after electroporation, Cas9-induced insertions and deletions (indels) in the Hpd locus were analyzed using TIDE16. The results show on-target indels of 47.4% in hepatocytes electroporated with CRISPR-Cas9 mRNA and 78.4% indels for the Cas9 RNP (Figure 4B). MTT and albumin assays were performed to evaluate hepatocyte viability and functionality after electroporating CRISPR-Cas9. The MTT results show a viability of 35.4% and 45.9% in hepatocytes treated with CRISPR-Cas9 mRNA and RNP, respectively (Figure 4C). Consistent with the MTT results, the normalized albumin levels were 31.8% in hepatocytes treated with Cas9 mRNA and 34.5% for Cas9 RNP (Figure 4D).
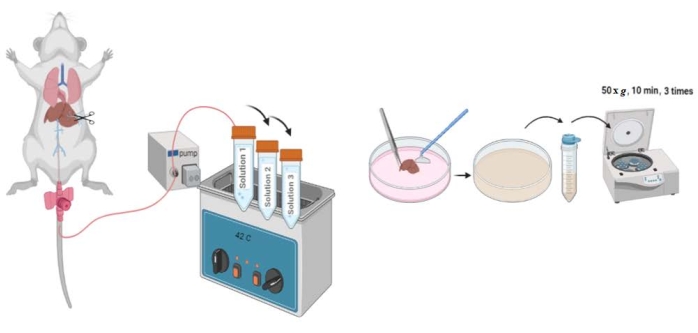
Figure 1: Schematic of the hepatocyte perfusion and isolation protocol. After cannulation of the inferior vena cava, the portal vein is cut, and Perfusion Solutions 1, 2, and 3 are pumped through the liver. The liver capsule is ruptured, and cells are released using a cell lifter. Released cells are strained and centrifuged. Please click here to view a larger version of this figure.
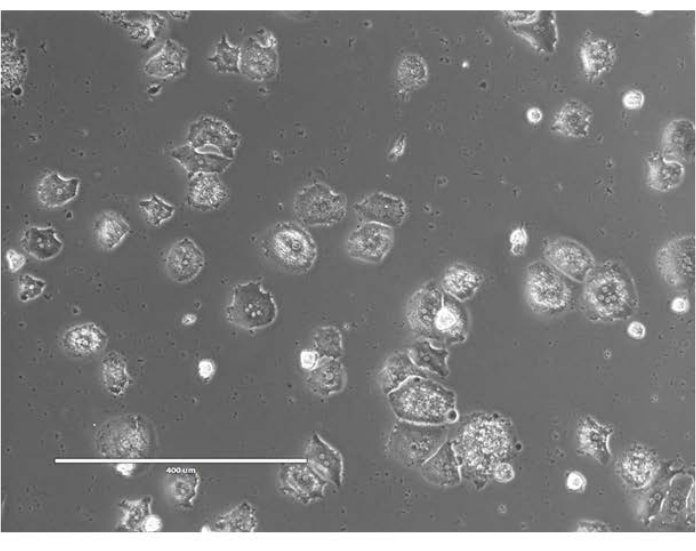
Figure 2: Freshly isolated primary hepatocytes. Hepatocytes isolated from C57BL/6J mice were plated on collagen I-coated 6-well plates with Hepatocyte Plating Media. Images taken at 24 h after plating. Scale bar = 400 µm. Please click here to view a larger version of this figure.
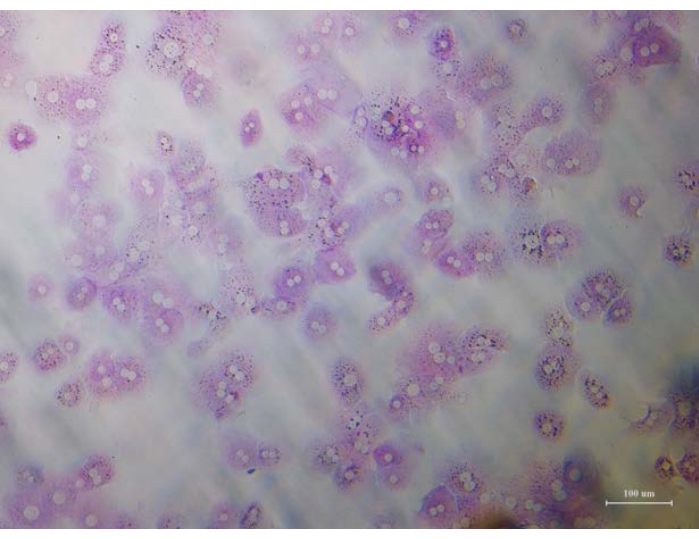
Figure 3: Glycogen staining. Freshly isolated primary mouse hepatocytes were stained for glycogen to confirm purity. Staining was done using Schiff Reagent at 24 h after plating. Cytoplasm of the stained hepatocytes appear magenta. Scale bar = 100 µm. Please click here to view a larger version of this figure.
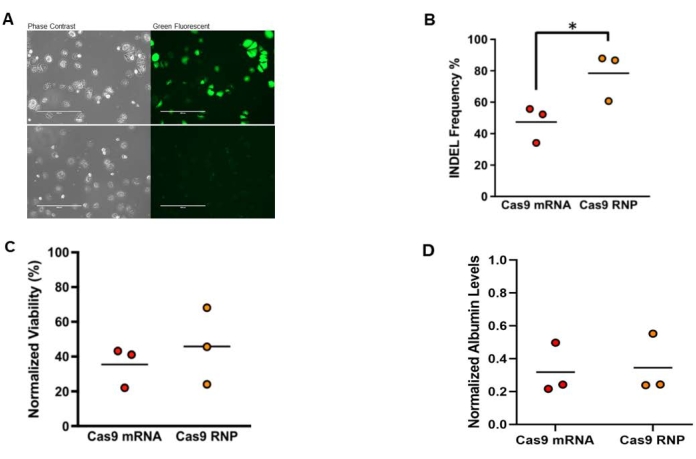
Figure 4: Electroporation of Hpd-targeting CRISPR-Cas9 into freshly isolated primary mouse hepatocytes. (A) Phase contrast and green fluorescence microscopy images of freshly isolated mouse hepatocytes at 24 h after electroporation. Images in the upper row are hepatocytes transfected with eGFP mRNA and the images in the lower row are untransfected control hepatocytes. Scale bars = 400 µm. (B) On-target indels for mouse hepatocytes electroporated with Cas9 mRNA combined with sgRNA or RNP. (C) Viability normalized to untransfected control hepatocytes in MTT assay at 24 h after electroporation. (D) Albumin levels in conditioned medium normalized to untransfected control hepatocytes measured at 24 h after electroporation(n = 3) with two technical replicates. Statistical analysis was performed by unpaired t-tests. This figure was modified from 14. Abbreviations: Hpd = 4-hydroxyphenylpyruvate dioxygenase; CRISPR = clustered regularly interspaced short palindromic repeats; Cas9 = CRISPR-associated protein 9; sgRNA = single guide RNA; RNP = ribonucleoprotein; INDEL = insertion-deletion; MTT = 3-[4,5-dimethylthiazol-2-yl]-2,5 diphenyl tetrazolium bromide. Please click here to view a larger version of this figure.
Table 1: Troubleshooting for the liver perfusion and hepatocyte isolation protocol. This table highlights common problems encountered during the protocol and suggests possible solutions. Please click here to download this Table.
Supplemental Figure S1: Mouse abdominal cavity during surgery. Catheter is inserted into the inferior vena cava (blue dot) before the renal branches (not seen). Abbreviations: L = Liver; K = Kidney; I = Small Intestines. Please click here to download this File.
Supplemental Table S1: Cas9 guide and PCR primer sequences. The guide sequence and PAM for sgRNA targeting Hpd, and PCR primer sequences for amplification of Hpd. Abbreviations: Hpd = 4-hydroxyphenylpyruvate dioxygenase; CRISPR = clustered regularly interspaced short palindromic repeats; Cas9 = CRISPR-associated protein 9; sgRNA = single guide RNA; PAM = protospacer-adjacent motif. Please click here to download this Table.
Discussion
The steps outlined in the protocol for hepatocyte isolation are challenging and require practice for proficiency. There are several key steps for the successful hepatocyte isolation from the liver. First, proper cannulation of the inferior vena cava is essential for complete liver perfusion. The absence of blanching in the liver after perfusion indicates displacement of the catheter (Table 1). The inferior vena cava (retrograde perfusion) was cannulated in the procedure because it is simpler and more accessible than the portal vein (antegrade perfusion)17,18. Some protocols use sutures to secure the catheter19; however, sutures complicate the process. Suturing the catheter can result in unwanted outcomes, such as the misplacement of the catheter and making it impossible to adjust its position. In addition, it is possible to perform the cannulation in the liver without removing the needle, which may further simplify the liver perfusion. Connecting the pump to the catheter is unnecessary with the needle in the vein. In addition, there is a reduced possibility of accidentally pulling out the catheter and forming blood clots. Inserting only the tip of the catheter with a flat angle relative to the vein enhanced the canulation. The inferior vena cava has several branches, so inserting the catheter in the wrong site will cause perfusion in another part of the body and not the liver (Table 1). Thus, it is vital to place the catheter in a site that avoids the branches. The vascular structure will vary slightly between different mice; thus, it is important to examine the veins before cannulating to determine the best site for injecting the catheter. The backflush of blood inside the catheter after removing the needle is an excellent indication of proper cannulation. Proper cannulation can also be checked by looking for liver swelling when pressure is momentarily applied to the portal vein (step 1.3.13).
When isolating hepatocytes, the second critical step is recognizing when complete liver digestion has occurred. The enzyme quality and concentration are essential for proper digestion. In contrast to the protocols found in 20,21, this protocol suggests using Liberase, consisting of two collagenase isoforms, because it provides better consistency in enzyme activity and reduced batch-to-batch variation than regular collagenase22. Variations in digestive enzyme quality and activity could cause inconsistencies in the isolation. It is essential to monitor the liver during digestion and stop the digestion immediately after the liver has softened. A digested liver will be flexible and show a loss in elasticity verified by depressing the liver using forceps or cotton swabs to confirm that indentations form without the tissue bouncing back. In this protocol, the maximum amount of Liberase solution recommended per liver is 50 mL; a higher volume would result in overdigestion. If the cannulation is perfectly performed, 30 mL of the Liberase solution is sufficient to achieve proper digestion. The final critical steps of the procedure are the isolation and washing steps. After dissecting the liver and transferring it to a sterile Petri dish containing ice-cold DMEM with FBS, it is crucial to avoid cutting the liver into pieces. At this stage, use a cell lifter to release the cells gently and maximize cell viability. Tilting the dish to submerge the liver in the DMEM facilitates the release of cells from the capsule into suspension and must be done slowly and on ice. It should be easy to disrupt the Glisson's capsule during the releasing step, and the medium should become cloudy and brown. Difficulty when disrupting the capsule is an indicator of poor digestion (Table 1).
This protocol is suitable for generating high levels of CRISPR-Cas9 edits in freshly isolated mouse hepatocytes. Indels generated by Cas9 RNP and mRNA were compared in the study. Higher levels of gene editing were observed in hepatocytes electroporated with the Cas9 RNP than mRNA, which is consistent with findings from other studies14,23,24. The advantage of Cas9 RNP over mRNA is that it provides lower off-target editing since the Cas9 protein exists in the cell for shorter periods than mRNA23,24. Because the efficiency of sgRNAs varies based on the design and target site, multiple sgRNAs should be designed and tested for editing efficiency at the gene of interest. Select the higher specificity scoring design when choosing between sgRNA having high gene efficiencies. If gene editing is consistently low, consider co-delivering a reporter, such as eGFP mRNA, to confirm delivery. Low levels of eGFP-positive cells after electroporation could indicate expired electroporation buffer, electroporation device issues, or air bubbles in the reaction. If there are high numbers of eGFP-positive cells but low editing efficiency, test the sgRNA design in a cell line to confirm editing activity and adjust the amounts of Cas9 and sgRNA electroporated into cells. Lastly, consider the method used to evaluate editing activity. Gel-based assays, such as T7 Endonuclease I, can under-report gene editing due to low sensitivity for 1 bp mismatches at the cut site. Deep sequencing is the most accurate method for quantifying Cas9 gene editing, particularly for low-editing events at off-target sites.
Another challenging aspect of the protocol is culturing freshly isolated hepatocytes after electroporation. Users must handle cells gently during each step of the protocol to obtain viable, plateable hepatocytes after electroporation. Avoid dispersing hepatocytes by vortex; instead, rock the vial containing cells gently until the cells become resuspended. When transferring hepatocytes, use wide-bore pipette tips to maintain viability. Incubating the cuvettes on ice for 15 min following electroporation allows the cell membrane to reseal and enhance viability. After plating, place the plate in the incubator and gently move the plate horizontally and vertically in a north-to-south and east-to-west motion to ensure the hepatocytes disperse evenly and maintain cell viability. If the hepatocytes fail to attach to the plate, consider increasing the number of cells to 1.3 × 106 in the electroporation reaction.
One promising application for the protocol is the generation of mouse models of human IMDs of the liver. Mouse hepatocytes that are positive for the gene encoding for fumarylacetoacetate hydrolase (Fah) have been shown to efficiently engraft and repopulate the liver after transplantation in Fah-deficient (Fah-/-) mice25. A novel approach for developing mouse models of IMDs of the liver is by electroporating CRISPR-Cas9 into wild-type mouse hepatocytes to introduce edits associated with a disease, followed by transplantation of the gene-edited hepatocytes in Fah-/- mice. The Cas9-edited hepatocytes would have a natural selective advantage after transplantation compared to the native Fah-deficient cells for repopulating the liver.
In conclusion, this protocol equips users with the capacity to isolate primary hepatocytes from the mouse liver, followed by gene-editing using CRISPR-Cas9 mRNA and RNPs. The protocol can be modified for use in different strains of mice to study various types of genetic diseases affecting the liver in vitro and in vivo and test therapeutic approaches.
Disclosures
The authors have nothing to disclose.
Acknowledgements
RNC received funding from South Carolina Bioengineering Center of Regeneration and Formation of Tissues Pilot grant supported by the National Institute of General Medical Sciences (NIGMS) of the National Institutes of Health, the American Association for the Study of Liver Diseases Foundation, and American Society of Gene & Cell Therapy under grant numbers P30 GM131959, 2021000920, and 2022000099, respectively. The content is solely the responsibility of the authors and does not necessarily represent the official views of the American Society of Gene & Cell Therapy or the American Association for the Study of Liver Diseases Foundation. The schematic for Figure 1 was created with BioRender.com.
Materials
| Equipment | |||
| 0.2 mL PCR 8-tube FLEX-FREE strip, attached clear flat caps, natural | USA Scientific | 1402-4700 | |
| 6-well Collagen Plates | Advanced Biomatrix | 5073 | |
| accuSpin Micro 17R | Fisher Scientific | 13-100-675 | |
| All-in-one Fluorescent Microscope | Keyence | BZ-X810 | |
| Analog Vortex Mixer | VWR | 97043-562 | |
| ART Wide BORE filtered tips 1,000 µL | ThermoFisher Scientific | 2079GPK | |
| Automated Cell Counter | Bio-Rad Laboratories | TC20 | |
| Blue Wax Dissection Tray | Braintree Scientific Inc. | DTW1 | 9" x 6.5" x 1/2"+F21 |
| Cell scraper/lifter | Argos Technologies | UX-04396-53 | Non-pyrogenic, sterile |
| Conical tubes (15 mL) | Fisher Scientific | 339650 | |
| Conical tubes (50 mL) | Fisher Scientific | 14-432-22 | |
| Cotton applicators | Fisher Scientific | 22-363-170 | |
| Curved scissors | Cooper Surgical | 62131 | |
| Disposable Petri Dishes | Falcon | 351029 | 100mm,sterile |
| Disposable Petri Dishes | VWR | 25373-100 | 35mm, sterile |
| Epoch Microplate Spectrophotometer | BioTek Instruments | 250082 | |
| Falcon Cell strainer (70 µm) | Fisher Scientific | 08-771-2 | |
| Forceps | Cooper Surgical | 61864 | Euro-Med Adson Tissue Forceps |
| IV catheters | BD | 382612 | 24 G x 0.75 in |
| IV infusion set | Baxter | 2C6401 | |
| MyFuge 12 Mini Centrifuge | Benchmark Scientific | 1220P38 | |
| Needles | Fisher Scientific | 05-561-20 | 25 G |
| Nucleofector 2b Device | Lonza | AAB-1001 | Program T-028 was used for electroporation in mouse hepatocytes |
| Peristaltic Pump | Masterflex | HV-77120-42 | 10 to 60 rpm; 90 to 260 VAC |
| Precision pump tubing | Masterflex | HV-96410-14 | 25 ft, silicone |
| Primaria Culture Plates | Corning Life Sciences | 353846 | Nitrogen-containing tissue culture plates |
| Serological Pipets (25 mL) | Fisher Scientific | 12-567-604 | |
| Syringes | BD | 329464 | 1 mL, sterile |
| T100 Thermal Cycler | Bio-Rad Laboratories | 1861096 | |
| Water bath | ALT | 27577 | Thermo Scientific Precision Microprocessor Controlled 280 Series, 2.5 L |
| Reagents | |||
| Alt-R S.p. Cas9 Nuclease V3 | IDT | 1081058 | |
| Beckman Coulter AMPure XP, 5 mL | Fisher Scientific | NC9959336 | |
| CleanCap Cas9 mRNA | Trilink Biotechnologies | L-7606-100 | |
| CleanCap EGFP mRNA | Trilink Biotechnologies | L-7201-100 | |
| Corning Matrigel Matrix | Corning Life Sciences | 356234 | |
| DMEM | ThermoFisher Scientific | 11885076 | Low glucose, pyruvate |
| Ethanol 70% | VWR | 71001-652 | |
| Fetal bovine serum | Thermoscientific | 26140-079 | |
| Hepatocyte Maintenance Medium (MM) | Lonza | MM250 | |
| Hepatocyte Plating Medium (PM) | Lonza | MP100 | |
| Mouse Albumin ELISA Kit | Fisher Scientific | NC0010653 | |
| Mouse/Rat Hepatocyte Nucleofector Kit | Lonza | VPL-1004 | |
| OneTaq HotStart DNA Polymerase | New England Biolabs | M0481L | |
| PBS 10x pH 7.4 | Thermoscientific | 70011-044 | No calcium or magnesium chloride |
| Percoll (PVP solution) | Santa Cruz Biotechnology | sc-500790A | |
| Periodic acid | Sigma-Aldrich | P7875-25G | |
| Permount Mounting Medium | VWR | 100496-550 | |
| QuickExtract DNA Extraction Solution | Lucigen Corporation | QE05090 | |
| Schiff’s fuchsin-sulfite reagent | Sigma-Aldrich | S5133 | |
| Trypsin-EDTA (0.25%) | ThermoFisher Scientific | 25200056 | Phenol red |
| Vybrant MTT Cell Viability Assay | ThermoFisher Scientific | V13154 | |
| Perfusion Solution 1 (pH 7.4, filter sterilized) | Stable at 4 °C for 2 months | ||
| EBSS | Fisher Scientific | 14155063 | Complete to 200 mL |
| EGTA (0.5 M) | Bioworld | 40520008-1 | 200 µL |
| HEPES (pH 7.3, 1 M) | ThermoFisher Scientific | AAJ16924AE | 2 mL |
| Perfusion Solution 2 (pH 7.4, filter sterilized) | Stable at 4 °C for 2 months | ||
| CaCl2·2H20 (1.8 mM) | Sigma | C7902-500G | 360 µL of 1 M stock |
| EBSS (no calcium, no magnesium, no phenol red) | Fisher Scientific | 14-155-063 | Complete to 200 mL |
| HEPES (1 M) | ThermoFisher Scientific | AAJ16924AE | 2 mL |
| MgSO4·7H20 (0.8 mM) | Sigma | 30391-25G | 160 µL of 1 M stock |
| Perfusion Solution 3 | Prepared fresh prior to use | ||
| Solution 2 | 50 mL | ||
| Liberase | Roche | 5401127001 | 0.094 Wunsch units/mL |
| Mouse Anesthetic Cocktail | |||
| Acepromzine | 0.25 mg/mL final concentration | ||
| Ketamine | 7.5 mg/mL final concentration | ||
| Xylazine | 1.5 mg/mL final concentration | ||
| Software | URL | ||
| Benchling | https://www.benchling.com/ | ||
| ImageJ | https://imagej.nih.gov/ij/ | ||
| TIDE: Tracking of Indels by Decomposition | https://tide.nki.nl/ |
References
- Pampols, T. Inherited metabolic rare disease. Advances in Experimental Medicine and Biology. 686, 397-431 (2010).
- Saudubray, J. M., Sedel, F., Walter, J. H. Clinical approach to treatable inborn metabolic diseases: an introduction. Journal of Inherited Metabolic Disease. 29 (2-3), 261-274 (2006).
- Arnon, R., et al. Liver transplantation in children with metabolic diseases: the studies of pediatric liver transplantation experience. Pediatric Transplantation. 14 (6), 796-805 (2010).
- Maiorana, A., et al. Preemptive liver transplantation in a child with familial hypercholesterolemia. Pediatric Transplantation. 15 (2), 25-29 (2011).
- . OPTN/SRTR 2019 Annual Data Report: Liver Available from: https://srtr.transplant.hrsa.gov/annual_reports/2019/Liver.aspx (2019)
- Fabris, L., Strazzabosco, M. Rare and undiagnosed liver diseases: challenges and opportunities. Translational Gastroenterology and Hepatology. 6, (2020).
- Haugabook, S. J., Ferrer, M., Ottinger, E. A. In vitro and in vivo translational models for rare liver diseases. Biochimica et Biophysica Acta (BBA) – Molecular Basis of Disease. 1865 (5), 1003-1018 (2019).
- Xue, Y., et al. Embryonic lethality and vascular defects in mice lacking the notch ligand Jagged1. Human Molecular Genetics. 8 (5), 723-730 (1999).
- Lima, A., Maddalo, D. SEMMs: Somatically engineered mouse models. a new tool for in vivo disease modeling for basic and translational research. Frontiers in Oncology. 11, 1117 (2021).
- Kim, S., Kim, D., Cho, S. W., Kim, J., Kim, J. S. Highly efficient RNA-guided genome editing in human cells via delivery of purified Cas9 ribonucleoproteins. Genome Research. 24 (6), 1012-1019 (2014).
- Ye, L., et al. Seamless modification of wild-type induced pluripotent stem cells to the natural CCR5Delta32 mutation confers resistance to HIV infection. Proceedings of the National Academy of Sciences. 111 (26), 9591-9596 (2014).
- Straub, C., Granger, A. J., Saulnier, J. L., Sabatini, B. L. CRISPR/Cas9-mediated gene knock-down in post-mitotic neurons. PLoS One. 9 (8), 105584 (2014).
- Hoban, M. D., et al. CRISPR/Cas9-mediated correction of the sickle mutation in human CD34+ cells. Molecular Therapy. 24 (9), 1561-1569 (2016).
- Rathbone, T., et al. Electroporation-mediated delivery of Cas9 ribonucleoproteins results in high levels of gene editing in primary hepatocytes. The CRISPR Journal. , (2022).
- . Benchling [Biology Software] Available from: https://benchling.com (2022)
- Brinkman, E. K., Chen, T., Amendola, M., van Steensel, B. Easy quantitative assessment of genome editing by sequence trace decomposition. Nucleic Acids Research. 42 (22), 168 (2014).
- Cabral, F., et al. Purification of hepatocytes and sinusoidal endothelial cells from mouse liver perfusion. Journal of Visualized Experiments: JoVE. (132), e56993 (2018).
- Huang, P., et al. Induction of functional hepatocyte-like cells from mouse fibroblasts by defined factors. Nature. 475 (7356), 386-389 (2011).
- Severgnini, M., et al. A rapid two-step method for isolation of functional primary mouse hepatocytes: cell characterization and asialoglycoprotein receptor based assay development. Cytotechnology. 64 (2), 187-195 (2012).
- Jung, Y., Zhao, M., Svensson, K. J. Isolation, culture, and functional analysis of hepatocytes from mice with fatty liver disease. STAR Protocols. 1 (3), 100222 (2020).
- Kim, Y., et al. Three-dimensional (3D) printing of mouse primary hepatocytes to generate 3D hepatic structure. Annals of Surgical Treatment and Research. 92 (2), 67-72 (2017).
- Li, W. C., Ralphs, K. L., Tosh, D. Isolation and culture of adult mouse hepatocytes. Methods in Molecular Biology. 633, 185-196 (2010).
- Lattanzi, A., et al. Optimization of CRISPR/Cas9 delivery to human hematopoietic stem and progenitor cells for therapeutic genomic rearrangements. Molecular Therapy. 27 (1), 137-150 (2019).
- Dever, D. P., et al. CRISPR/Cas9 beta-globin gene targeting in human haematopoietic stem cells. Nature. 539 (7629), 384-389 (2016).
- Grompe, M., Tanguay, R. M. . Hereditary Tyrosinemia: Pathogenesis, Screening and Management. , 215-230 (2017).

