Rapid, Enzymatic Methods for Amplification of Minimal, Linear Templates for Protein Prototyping using Cell-Free Systems
Summary
The study describes a protocol for creating large (µg-mg) quantities of DNA for protein screening campaigns from synthetic gene fragments without cloning or using living cells. The minimal template is enzymatically digested and circularized and then amplified using isothermal rolling circle amplification. Cell-free expression reactions could be performed with the unpurified product.
Abstract
This protocol describes the design of a minimal DNA template and the steps for enzymatic amplification, enabling rapid prototyping of assayable proteins in less than 24 h using cell-free expression. After receiving DNA from a vendor, the gene fragment is PCR-amplified, cut, circularized, and cryo-banked. A small amount of the banked DNA is then diluted and amplified significantly (up to 106x) using isothermal rolling circle amplification (RCA). RCA can yield microgram quantities of the minimal expression template from picogram levels of starting material (mg levels if all starting synthetic fragment is used). In this work, a starting amount of 20 pg resulted in 4 µg of the final product. The resulting RCA product (concatemer of the minimal template) can be added directly to a cell-free reaction with no purification steps. Due to this method being entirely PCR-based, it may enable future high-throughput screening efforts when coupled with automated liquid handling systems.
Introduction
Cell-free gene expression (CFE) has emerged as a powerful tool with many applications. Such applications include disease detection1,2,3,4,5,6, micronutrient and small molecule detection7,8,9,10,11,12, biomanufacturing13,14,15,16,17,18, education19,20,21, manufacturing difficult proteins17,22,23,24,25,26,27, and variant screening23,28,29,30,31,32,33. This is due to the open nature of cell-free systems and the flexibility they confer. Many great review articles offer historical education and future perspectives on the technology34,35,36,37,38,39,40,41,42,43,44.
A typical cell-free reaction consists of three major components: cell extract, energy mix, and genetic template. Active cell extract contains all the necessary machinery for transcription and translation (TXTL) and can be processed in a variety of ways36. Glycolytic intermediates, electrolytes, amino acids, and cofactors in the energy mix support the TXTL process. It is a major source of variability in cell-free experiments45 and can be prepared in many ways34,46. Preparation of the genetic template has seen fewer improvements since traditional cloning methods result in plasmids with excellent expression characteristics. The downside to these traditional methods is the turnaround time and amount of biological expertise needed to construct and propagate them. Recent optimization efforts have resulted in simple 24-hour workflows for cell extract preparation47,48 that can be performed in parallel with energy mix preparation49,50. However, traditional cloning adds multiple days to the CFE prototyping timeline (Table 1)23. Quickly amplified PCR products from the commercial gene fragment can be used directly51, but this limits the number of prototyping experiments as only 1 µg of DNA is produced, which corresponds to approximately five reactions (traditional 15 µL volumes). With these additional steps of circularization and isothermal amplification, greater than milligram quantities of the DNA is possible (~5,000 reactions for 1 mg). This dramatically increases the number of tests that can be made in high-throughput screening of proteins or combinatorial enzyme networks (cell-free metabolic engineering); it also allows for effective preservation of the linear template library as high concentration DNA. Furthermore, an increased amount of template would be necessary to prototype larger quantities of protein needed for material science applications (protein-based fibers and hydrogels). Some limitations of linear templates can be overcome by using an extract from BL21 DE3 Star or using recently discovered methods to protect linear templates from degradation52,53,54. However, this does not address having limited stocks of vendor-produced DNA for PCR amplification or the issue of biological expertise and equipment needed for cloning.
This work presents a protocol explicitly designed to increase the amount of expression template that can be obtained from small quantities of vendor-produced gene fragments (typically 500-1000 ng of lyophilized powder). The described method does not require the skills necessary to perform traditional cloning in plasmids or transforming and propagating in living cells. Upon receiving a gene fragment in the mail, a user can produce enough templates for many cell-free reactions by employing isothermal rolling circle amplification (RCA) (Figure 1)23. While the amount of DNA received from the vendor may be enough for limited screening efforts, it is quickly depleted, and re-purchasing gene fragments is time consuming and costly. The method is also especially well-suited for genes that are toxic and difficult to clone in E. coli.
Protocol
1. Designing the gene fragment
NOTE: The gene fragment should have all the necessary genetic elements for transcription/translation, including promoter, ribosome binding site (RBS), start codon, the gene of interest, and terminator. While the terminator is not necessary for a linear expression template (LET), it will be important if the user decides to insert the sequence into a plasmid. These sequences were lifted from the pJL1-sfGFP plasmid55 (gift from Michael Jewett's lab), which uses a T7 promoter. In addition to these necessary genetic elements, a restriction enzyme cut site is added six base pairs before the promoter (5' cut site) and another six base pairs after the terminator (3' cut site), in this case using HindIII (other restriction enzymes can be used, but it is helpful to standardize the sequences with one high fidelity restriction enzyme to reduce the number needed to keep in the library). Primer sites are added ten base pairs upstream of the 5' cut site and ten base pairs downstream of the 3' cut site, in this case using standardized M13 primer sequences (primers are inexpensive stock items). The restriction enzyme site and primers used are at the discretion of the user. However, the user must ensure the sequences are not present anywhere else in the template (do not want to create unwanted cuts or sites of amplification initiation). The sequences for the templates used in this work are detailed in the supplemental material. These steps are used to modify from this base template.
- Determine the desired gene to be expressed and obtain the amino acid sequence or the genetic sequence if it has been expressed in E. coli.
- If it is an amino acid sequence, perform codon optimization for E. coli using one of many standard vendor tools56. If using the template provided in the supplement, ensure the optimized sequence has no HindIII restriction sites (AAGCTT). In the case that it does, continue to optimize the sequence until there is no longer a HindIII site.
- Copy the sequence and paste it into the provided template for Supplementary Sequence #1 where the gene of interest is indicated. If expressing sfGFP, use Supplementary Sequence #1 as is. If expressing subtilisin, use Supplementary Sequence #2 as is.
- Order the minimal template and the necessary primers from the preferred DNA synthesis service.
2. Resuspending the gene fragment and the primers
NOTE: Upon receipt of the gene fragment, follow the manufacturer's protocols for resuspension or use this simple guide to create a DNA stock.
- Centrifuge the tube (300 x g for 5 s) to collect the DNA pellet at the bottom.
- Add double distilled water (ddH2O) to make a final concentration of 10 ng/µL of DNA template.
- Vortex the solution on a medium setting for 5-10 s.
- Dissolve the entire pellet by incubating at 50 °C for 20 min.
- Briefly vortex again
- Centrifuge at 300 x g for 5 s to collect the solution at the bottom of the tube.
- Store at -20 °C or use in PCR.
- Prepare a 100 µM primer stock by resuspending the primers in nuclease-free water. To determine the amount of water to add, multiply the number of nanomoles of lyophilized primer by 10. For example, if the tube contains 45 nM of lyophilized primer, add 450 µL of ddH2O and vortex the solution.
- Store the primer stock solutions at -20 °C or continue to perform the amplification.
3. Amplifying the gene fragment via PCR
NOTE: Decide which PCR kit is right for the gene of interest. Smaller genes (<1,000 kb) may be more amenable to a cheaper Taq polymerase, while larger genes (≥1,000 kb) may benefit from high fidelity polymerase to reduce errors. It is important to note that this initial PCR amplification is not necessary if the user is not concerned with preserving the initial gene fragment (It provides multiple attempts at circularization and allows for comparative studies of LET vs. RCA product). It is also important to note that this PCR amplified LET can be used directly in reactions; however, as mentioned in the introduction, it would only allow for a limited number of reactions if the further amplification steps were disregarded. Digestion and ligation can be performed on the resuspended gene fragment directly57 (if one is certain, they will not need more LET to perform additional circularization stocks). If this is the case, skip section 3 and continue to section 4. For performing PCR, follow these steps.
- Use the 100 µM stocks from step 2.8 to create 10 µM working solutions. Many PCR kit protocols call for 10 µM solutions of primers.
- Program the thermal cycler to conduct the reaction according to the kit manufacturer's protocols. Different kits call for slightly varied cycling parameters. For the kit listed in the Table of Materials, the conditions are 94 °C for 30 s of initial denaturation; 30 cycles of 94 °C for 30 s of denaturing, 45 °C for 30 s of primer annealing, and 68 °C for 60 s of extension; with a final extension at 68 °C for 5 min; and finally, a 10 °C indefinite hold.
- Ensure to select the correct elongation time (variable depending on the length of the gene to be amplified). Have an elongation time of 1 min for every 1,000 bp.
- Ensure to enter the correct annealing temperature for the primers. Use an online Tm calculator that uses both primers as inputs to determine the best annealing temperature58. An annealing temperature of 45 °C is sufficient when using M13 primers.
- When determining the number of cycles, refer to the manufacturer's protocol, but 30 cycles will most often result in sufficient amplification.
- If performing PCR, thaw and vortex the dNTPs. Use the PCR buffer provided in the kit.
- In a single PCR tube, combine all the kit components as directed in the manufacturer's protocol. To ensure successful amplification, add 1 µL of resuspended DNA stock (step 2.6).
- Gently homogenize the mixture by vortexing on medium setting for 5-10 s. Alternatively, pipette half the volume up and down 10-20 times to vortex.
- Perform the PCR reaction.
- If the PCR protocol did not include a final cooling step, allow the reaction to cool for 5 min at 10 °C before removing to drive condensation to the bottom of the tube.
- Purify the reaction using a PCR clean-up kit following the vendor's instructions.
- In a 1.5 mL tube, add DNA binding buffer and PCR sample at a ratio of 5:1, respectively.
- Transfer this mixture to the spin column and centrifuge at 16,000 x g for 1 min. Discard the flow-through.
- Add 200 µL of DNA wash buffer to the column and incubate at room temperature for 1 min.
- Centrifuge for 1 min at 16,000 x g and discard the flow-through.
- Repeat steps 2.8.3 and 2.8.4 without the 1 min incubation step.
- Centrifuge for an additional 1-2 min at 16,000 x g to remove any remaining buffer.
- Elute the DNA in 46 µL of ddH2O.
- Quantify the purified DNA using a spectrophotometer.
- Store the purified DNA at -20 °C or proceed to the next step.
4. Digestion and circularization
NOTE: Further amplification can be achieved by circularizing the DNA followed by RCA. Digest the DNA to prepare the template for circularization. This will remove the primer sequences and create sticky ends at both the 5' and 3' ends of the template. Reattach these ends via ligation reaction.
- In a PCR tube, combine 5 µL of the necessary buffer, 20 U of HindIII, and 45 µL of the purified DNA from step 3.8.
- Gently homogenize this mixture with a pipette.
- Incubate the mixture in a thermal cycler for 15 min at 37 °C.
- Heat inactivate HindIII by incubating for 20 min at 80 °C.
- Allow the reaction to cool to 10 °C before removing to drive condensation to the bottom of the tube.
- Add 5 µL of T4 ligase buffer and 800 U of T4 ligase to the newly digested DNA.
- Use T7 ligase, if desired.
- Gently homogenize this mixture with a pipette.
- Incubate the mixture for 1 h at 25 °C to perform the circularization reaction.
- Purify the reaction using a PCR clean-up kit following the vendor's instructions. Use the same protocol detailed in step 3.8.
- Quantify the DNA using a spectrophotometer. The expected values are ~20 ng/µL.
- Store at -20 °C or proceed to the next step.
5. Isothermal rolling circle amplification
NOTE: The Rolling circle amplification (RCA) can be performed using a commercial kit or with individually purchased components. Following the manufacturer's protocol will ensure a successful amplification. Kits typically contain a sample buffer, reaction buffer, and strand displacing polymerase, such as φ29 polymerase. Multiple reaction tubes can be combined to produce a large amount of DNA for cell-free expression (4 µg from 20 pg of starting material). The following protocol works efficiently.
- In a single tube, combine 20 µL of the sample buffer, 20 µL of the reaction buffer, 0.8 µL of the enzyme, and 1 µL of the circular expression template (CET) from step 4.9.
NOTE: This will have a total DNA mass of ~20 ng, but RCA can work with picogram amounts, thus allowing the dilution of the CET and extreme enzymatic amplification if there is a significant need for material in the high throughput screening. - Homogenize the mixture with a pipette and aliquot 10 µL of the mixture into four separate tubes.
- Incubate at 30 °C for 4-18 h.
- Heat inactivate the enzyme by incubating at 65 °C for 10 min. Reduce the temperature to 12 °C for 5 min to encourage condensation at the bottom of the tube.
NOTE: It's easiest to combine all temperature steps in an automated protocol on a thermal cycler. - Dilute the resulting solution by adding 15 µL of ddH2O to each tube.
- Combine all tubes and add directly to a cell-free reaction.
- If desired, use a PCR clean-up kit to purify the product and elute it in 36 µL of ddH2O to quantify. Ensure that the template concentration is ~100 ng/µL.
6. Cell-free reaction
NOTE: Perform cell-free expression by combining energy buffer, extract, and RCA template. A typical cell-free reaction using the PANOx-SP energy buffer consists of 1.2 mM ATP, 0.85 mM each of GMP, UMP, and CMP, 30 mM phosphoenolpyruvate, 130 mM potassium glutamate, 10 mM ammonium glutamate, 12 mM magnesium glutamate, 1.5 mM spermidine, 1 mM putrescine, 34 µg/mL of folinic acid, 171 µg/mL of E. coli tRNA mixture, 2 mM each of 20 unlabeled amino acids, 0.33 mM NAD, 0.27 mM Coenzyme A (CoA), 4 mM potassium oxalate, 57 mM HEPES-KOH buffer (pH 7.5), 0.24% volume of the E. coli extract, and variable amounts of DNA23,49. The volume of reaction can vary but 15 µL reactions can save on reagent usage and are small enough for use in a 384 black-walled microplate49,50.
- If expressing a fluorescent protein such as sfGFP, prepare a plate reader to read at the desired excitation/emission, temperature, and agitation.
- If using a 384-well plate, aliquot 60 µL of H2O into the wells bordering an empty sample well to maintain humidity and reduce the edge effect.
- Add the various required components into a tube for each sample. Add enough to perform triplicates. Replicates within the plate can help identify causes of variability.
- Add the extract, the energy buffer and then the DNA.
- Dilute to the final desired volume with ddH2O.
- Mix this solution thoroughly by pipetting half of the solution volume up and down 10-20 times.
- Transfer the reaction mixture in 15 µL aliquots to the desired wells in the microtiter plate.
- Seal the plate with a colorless sealing film to maintain humidity and prevent evaporation.
- Place the sealed plate in the plate reader and allow the reaction to complete.
- If expressing a protein that does not have the capability of being monitored live, use another temperature-controlled apparatus such as a thermoblock to incubate the plate.
7. Subtilisin assay
NOTE: If expressing the subtilisin BPN' (SBT(n)) gene in Supplementary Sequence #2, follow this protocol to assay the activity.
- Prepare a 10 µM stock solution of N-succinyl-Ala-Ala-Pro-Phe p-nitroanilide in dimethylformamide (DMF).
- Set a plate reader to measure absorbance at 410 nm every 20 s for 10 min while maintaining a temperature of 25 °C.
- In a flat bottom, colorless 96-well plate, aliquot 94 µL of ddH2O and 1 µL of N-succinyl-Ala-Ala-Pro-Phe p-nitroanilide from step 7.1.
- Add 5 µL of the finished cell-free reaction from step 6.7 and read using a plate reader set to the protocol described in step 7.2.
Representative Results
Expression of sfGFP from RCA templates was comparable to that of the pJL1 plasmid when using only 0.30 µL of unpurified RCA DNA in a 15 µL reaction (Figure 2A). In fact, doubling and tripling the amount of template appears to offer no benefit in BL21 DE3 Star extract, suggesting already saturated levels of the template at 0.30 µL per reaction. Conversely, there appears to be a benefit to increasing the amount of RCA template when added to cell extract sourced from the SHuffle strain (Figure 2B)28. For some proteins, results can be observed very quickly, which compresses the entire workflow (amplification and assaying) to under 24 h. However, some proteins require lower temperature or have slower folding times, which will increase the time until results are obtained but will affect the workflow presented here. This can be observed when expressing subtilisin (SBT(n)) where assaying after 4 h of expression was not long enough for SBT(n) maturation (Figure 3A, example of a failed result). Allowing the reaction to continue to 16 h can lead to detectable levels of SBT(n) (Figure 3B). This improvement may be temperature-dependent, as observed in literature where optimized temperature conditions were explored59,60.
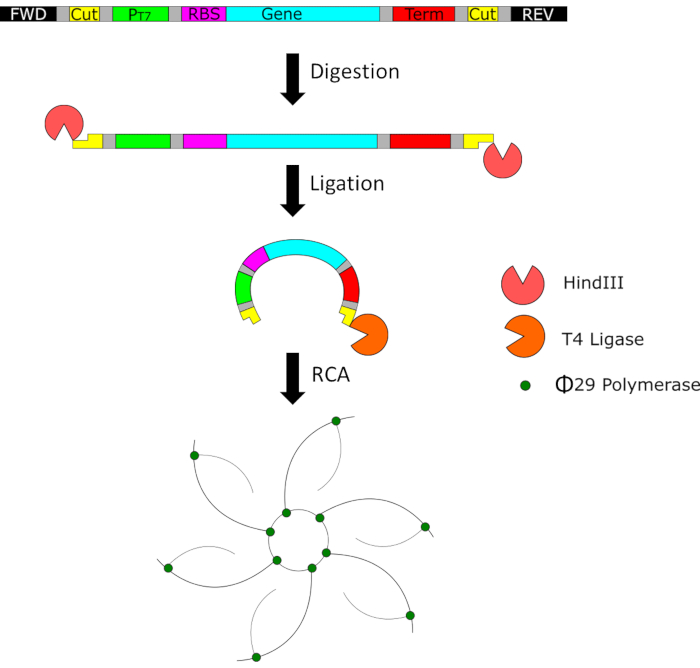
Figure 1: A representative schematic of the minimal genetic template and the process it undergoes after the initial PCR amplification step. The templates are digested with HindIII, circularized with T4 ligase, and further amplified with φ29 polymerase to create large concatemers. Please click here to view a larger version of this figure.
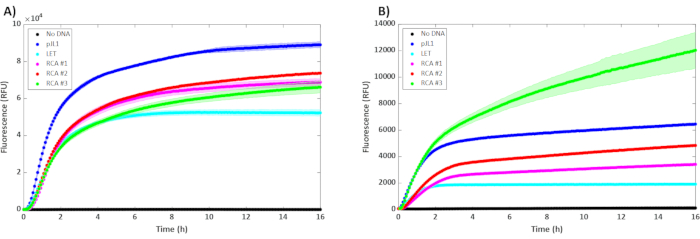
Figure 2: Results of the cell-free reactions with unpurified 5 nM of plasmid (pJL1), 5 nM of linear template (LET), and varying concentrations of unpurified RCA. RCA #1, #2, and #3 contained 0.3 µL, 0.6 µL, and 0.9 µL (respectively) of unpurified RCA product in a 15 µL reaction incubated at 30 °C (n = 3). Error bars represent ± 1 SD from the mean. The y-axis is fluorescence, and the x-axis is the time that has passed during the reaction. The kinetics of sfGFP expression are represented in (A) BL21 DE3 Star and (B) T7 SHuffle. Please click here to view a larger version of this figure.
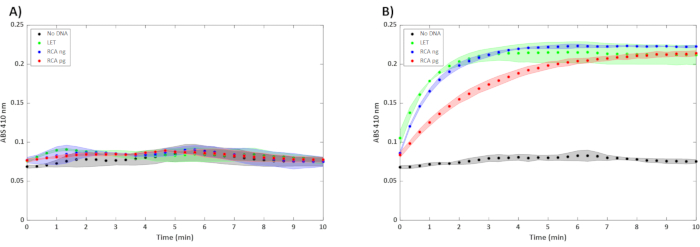
Figure 3: Cell-free reactions with 5 nM of SBT(n) LET and varying concentrations of unpurified RCA product. RCA ng and pg correspond to the concentration of the DNA used to perform rolling circle amplification. Unpurified RCA product was used in a 15 µL reaction incubated at 30 °C (n = 3). Error bars represent ± 1 SD from the mean. The y-axis is the absorbance at 410 nm, and the x-axis is the amount of time that has passed in the assay. Reactions were performed for (A) 4 h and (B) 16 h. Please click here to view a larger version of this figure.
| Cloning Method | |
| PCR | 2 -4 h |
| Plasmid Digestion | 35 min |
| Ligation | 1 h |
| Transformation | 2 h |
| Overnight Incubation | 16 h |
| Sequencing | 24 – 48 h |
| Glycerol Stock Prep | 16 h |
| Growth and Purification | 16 h |
| Total Time | 46 – 72 h |
| RCA Method | |
| PCR | 2 – 4 h |
| Digestion | 35 min |
| Ligation | 1 h |
| RCA | 4 – 18 h |
| CFE | 4 – 16 h |
| Total Time | 12 – 40 h |
Table 1: A comparison of the timeline between a simplified traditional cloning protocol and the RCA protocol covered herein.
Supplemental File: The supplemental file lists the sequences. Sequence #1 is sfGFP (999 base pairs) and sequence #2 is subtilisin BPN' (1344 bp). Please click here to download this File.
Discussion
The gene of interest can be any desired protein, but it is best to start with a fluorescent protein as a convenient reporter for real-time or end-point readout on a well plate reader for new adopters of this method. For new protein sequences, copy the amino acid sequence of the desired protein and paste it into the desired codon optimization tool61,62. There are usually many available organisms and strains of E. coli in the codon optimization tool, but choosing the general Escherichia coli option will be suitable. After optimization, double-check the gene to ensure the previously chosen cut site and primer sequences are not present. If so, the sequence can be optimized until the sequences are no longer present. In some situations, it may be necessary to replace the HindIII cut sites with a different restriction site if repeated optimization consistently results in an internal HindIII cut site. It is best to use the same cut site as often as possible to keep the process standardized and reduce the cost of keeping multiple restriction enzymes on hand.
Before beginning amplification, choose the PCR kit that best suits the LET. An LET with <1,000 base pairs can be amplified with a simple Taq polymerase while an LET with ≥ 1,000 base pairs may require a higher-fidelity polymerase63. Set up the protocol for amplification according to the kit manufacturer and the annealing temperature of the desired primers. The annealing temperature is critical to a successful amplification. A general rule of thumb is to select an annealing temperature that is 5 °C lower than the Tm of the primers. There are free online tools that will provide an optimized annealing temperature based on the sequence of the primers58. Using the correct annealing temperature is critical to producing high-quality DNA template.
Cell-free gene expression has seen a renaissance in recent years due to its speed, simplicity, and utility for synthetic biology prototyping. This work has outlined a method to increase the speed and ease when testing a large library of new, functional proteins. While traditional cloning methods can take days or weeks, this protocol can be done in less than 24 h. Table 1 outlines typical time ranges for both traditional cloning and the RCA protocol. Note that the cloning protocol has been simplified, and some optional steps have been left out, such as gel isolation. Table 1 also refers only to the common restriction digestion protocols; there are many other cloning methods, but these require a similar amount of time. The upper range of this enzymatic amplification protocol requires less time than the lower range of the cloning method, especially when considering an 8-hour workday. This is largely due to the removal of overnight incubations and lack of sequencing confirmation. The RCA protocol is based entirely on PCR and RCA, which require less specialized skills than traditional cloning, transformation, and cell culture techniques. This method makes cell-free expression accessible for labs that have no prior experience in cloning or access to capital necessary to transform and grow cells. This method is also well suited for projects focused on cytotoxic proteins, where cloning and propagation of the genes with traditional cloning is difficult due to toxicity in the living cell. This RCA protocol is also capable of amplifying very small amounts of circular DNA (picograms of DNA) to the levels necessary for CFE. In Figure 3B, the circularized SBT(n) template was diluted to 20 ng/µL and 20 pg/uL prior to RCA. While the observed rates of degradation were different, both reactions resulted in the same amount of substrate degradation within 10 min. The proposed method is not meant to replace cloning; plasmid propagation can produce quantities of DNA for non-toxic genes that cannot be matched. Rather, this is a convenient prototyping tool at a massive library screening phase (enzymatic steps are amenable to automation with standard fluid handler) that would help identify which sequences should be cloned for archiving and further propagation.
When designing a cell-free reaction, there is much flexibility, but there are some factors that must be taken into consideration to ensure success. Smaller reactions have a greater surface area to volume ratio, which is great for gas exchange, but the reaction needs to completely cover the bottom of the well for accurate fluorescence measurements. The temperature of the reaction can also vary depending on the gene of interest59. There are multiple choices for users when it comes to the selection of an energy buffer34,46. Some recipes are more expensive than others, but the decision is left to the user. There are also many potential sources for E. coli extract36. Users should familiarize themselves with extract production protocols and decide which is best for their purposes28,48,50,64,65,66,67.
When using RCA to amplify templates for cell-free expression, the choice of bacterial strain is very important. The expression profile tends to be better than that of LETs. The popular BL21 DE3 Star strain handles this well, with RCA performing ~1.4x better than LET and the pJL1 plasmid performing ~1.2x better than the best RCA concentration (Figure 2A). On the other hand, some strains exhibit template degradation and perform poorly due to the presence of native nucleases23,28. In this case, it appears that the SHuffle strain benefits from an increased RCA template. Previous literature has shown that SHuffle extract does not perform well with PCR products, but the highest concentration of unpurified RCA product used in this study outperformed the pJL1 plasmid (Figure 2B). The expression of functional SBT(n) (Figure 3) is an example of a cytotoxic protein that is conveniently made by cell-free but difficult to prototype in living cells due to toxicity (unable to clone this expression template in plasmid and propagate in E. coli). Unlike sfGFP, SBT(n) activity cannot be observed after only 4 h of expression (Figure 3A). The signal is detectible after 16 h of expression and maturation (Figure 3B). The unpurified DNA used in these examples came from 100 µL, which was four 10 µL RCA reactions diluted and combined. This one stock can support 333 cell-free reactions if only using 0.30 µL per reaction.
The compatibility of RCA templates with CFE needs to be further explored with various extracts. Potential complications with linear products may be alleviated by adding short oligos containing the E. coli chi sequence or the GamS protein52,53. Literature suggests that the addition of chi oligos can provide greater protection to linear templates than the GamS protein53. If using an extract known to provide low yields from linear templates, the user should analyze the cost of each protective agent and use the one that best suits their purposes. Another limitation is the inability to convert RCA template concentrations to molarity due to the nature of the resulting concatemer. This means one can add the same number of minimal templates based on mass, but they will have varying concatemer lengths, which might affect expression levels. The authors have not found this to be an issue in screening/prototyping as the averaged expression levels from each well are the same (low variance) but could be an issue if expressing in smaller volumes (e.g., microdroplets).
Cell-free gene expression has the potential to be used as a significant tool in protein prototyping and design-build-test workflows. However, most cell-free expression workflows still rely on traditional plasmids as the genetic template. This slows down the process and keeps cell-free expression from being utilized to its fullest extent for screening/prototyping purposes. Amplifying templates and subsequently performing RCA can quickly produce enough genetic templates for many reactions, producing enough protein for downstream characterization and functional testing.
Divulgaciones
The authors have nothing to disclose.
Acknowledgements
The authors acknowledge NIH 1R35GM138265-01 and NSF 2029532 for partial support of this project.
Materials
| Alaline | Formedium | DOC0102 | |
| Ammonium glutamate | MP Biomedicals | MP21805951 | |
| Arginine | Formedium | DOC0106 | |
| Asparagine | Formedium | DOC0114 | |
| Aspartic Acid | Formedium | DOC0118 | |
| ATP | Sigma | A2383 | |
| Axygen Sealing Film | Corning | PCR-SP | |
| CMP | Sigma | C1006 | |
| Coenzyme A | Sigma | C3144 | |
| CutSmart Buffer | NEB | B7204S | Provided with HindIII |
| Cysteine | Formedium | DOC0122 | |
| DNA Clean and Concentrator Kit | Zymo Research | D4004 | Used for purifying DNA |
| dNTPs | NEB | N0447 | |
| E. coli tRNA | Sigma (Roche) | 10109541001 | |
| Folinic Acid | Sigma | 47612 | |
| Gene Fragment | IDT | ||
| Glutamic Acid | Formedium | DOC0134 | |
| Glutamine | Formedium | DOC0130 | |
| Glycine | Formedium | DOC0138 | |
| GMP | Sigma | G8377 | |
| HEPES | Sigma | H3375 | |
| HindIII-HF | NEB | R3104L | |
| Histidine | Formedium | DOC0142 | |
| Isoleucine | Formedium | DOC0150 | |
| Leucine | Formedium | DOC0154 | |
| Lysine | Formedium | DOC0158 | |
| Magnesium glutamate | Sigma | 49605 | |
| Methionine | Formedium | DOC0166 | |
| Microtiter Plate (384 well) | Greiner | 781906 | |
| Microtiter Plate (96 well) | Greiner | 655809 | |
| Multimode Plate Reader | BioTek | Synergy Neo2 | |
| NAD | Sigma | N8535 | |
| NanoPhotometer | Implen | NP80 | |
| OneTaq DNA Polymerase | NEB | M0480 | |
| PCR Tube | VWR | 20170-012 | |
| Phenylalanine | Formedium | DOC0170 | |
| Phosphoenolpyruvate | Sigma (Roche) | 10108294 | |
| Potassium glutamate | Sigma | G1501 | |
| Potassium oxalate | Fisher Scientific | P273 | |
| Proline | Formedium | DOC0174 | |
| Putrescine | Sigma | P5780 | |
| Serine | Formedium | DOC0178 | |
| Spermidine | Sigma | S0266 | |
| T4 DNA Ligase | NEB | M0202S | |
| T4 DNA Ligase Reaction Buffer | NEB | B0202S | Provided with T4 DNA Ligase |
| TempliPhi Amplification Kit | Cytiva | 25640010 | Used for RCA |
| Thermal Cycler | Biorad | C1000 Touch | |
| Thermoblock | Eppendorf | ThermoMixer FP | |
| Threonine | Formedium | DOC0182 | |
| Tryptophan | Formedium | DOC0186 | |
| Tyrosine | Formedium | DOC0190 | |
| UMP | Sigma | U6375 | |
| Valine | Formedium | DOC0194 |
Referencias
- Sun, Q., et al. A simple and low-cost paper-based colorimetric method for detecting and distinguishing the GII.4 and GII.17 genotypes of norovirus. Talanta. 225, 121978 (2021).
- Pardee, K., et al. Low-Cost Detection of Zika Virus Using Programmable Biomolecular Components. Cell. 165 (5), 1255-1266 (2016).
- Pardee, K., et al. Paper-based synthetic gene networks. Cell. 159 (4), 940-954 (2014).
- Ma, D., Shen, L., Wu, K., Diehnelt, C. W., Green, A. A. Low-cost detection of norovirus using paper-based cell-free systems and synbody-based viral enrichment. Synthetic Biology. 3 (1), (2018).
- Park, S., Lee, J. W. Detection of coronaviruses using rna toehold switch sensors. International Journal of Molecular Sciences. 22 (4), 1772 (2021).
- Cao, M., Sun, Q., Zhang, X., Ma, Y., Wang, J. Detection and differentiation of respiratory syncytial virus subgroups A and B with colorimetric toehold switch sensors in a paper-based cell-free system. Biosensors and Bioelectronics. 182, 113173 (2021).
- Mcnerney, M. P., et al. Point-of-care biomarker quantification enabled by sample-specific calibration. Science Advances. 5 (9), (2019).
- Silverman, A. D., Akova, U., Alam, K. K., Jewett, M. C., Lucks, J. B. Design and optimization of a cell-free atrazine biosensor. ACS Synthetic Biology. 9 (3), 671-677 (2020).
- Salehi, A. S. M., et al. Cell-free protein synthesis approach to biosensing hTRβ-specific endocrine disruptors. Analytical Chemistry. 89 (6), 3395-3401 (2017).
- Garamella, J., Majumder, S., Liu, A. P., Noireaux, V. An adaptive synthetic cell based on mechanosensing, biosensing, and inducible gene circuits. ACS Synthetic Biology. 8 (8), 1913-1920 (2019).
- Glasscock, C. J., et al. Dynamic control of pathway expression with riboregulated switchable feedback promoters. ACS Synthetic Biology. 16, (2019).
- Thavarajah, W., et al. Point-of-use detection of environmental fluoride via a cell-free riboswitch-based biosensor. ACS Synthetic Biology. 9 (1), 10-18 (2020).
- Pardee, K., et al. Portable, on-demand biomolecular manufacturing. Cell. 167 (1), 248-259 (2016).
- Nelson, J. A. D., et al. Hydrofoam and oxygen headspace bioreactors improve cell-free therapeutic protein production yields through enhanced oxygen transport. Biotechnology Progress. 37 (2), 3079 (2020).
- Cai, Q., et al. A simplified and robust protocol for immunoglobulin expression in Escherichia coli cell-free protein synthesis systems. Biotechnology Progress. 31 (3), 823-831 (2015).
- Ogonah, O. W., Polizzi, K. M., Bracewell, D. G. Cell free protein synthesis: a viable option for stratified medicines manufacturing? A brief history of cell free synthesis systems. Current Opinion in Chemical Engineering. 18, 77-83 (2017).
- Zawada, J. F., et al. Microscale to manufacturing scale-up of cell-free cytokine production-a new approach for shortening protein production development timelines. Biotechnology and Bioengineering. 108 (7), 1570-1578 (2011).
- Stark, J. C., et al. On-demand biomanufacturing of protective conjugate vaccines. Science Advances. 7 (6), (2021).
- Huang, A., et al. BioBitsTM Explorer: A modular synthetic biology education kit. Science Advances. 4 (8), 1-11 (2018).
- Stark, J. C., et al. BioBits health: classroom activities exploring engineering, biology, and human health with fluorescent readouts. ACS Synthetic Biology. 8 (5), 1001-1009 (2019).
- Stark, J. C., et al. BioBitsTM Bright: A fluorescent synthetic biology education kit. Science Advances. 4 (8), 33 (2018).
- Shinoda, T., et al. Cell-free methods to produce structurally intact mammalian membrane proteins. Scientific Reports. 6, (2016).
- Dopp, J. L., Rothstein, S. M., Mansell, T. J., Reuel, N. F. Rapid prototyping of proteins: Mail order gene fragments to assayable proteins within 24 hours. Biotechnology and Bioengineering. 116 (3), 667-676 (2019).
- Sachse, R., Dondapati, S. K., Fenz, S. F., Schmidt, T., Kubick, S. Membrane protein synthesis in cell-free systems: From bio-mimetic systems to bio-membranes. FEBS Letters. 588 (17), 2774-2781 (2014).
- Salehi, A. S. M., et al. Cell-free protein synthesis of a cytotoxic cancer therapeutic: Onconase production and a just-add-water cell-free system. Biotechnology Journal. 11 (2), 274-281 (2016).
- Georgi, V., et al. On-chip automation of cell-free protein synthesis: New opportunities due to a novel reaction mode. Lab on a Chip. 16 (2), 269-281 (2016).
- Thoring, L., et al. Cell-free systems based on CHO cell lysates: Optimization strategies, synthesis of “difficult-to-express” proteins and future perspectives. PLoS One. 11 (9), (2016).
- Dopp, J. L., Reuel, N. F. Simple, functional, inexpensive cell extract for in vitro prototyping of proteins with disulfide bonds. Biochemical Engineering Journal. 164, 107790 (2020).
- Isaksson, L., Enberg, J., Neutze, R., Göran Karlsson, B., Pedersen, A. Expression screening of membrane proteins with cell-free protein synthesis. Protein Expression and Purification. 82 (1), 218-225 (2012).
- Techner, J. M., et al. High-throughput synthesis and analysis of intact glycoproteins using SAMDI-MS. Analytical Chemistry. 92 (2), 1963-1971 (2020).
- Kim, H. C., et al. Implementing bacterial acid resistance into cell-free protein synthesis for buffer-free expression and screening of enzymes. Biotechnology and Bioengineering. 112 (12), 2630-2635 (2015).
- Rolf, J., Siedentop, R., Lütz, S., Rosenthal, K. Screening and identification of novel cGAS homologues using a combination of in vitro and in vivo protein synthesis. International Journal of Molecular Sciences. 21 (1), (2020).
- Haslinger, K., Hackl, T., Prather, K. L. J. Rapid in vitro prototyping of O-methyltransferases for pathway applications in Escherichia coli. bioRxiv. , (2020).
- Dopp, J. L., Tamiev, D. D., Reuel, N. F. Cell-free supplement mixtures: Elucidating the history and biochemical utility of additives used to support in vitro protein synthesis in E. coli extract. Biotechnology Advances. 37 (1), 246-258 (2018).
- Gregorio, N. E., Levine, M. Z., Oza, J. P. A user’s guide to cell-free protein synthesis. Methods and Protocols. 2 (1), 24 (2019).
- Cole, S. D., Miklos, A. E., Chiao, A. C., Sun, Z. Z., Lux, M. W. Methodologies for preparation of prokaryotic extracts for cell-free expression systems. Synthetic and Systems Biotechnology. 5 (4), 252-267 (2020).
- Chiba, C. H., Knirsch, M. C., Azzoni, A. R., Moreira, A. R., Stephano, M. A. Cell-free protein synthesis: advances on production process for biopharmaceuticals and immunobiological products. BioTechniques. 70, (2021).
- Laohakunakorn, N. Cell-free systems: A proving ground for rational biodesign. Frontiers in Bioengineering and Biotechnology. 8, 788 (2020).
- Dondapati, S. K., Stech, M., Zemella, A., Kubick, S. Cell-free protein synthesis: A promising option for future drug development. BioDrugs. , 1-22 (2020).
- Noireaux, V., Liu, A. P. The new age of cell-free biology. Annual Review of Biomedical Engineering. 22, 51-77 (2020).
- Khambhati, K., et al. Exploring the Potential of Cell-Free Protein Synthesis for Extending the Abilities of Biological Systems. Frontiers in Bioengineering and Biotechnology. 7, (2019).
- Carlson, E. D., Gan, R., Hodgman, C. E., Jewett, M. C. Cell-free protein synthesis: Applications come of age. Biotechnology Advances. 30 (5), 1185-1194 (2012).
- Rosenblum, G., Cooperman, B. S. Engine out of the chassis: Cell-free protein synthesis and its uses. FEBS Letters. 588 (2), 261-268 (2014).
- Swartz, J. R. Transforming biochemical engineering with cell-free biology. AIChE Journal. 58 (1), 5-13 (2012).
- Cole, S. D., et al. Quantification of interlaboratory cell-free protein synthesis variability. ACS Synthetic Biology. 8 (9), 2080-2091 (2019).
- Caschera, F., Noireaux, V. A cost-effective polyphosphate-based metabolism fuels an all E. coli cell-free expression system. Metabolic Engineering. 27, 29-37 (2015).
- Levine, M. Z., et al. Activation of energy metabolism through growth media reformulation enables a 24-hour workflow for cell-free expression. ACS Synthetic Biology. 9 (10), 2765-2774 (2020).
- Hunt, J. P., et al. Streamlining the preparation of “endotoxin-free” ClearColi cell extract with autoinduction media for cell-free protein synthesis of the therapeutic protein crisantaspase. Synthetic and Systems Biotechnology. 4 (4), 220-224 (2019).
- Dopp, J. L., Jo, Y. R., Reuel, N. F. Methods to reduce variability in E. Coli-based cell-free protein expression experiments. Synthetic and Systems Biotechnology. 4 (4), 204-211 (2019).
- Sun, Z. Z., et al. Protocols for Implementing an Escherichia coli Based TX-TL Cell-Free Expression System for Synthetic Biology. Journal of Visualized Experiments: JoVE. , e50762 (2013).
- Schinn, S. M., Broadbent, A., Bradley, W. T., Bundy, B. C. Protein synthesis directly from PCR: Progress and applications of cell-free protein synthesis with linear DNA. New Biotechnology. 33 (4), 480-487 (2016).
- Sitaraman, K., et al. A novel cell-free protein synthesis system. Journal of Biotechnology. 110 (3), 257-263 (2004).
- Marshall, R., Maxwell, C. S., Collins, S. P., Beisel, C. L., Noireaux, V. Short DNA containing χ sites enhances DNA stability and gene expression in E. coli cell-free transcription-translation systems. Biotechnology and Bioengineering. 114, 2137-2141 (2017).
- Sun, Z. Z., Yeung, E., Hayes, C. A., Noireaux, V., Murray, R. M. Linear DNA for rapid prototyping of synthetic biological circuits in an escherichia coli based TX-TL cell-free system. ACS Synthetic Biology. 3 (6), 387-397 (2014).
- . Addgene: pJL1 Available from: https://www.addgene.org/69496/ (2021)
- . IDT Codon Optimization Tool Available from: https://www.idtdna.com/pages/tools/codon-optimization-tool (2021)
- Hadi, T., et al. Rolling circle amplification of synthetic DNA accelerates biocatalytic determination of enzyme activity relative to conventional methods. Scientific Reports. 10 (1), 10279 (2020).
- . New England Biolabs Tm Calculator Available from: https://tmcalculator.neb.com/#!/main (2021)
- Shin, J., Noireaux, V. Efficient cell-free expression with the endogenous E. Coli RNA polymerase and sigma factor 70. Journal of Biological Engineering. 4, (2010).
- Colant, N., et al. A rational approach to improving titer in Escherichia coli-based cell-free protein synthesis reactions. Biotechnology Progress. 37 (1), 3062 (2021).
- Burgess-Brown, N. A., et al. Codon optimization can improve expression of human genes in Escherichia coli: A multi-gene study. Protein Expression and Purification. 59, 94-102 (2008).
- Maertens, B., et al. Gene optimization mechanisms: A multi-gene study reveals a high success rate of full-length human proteins expressed in Escherichia coli. Protein Science. 19 (7), 1312-1326 (2010).
- Eckert, K. A., Kunkel, T. A. DNA polymerase fidelity and the polymerase chain reaction. Genome Research. 1 (1), 17-24 (1991).
- Dopp, J. L., Reuel, N. F. Process optimization for scalable E. coli extract preparation for cell-free protein synthesis. Biochemical Engineering Journal. 138, 21-28 (2018).
- Liu, D. V., Zawada, J. F., Swartz, J. R. Streamlining Escherichia Coli S30 extract preparation for economical cell-free protein synthesis. Biotechnology Progress. 21 (2), 460-465 (2005).
- Levine, M. Z., Gregorio, N. E., Jewett, M. C., Watts, K. R., Oza, J. P. Escherichia coli-Based Cell-Free Protein Synthesis: Protocols for a robust, flexible, and accessible platform technology. Journal of Visualized Experiments: JoVE. (144), e58882 (2019).
- Kwon, Y. C., Jewett, M. C. High-throughput preparation methods of crude extract for robust cell-free protein synthesis. Scientific Reports. 5, (2015).

