Chromatin Immunoprecipitation Assay for Tissue-specific Genes using Early-stage Mouse Embryos
Summary
We demonstrate a chromatin immunoprecipitation (ChIP) method to identify factor interactions at tissue-specific genes during or after the onset of tissue-specific gene expression in mouse embryonic tissue. This protocol should be widely applicable for the study of tissue-specific gene activation as it occurs during normal embryonic development.
Abstract
Chromatin immunoprecipitation (ChIP) is a powerful tool to identify protein:chromatin interactions that occur in the context of living cells 1-3. This technique has been widely exploited in tissue culture cells, and to a lesser extent, in primary tissue. The application of ChIP to rodent embryonic tissue, especially at early times of development, is complicated by the limited amount of tissue and the heterogeneity of cell and tissue types in the embryo. Here we present a method to perform ChIP using a dissociated embryonic day 8.5 (E8.5) embryo. Sheared chromatin from a single E8.5 embryo can be divided into up to five aliquots, which allows the investigator sufficient material for controls and for investigation of specific protein:chromatin interactions.
We have utilized this technique to begin to document protein:chromatin interactions during the specification of tissue-specific gene expression programs. The heterogeneity of cell types in an embryo necessarily restricts the application of this technique because the result is the detection of protein:chromatin interactions without distinguishing whether the interactions occur in all, a subset of, or a single cell type(s). However, examination of tissue-specific genes during or following the onset of tissue-specific gene expression is feasible for two reasons. First, immunoprecipitation of tissue specific factors necessarily isolates chromatin from the cell type where the factor is expressed. Second, immunoprecipitation of coactivators and histones containing post-translational modifications that are associated with gene activation should only be found at genes and gene regulatory sequences in the cell type where the gene is being or has been activated. The technique should be applicable to the study of most tissue-specific gene activation events.
In the example described below, we utilized E8.5 and E9.5 mouse embryos to examine factor binding at a skeletal muscle specific gene promoter. Somites, which are the precursor tissues from which the skeletal muscles of the trunk and limbs will form, are present at E8.5-9.54,5. Myogenin is a regulatory factor required for skeletal muscle differentiation 6-9. The data demonstrate that myogenin is associated with its own promoter in E8.5 and E9.5 embryos. Because myogenin is only expressed in somites at this stage of development 6,10, the data indicate that myogenin interactions with its own promoter have already occurred in skeletal muscle precursor cells in E8.5 embryos.
Protocol
1. Isolation of Embryos
Note: All operations involving mice should be performed in accordance with the appropriate animal care and usage policies and protocols
- Check for the presence of a mating plug in the female mouse the morning after mating and separate the mated females from the stud males by placing them in a different cage. Noon of the day that the mating plug is observed is considered embryonic day 0.5 (E0.5) of development.
- At E8.5, (or the desired stage, if different), sacrifice the mouse using an approved protocol.
- Wet the belly of the euthanized animal with 70% ethanol and open the abdominal cavity. The implantation sites indicating the presence of developing embryos are seen as a “beads on a string” arrangement along the length of each uterine horn (Figure 1A). Using scissors, remove both uterine horns, cut each implantation site individually to separate (Figure 1B) and place them in a 100 mm petri dish containing room temperature (RT) dissection medium (DMEM, 10% fetal bovine serum, 20 mM HEPES pH 7.4, 60 μg/ml penicillin, 2 mM streptomycin).
- Using forceps, remove each embryo and the surrounding membranes from the uterine tissue (Figure 1C) and place them in fresh dissection medium.
- Isolate individual embryos under a dissecting microscope using forceps. At this stage in development, the embryos are surrounded by the parietal yolk sac, which contacts the deciduum tissue, the visceral yolk sac and the amnion. The amnion is a transparent membrane that is in direct contact with the embryo. The visceral yolk sac is located between the amnion and the parietal yolk sac and is readily distinguished in E8.5-E9.5 embryos by the presence of prominent blood vessels that nurture the embryo. Remove each embryo from its surrounding membranes and transfer individually into a new plate containing dissection media to remove excess blood. The developmental stage can vary from embryo to embryo and between litters; representative E8.5 embryos and a representative E9.5 embryo are shown in Figure 1D.
- Transfer each embryo to a 1.5 ml eppendorf tube containing 200 μl of dissection media (described in step 4 above).
2. Homogenization of the Isolated Embryo
- Add 20 units of Collagenase type II in a volume of 200 ul (diluted in 1X Dubbecco’s Phosphate Buffered Saline; DPBS) to each eppendorf tube.
- Gently shake samples in a 37°C shaker at 100 rpm for 20 min.
- Resuspend well by pipetting to disrupt clumps of cells.
- Apply the cell suspension on the top of sterile cell strainer (40 μm mesh size) placed over a 1.5 ml eppendorf tube.
- Immediately apply 600 μl of room temperature 1X DPBS on top of the cell strainer to complete the separation. Discard the strainer.
- Centrifuge samples at 4°C at 4000 x g for 5 min.
- Discard the supernatant.
- Resuspend the pellet in 1 ml of RT 1X DPBS.
- Centrifuge the samples at 4000 x g for 1 min at RT.
- Discard the supernatant.
- Resuspend the pellet in 200 μl of RT dissection media.
- Count the embryonic cells. There is an average 3-5 x 106 cells/E8.5 embryo.
3. Cross-link Chromatin
- Add 5.6 μl of 37% formaldehyde to the 200 μl samples for a final concentration of 1% formaldehyde.
- Incubate samples for 10 min at RT.
- Centrifuge the samples at 4°C at 4000 x g for 3 min.
- Discard the supernatant.
- Resuspend the cells with cold 1X DPBS containing 80 μl/ml protease inhibitor cocktail (PIC).
- Centrifuge samples at 4°C at 4000 x g for 3 min.
- Discard the supernatant. At this step, the samples can be frozen at -80°C. The frozen, crosslinked samples are stable for several months.
4. Sonication
- Resuspend the cell pellet in 100 μl of RT SDS lysis buffer (50 mM Tris-HCl (pH 8.1), 10 mM EDTA and 1% SDS with freshly added PIC at 1 μl PIC/800 μl total volume). When resuspended, add an additional 100 ul of RT SDS lysis buffer to promote resuspension. Mix well by pipetting and inversion.
- Incubate the cell suspension on ice for 10 min.
- Sonicate lysed cells to shear DNA to between 200 and 500 basepairs using a Diagenode Bioruptor™ UCD200 (5 min x 8 times: amplitude setting of 30 sec on/30 sec off on high (320 watts) power). The samples should be surrounded by an ice slurry. Do not allow the samples to warm above 4°C or to freeze. There are many types of sonicators available. Sonication conditions should be optimized for each sonicator to result in shearing of the cross-linked DNA to a length of approximately 500 bp.
- Centrifuge sonicated samples at 4°C at 14000 x g for 10 min.
- Transfer the supernatant into a fresh eppendorf tube pre-cooled on ice.
- Divide the sonicated sample into a maximum of 5 aliquots. We have empirically determined that one E8.5 embryo can be divided into 5 aliquots. Smaller or larger sample sizes can be scaled accordingly. At this step, the sample can be stored at -80°C until further use.
- Reserve one aliquot for use as input and store at -20°C until step 6 when the cross-links will be reversed. Dilute the other aliquots 10-fold in ChIP dilution buffer (16.7 mM Tris-HCl (pH 8.1), 1.2 mM EDTA, 1.1% Triton X-100, 167 mM NaCl and 0.01% SDS) containing freshly added PIC at 1 μl/800 μl buffer.
5. Pre-clearing and Immunoprecipitation
- Pre-clear the diluted supernatant with 75 μl of Salmon-Sperm DNA/Protein A or G Agarose-50% slurry at 4°C, with rotation, for 1 hr. Use of either Protein A or Protein G beads should be determined by the antibody to be used for the ChIP. For example, protein A beads are recommended for rabbit antibodies, whereas protein G beads are recommended for goat antibodies or mouse IgG1 antibodies. For more detailed information, see the recommendations made by the bead manufacturer.
- Centrifuge at 4°C at 2500 x g for 1 min.
- Transfer the supernatant to a new, pre-cooled eppendorf tube.
- Add antibody (4 μg/sample) to the pre-cleared aliquot and incubate overnight at 4°C with rotation.
6. Washing the Chromatin-protein-bead Complex
- Add 60 μl of Salmon Sperm DNA/Protein A or G agarose slurry into each vial and incubate at 4°C with rotation for 1 hr. Use the same bead type used for pre-clearing in step 3 above.
- Centrifuge at 4°C at 1000 x g for 1 min.
- Carefully remove the supernatant and discard. Keep the DNA-protein-bead complex on ice.
- Resuspend and wash the pellet as directed below with wash buffers 1, 2, 3, and 4. Wash buffers 1-3 should be chilled to 4°C and washes with these buffers should be performed at 4°C. Wash buffer 4 should be at room temperature and washes with this buffer should be performed at room temperature. Each wash is for 5 minutes with rotation at the appropriate temperature. Following each wash, centrifuge at 4°C (room temperature for wash buffer 4 washes) at 1000 x g for 1 min, then carefully aspirate or remove the supernatant.
Buffer 1:Low salt wash buffer (20 mM Tris-HCl (pH 8.1), 2 mM EDTA, 1% Triton X-100,
150 mM NaCl and 0.1% SDS), 1 ml x 2 washes.
Buffer 2:High salt wash buffer (20 mM Tris-HCl (pH 8.1), 2 mM EDTA, 1% Triton X-100,
500 mM NaCl and 0.1% SDS), 1 ml x 2 washes.
Buffer 3:LiCl salt wash buffer (10 mM Tris-HCl (pH 8.1), 1 mM EDTA, 1% IGEPAL-CA630,
0.25 M LiCl and 1% deoxycholic acid (sodium salt)), 1 ml x 1 wash.
Buffer 4:TE buffer (10 mM Tris-HCl (pH 8.1), 1 mM EDTA), 1 ml x 2 washes.
7. Elution of the Chromatin-antibody Complex
- Resuspend the washed chromatin-protein-bead complex in 250 μl of freshly prepared elution buffer (1% SDS, 0.1 M NaHCO3). Vigorously vortex the sample for 5 sec, then incubate at RT for 15 min with rotation.
- Centrifuge at 2500 x g for 1 min at RT and transfer the eluate into a new eppendorf tube.
- Repeat steps 1 and 2 and combine the eluates in the same eppendorf tube. The eluates should be kept at room temperature. Upon completion of this step, the eluates can be stored at -20°C until further use.
8. Reverse Cross-link and Recover DNA
- Add 20 μl 5 M NaCl for 500 μl eluate (40 μl/ml for input control).
- Heat the eluates at 65°C in a water bath for 4 hr to overnight.
- 10 μl of each sample (including the input control) can be run on a 2% agarose gel next to size markers spanning 0.1 to 1 kbp to check the DNA fragment size. The DNA will appear as a smear and not a sharp band. The rest of the samples can be left at 65°C while the gel is running.
- Recover the DNA from each sample using a QIAquick Gel Extraction Kit (Qiagen). Add 600 μl QG buffer (supplied in the kit) to optimize DNA adsorption with the silica membrane in a QIAquick spin column and 200 μl isopropanol into a 500 μl sample. Note that QG buffer contains a pH indicator allowing easy determination of the pH of the sample. If the color of the sample turns from yellow to purple (pH > 7.5) after mixing, add 10 μl of 3 M NaAcetate (pH 5.2) to the sample and mix. This reduces the pH of the mixture to maximize DNA binding to the beads in the column.
- Prior to loading the column, check the mixture to determine whether any SDS precipitate is present. If SDS precipitation has occurred, the sample should be warmed in a 42°C water bath until the precipitate disappears.
- Load 700 μl of the mixture into the QIAquick spin column (purple column in the kit) and centrifuge at RT at 14000 x g for 1 min. Discard the flow through from the column before proceeding and repeat this step with the remainder of the sample.
- Add 500 μl QG buffer to wash the column. Centrifuge at RT at 14000 x g for 2 min. Discard the flow through from the column before proceeding.
- Wash the column with 700 μl of PE wash buffer (supplied in the kit) to which ethanol has been added as instructed by the manufacturer. Centrifuge at RT at 14000 x g for 1 min. Discard the flow through from the column before proceeding.
- Repeat the centrifugation step to completely remove the remainder of the PE buffer from the column. Discard the flow through and the collection tube.
- Elute DNA from the column into a fresh eppendorf tube by adding 60 μl EB buffer (elution buffer supplied in the kit).
- Centrifuge at room temperature at 14000 x g for 1 min. The eluted DNA can be stored at -20°C until detection. A260 readings of the recovered material typically indicate concentrations of 90-120 ng/ul, for a total recovery of ~6-7 ug per aliquot. However, the A260/280 ratio of these samples is typically >1.8, suggesting that RNA is present in the sample. Thus the exact amount of recovered DNA may be lower.
9. Analysis of Recovered DNA
- SDS can interfere with the activity of PCR Taq polymerase. SDS should be removed completely before performing PCR. Incubate the DNA on ice to precipitate any residual SDS and centrifuge at 4°C at 14000 x g for 1 min. Transfer the supernatant to a new eppendorf tube.This step is strongly recommended.
- The DNA eluates can be detected either by conventional PCR or by quantitative real-time PCR (Q-PCR) with primers specific for each sequence to be analyzed. 5-10 μl from total DNA eluate may be used for one PCR or Q-PCR reaction. PCR with the input control can be performed with a 5-10 fold dilution of the DNA compared with the amount of other samples. PCR and Q-PCR conditions will vary, however, the limited amount of starting material requires additional PCR cycles. For conventional PCR detection, we recommend starting with 40 cycles and adjusting as needed.
10. Representative Results
We have used this protocol to perform ChIP from both E8.5 and E9.5 embryos (Figure 2). The results demonstrate that myogenin is present on the myogenin promoter in E8.5 and E9.5 embryos. ChIP purified DNAs were analyzed by conventional PCR (Figure 2A) and by quantitative real-time PCR (Figure 2B). In contrast, there was no indication of myogenin binding to the myogenin promoter in the yolk sac, where myogenin is not expressed. The interferon-γ (IFNγ) promoter, which contains sequences matching the myogenin binding site, was used as a negative sequence control. As expected, myogenin was not bound to the IFNγ promoter in any of the tissue samples tested. In E8.5 and 9.5 embryos, myogenin is specifically expressed in the somites (Figure 3; 6,10), which are the precursors to skeletal muscle. Thus the results indicate that myogenin is bound to the myogenin promoter in the somites at E8.5 and E9.5.
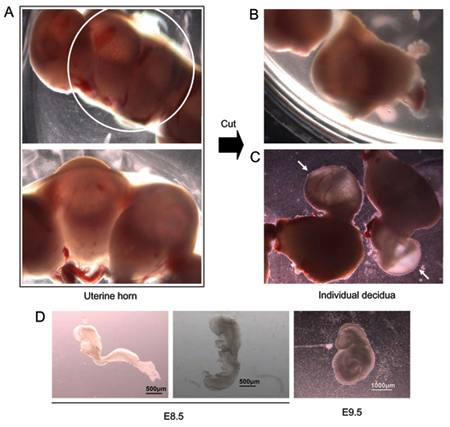
Figure 1. E8.5 embryo dissection. (A) Isolated uterine horn (top panel; higher magnification shown in lower panel). (B) Uterine horn cut with scissors to separate individual implantation sites containing individual embryos. (C) Embryos protruding from uterine tissue during dissection. Arrows mark embryos still covered by extra-embryonic tissue. (D) Two E8.5 embryos from the same litter. The embryo on the left has not begun the process of turning. The embryo on the right is undergoing turning. Far right – representative E9.5 embryo.
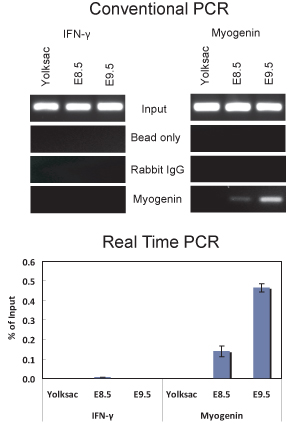
Figure 2. ChIP assay demonstrating myogenin binding to the myogenin promoter in E8.5 and E9.5 embryos. (Top) Conventional PCR analysis of 5 ul of DNA purified from ChIP experiments using a myogenin antibody or non-specific IgG was performed with primers that amplify a portion of the myogenin promoter from -79 to +69 relative to the start site of transcription that contains a myogenin binding site located at -12 or primers that amplify a portion of the IFNγ promoter that contains a sequence matching the myogenin binding site located ~ 1075 bp upstream of the transcription start site. The IFNγ primer sequences used were 5′-GCT GAC TCA AGA CCC CGA GGC-3′ and 5′-TGA GGA TGG GGC AGG AGG CC-3′. (Bottom) Quantitative Real-time PCR analysis of the same samples used in (A). The data are plotted as % of input +/- standard deviation.
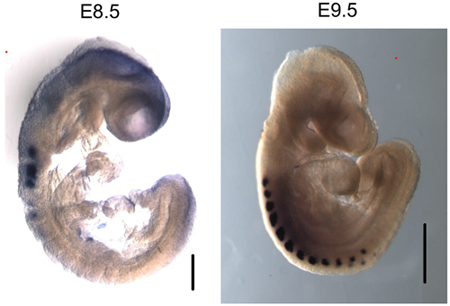
Figure 3. Myogenin is specifically expressed in the somites of E8.5 and E9.5 embryos. Whole mount in situ hybridization of myogenin shows specific mRNA expression in the somites. Size bar in E8.5 image – 200 μm. Size bar in E9.5 image – 500 μm.
Discussion
In the described ChIP protocol, we show that the myogenic regulator myogenin is associated with the myogenin promoter in skeletal muscle precursor tissue present in single E8.5 and E9.5 embryos. Prior studies have extensively characterized myogenin binding to E box containing sequences, beginning with the initial in vitro gel shift experiments utilizing in vitro translated or bacterially produced myogenin and radiolabeled DNA encoding the relevant portion of target gene regulatory sequences 11-20. Conventional ChIP studies have demonstrated myogenin binding to the myogenin promoter in tissue culture models for myogenesis 21-24. However, there is no evidence that demonstrates that myogenin binds to the myogenin promoter during embryonic skeletal muscle development, though one might predict this to be the case later in embryogenesis based on the down regulation of myogenin expression observed in E15.5 tongue tissue from myogenin deficient mice 25. Thus the data provides evidence that the interaction between myogenin and the myogenin promoter interaction occurs in vivo. An obvious application of this technique is to use it to verify the physiological relevance of prior studies characterizing tissue-specific gene activation tissue performed using culture models. The more interesting application is to use this technique as part of the initial characterization of a new or previously uncharacterized factor to determine the physiological relevance of a specific interaction prior to performing tissue culture studies designed at understanding functional mechanisms.The protocol can also be used to directly compare protein:chromatin interactions occurring at specific developmental stages in mouse models containing an engineered genetic manipulation.
We anticipate that this method will also be useful for time course studies of tissue-specific gene regulation during development. By assessing specific factor:chromatin interactions at different embryonic stages, one could identify the time point at which the factor interaction first occurs, could determine whether the interaction was maintained throughout and beyond the differentiation process or was more transient in nature, and could determine the order of factor binding if multiple factors interact with a specific sequence. Finally, we envision that the protocol could be extended to a re-ChIP procedure 26 whereby the chromatin recovered from one immunoprecipitation is subjected to a second immunoprecipitation with an antibody against a different factor believed to be co-occupying the same piece of chromatin. The re-ChIP protocol provides more conclusive evidence that two distinct factors are present on the same pieces of chromatin; the likely requirement for this modification would be an increase in the amount of starting material.
A remarkable realization that came from our efforts was that the limited amount of starting material was not the impediment to success that one might have predicted, especially given that tissue culture cell based protocols often suggest starting with a homogeneous population of millions or tens of millions of cells 26,27. The success of our efforts speaks to the primary (and often uncontrollable) requirement for antibodies capable of specific immunoprecipitation of the factor under investigation. Once an antibody is identified as capable of immunoprecipitation, the investigator can optimize the ChIP assay by titrating the amount of antibody used. It is important to note that more is not always better; we frequently find that too much antibody, especially with samples of limited quantity, results in higher background binding to irrelevant sequences. In addition, use of immunoglobulin fractions or affinity-purified antibodies often result in lower background than does use of antisera. Background from the control antibody can often be gauged by including a sample containing beads without any antibody.
We conclude that using early stage embryos for ChIP analysis of protein:chromatin interactions that occur during the activation of tissue-specific genes has the potential to provide significant insight into the induction and maintenance of tissue-specific gene expression programs directly within the context of development.
Divulgaciones
The authors have nothing to disclose.
Acknowledgements
This work was supported by NIH R01 GM56244 to ANI, which includes funds awarded through the American Recovery and Reinvestment Act of 2009, and by NIH R01 GM87130 to JARP
Materials
| Material Name | Tipo | Company | Catalogue Number | Comment |
|---|---|---|---|---|
| ChIP Assay Kit | Upstate Cell Signaling Solutions, Millipore | 17-295 | ||
| Collagenase Type II | Invitrogen | 17101015 | Dilution by 1 x PBS | |
| Dulbecco’s modified eagle medium (DMEM) | Gibco Labs, Invitrogen | 12100-061 | High glucose content | |
| Dulbecco’s phosphate buffered saline 1X (DPBS) | Gibco Labs, Invitrogen | 14190-144 | Calcium chloride free, Magnesium chloride free | |
| Fetal bovine serum (FBS) | Mediatech, Inc. | 35-010-CV | ||
| Gel extraction kit | QIAquick | 28704 | 50 reaction kit | |
| Penicillin/streptomycin stock solution | Gibco Labs, Invitrogen | 5000 μg/ml concentration | ||
| Protease Inhibitor Cocktail | Sigma-Aldrich | P8340 | ||
| Salmon sperm DNA /Protein A agarose | Millipore | 16-157 | ||
| myogenin antibody | Santa Cruz Biotechnology, Inc. | sc-576 | ||
| Normal rabbit IgG | Millipore | 12-370 | ||
| Platinum PCR Supermix | Invitrogen | 11306-016 | ||
| GoTaq Q-PCR master mix | Promega | A6001 |
Referencias
- Minard, M. E., Jain, A. K., Barton, M. C. Analysis of epigenetic alterations to chromatin during development. Genesis. 47, 559-572 (2009).
- Kuo, M. H., Allis, C. D. In vivo cross-linking and immunoprecipitation for studying dynamic Protein:DNA associations in a chromatin environment. Methods. 19, 425-433 (1999).
- Johnson, K. D., Bresnick, E. H. Dissecting long-range transcriptional mechanisms by chromatin immunoprecipitation. Methods. 26, 27-36 (2002).
- Yusuf, F., Brand-Saberi, B. The eventful somite: patterning, fate determination and cell division in the somite. Anat Embryol (Berl). 211, 21-30 (2006).
- Buckingham, M., Bajard, L., Chang, T., Daubas, P., Hadchouel, J., Meilhac, S., Montarras, D., Rocancourt, D., Relaix, F. The formation of skeletal muscle: from somite to limb. J Anat. 202, 59-68 (2003).
- Wright, W. E., Sassoon, D. A., Lin, V. K. Myogenin, a factor regulating myogenesis, has a domain homologous to MyoD. Cell. 56, 607-617 (1989).
- Edmondson, D. G., Olson, E. N. A gene with homology to the myc similarity region of MyoD1 is expressed during myogenesis and is sufficient to activate the muscle differentiation program. Genes Dev. 3, 628-640 (1989).
- Nabeshima, Y., Hanaoka, K., Hayasaka, M., Esumi, E., Li, S., Nonaka, I. Myogenin gene disruption results in perinatal lethality because of severe muscle defect. Nature. 364, 532-535 (1993).
- Hasty, P., Bradley, A., Morris, J. H., Edmondson, D. G., Venuti, J. M., Olson, E. N., Klein, W. H. Muscle deficiency and neonatal death in mice with a targeted mutation in the myogenin gene. Nature. 364, 501-506 (1993).
- Sassoon, D., Lyons, G., Wright, W. E., Lin, V., Lassar, A., Weintraub, H., Buckingham, M. Expression of two myogenic regulatory factors myogenin and MyoD1 during mouse embryogenesis. Nature. 341, 303-307 (1989).
- Brennan, T. J., Olson, E. N. Myogenin resides in the nucleus and acquires high affinity for a conserved enhancer element on heterodimerization. Genes Dev. 4, 582-595 (1990).
- Rosenthal, N., Berglund, E. B., Wentworth, B. M., Donoghue, M., Winter, B., Bober, E., Braun, T., Arnold, H. H. A highly conserved enhancer downstream of the human MLC1/3 locus is a target for multiple myogenic determination factors. Nucleic Acids Res. 18, 6239-6246 (1990).
- Braun, T., Gearing, K., Wright, W. E., Arnold, H. H. Baculovirus-expressed myogenic determination factors require E12 complex formation for binding to the myosin-light-chain enhancer. Eur J Biochem. 198, 187-193 (1991).
- Chakraborty, T., Brennan, T., Olson, E. Differential trans-activation of a muscle-specific enhancer by myogenic helix-loop-helix proteins is separable from DNA binding. J Biol Chem. 266, 2878-2882 (1991).
- French, B. A., Chow, K. L., Olson, E. N., Schwartz, R. J. Heterodimers of myogenic helix-loop-helix regulatory factors and E12 bind a complex element governing myogenic induction of the avian cardiac alpha-actin promoter. Mol Cell Biol. 11, 2439-2450 (1991).
- Brennan, T. J., Chakraborty, T., Olson, E. N. Mutagenesis of the myogenin basic region identifies an ancient protein motif critical for activation of myogenesis. Proc Natl Acad Sci U S A. 88, 5675-5679 (1991).
- Lassar, A. B., Davis, R. L., Wright, W. E., Kadesch, T., Murre, C., Voronova, A., Baltimore, D., Weintraub, H. Functional activity of myogenic HLH proteins requires hetero-oligomerization with E12/E47-like proteins in vivo. Cell. 66, 305-315 (1991).
- Chakraborty, T., Brennan, T. J., Li, L., Edmondson, D., Olson, E. N. Inefficient homooligomerization contributes to the dependence of myogenin on E2A products for efficient DNA binding. Mol Cell Biol. 11, 3633-3641 (1991).
- Cserjesi, P., Olson, E. N. Myogenin induces the myocyte-specific enhancer binding factor MEF-2 independently of other muscle-specific gene products. Mol Cell Biol. 11, 4854-4862 (1991).
- Braun, T., Arnold, H. H. The four human muscle regulatory helix-loop-helix proteins Myf3-Myf6 exhibit similar hetero-dimerization and DNA binding properties. Nucleic Acids Res. 19, 5645-5651 (1991).
- Serna, d. e. l. a., L, I., Ohkawa, Y., Berkes, C. A., Bergstrom, D. A., Dacwag, C. S., Tapscott, S. J., Imbalzano, A. N. MyoD targets chromatin remodeling complexes to the myogenin locus prior to forming a stable DNA-bound complex. Mol Cell Biol. 25, 3997-4009 (2005).
- Blais, A., Tsikitis, M., Acosta-Alvear, D., Sharan, R., Kluger, Y., Dynlacht, B. D. An initial blueprint for myogenic differentiation. Genes Dev. 19, 553-569 (2005).
- Cao, Y., Kumar, R. M., Penn, B. H., Berkes, C. A., Kooperberg, C., Boyer, L. A., Young, R. A., Tapscott, S. J. Global and gene-specific analyses show distinct roles for Myod and Myog at a common set of promoters. EMBO J. 25, 502-511 (2006).
- Ohkawa, Y., Yoshimura, S., Higashi, C., Marfella, C. G., Dacwag, C. S., Tachibana, T., Imbalzano, A. N. Myogenin and the SWI/SNF ATPase Brg1 maintain myogenic gene expression at different stages of skeletal myogenesis. J Biol Chem. 282, 6564-6570 (2007).
- Davie, J. K., Cho, J. H., Meadows, E., Flynn, J. M., Knapp, J. R., Klein, W. H. Target gene selectivity of the myogenic basic helix-loop-helix transcription factor myogenin in embryonic muscle. Dev Biol. 311, 650-664 (2007).
- Metivier, R., Penot, G., Hubner, M. R., Reid, G., Brand, H., Kos, M., Gannon, F. Estrogen receptor-alpha directs ordered, cyclical, and combinatorial recruitment of cofactors on a natural target promoter. Cell. 115, 751-763 (2003).
- Ausubel, F. M., Brent, R., Kingston, R. E., Moore, D. D., Seidman, J. G., Smith, J. A., Struhl, K. . Current Protocols in Molecular Biology. , (2010).

