Size Exclusion Chromatography for Separating Extracellular Vesicles from Conditioned Cell Culture Media
Summary
The protocol here demonstrates that extracellular vesicles can be adequately separated from conditioned cell culture media using size exclusion chromatography.
Abstract
Extracellular vesicles (EVs) are nano-sized lipid-membrane bound structures that are released from all cells, are present in all biofluids, and contain proteins, nucleic acids, and lipids that are reflective of the parent cell from which they are derived. Proper separation of EVs from other components in a sample allows for characterization of their associated cargo and lends insight into their potential as intercellular communicators and non-invasive biomarkers for numerous diseases. In the current study, oligodendrocyte derived EVs were isolated from cell culture media using a combination of state-of-the-art techniques, including ultrafiltration and size exclusion chromatography (SEC) to separate EVs from other extracellular proteins and protein complexes. Using commercially available SEC columns, EVs were separated from extracellular proteins released from human oligodendroglioma cells under both control and endoplasmic reticulum (ER) stress conditions. The canonical EV markers CD9, CD63, and CD81 were observed in fractions 1-4, but not in fractions 5-8. GM130, a protein of the Golgi apparatus, and calnexin, an integral protein of the ER, were used as negative EV markers, and were not observed in any fraction. Further, when pooling and concentrating fractions 1-4 as the EV fraction, and fractions 5-8 as the protein fraction, expression of CD63, CD81, and CD9 in the EV fraction was observed. The expression of GM130 or calnexin was not observed in either of the fraction types. The pooled fractions from both control and ER stress conditions were visualized with transmission electron microscopy and vesicles were observed in the EV fractions, but not in the protein fractions. Particles in the EV and protein fractions from both conditions were also quantified with nanoparticle tracking analysis. Together, these data demonstrate that SEC is an effective method for separating EVs from conditioned cell culture media.
Introduction
The explosion of interest in studying extracellular vesicles (EVs) has been accompanied by major advancements in the technologies and techniques used to separate and study these nano-sized, heterogeneous particles. In the time since their discovery nearly four decades ago1,2, these small membranous structures have been found to contain bioactive lipids, nucleic acids, and proteins, and play major roles in intercellular communication3,4. EVs are released from all cell types and are therefore present in all biological fluids, including blood plasma and serum, saliva, and urine. EVs within these fluids hold great promise to serve as non-invasive biomarkers for various diseases, including neuroinflammatory and neurodegenerative diseases, cancers, and autoimmune disorders5,6,7. Further, in vitro mechanistic studies can be performed through cell culture techniques by separating EVs released into the culture medium3,8,9.
To understand the role of EVs in disease pathophysiology, adequate separation from the fluid in which they are found is paramount. The gold standard for EV separation has long been differential ultracentrifugation (dUC)10, however more sophisticated techniques have arisen to achieve better separation of EVs from other extracellular components. Some of these techniques include density gradients, asymmetrical-flow field-flow fraction (A4F), flow cytometry, immunocapture, polyethylene glycol precipitation, and size exclusion chromatography (SEC)11,12,13. Each technique has its own set of advantages and disadvantages; however, SEC in particular has been shown to separate EVs from both biological fluids and cell culture supernatants quite effectively8,14,15. SEC also has the added bonus of being relatively straightforward and user-friendly.
SEC is a method that separates components of a fluid based on size. With this technique, a column of resin (either made in-house or purchased commercially) is used to fractionate a sample. Small particles in the sample get trapped between the beads within the resin, while larger particles are able to pass through the resin more freely, and thus elute earlier in the process. Because EVs are larger in size than many extracellular proteins and protein aggregates, EVs pass through the column faster and elute in earlier fractions than extracellular proteins14.
In this methods paper, the use of SEC for separation of EVs from cell culture media (CCM) from human oligodendrocytes under both control and endoplasmic reticulum (ER) stress conditions is outlined. Using this protocol, it is shown that EVs separated with this technique are found within specific fractions that can be pooled together and concentrated for downstream characterization, and that the separated EVs are derived from cells and not from an exogenous source such as fetal bovine serum (FBS) used to supplement the CCM. The presence of the canonical EV markers, CD63, CD81, and CD916,17,18,19 in the EV fractions, and their absence in the protein fractions is demonstrated with western blotting. Using transmission electron microscopy (TEM), EVs are visualized and display the expected morphology and are only observed in the EV fraction. Particles are also counted in the EV and protein fractions of both control and ER stress conditions, and a large number of particles within the expected size range of 50-200 nm in diameter are observed in the EV samples. Together, these data support the notion that SEC is an efficient and effective method for separating EVs from cell culture media.
Protocol
1. Preparation of buffers and reagents
NOTE: Make cell culture reagents in cell culture hood to maintain sterility.
- Preparation of cell culture reagents
- Prepare normal high glucose DMEM by adding 50 mL of FBS and 5 mL of penicillin-streptococcus (Pen-Strep) into 500 mL of high glucose DMEM and store at 4 °C. Use this media for culturing and expanding cells.
- Prepare exosome-depleted high glucose DMEM by adding 50 mL of exosome-depleted FBS and 5 mL of Pen-Strep into 500 mL of high glucose DMEM and store at 4 °C. Use this media for cell treatments prior to EV isolation.
- Prepare tunicamycin by adding 10 mg of tunicamycin to 1 mL of dimethyl sulfoxide (DMSO). Make 50 µL aliquots in 0.2 mL tubes and store at -20 °C.
- Preparation of EV separation reagents
- Prepare 1x phosphate buffered saline (PBS) by adding 9.89 g of PBS powder to 1 L of deionized (DI) water in a 1 L graduated cylinder. Stir the solution on a stir plate until the PBS dissolves.
- Use the autoclave to sterilize the glassware before adding PBS. Put the glassware into a bin, close the door, and sterilize for 30 min at 121 °C.
- Inside of a laminar flow cabinet, connect a vacuum to the 0.22 µm filter and place filter on top of the autoclaved bottle. Slowly pour the PBS onto the filter and collect the filtered PBS in the autoclaved bottle.
- Prepare 0.5 M sodium hydroxide by adding 9.99 g of sodium hydroxide to 500 mL of 1x PBS in a 500 mL graduated cylinder. Stir the solution on a stir plate until the sodium hydroxide dissolves into the PBS and then filter using a 0.22 µm filter into an autoclaved 500 mL bottle.
- Prepare 0.05% sodium azide by adding 0.25 g of sodium azide to 500 mL of 1x PBS in a 500 mL graduated cylinder. Stir the solution on a stir plate until sodium azide dissolves into the PBS, and then filter using a 0.22 µm filter into an autoclaved 500 mL bottle.
- Preparation of protein isolation, quantification, and identification reagents
- Prepare 100 mL of RIPA lysis buffer by combining 1 mL of 1 M Tris (pH 7.4), 1 mL of 10% sodium dodecyl sulfate (SDS), 1 g of deoxycholate, 1 mL of NP-40, and 0.89 g of sodium chloride (NaCl). Bring the volume to 100 mL with distilled H2O and store at 4 °C. To make RIPA plus phosphatase and protease inhibitor solution, add 10 µL of phosphatase and protease inhibitor for every 1 mL of RIPA buffer in a 15 mL tube and store the solution at 4 °C.
- Immediately before use, prepare the BCA assay working reagent by adding 10 mL of reagent A and 200 µL of reagent B provided by the BCA assay kit into a 15 mL tube. Immediately before use, prepare the microBCA assay working reagent by adding 25 parts of reagent A, 24 parts of reagent B, and 1 part of reagent C into a 15 mL tube.
- Prepare 10x gel electrophoresis running buffer by adding 30.3 g of Tris base, 144 g of glycine, and 10 g of SDS in 1 L of DI water. Store the running buffer at 4 °C. Prepare 1x transfer buffer by adding 3.03 g of Tris base, 14.4 g of glycine, and 200 mL of methanol and adjust the volume to 1 L with DI water. Store the transfer buffer at 4 °C.
- Prepare Tris buffered saline with Tween-20 (TBS-Tween) by adding 2.4 g of Tris base, 8 g of NaCl, and 1 mL of Tween-20 into 1 L of DI water. Store the TBS-Tween at 4 °C.
- Prepare 5% blocking solution by adding 5 g of powdered non-fat milk to 100 mL of TBS-Tween. Store the blocking buffer at 4 °C.
- Immediately before use, prepare chemiluminescent substrate by adding 4 mL of peroxide solution and 4 mL of luminol/enhancer solution into a 15 mL tube and invert gently to mix.
2. Culturing and treating cells
- Seed two T175 cell culture flasks with 2.3 x 106 human oligodendroglioma (HOG) cells each. Bring final volume of normal high glucose DMEM to 25 mL and incubate at 37 °C with 5% CO2 in a cell culture incubator for 48 h.
- At the end of the 48 h, warm exosome-depleted high glucose DMEM in a 37 °C water bath. For treatment, add 25 µL of 10 mg/mL of tunicamycin to 25 mL of exosome-depleted media, final concentration of 10 µg/mL of tunicamycin to induce ER-stress. For the control, add 25 µL of DMSO to 25 mL of exosome-depleted media.
- Remove and dispose of the normal high glucose DMEM from cells and gently rinse cells and flask with 10 mL of 1x PBS. Aspirate and discard PBS.
- Add 25 mL of control media into the flask labeled control, and 25 mL of 10 µg/mL tunicamycin in media into the flask labeled treatment. Return flasks to cell culture incubator for 24 h.
3. Collection and concentration of conditioned CCM (Figure 1)
- Place PBS in a 37 °C water bath. Observe flasks under a compound microscope with 10x and 100x objective lens. Take notes of any differences between the flasks. Perform the following steps for both control and treated flasks.
- Remove media from flask and place in a 50 mL tube. Centrifuge the tube at 500 x g for 5 min at 4 °C. Transfer the supernatant to a new 50 mL tube.
- Centrifuge the tube at 2,000 x g for 20 min at 4 °C. Transfer the supernatant to a new 50 mL tube and place on ice.
- Obtain four 15 mL 3 kDa cutoff ultrafiltration units and add 5 mL of PBS to each tube. Prime the filter by centrifuging the ultrafiltration units at 4,000 x g for 10 min at 4 °C.
NOTE: 3 kDa molecular weight cutoff ultrafiltration units were used in the current protocol to retain all extracellular proteins released from cells. Larger molecular weight cutoff ultrafiltration units (e.g., 30 kDa) would also work with this protocol, however extracellular proteins smaller than 30 kDa would be filtered out and removed from the samples20,21. - Remove PBS from the ultrafiltration units. Divide the supernatant from each condition (control and tunicamycin treated) into two ultrafiltration units each, approximately 12.5 mL of media in each.
- Centrifuge the tubes at 4,000 x g for 1 h 45 min at 4 °C, or until the concentrated media (referred to as the retentate) in each ultrafiltration unit is concentrated to 250 µL or less.
- Transfer the retentate from both control ultrafiltration units into one labeled 1.5 mL microcentrifuge tube. Transfer the retentate from both treated ultrafiltration units into another labeled 1.5 mL microcentrifuge tube.
- Measure the total volume of the retentate in each microcentrifuge tube to be 500 µL. If the sample is less than 500 µL, add 0.22 μm filtered 1x PBS until the final volume is 500 µL. Place tubes in a -80 °C freezer.
4. Cell collection
- Place trypsin, PBS, and exosome-depleted DMEM in a 37 °C water bath. Add 7 mL of trypsin to both control and treated flasks and place in the incubator for 5 min.
- View under the microscope to see if the cells are lifted. If the cells are not lifted, firmly tap the flask against palm of the hand to dislodge cells.
- Wash the flask with 7 mL of exosome-depleted DMEM. Collect the cell suspension from the flask and place into 15 mL tubes.
- Centrifuge the tubes for 10 min at 500 x g at 4 °C. Remove the supernatant and add 5 mL of 1x PBS to cell pellet. Gently pipette mix to break up the pellet.
- Centrifuge the tubes at 500 x g for 5 min at 4 °C. Remove the supernatant and add 200 µL of RIPA plus phosphatase and protease inhibitor solution to cell pellet, pipette to mix.
- Transfer the cells in the RIPA solution into 1.5 mL tubes. Let the tubes sit on ice for 5 min. Centrifuge the tubes at 14,000 x g for 10 min at 4 °C.
- Transfer the supernatant to new 1.5 mL tubes. Label the tubes with treatment condition and date. Place tubes into a -80 °C freezer.
NOTE: The protocol can be paused here.
5. EV separation using size exclusion chromatography (Figure 2)
NOTE: Perform the following steps twice, once for control retentate and once for tunicamycin treated retentate.
NOTE: The PBS utilized in this section is 0.22 μm filtered 1x PBS.
- Turn on the automatic fraction collector (AFC) and adjust settings to collect eight fractions, 0.5 mL per fraction, and buffer (void) volume to default.
NOTE: The AFC operating conditions are as follows: column/bed volume = 10 mL; void volume = 2.70 mL; flush volume = 15 mL; optimal fraction size = 500 µL; buffer required per run = 65 mL; flow rate at 20 °C = 1.0 mL/min; reusability of the column = five uses. - Remove the column (see Table of Materials) from 4 °C and let the column warm up to room temperature before beginning (usually ~30 min).
- Remove retentate samples from the -80 °C freezer and place on ice to thaw. While waiting, set eight labeled 1.5 mL tubes in the AFC carousel with lids facing inwards.
- Insert column into the AFC with the barcode facing the reader. Start the collection by following the instructions on screen to verify there are tubes in the carousel and the waste collector is functioning properly.
- Begin flushing the column by adding 15 mL of 1x PBS to column after the storage buffer is completely absorbed into the top portion of the column matrix. Do not add PBS too early as this will dilute the storage buffer and prevent adequate flushing.
- Just before all of the 15 mL of PBS is absorbed by the matrix, click OK to stop the flushing and remove any residual PBS from the top of the column matrix with a micropipette. Do not touch the matrix.
- Add 500 µL of sample to the center of the column. Click OK to begin the run. Watch the sample absorb into the matrix and quickly add 8 mL of PBS to the column immediately after the sample is fully absorbed.
- AFC will automatically dispel the void volume into the center of the carousel and then collect all eight fractions of 500 µL each. At the conclusion of the run, remove fractions from the carousel and place on ice. Fractions 1-4 contain EVs while fractions 5-8 contain extracellular proteins. Remove void volume from the center of carousel.
- Add 10 mL of PBS to the column after the eight fractions have been collected to wash any residual sample from the column. After the retentate sample has been fully flushed through the column, add 15 mL of 1x PBS to the column.
- As the final PBS is being absorbed by the matrix, add 500 µL of 0.5 M sodium hydroxide to the center of the column's matrix. As the sodium hydroxide is absorbed, add 30 mL of 1x PBS to the column and let it flush through.
- If running another sample, add eight labeled 1.5 mL microcentrifuge tubes to carousel and repeat steps 5.7-5.10.
- Once done with all samples, complete the following steps to preserve the column for later use. After flushing 30 mL of PBS through the column as outlined in step 5.10, add 15 mL of 0.05% sodium azide to the column.
- Leave approximately 5 mL of sodium azide on the top of the column matrix. Remove the column from the AFC and attach the top and bottom caps. Place column back at 4 °C for storage (column can be used up to five times).
6. Ultrafiltration of EV and protein fractions
- Obtain two 2 mL, 3 kDa cutoff ultrafiltration units per sample. Use one unit to concentrate the EV fractions (fractions 1-4), and the other to concentrate the protein fractions (fractions 5-8). Each unit is comprised of three parts: cone, filter, and flow-through cylinder; label each part properly.
- Construct the ultrafiltration unit and add 1 mL of 0.22 μm filtered 1x PBS to the filter portion and cap with the cone piece. Centrifuge the ultrafiltration unit, cone-side up at 3,500 x g for 10 min at 4 °C.
- Remove the filtered PBS from the flow-through cylinder. Flip the ultrafiltration unit upside down (cone-side down) and centrifuge for 2 min at 1,000 x g at 4 °C. Remove any remaining PBS from the cone.
- Add the appropriate fractions (the entire 500 µL volume) to each ultrafiltration unit (total volume will be 2 mL). Centrifuge columns cone-side up for 1 h 45 min at 3,500 x g at 4 °C, or until the retentate level is at 100 µL.
- Remove the flow-through cylinder and discard flow through. Flip the ultrafiltration unit upside down (cone-side down) and centrifuge for 2 min at 1,000 x g at 4 °C.
- Transfer the retentate in the cone to a labeled 1.5 mL microcentrifuge tube and bring each sample volume to 150 µL with 0.22 µm filtered 1x PBS. Place tubes in -80 °C freezer.
NOTE: The protocol can be paused here.
7. Media controls
- Obtain two T175 cell culture flasks. Label one flask control and the other treatment. Warm exosome-depleted high glucose DMEM in a 37 °C water bath.
- Add 25 µL of tunicamycin stock solution (10 mg/mL) to 25 mL of media and put in the flask labeled treatment. Add 25 µL of DMSO to 25 mL of media and put in the flask labeled control. Keep flasks in a cell culture incubator for 24 h.
- Collect media and process to separate EVs as outlined in steps 3.2-3.8. Subsequently, perform all steps described in sections 5 and 6.
8. Western blotting
- Protein quantification from cell lysates and protein fractions with BCA assay
NOTE: Protein quantification from both control and tunicamycin treated samples is performed with the following steps.- Thaw lysed cells and protein fractions on ice. Make three dilution tubes for each sample; 1:2, 1:10, and 1:100.
- Load in triplicate on a 96-well clear-bottomed assay plate, 25 µL of bovine serum albumin (BSA) protein standards and 25 µL of each sample dilution. Add 200 µL working reagent to each well containing standard or sample, using a multichannel pipette.
- Incubate the plate at 37 °C for 30 min, then read plate with a plate reader at an absorbance of 562 nm. Calculate protein concentration in each sample and determine the volume of sample needed to obtain 20 µg of protein for cell lysates and 15 µg of protein for protein fractions.
- Protein quantification from EV fractions with microBCA assay
- Thaw EV samples on ice. Make two dilutions for each sample: 1:50 and 1:100.
- Load in triplicate on a 96-well clear-bottomed assay plate, 100 µL of BSA protein standards, and 100 µL of each sample dilution. Add 100 µL of microBCA working reagent to each well containing standard or sample, using a multichannel pipette.
- Incubate the plate at 37 °C for 2 h, then read plate with a plate reader at an absorbance of 562 nm. Calculate protein concentration in each sample and determine the volume of sample needed to obtain 15 µg of protein for EV fractions.
- Western blotting
NOTE: Western blotting protocol was performed similar to as described in22, and modified as outlined below.- Make 10% polyacrylamide gels with a gel making kit, using the manufacturer's instructions.
- For gel electrophoresis, prepare cell lysate samples by combining in a 1.5 mL tube, the volume of cell lysate necessary for 20 µg of protein, 5 µL of 4x Laemmli buffer, and the volume of RIPA buffer necessary to bring the total volume to 20 µL.
- Prepare EV and protein fraction samples by combining in a 1.5 mL tube the volume of sample necessary for 15 µg of protein, 5 µL of 4x Laemmli buffer, and RIPA buffer to a volume of 20 µL.
- Prepare media control samples by combining in a 1.5 mL tube 15 µL of sample and 5 µL of 4x Laemmli buffer.
NOTE: Depending on the antibodies to be used, Laemmli buffer may or may not need to contain a reducing agent, such as 50 mM dithiothreitol (DTT). - Boil all samples at 95 °C for 5 min, transfer samples to ice to cool, and briefly centrifuge for 30 s at 10,000 x g, then return samples to ice.
- Add gels to electrophoresis tank and add 1x gel electrophoresis running buffer. Load 15 µL of protein ladder and 20 µL of sample. Run the gel at 200 V for 30-50 min, or until samples have reached the bottom of the gel, just prior to running off.
- Transfer the protein to a PVDF membrane with 1x transfer buffer at 4 °C for 30 min at 100 V, or until the transfer is complete. Supplementary Figure 1, Supplementary Figure 2, and Supplementary Figure 3 show stain-free images of each blot after the transfer is completed.
- Block the membrane in 5% milk/TBS-Tween for 1 h at room temperature on a shaker. Incubate membrane in primary antibody solution overnight on a shaker at 4 °C. See Table 1 for antibody dilution information.
- The following day, remove the primary antibody and wash the membrane 3x with TBS-Tween for 5 min each at room temperature on a shaker.
- Prepare appropriate secondary antibodies in 5% milk/TBS-Tween. See Table 1 for dilution information. Add secondary antibody to membrane and incubate on a shaker at room temperature for 1 h, then wash 3x with TBS-Tween for 5 min each at room temperature on a shaker.
- Add chemiluminescent substrate to membrane and incubate for 5 min at room temperature on a shaker. Image blot by using colorimetric and chemiluminescent analysis. Supplementary Figure 4, Supplementary Figure 5, and Supplementary Figure 6 show uncropped images of each blot.
9. TEM imaging
- Glow discharge23 400 mesh carbon/formvar coated grids to remove hydrocarbons and make the grids hydrophilic.
- Add 5 µL of EV or protein fraction sample to grids. Incubate grids at room temperature for 3-5 min.
- Remove excess solution by wicking with filter paper. Wash the grids with 5 µL of nanopure water and remove the excess by wicking.
- Apply 5 µL of 1% aqueous uranyl acetate and immediately wick off. Allow grids to dry. Image grids at 120 kV on a transmission electron microscope and capture images with a charged-couple device camera.
10. Nanoparticle tracking analysis (NTA)
- Obtain EV and protein fraction samples from -80 °C and thaw on ice.
- Turn on the computer and the NTA instrument. Open the NTA software and it will begin to initialize automatically.
- Select the appropriate pump that contains particle-free water and click Rinse The Instrument to rinse the cell. While rinsing, inject 10-20 mL of particle-free water into injection port on the front of the instrumentusing a syringe and avoid air bubbles.
- Cell quality check screen will pop up; click OK and cell quality check will begin. After the check is successfully passed, the autoalignment pop-up screen will appear stating: Please fill the cell with 100 nm alignment suspension (dilution 1:250,000).
- Prepare dilution 1 of alignment suspension by adding 10 mL of particle-free water and 10 µL of 100 nm polystyrene beads to a tube (1:1,000 dilution). Gently invert to mix. Dilution 1 has a shelf life of 2-3 days at 4 °C.
- Create dilution 2 of alignment suspension by adding 20 mL of particle-free water and 80 µL of dilution 1 (1:1,000) into a tube to create the dilution of 1:250,000. Gently invert the tube to mix. Dilution 2 has a shelf life of 30-60 min at room temperature.
- Fill cell with 2 mL of dilution 2 (1:250,000). Click OK and the program will complete autoalignment (focus) and focus optimization. The analysis tab will open up creating a profile that provides results of the cleanliness and symmetry of the measuring cell in a parabola. A pop-up screen stating: View Video Microscope is now ready for experiments, will appear; click OK and the program will return to the cell check tab.
- Rinse the polystyrene beads from the cell by injecting particle-free water into the injection port to wash the cell. Continue washing until the number of detected particles is between 0-10 (usually between 5-10 mL). Add 1 mL of 0.22 µm filtered PBS to prime the cell for the first sample.
- Dilute the first sample with 0.22 µm filtered PBS (start with 1:1000 dilution) and enter the dilution factor into the program. Insert 1 mL of sample into the cell and wait 5 min for particle movement to slow, as they will initially move very quickly across the screen. Set the sensitivity and shutter at 75.0 and 100, respectively.
- The number of detected particles for ideal sample dilution is 50-200 particles. If below 50 or above 200 particles are reported, adjust the dilution of the sample. If a new dilution is required, wash the cell with particle-free water until there are 0-10 detected particles. Repeat adding diluted sample and washing until the ideal dilution is found.
- Check all 11 camera positions to visualize that particle movement has slowed and ensure that the number of detected particles is similar between all camera positions. If the number of particles differs greatly across camera positions, add more sample.
- Click on the Measurement tab and Run Video Acquisition. Under the video acquisition tab, enter custom name for sample and identify a safe location. Check the autosave.txt and autosave.pdf boxes to save data.
- Set the temperature to 25 °C, click on 488 nm to set laser, and click on 11 for camera positions. Click OK to run data collection. The program will begin analyzing the number of particles and size in all 11 positions. Under the analysis tab, a bar graph will appear with diameter/nm on the x-axis and particles/mL on the y-axis.
- After data is collected, a pop-up screen titled Positions Overview will appear providing data for each of the 11 positions. If there are more than two positions removed from analysis, repeat the procedure with more sample. If not, click on the Weitermachen button. A PDF and txt file will open containing the results. Save the files in a safe location.
- To run another sample, go to Cell Check tab and repeat steps 10.6 through 10.12. If done, wash cell with particle-free water until the number of detected particles is 0-10 (about 5-10 mL). Rinse cell with 1 mL of 0.22 µm filtered PBS, close the program, and turn off the computer.
Representative Results
Western blotting reveals adequate separation of EVs from CCM
To evaluate the effectiveness of SEC for separating EVs from cell culture media, a western blot was run using each individual fraction from the control samples to probe expression of the three canonical EV markers, CD9, CD63 and CD81, as well as GM130 and calnexin18, which were used as negative controls (Figure 3). Albumin expression18 was also probed to ensure that extracellular proteins in the CCM can be adequately separated from EVs. Strong expression of CD9, CD63, and CD81 was observed in fractions 1-4, with little to no expression in fractions 5-8; neither GM130 nor calnexin were observed in any fraction. Albumin was only present in fractions 6-8. Together, these data indicate that vesicles elute predominately into fractions 1-4, leaving fractions 5-8 to contain extracellular proteins. Because of this, fractions 1-4 were combined and concentrated and deemed the EV fraction, while fractions 5-8 were combined and concentrated and referred to as the protein fraction.
Next, additional western blots were run to evaluate the effectiveness of concentrating the EV and protein fractions via ultrafiltration in both control and treated samples. Expression of CD9, CD63, and CD81 was observed in the cell lysates and EV fractions of both control and tunicamycin treated samples, but not in the protein fractions (Figure 4). Tumor susceptibility gene 101 (TSG101) is associated with the endosomal sorting complex required for transport (ESCRT)24 and is commonly used as a marker for exosomes25. This protein was present in cell lysates but not in EV or protein fractions. Additionally, GM130 and calnexin were only observed in the cell lysate samples. Therefore, the data indicates that EVs were effectively separated from cell culture media.
To ensure that the positive signal observed in the EV fractions was coming from EVs released by the cells and not the media itself, the exosome-depleted media underwent the same ultrafiltration and SEC protocol as conditioned CCM and was then analyzed via western blot (Figure 5). Both control and tunicamycin containing medias were processed and cell lysates were used as a positive control on the western blots. No EV markers (CD9, CD63, CD81, or TSG101) were observed in the media samples, but were present in the cell lysates. Albumin expression was observed within the protein fractions and minimal expression in the cell lysates, likely residual from the media. As no signal is observed for any of the EV markers in the EV fractions, these data demonstrate that exosome-depleted media does not contain EVs that mask the signal of cell-derived EVs.
TEM demonstrates expected morphology of EVs
TEM images were taken of both the control and tunicamycin treated EV and protein fractions (Figure 6). Spherical structures can be observed in the EV fractions of both the control and tunicamycin treated samples, indicating the presence of EVs. These structures are not observed in the protein fractions of either sample types, which instead have significant dark staining, indicative of protein in EM imaging.
NTA reveals differences in concentrations in control and treated factions
Particle concentrations of control and tunicamycin treated EV and protein fractions (n = 3) were quantified with nanoparticle tracking analysis (NTA; Figure 7). Figure 7A shows particle concentrations for the control and tunicamycin treated EV fractions, with slightly more particles present in the tunicamycin treatment relative to control. Peak particle concentrations were between 105-165 nm. Figure 7B shows concentrations for particles detected in the protein fraction of both control and tunicamycin treated protein fractions. In the protein fraction, fewer particles overall were detected relative to EVs (109 vs. 1013), and peak concentrations were also smaller, between 75-135 nm. Interestingly, the tunicamycin protein fraction had more particles than the control. The majority of EV-sized particles (approximately 50-200 nm) are observed in the EV fraction of both control and tunicamycin treated cells further supporting the use of SEC as an effective means of EV separation from conditioned cell culture media.
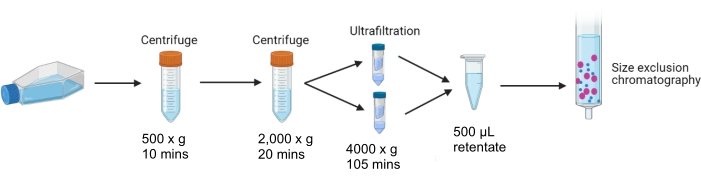
Figure 1: Differential centrifugation and ultrafiltration of conditioned cell culture media. Schematic outlining of the steps of differential centrifugation and ultrafiltration of media collected from cultured cells 24 h after control or 10 µg/mL tunicamycin treatment. Media is centrifuged and concentrated, leaving 500 µL of concentrated retentate suitable for SEC. Please click here to view a larger version of this figure.
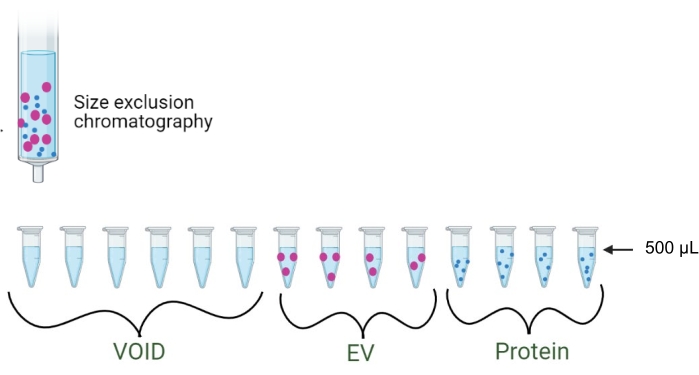
Figure 2: Size exclusion chromatography. Schematic of the separation of EVs and proteins via SEC. The first 3 mL (six fractions of 500 µL each) are discarded as the void volume. Fractions 1-4 elute as EVs, while fractions 5-8 elute as proteins. Please click here to view a larger version of this figure.
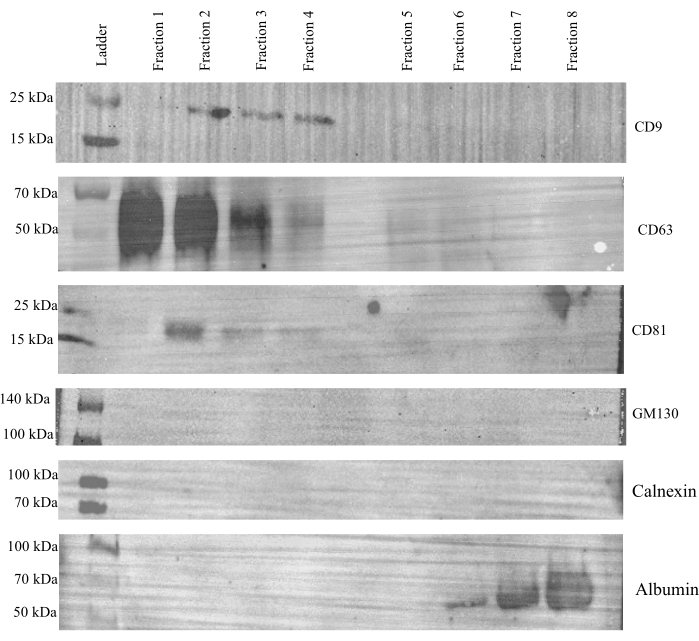
Figure 3: Representative western blot analysis of individual SEC fractions. A volume of 15 µL is used from each individual SEC fraction of the control cells and probed for CD9, CD63, CD81, GM130, calnexin, and albumin expression. CD9, CD63, and CD81 were most strongly observed in fractions 1-4 while GM130 and calnexin were not observed in any fraction. Albumin was only observed in fractions 6-8. Please click here to view a larger version of this figure.
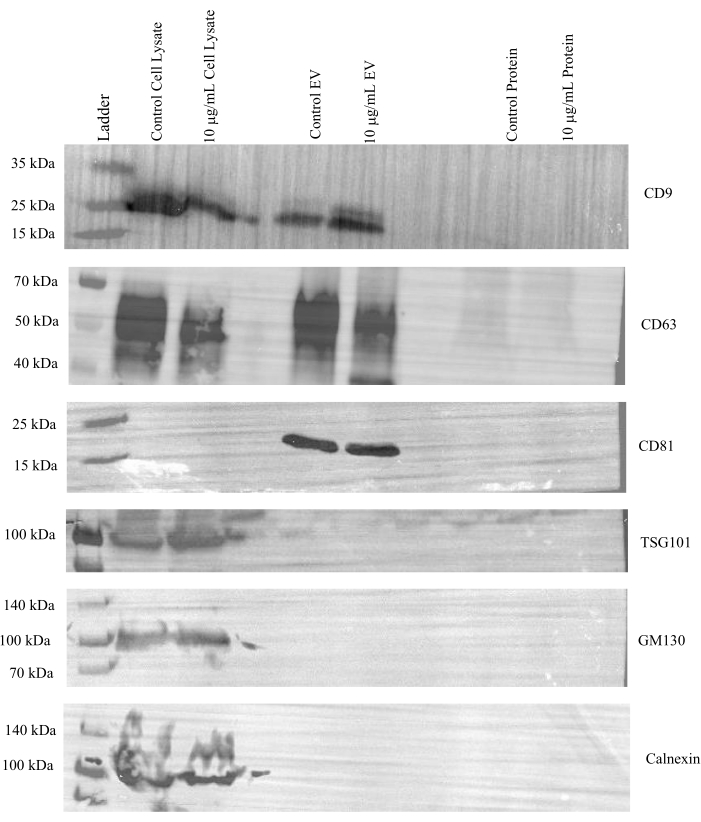
Figure 4: Representative western blot images for canonical EV makers. A final concentration of 20 µg of protein from cell lysates and 15 µg of protein from the pooled and concentrated SEC EV (1-4) and protein (5-8) fractions from control and 10 µg/mL tunicamycin treated cells were probed for CD9, CD63, CD81, TSG101, calnexin, and GM130. All markers were present in the cell lysates. CD9, CD63, and CD81 were observed in all the EV fractions, while TSG101 was not. The negative EV markers calnexin and GM130 were absent in the EV and protein fractions. Control cell lysate refers to lysates of cells that underwent control treatment, while 10 µg/mL cell lysate refers to lysates of cells that underwent tunicamycin treatment. Control EV indicates EVs from control samples. 10 µg/mL EV represents EVs from tunicamycin treated samples. Control protein implies extracellular proteins were from control samples, while 10 µg/mL protein signifies extracellular proteins from tunicamycin treated samples. Please click here to view a larger version of this figure.
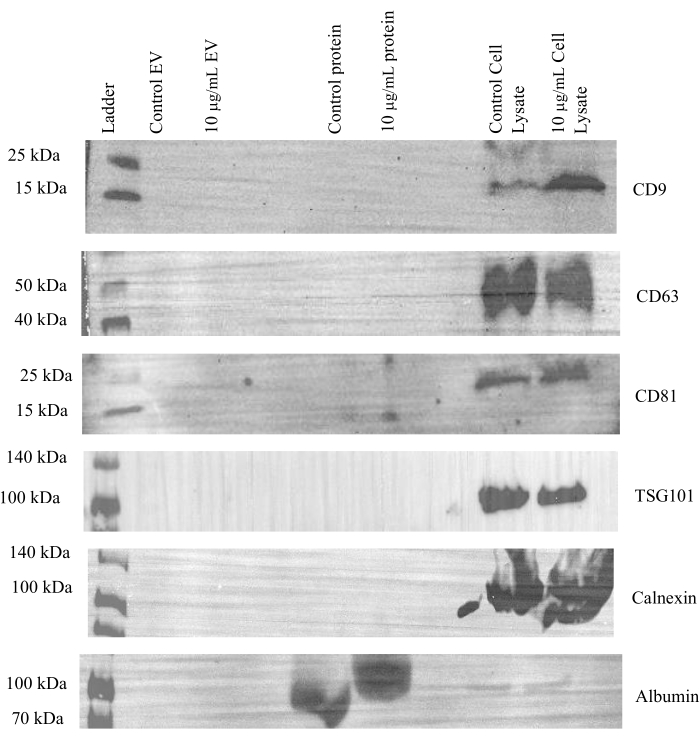
Figure 5: Representative western blot images of exosome-depleted cell culture media. Control and 10 µg/mL tunicamycin exosome-depleted cell culture media underwent SEC and ultrafiltration. A total volume of 15 µL of SEC EV and protein fractions, and 20 µg of protein from cell lysates were probed for CD9, CD63, CD81, TSG101, calnexin, and albumin. No EV markers were observed in the EV or protein fractions but were observed in the cell lysates used as positive controls. Albumin was observed in the protein fractions but not the EV fraction. Control EV indicates EVs from control media samples. 10 µg/mL EV represents EVs from tunicamycin treated samples. Control protein implies extracellular proteins were from control samples. 10 µg/mL protein signifies extracellular proteins from tunicamycin treated samples. Control cell lysate stands for cells in the control treatment that were lysed, while 10 µg/mL cell lysate means lysed cells came from tunicamycin treatment. Please click here to view a larger version of this figure.
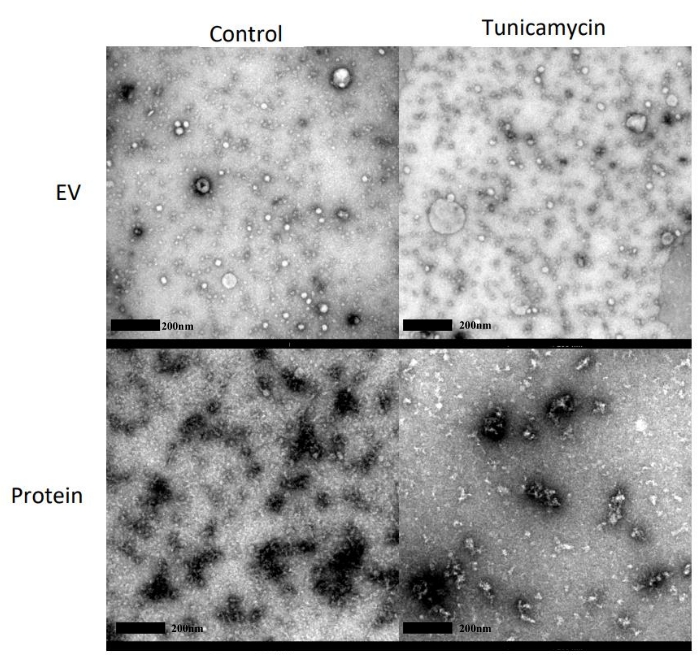
Figure 6: Representative TEM images for morphology of EVs. SEC fractions 1-4 and 5-8 were pooled and concentrated as the EV and protein fractions, respectively, from both control and 10 µg/mL tunicamycin treated cells. EVs are observed in the EV fractions of both the control and tunicamycin samples as small spherical particles less than 200 nm in diameter and are not observed in the protein samples. Scale bars = 200 nm. Please click here to view a larger version of this figure.
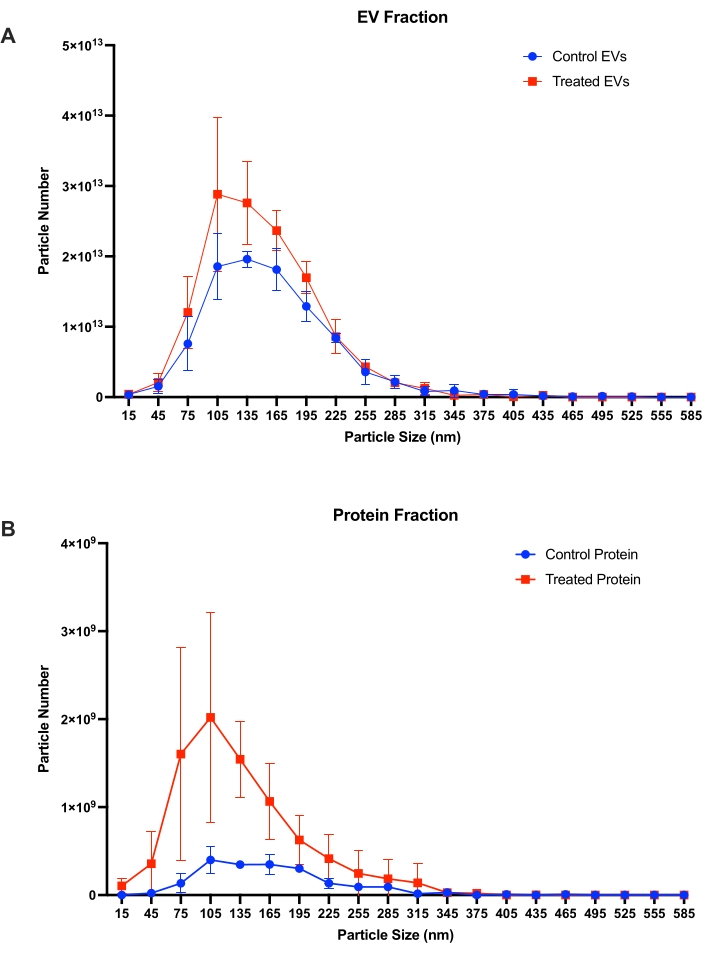
Figure 7: Particle size and concentrations from nanoparticle tracking analysis. Particle size and quantity in (A) EV fractions and (B) protein fractions from control and tunicamycin treated cells (n = 3). Control EVs indicates EV from control samples, while treated EVs represents EVs from tunicamycin treated samples. Control protein implies extracellular proteins from control samples, and treated protein signifies extracellular proteins from tunicamycin treated samples. Error bars are ± one standard deviation. Please click here to view a larger version of this figure.
Supplementary Figure 1: Stain-free image of PVDF membranes from Figure 3. Each PVDF membrane was imaged to visualize successful protein transfer at the conclusion of step 8.3.7. The individual SEC fractions (1-8) were visualized for blots that were later probed for (A) CD9, (B) CD63, (C) CD81, (D) GM130, (E), calnexin, and (F) albumin. Please click here to download this File.
Supplementary Figure 2: Stain-free image of PBDF membranes from Figure 4. Each PVDF membrane was imaged to visualize successful protein transfer at the conclusion of step 8.3.7. Blots were later probed for (A) CD9, (B) CD63, (C) CD81, (D) TSG101, (E) GM130, and (F) calnexin. Control cell lysate refers to cell lysates from control treated cells, while 10 µg/mL cell lysate refers to cell lysates from the tunicamycin treatment. Control EV indicates EVs from control samples and 10 µg/mL EV represents EVs from tunicamycin treated samples. Control protein refers to extracellular proteins from control samples, and 10 µg/mL protein signifies extracellular proteins from tunicamycin treated samples. Please click here to download this File.
Supplementary Figure 3: Stain-free image of PVDF membranes from Figure 5. Each PVDF membrane was imaged to visualize successful protein transfer at the conclusion of step 8.3.7. Blots were later probed for (A) CD9, (B) CD63, (C) CD81, (D) TSG101, (E) calnexin, and (F) albumin. Control EV indicates EVs from control media and 10 µg/mL EV represents EVs from tunicamycin treated media. Control protein refers to extracellular proteins from control media, and 10 µg/mL protein signifies extracellular proteins from tunicamycin treated media. Control cell lysate refers to cell lysates from control treated cells, while 10 µg/mL cell lysate refers to cell lysates from the tunicamycin treatment. Please click here to download this File.
Supplementary Figure 4: Uncropped western blot images used to create Figure 3. Uncropped composite chemiluminescent and colorimetric images of western blots probed for (A) CD9, (B) CD63, (C) CD81, (D) GM130, (E) calnexin, and (F) albumin for each individual SEC fraction (fractions 1-8). Please click here to download this File.
Supplementary Figure 5: Uncropped western blot images from Figure 4. Uncropped composite chemiluminescent and colorimetric images of western blots probed for (A) CD9, (B) CD63, (C) CD81, (D) TSG101, (E) GM130, and (F) calnexin. Control cell lysate refers to cell lysates from control treated cells, while 10 µg/mL cell lysate refers to cell lysates from the tunicamycin treatment. Control EV indicates EVs from control samples and 10 µg/mL EV represents EVs from tunicamycin treated samples. Control protein implies extracellular proteins were from control samples, while 10 µg/mL protein signifies extracellular proteins from tunicamycin treated samples. Please click here to download this File.
Supplementary Figure 6: Uncropped western blot images from Figure 5. Uncropped composite chemiluminescent and colorimetric images of western blots probed for (A) CD9, (B) CD63, (C) CD81, (D) TSG101, (E) calnexin, and (F) albumin. Control EV indicates EVs from control media while 10 µg/mL EV represents EVs from tunicamycin treated media. Control protein implies extracellular proteins were from control media and 10 µg/mL protein signifies extracellular proteins from tunicamycin treated media. Control cell lysate refers to cell lysates from control treated cells, while 10 µg/mL cell lysate refers to cell lysates from the tunicamycin treatment. Please click here to download this File.
| Antibody | Host Species | Dilution |
| CD9 | Mouse | 1 : 500 |
| CD63 | Mouse | 1 : 1000 |
| CD81 | Mouse | 1 : 500 |
| GM130 | Rabbit | 1 : 500 |
| Albumin | Rabbit | 1 : 1000 |
| TSG101* | Rabbit | 1 : 1000 |
| Calnexin* | Rabbit | 1 : 1000 |
| Anti-mouse^ | Horse | 1 : 1000 |
| Anti-rabbit^ | Goat | 1 : 1000 |
Table 1: Antibody dilutions. Dilutions used for antibodies. Stock antibodies were diluted in 5% milk in TBS-Tween. *Represents antibodies that require samples to be run under reducing conditions; ^ represents secondary antibodies.
Discussion
SEC is a user-friendly method for adequately separating EVs from conditioned CCM. In order to specifically isolate cell derived EVs, careful consideration of the type of CCM and its supplements must be taken into account. Many cell culture medias need to be supplemented with FBS, which contains EVs derived from the animal in which the serum was harvested. These serum EVs may saturate and mask any signal produced by EVs derived from cells in culture26. Therefore, when performing experiments, EV-depleted FBS should be used whenever possible to ensure that the EVs found can be attributed to cell derived, and not bovine derived, EVs This can either be purchased commercially or made in-house through various techniques reviewed elsewhere26,27.
When running samples through a SEC column, it is imperative that filtered PBS is used. Otherwise, each fraction will be contaminated by particles from the PBS instead of solely cell derived EVs and extracellular proteins. By filtering PBS through a 0.22 µm filter prior to its utilization for SEC, one can ensure that any particle in the fractions is from the cells themselves and not contaminated PBS.
The protocol outlined here can also be modified to alter the number and size of the collected fractions. For example, fractions could be made larger (1 mL vs. 500 µL), or more fractions could be collected as there are likely extracellular proteins that elute past what is deemed fraction 8 in this protocol. This is especially true for other sample types like blood plasma, or other column compositions28,29. These small extracellular proteins that potentially elute in the later fractions may contain important signaling molecules that are not collected or assayed with the current methodology. In addition, the use of the AFC to collect fractions alleviates any potential user errors that may be introduced through manual fraction collection. As the fractions elute droplet by droplet, great care must be taken to ensure that each droplet is collected into the appropriate fraction, which can be challenging when performing manually.
One major limitation of SEC is the limited volume of sample that can be run through the column. With the commercially available columns used in this protocol, 500 µL of sample is the maximum volume that can be used. Other commercially available columns or in-house made columns may accommodate other volumes of sample, however these are still relatively small volumes29,30. Other methods, like dUC for example, can accommodate larger sample volumes, however this technique cannot easily separate EVs from other extracellular components. Gradients of sucrose or iodixanol can be useful for separating particles based on density, but this technique is time consuming and requires steady hands and very strong pipetting skills31,32.
Overall, SEC is an important method for EV research that allows for sufficient separation of EVs from conditioned CCM. This is especially true for commercial columns as they allow for greater reproducibility between biological replicates, and fractionation can be automated, reducing the chances of user error. The protocol outlined here demonstrates that EVs elute predominately in early fractions (1-4), while later fractions contain extracellular proteins. These fractions can then be combined and undergo ultrafiltration to effectively concentrate the sample for downstream analyses including western blotting, TEM imaging, and NTA particle sizing and quantification.
Offenlegungen
The authors have nothing to disclose.
Acknowledgements
The authors would like to thank Penn State Behrend and the Hamot Health Foundation for funding, as well as the Penn State Microscopy Facility in University Park, PA.
Materials
| 2-Mercaptoethanol | VWR | 97064-588 | |
| 4X Laemmli Sample Buffer | BioRad | 1610747 | |
| Amicon Ultra-15 Centrifugal Filter Unit, Ultracel, 3 KDa, 15mL | Sigma-Aldrich | UFC900308 | 3 kDa cutoff |
| Amicon Ultra-2 Centrifugal Filter Unit with Ultracel-3 membrane | Sigma-Aldrich | UFC200324 | 3 kDa cutoff |
| Ammonium Persulfate | Sigma-Aldrich | A3678-100G | |
| Anti rabbit IgG, HRP linked Antibody | Cell Signaling Technology | 7074V | 1:1000 Dilution |
| Anti-Calnexin antibody | Abcam | ab22595 | 1:500 Dilution |
| Anti-CD9 Mouse Monoclonal Antibody | BioLegend | 312102 | 1:500 Dilution |
| Anti-GM130 antibody [EP892Y] – cis-Golgi Marker | Abcam | ab52649 | 1:500 Dilution |
| Anti-mouse IgG, HRP-linked Antibody | Cell Signaling Technology | 7076V | 1:1000 Dilution |
| Automatic Fraction Collector | Izon Science | ||
| BCA assay Kit | Bio-Rad | ||
| CCD camera | Gatan Orius SC200 | ||
| Cd63 Mouse anti Human | BD | 556019 | 1:1000 Dilution |
| CD81 Antibody | Santa Cruz Biotechnology | sc-23962 | 1:1000 Dilution |
| Cellstar Filter Cap Cell Culture Flasks | Greiner Bio-One | 660175 | |
| ChemiDoc MP Imager | BioRad | ||
| Clarity Western ECL Substrate | BioRad | 1705061 | |
| deoxycholate | Sigma-Aldrich | D6750-10G | |
| dithiothreitol | Sigma | 3483-12-3 | |
| DMEM/High glucose with L-glutamine; without sodium | Cytiva | SH300022.FS | |
| Fetal Bovine Serum Premium grade | VWR | 97068-085 | |
| Fetal Bovine Serum, exosome-depleted | Thermo Scientific | A2720801 | |
| Glycine | BioRad | 1610718 | |
| Great Value Nonfat Dry Milk | Amazon | B076NRD2TZ | |
| HOG Human Oligodendroglioma Cell Line | Sigma-Aldrich | SCC163 | |
| Izon Science Usa Ltd qev Size Exclusion Columns 5pk | Izon Science | ||
| Methanol >99.8% ACS | VWR | BDH1135-4LP | |
| Mini-PROTEAN Glass plates | BioRad | 1653310 | with 0.75mm spacers |
| Mini-PROTEAN Short plates | BioRad | 1653308 | |
| NP-40 | Sigma-Aldrich | 492016 | |
| Penicillin-Streptomycin,Solution | Sigma-Aldrich | P4458-100mL | |
| Phosphate Buffered Saline PBS | Fisher Scientific | BP66150 | |
| Pierce BCA Protein Assay Kits and Reagents | Thermo Fisher Scientific | 23227 | |
| Pierce PVDF Transfer Membranes | Thermo Scientific | 88518 | |
| Pierce Western Blotting Filter Paper | Thermo Scientific | 84783 | |
| Polyoxyethylene-20 (TWEEN 20), 500mL | Bio Basic | TB0560 | |
| Protease/phosphatase Inhibitor Cocktail (100X) | Cell Signaling Technology | 5872S | |
| Recombinant Anti-TSG101 antibody [EPR7130(B)] | ABCam | ab125011 | 1:1000 dilution |
| Slodium hydroxide | Sigma-Aldrich | SX0603 | |
| Sodium azide | Fisher Scientific | BP922I-500 | |
| Sodium Chloride | Sigma-Aldrich | S9888-500G | |
| Sodium dodecyl sulfate,≥99.0% (GC), dust-free pellets | Sigma-Aldrich | 75746-1KG | |
| Tetramethylethylenediamine | Sigma-Aldrich | T9281-25ML | |
| TGX Stain-Free FastCast Acrylamide Kit, 10% | BioRad | 1610183 | |
| Transmission Electron Microscope | FEI Tecnai 12 Biotwin | ||
| Tris | BioRad | 1610716 | |
| Trypsin 0.25% protease with porcine trypsin, HBSS, EDTA; without calcium, magnesium | Cytiva | SH30042.01 | |
| Tunicamycin | Tocris | 3516 | |
| Zeta View software | Analytik | NTA software |
Referenzen
- Pan, B. T., Teng, K., Wu, C., Adam, M., Johnstone, R. M. Electron microscopic evidence for externalization of the transferrin receptor in vesicular form in sheep reticulocytes. The Journal of Cell Biology. 101 (3), 942-948 (1985).
- Harding, C., Heuser, J., Stahl, P. Receptor-mediated endocytosis of transferrin and recycling of the transferrin receptor in rat reticulocytes. The Journal of Cell Biology. 97 (2), 329-339 (1983).
- Valadi, H., et al. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nature Cell Biology. 9 (6), 654-659 (2007).
- Raposo, G., Stoorvogel, W. Extracellular vesicles: Exosomes, microvesicles, and friends. The Journal of Cell Biology. 200 (4), 373-383 (2013).
- Rontogianni, S., et al. Proteomic profiling of extracellular vesicles allows for human breast cancer subtyping. Communications Biology. 2 (1), 1-13 (2019).
- Lane, R. E., Korbie, D., Hill, M. M., Trau, M. Extracellular vesicles as circulating cancer biomarkers: opportunities and challenges. Clinical and Translational Medicine. 7 (1), 14 (2018).
- Thompson, A. G., et al. Extracellular vesicles in neurodegenerative disease-pathogenesis to biomarkers. Nature Reviews Neurology. 12 (6), 346-357 (2016).
- O’Brien, K., Ughetto, S., Mahjoum, S., Nair, A. V., Breakefield, X. O. Uptake, functionality, and re-release of extracellular vesicle-encapsulated cargo. Cell Reports. 39 (2), 110651 (2022).
- Joshi, B. S., de Beer, M. A., Giepmans, B. N. G., Zuhorn, I. S. Endocytosis of extracellular vesicles and release of their cargo from endosomes. ACS Nano. 14 (4), 4444-4455 (2020).
- Théry, C., Amigorena, S., Raposo, G., Clayton, A. Isolation and characterization of exosomes from cell culture supernatants and biological fluids. Current Protocols in Cell Biology. 30 (1), 3-22 (2006).
- Willms, E., Cabañas, C., Mäger, I., Wood, M. J. A., Vader, P. Extracellular vesicle heterogeneity: Subpopulations, isolation techniques, and diverse functions in cancer progression. Frontiers in Immunology. 9, 738 (2018).
- Tzaridis, T., et al. Extracellular vesicle separation techniques impact results from human blood samples: Considerations for diagnostic applications. International Journal of Molecular Sciences. 22 (17), 9211 (2021).
- Liangsupree, T., Multia, E., Riekkola, M. L. Modern isolation and separation techniques for extracellular vesicles. Journal of Chromatography A. 1636, 461773 (2021).
- Gámez-Valero, A., et al. Size-exclusion chromatography-based isolation minimally alters extracellular vesicles’ characteristics compared to precipitating agents. Scientific Reports. 6 (1), 1-9 (2016).
- Mol, E. A., Goumans, M. J., Doevendans, P. A., Sluijter, J. P. G., Vader, P. Higher functionality of extracellular vesicles isolated using size-exclusion chromatography compared to ultracentrifugation. Nanomedicine: Nanotechnology, Biology, and Medicine. 13 (6), 2061-2065 (2017).
- Campos-Silva, C., et al. High sensitivity detection of extracellular vesicles immune-captured from urine by conventional flow cytometry. Scientific Reports. 9 (1), 2042 (2019).
- Norman, M., et al. L1CAM is not associated with extracellular vesicles in human cerebrospinal fluid or plasma. Nature methods. 18 (6), 631-634 (2021).
- Théry, C., et al. Minimal information for studies of extracellular vesicles 2018 (MISEV2018): a position statement of the International Society for Extracellular Vesicles and update of the MISEV2014 guidelines. Journal of Extracellular Vesicles. 7 (1), 1535750 (2018).
- Mathieu, M., et al. Specificities of exosome versus small ectosome secretion revealed by live intracellular tracking of CD63 and CD9. Nature Communications. 12 (1), 1-18 (2021).
- Guo, J., et al. Establishment of a simplified dichotomic size-exclusion chromatography for isolating extracellular vesicles toward clinical applications. Journal of Extracellular Vesicles. 10 (11), 12145 (2021).
- Benedikter, B. J., et al. Ultrafiltration combined with size exclusion chromatography efficiently isolates extracellular vesicles from cell culture media for compositional and functional studies. Scientific Reports. 7 (1), 1-13 (2017).
- Eslami, A., Lujan, J. Western blotting: sample preparation to detection. Journal of Visualized Experiments: JoVE. (44), e2359 (2010).
- Grassucci, R. A., Taylor, D. J., Frank, J. Preparation of macromolecular complexes for cryo-electron microscopy. Nature Protocols. 2 (12), 3239 (2007).
- Williams, R. L., Urbé, S. The emerging shape of the ESCRT machinery. Nature Reviews Molecular Cell Biology. 8 (5), 355-368 (2007).
- Willms, E., et al. Cells release subpopulations of exosomes with distinct molecular and biological properties. Scientific Reports. 6 (1), 1-12 (2016).
- Lehrich, B. M., Liang, Y., Fiandaca, M. S. Foetal bovine serum influence on in vitro extracellular vesicle analyses. Journal of Extracellular Vesicles. 10 (3), 12061 (2021).
- Kornilov, R., et al. Efficient ultrafiltration-based protocol to deplete extracellular vesicles from fetal bovine serum. Journal of Extracellular Vesicles. 7 (1), 1422674 (2018).
- Böing, A. N., et al. Single-step isolation of extracellular vesicles by size-exclusion chromatography. Journal of Extracellular Vesicles. 3 (1), (2014).
- Zhang, X., Borg, E. G. F., Liaci, A. M., Vos, H. R., Stoorvogel, W. A novel three step protocol to isolate extracellular vesicles from plasma or cell culture medium with both high yield and purity. Journal of Extracellular Vesicles. 9 (1), 1791450 (2020).
- Guerreiro, E. M., et al. Efficient extracellular vesicle isolation by combining cell media modifications, ultrafiltration, and size-exclusion chromatography. PLoS ONE. 13 (9), 02024276 (2018).
- Onódi, Z., et al. Isolation of high-purity extracellular vesicles by the combination of iodixanol density gradient ultracentrifugation and bind-elute chromatography from blood plasma. Frontiers in Physiology. 9, 1479 (2018).
- Yuana, Y., Levels, J., Grootemaat, A., Sturk, A., Nieuwland, R. Co-isolation of extracellular vesicles and high-density lipoproteins using density gradient ultracentrifugation. Journal of Extracellular Vesicles. 3 (1), (2014).

