Measurement of Microtubule Dynamics by Spinning Disk Microscopy in Monopolar Mitotic Spindles
Summary
Here we present a robust and detailed method of microtubule dynamics analysis in cells synchronized in prometaphase using live-cell spinning disk confocal microscopy and MATLAB-based image processing.
Abstract
We describe a modification of an established method to determine microtubule dynamics in living cells. The protocol is based on the expression of a genetically encoded marker for the positive ends of microtubules (EB3 labelled with tdTomato fluorescent protein) and high-speed, high-resolution, live-cell imaging using spinning disk confocal microscopy. Cell cycle synchronization and increased density of microtubules are achieved by inhibiting centrosomal separation in mitotic cells, and analysis of growth is performed using open-source U-Track software. The use of a bright and red-shifted fluorescent protein, in combination with the lower laser power and reduced exposure time required for spinning disk microscopy reduce phototoxicity and the probability of light-induced artifacts. This allows for imaging a larger number of cells in the same preparation while maintaining the cells in a growth medium under standard culture conditions. Because the analysis is performed in a supervised automatic fashion, the results are statistically robust and reproducible.
Introduction
Microtubules (MTs) are highly dynamic structures found in virtually all eukaryotic cells and in some bacteria1. Together with actin and intermediate filaments, they sculpt the cytoskeleton2,3. Cell division4, molecule transport5, flagellar beating6, the sensation of the surrounding environment through primary cilium7, hearing (kinocilium)8,9, embryogenesis10,11,12, invasion and metastasis13,14, and even memory formation15,16,17,18, and many other processes primarily rely on MTs. Participation of MTs in all these events would be impossible without their remarkable ability to rapidly switch between growth (polymerization) and shrinkage (depolymerization). This property is described as dynamic instability19. MT dynamicity is altered in many pathological conditions20,21,22. Hence, determining the nature of this property can help to understand disease mechanisms and subsequently their treatment.
A long list of methods has been developed for MT dynamics analysis, most of which are based on imaging techniques23. Initially, wide field light microscopes were used for observing the formation of tubulin polymers in vitro24. The discovery of end-binding (EB)-proteins that collect at MT plus-ends and the development of methods to fluorescently label proteins made it possible to observe the behavior of MTs directly in living cells with wide field and confocal fluorescence microscopes25,26,27. One EB-protein is end-binding protein 3 (EB3)28; by overexpressing and tracking EB3 fused to a fluorescent protein, MT plus-end assembly rates can be determined29,30.
Confocal laser scanning fluorescence microscopy (CLSM) is frequently used to follow MT dynamics. However, this imaging technique poses a high risk of phototoxicity and photobleaching, two undesirable processes for live cell and dim sample imaging31. In order to obtain a better signal-to-noise ratio, the laser power and the exposure duration should be high enough while not damaging the samples, and this requires sacrificing resolution in exchange for speed. A suitable alternative to CLSM is spinning disk microscopy32. This imaging modality is based on the use of a Nipkow disk33, which consists of a moving disk bearing an array of pinholes, and works equivalently to many CLS microscopes imaging the same sample simultaneously34. Therefore, the light from the laser will illuminate several regions in the sample simultaneously but retain the confocal nature. The Nipkow disk, therefore, allows obtaining images similar to CLSM but faster and using less laser power. The Nipkow disk was further improved by Yokogawa Electric, which introduced a second disk with an array of microlenses on it that individually direct light into a respective pinhole, further reducing phototoxicity and photobleaching35. Thus, spinning disk laser scanning microscopy became a method of choice for live cell imaging, and it makes it possible to obtain images with high signal-to-noise ratio at a high speed31,36, which is crucial for resolving signals such as those from the fast-growing MT ends.
MT dynamics differ temporarily. For example, the mitotic MTs are more dynamic than the interphase ones37,38. Similarly, differences in the growth rate and shrinkage have been observed even within the same cell cycle phase, such as mitosis39,40. Therefore, to avoid false data collection, the measurement of MT dynamics should be limited to a narrow time-window during the cell cycle. For example, measurement of MT dynamics in prometaphase can be achieved by treating the cells with dimethylenastron (DME), a monastrol analogue that inhibits the motor kinesin Eg541 and prevents the formation of the bipolar mitotic spindle42. Inhibition of cells at prometaphase with Eg5 inhibitor DME and other monastrol derivatives does not affect the MT dynamics43,44,45, which makes DME a useful tool for studying MT dynamics both in fixed and live cells44.
Here we combine the method of MT dynamics analysis in prometaphase cells described by Ertych et al.44 with dual spinning disk imaging. This method allows measurement of the MT dynamics in prometaphase cells collected from a single focal plane with a higher imaging rate, yet without photobleaching and minimal phototoxicity. Furthermore, as a fluorescent reporter, we use tandem dimer Tomato fluorescent protein (tdTomato) which has improved brightness and photostability in comparison to the green fluorescent protein (EGFP) and is excited with lower energy light46. Therefore, tdTomato requires less laser power for excitation and is less phototoxic. Altogether, we further improve the method by reducing the phototoxicity and improving the resolution and postprocessing required for the MT dynamics analysis. Additionally, we create a basis for future modifications of the method by combining it with other synchronization techniques.
Protocol
1. Seeding of HeLa Cells
- Prepare 2 mL of 5 μg/mL fibronectin solution in phosphate buffered saline (PBS) and add 450 μL of it into each well of a 4 well chambered coverslip (#1.5). Incubate the slide for 15 min at 37 °C and 5% CO2.
- Rinse asynchronously growing HeLa cells with Dulbecco’s Phosphate Buffered Saline (DPBS) and incubate with trypsin-EDTA (0.05%: 0.02%; w:v) for 5 min at 37 °C. Stop the enzymatic reaction by the addition of Roswell Park Memorial Institute (RPMI) 1640 medium supplemented with 10% heat-inactivated fetal calf serum (FCS) at 3:1 (v:v) ratio of added trypsin-EDTA.
NOTE: HeLa cells were maintained in RPMI 1640 medium supplemented with 10% heat-inactivated FCS at 37 °C and 5% CO2 and were routinely passaged once they reached 80–90% confluency as described above. - Determine the cell concentration using a Neubauer chamber. Mix a 50 μL aliquot of the cell suspension with trypan blue at 1:1 (v:v) ratio, resuspend, and transfer 10 μL of the suspension into the chamber. Count only the trypan blue-negative cells inside of the four large squares (for details see Phelan et al.47). Derive the cell concentration from the counted cell number using the following formula:
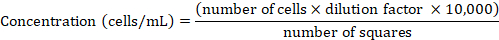
- Pellet the cells by centrifugation at 300 x g for 2 min. Resuspend with fresh RPMI 1640 in order to obtain 1 x 106 cells/mL.
- Remove the fibronectin from the chambered coverslip, wash the wells twice with DPBS, and seed 50,000 cells per well.
- Return the chambered coverslip with the cells to the incubator and grow them for 24 h at 37 °C and 5% CO2.
2. Expression of pEB3-tdTomato in HeLa Cells
- Prepare a 1.5 mL microcentrifuge tube. For each tube, dilute 2 μg of pEB3-tdTomato48 with transfection buffer (synthetic product in aqueous solution) to a final volume of 396 μL.
- Add 4 μL of transfection reagent (non-lipidic, containing polyethylenimine) to the first tube, and vortex the mixture immediately for exactly 10 s.
- Briefly spin down the tube with a microcentrifuge and incubate at room temperature (RT) for 10 min.
- Remove the HeLa cells from the incubator. Dropwise, add 100 μL of the transfection mixture to each well of a 4 well chambered coverslip, and return the cells to the incubator.
- After 4 h of incubation at 37 °C and 5% CO2, supplement the cells with fresh growth medium and incubate for at least 24 h at 37 °C and 5% CO2.
NOTE: It is necessary to optimize transfection conditions for each cell type. The expression levels need to be low enough to allow the identification of single MT growing ends. Alternatively, a cell line stably expressing EB3-tdTomato can be used in the experiments; this would reduce variability in expression levels of EB3-tdTomato between preparations and between cells from the same preparation49.
3. Synchronization and Live-cell Imaging of pEB3-tdTomato–expressing HeLa Cells
- Prepare a 2.5 μM solution of dimethylenastron (DME) in phenol-red free Dulbecco's Modified Eagle Medium (DMEM) supplemented with 10% FCS and 2 mM L-glutamine or an alternative glutamine supply.
- Replace the growth medium in the chambered coverslip with 500 μL of the growth medium containing 2.5 μM DME and incubate the cells at 37 °C and 5% CO2.
- After 3.5 h of incubation with DME, transfer the cells to the microscope, mount the chambered coverslip into an environmental chamber with dark panels for imaging at 37 °C and 5% CO2, and further incubate until the total incubation time is 4 h.
NOTE: The maintenance of temperature at 37 °C without fluctuation is crucial for the experiment. - Perform the time-lapse imaging on an inverted microscope equipped with a 100x 1.49 N.A. oil immersion objective, a dual spinning disk confocal system, and a reliable autofocus system for continuous maintenance of the focal plane. Define the imaging parameters as follows.
NOTE: We use an Electron Multiplying Charge-Coupled Device camera (EM-CCD).- For EB3-tdTomato excitation, use a 561 nm laser line with 200 ms exposure time. Collect the emitted light through a quadruple bandpass (405, 488, 561, 640 nm) dichroic mirror and a 600/52 nm emission filter.
NOTE: Laser power can be adjusted for each imaged cell to prevent image saturation. In all time-lapse movies given here the laser power was set to 5.3 mW. - Find a cell in prophase and focus in the Z-plane corresponding to the center of the monopolar mitotic spindle. Acquire images every 0.5 s over a total of 1 min with no binning and no illumination between the exposures.
- For EB3-tdTomato excitation, use a 561 nm laser line with 200 ms exposure time. Collect the emitted light through a quadruple bandpass (405, 488, 561, 640 nm) dichroic mirror and a 600/52 nm emission filter.
4. Analysis of the MT Dynamics Using U-Track v2.2.0
- To analyze the MT dynamics a numerical computing environment software is required (e.g., MATLAB).
NOTE: Basic understanding of the software is sufficient for the analysis. Comprehensive help material and tutorials are available on the developer’s website (https://uk.mathworks.com/products/matlab/getting-started.html). - Download (https://github.com/DanuserLab/u-track) and install the open-source U-Track v2.2.0 software following the detailed instructions given in the "Readme_u-track.pdf" file50,51,52.
- Launch the numerical-analysis software and add U-Track v2.2.0 folder with subfolders into the software search path.
- From the command window call "movieSelectorGUI". This opens a dialogue window from which the raw files generated by the image acquisition software at the microscope can be imported (Supplementary Figure 1, Figure 2, Figure 3, Figure 4).
NOTE: The U-Track software is compatible with other image data formats. It uses Bio-Formats, which recognizes different life science data formats53. - The size of each image is read from the metadata automatically. Manually enter the numerical aperture of the objective (in this case 1.49) and the time interval (0.5 s) used for imaging (Supplementary Figure 1B). Additionally, information on the excitation wavelength, the fluorophore, and the exposure time can also be provided, but they are not critical for further analysis.
- Once all the images are loaded, save the entered time-lapse series as a movie list by selecting the "Save As Movie List". On the right side of the dialogue window select the "U-Track" option and press "Continuer" (Supplementary Figure 1C).
NOTE: The values are optimized for HeLa cells. If switching to a different cell line, the values should be defined again. Alternatively, use the settings recommended by the software developers. The detailed explanation of each of the parameters and how they should be defined can be found in the technical report provided with the previous version of the software, plusTipTracker50. - From the pop-up window select "Microtubule Plus-Ends" and press "Ok" (Supplementary Figure 1C). The new dialogue window allows determining the parameters for the three steps of the analysis (Supplementary Figure 1D), which are detection, tracking, and track analysis.
- In step 1 choose "Settings" and from a drop-down menu select "Comet Detection" as a detection method (Supplementary Figure 2B).
- From the new dialogue window define the parameters for the difference of Gaussians filter and the watershed segmentation as follows (Supplementary Figure 2C): Mask process to be used for the detection = None; Low-pass Gaussian standard deviation = 1 pixel; High-pass Gaussian standard deviation = 3 pixels; Minimum threshold = 3 standard deviations; Threshold step size = 0.25 standard deviations. Select "Apply Settings to All Movies" and "Apply".
- In step 2, the parameters for linking, gap closing, merging and splitting, and Kalman filter functions are defined in three steps as highlighted in pink, green, and blue, accordingly (Supplementary Figure 3B). For these steps, select the "Microtubule Plus-end Dynamics" and from the "Setting" option define the values as indicated in Supplementary Figure 3C–E, respectively.
- For problems with dimensionality, choose "2" from the drop-down menu. Use Maximum Gap to Close = 5 frames; Minimum Length of Track Segments from First Step = 3 frames. As before, select "Apply Settings to All Movies" and click on "Apply".
- In step 3 of the analysis, the detected MT tracks are classified (Supplementary Figure 4). As a track analysis method, choose "Microtubule Dynamics Classification" and define the parameters through the "Setting" button as indicated in Supplementary Figure 4B,C. After that, choose the "Apply Settings to All Movies" box and click on "Apply".
- Once all the parameters are defined, from the "Control Panel–U-Track" window (Supplementary Figure 1D) select the "Apply Check/Uncheck to All Movies" and "Run All Movies" boxes and press "Run". This will initiate the MT analysis of the time-lapse series.
- Once the movie processing is completed, a message "Your movie(s) have been processed successfully" is displayed. Press "Ok", then "Save".
- Now it is safe to quit the numerical-analysis software. The results from the movie processing are stored in subfolder structures as m-files in the folder where the raw files are stored.
5. Statistical Analysis of the MT Dynamics
- Import the m-files into a preferred statistic analysis program.
NOTE: In our case, we first import the files in a standard spreadsheet to make them readable. The m-files contain statistical information (median, mean, and standard deviation) on different parameters (e.g., growth speed, MT dynamicity). The detailed list of the parameters is given in the technical report provided with the previous version of the software, plusTipTracker50,52. The generated m-files can also be imported into other data processing software. - Choose the "growth speed mean" parameter and import it into a table for statistics and display. Enter the information on other parameters, (e.g., "dynamicity") either in a new table or in a new column of the same grouped table and plot.
Representative Results
Following the given protocol outlined in Figure 1A, the pEB3-tdTomato plasmid was transiently expressed in asynchronously growing HeLa cells. The cells were synchronized 48 h after the transfection at prometaphase through DME treatment (Figure 1B). This step ensured that the measurement of MT dynamics was always done at the same phase of the cell cycle. The time-lapse movies were further processed and analyzed with U-Track v2.2.0 as described in its supplementary documentation50,51,52. Although the plus-end binding proteins trace only MT growth phases, the U-Track v2.2.0 extrapolates the information on the pause and shrinkage events by linking sequential growth phases and reconstructing the full trajectories26,50. The algorithm is based on the spatially and temporally global tracking framework described by Jaqaman et al.51.
It is important to note that the sensitivity and accuracy of the analysis are strongly dependent on several analysis parameters. As an example, the time-lapse movies were analyzed as described in the protocol (Figure 1C, Video 1 "Before", and Video 2 "After" the analysis), and the resulting growth speed and dynamicity (collective displacement of gap-containing tracks over their entire lifetime) are plotted in Figure 1E,F, respectively (black circles). Then the parameters described to greatly affect the analysis50, such as "Maximum Gap Length" and "Maximum Shrinkage Factor" were modified for the same set of time-lapse movies (Videos 3 and 4, respectively). The corresponding values of growth speed and dynamicity are given in Figure 1E,F as red squares and blue triangles, respectively. The resulting growth speed was not deeply affected. However, the values obtained for dynamicity were significantly different when "Maximum Gap Length" was modified, while it remained unchanged upon altering the "Maximum Shrinkage Factor". As shown in Figure 1D, in all three cases the detection of the MT subtracks was similarly robust. Yet, the reconstruction of the full MT trajectories was mostly affected when "Maximum Gap Length" was set to 15 (Figure 1D, inset images). Further, in order to assess whether the imaging conditions interfered with the MT behavior, the first (1–61 frames) and the second (61–121 frames) halves of the time-lapse series were analyzed separately and the corresponding growth speed and dynamicity values were compared (Figure 1G,H, respectively). As expected, no significant differences were detected between the two parts of the time-lapse series. In videos 1–8 time-lapse images of a mitotic cell synchronized in prophase and expressing EB3-tdTomato are given (duration = 1 min; interval = 0.5 s).
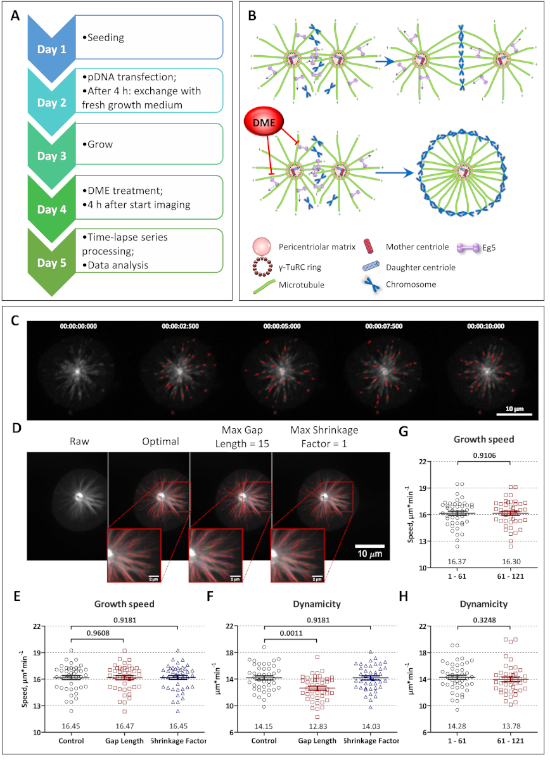
Figure 1: Analysis of the MT dynamics in HeLa cells synchronized in prometaphase. (A) An outline of the steps of the protocol. (B) The schematic representation of the mechanism of DME mediated formation of a monopolar mitotic spindle. (C) A montage of the first 10 frames of the time-lapse movie processed with U-Track software with every second frame shown. The detected trajectories of the MT growth are marked with red. (D) The time series projection of the raw image file and after MT tracking using the settings described in the protocol ("optimal"), and when changing either "Maximum Gap Length" or "Maximum Shrinkage Factor" are given. The insets represent the full MT trajectories, which consist of the growth (red), pause (light blue), shrinkage (yellow), fgap reclassified as growth (green) and bgap reclassified as pause (dark blue) events. The growth speed means (E) and the dynamicity (F) values are shown, and the results using either of the suggested optimal criteria (black circles), maximal gap length set to 15 (red squares), or the maximal shrinkage factor set to 1.0 (blue triangles) are plotted (n = 45 cells; mean ± SEM; one-way ANOVA analysis with Tukey post hoc test for multiple comparison). The growth speed means (G) and the dynamicity (H) values are shown for the first (1–61 frames) and second (61–121 frames) halves of the time-lapse movies (n = 45 cells; mean ± SEM; unpaired t-test with Welch's correction). Please click here to view a larger version of this figure.
Video 1: A representative time-lapse raw image of a prometaphase cell before the analysis. Please click here to view this video. (Right-click to download.)
Video 2: Detection and tracking of the MTs in a cell in Video 1 using the suggested settings for U-Track software. The same time-lapse image in Video 1 processed with U-Track v2.2.0 software using the described settings, and the detected growth tracks are marked in red. Please click here to view this video. (Right-click to download.)
Video 3: Detection and tracking of the MTs in a cell in Video 1 using a nonoptimal value for the "Maximum Gap Length". The same time-lapse image in Video 1 processed with U-Track v2.2.0 software using the same settings as before, but with the "Maximum Gap Length" set to 15. The rest of the parameters were not altered. Please click here to view this video. (Right-click to download.)
Video 4: Detection and tracking of the MTs in a cell in Video 1 using a nonoptimal value for the "Maximum Shrinkage Factor". The same time-lapse image in Video 1 processed with U-Track v2.2.0 software using the same settings as before, but with the "Maximum Shrinkage Factor" set to 1. The rest of the parameters were not altered. Please click here to view this video. (Right-click to download.)
Video 5: An example of a time-lapse series of a cell with cell debris. After the analysis, some cell debris was also detected by the software during MT tracking. Please click here to view this video. (Right-click to download.)
Video 6: Raw data corresponding to Video 5. Please click here to view this video. (Right-click to download.)
Video 7: An example of a time-lapse series of a cell with a high expression of EB3-tdTomato resulting in poor definition of growing tips. Please click here to view this video. (Right-click to download.)
Video 8: Raw data corresponding to Video 7. Please click here to view this video. (Right-click to download.)
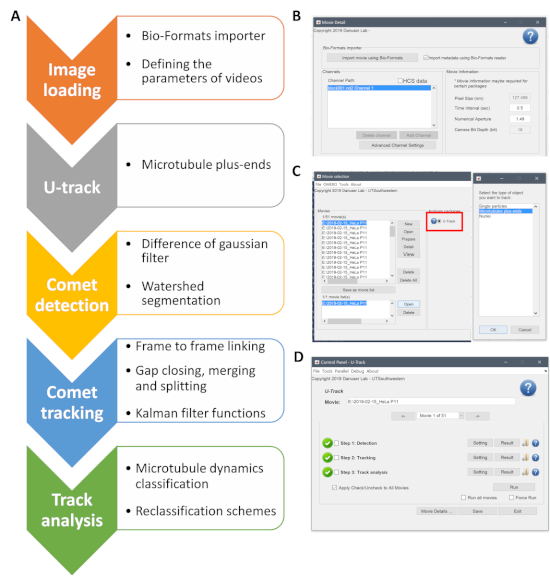
Supplementary Figure 1: The workflow of the analysis using U-Track software. (A) A schematic of the steps employed by the software. (B) A screenshot of the Bio-Formats importer showing how to import the time-lapse files. (C) After uploading the files, U-Track with the MT plus-ends module is selected. (D) A screenshot of the control panel of U-Track where the settings for comet detection, tracking, and track analysis are defined. Please click here to view a larger version of this figure.
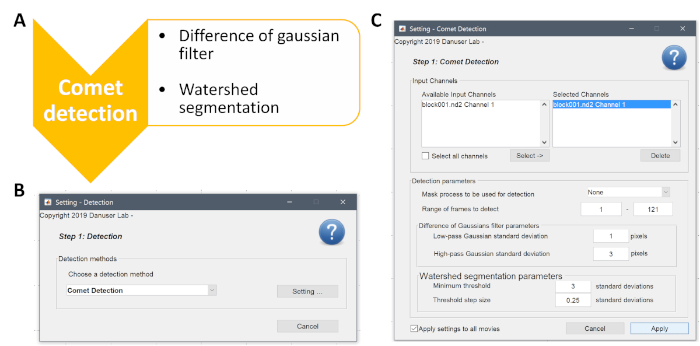
Supplementary Figure 2: Description of the first step of the analysis, the comet detection. (A) An outline of the major events performed by the algorithm. (B, C) Screenshots from the software are given with the optimal values indicated. Please click here to view a larger version of this figure.
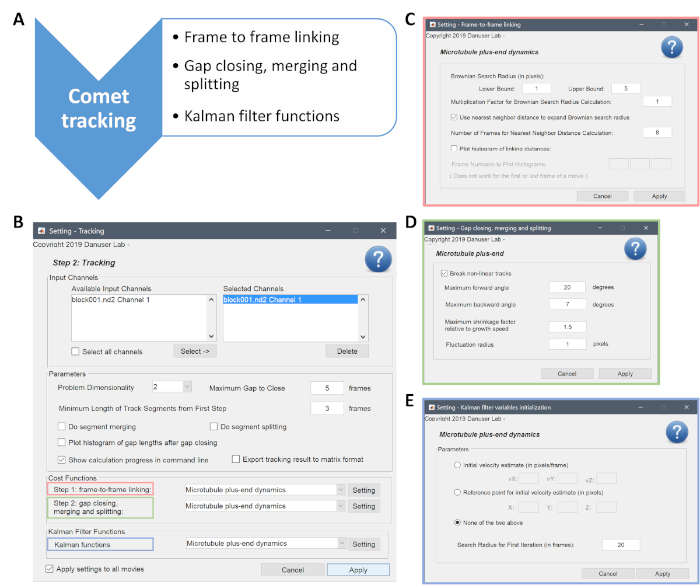
Supplementary Figure 3: Description of the second step of the analysis, the comet tracking. (A) The main steps performed by the algorithm are outlined. (B) A screenshot of the "Tracking" panel is given. The Maximum Gap Close corresponds to the Maximum Gap Length and is set to 5. The tracking of three substeps highlighted with red, green, and blue rectangles. (C,D,E) The numerical values necessary for each substep are entered here. The Maximum Shrinkage Factor is set to 1.5 as indicated in (D). Please click here to view a larger version of this figure.
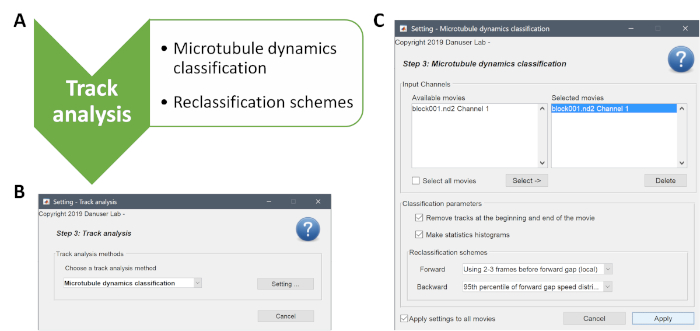
Supplementary Figure 4: Description of the last step of the analysis, track analysis. (A) The MT dynamics classification and reclassification of the compound tracks is performed during this step. (B,C) Screenshots of the track analysis and the corresponding settings are shown. Please click here to view a larger version of this figure.
Discussion
Here, we describe a modification of a method first established by Ertych et al.44. Along with several other modifications, we combine this technique of MT dynamics analysis with dual spinning disk confocal imaging. The use of the dual spinning disk improves the resolution of growing MTs while reducing phototoxicity36. We further reduce the photobleaching and laser light-induced damage of the cells by switching to a longer wavelength fluorescent reporter. The tdTomato fluorescent protein has a higher coefficient of photostability and brightness in comparison to an EGFP46. Finally, the measurement of MT dynamics is limited to only one Z-plane due to the limitations of the follow-up analysis with U-Track. The U-Track software is designed to detect the fluorescently-labelled MT tips in an XYZ-axis50,51. Therefore, taking a Z-stack time-lapse series and creating maximal projection time-lapse series is prone to generate false results. Signals detected in different Z-planes and not belonging to the same growing MT are brought together in the maximal projection, thus creating a false trajectory of MT growth.
The synchronization protocol used here induces a high density of MTs by restricting the mitotic spindle to a monopolar structure. The mitotic MTs are highly dynamic structures with phases of growth and shrinkage, with a pause at the transition between them19,54,55. Due to the high density of the growing MTs, detection of the pause events followed by either shrinkage or growth is prone to false results if the tracking parameters are set incorrectly. The U-Track software tracking modules detect so-called subtracks (episodes of continuous growth) and then classifies them as compound tracks with pause events termed "gaps". Applegate et al. discuss two parameters critical for the tracking and subtrack linking50. These are "Maximum Gap Length" and "Maximum Shrinkage Factor". If the subtrack being followed reappears in the growing direction of the MT in the subsequent time-steps, then it is classified as a forward gap. On the other hand, if the subtrack reappears opposite to the growth direction, it will be classified as a backward gap. The Maximum Gap Length defines the number of the frames to be searched for the forward and backward gaps. As mentioned, the high density and by nature high dynamicity of the mitotic MTs sets the limitation, and smaller values should be defined. As shown in Figure 1E the dynamicity is affected the most. The dynamicity is calculated as collective displacement of all gap-containing tracks over their global lifespans. The second parameter, the Maximum Shrinkage Factor, has little to no effect on either dynamicity or growth speed (Figure 1D,E).
In general, when studying MT growth properties, careful attention should be paid to the imaging conditions. First, the MTs are very sensitive to temperature and depolymerize when exposed to cold growth medium56,57,58,59. Therefore, to avoid the collection of false results, the temperature should be strictly controlled throughout the entire experiment. Second, the ionic composition of the medium used during the experiments can affect MT growth58,60. For example, exposure to calcium ions affects the MT dynamics in different ways61,62. Hence, the composition of the growth medium used in all experiments should be the same. Similarly, the parameters of the analysis should be defined once and maintained constant for all the repetitions. Additionally, the time-lapse movies generated after the analysis should be visually inspected, and any movie with background noise giving rise to false positives (Videos 5 and 6) or with high expression of tdTomato resulting in poor resolution of the MT growing tips (Videos 7 and 8) should be excluded from further statistical analysis.
Recently, the combination of lattice light-sheet microscopy of a mitotic spindles at subsecond intervals, together with sophisticated image processing allows the analysis of MT assembly rates in three dimensions63,64. This has obvious advantages over CLSM, but further improvements will be required before the method becomes of general use, such as the expansion of strategies used in U-Track to the third dimension26,50,63.
The protocol of MT dynamics detection we describe here can be a method of choice for drug screening. The method is robust, and it successfully removes human bias compared to the analysis performed manually. The automation of the movie processing allows the analysis of thousands of MT tracks within each cell, thus increasing the statistical power of the analysis. Furthermore, the method can be modified by changing, for instance, the synchronization protocol and obtaining cells from different phases of the cell cycle. This can, for example, be a useful tool for screening MT targeting chemotherapeutic drugs when the effect on interphase and dividing cells should be distinguished.
Divulgations
The authors have nothing to disclose.
Acknowledgements
We thank the members of the Light Microscopy Facility, Max-Planck Institute of Experimental Medicine, for their expert advice and support.
Materials
| Dimethylenastron | Merck | 324622 | |
| DMEM w/o phenol red | Gibco | 31053-28 | |
| DPBS | Gibco | 14190-094 | |
| Fetal bovine serum | Biochrom | S0415 | |
| Fibronectin Bovine Plasma | Merck | F4759 | Sterile powder |
| GlutaMAX | Gibco | 35050-038 | Stable glutamine substitutive |
| jetPRIME | Polyplus | 114-15 | |
| EB3-TdTomato | Addgene | plasmid #50708 | |
| RPMI 1640 | Gibco | 61870-010 | |
| Trypan Blue | Merck | T8154-20ML | |
| Trypsin/EDTA solution | Biochrom | L2143 | 0.05% / o.02 % w/o calcium and magnesium |
| µ-slide | Ibidi | 80426 | 4-well slide with #1.5 coverslip |
| Eclipse Ti Inverted microscope | Nikon | NA | |
| Objective | Nikon | MRD01991 | CFI Apo TIRF 100XC Oil |
| ACAL Laser Excahnger | Nikon | Laser box. 405, 458, 488, 514, 561 and 647nm | |
| Spinning disk module | Andor | CSU-W | |
| Camera | Andor | iXon Ultra 888 | |
| Environmental Chamber | Okolab | Dark chamber equipped with CO2 supply, tmeperature control and humidifier | |
| HeLa Cells | DSMZ | ACC-57 | |
| NIS Elements v4 | Nikon | Spinning disk microscope. Acquisition Software | |
| MATLAB | Mathworks | Computing environment | |
| Prism 8 | GraphPad | Statistical analysis and display software |
References
- Erickson, H. P. Evolution of the cytoskeleton. Bioessays. 29 (7), 668-677 (2007).
- Pollard, T. D., Goldman, R. D. Overview of the Cytoskeleton from an Evolutionary Perspective. Cold Spring Harbor Perspectives in Biology. 10 (7), (2018).
- Wade, R. H. On and around microtubules: an overview. Molecular Biotechnology. 43 (2), 177-191 (2009).
- Forth, S., Kapoor, T. M. The mechanics of microtubule networks in cell division. Journal of Cell Biology. 216 (6), 1525-1531 (2017).
- Franker, M. A., Hoogenraad, C. C. Microtubule-based transport – basic mechanisms, traffic rules and role in neurological pathogenesis. Journal of Cell Science. 126, 2319-2329 (2013).
- Lindemann, C. B., Lesich, K. A. Flagellar and ciliary beating: the proven and the possible. Journal of Cell Science. 123, 519-528 (2010).
- Wheway, G., Nazlamova, L., Hancock, J. T. Signaling through the Primary Cilium. Frontiers in Cell and Developmental Biology. 6, 8 (2018).
- Falk, N., Losl, M., Schroder, N., Giessl, A. Specialized Cilia in Mammalian Sensory Systems. Cells. 4 (3), 500-519 (2015).
- Spoon, C., Grant, W. Biomechanical measurement of kinocilium. Methods in Enzymology. 525, 21-43 (2013).
- Zenker, J., et al. A microtubule-organizing center directing intracellular transport in the early mouse embryo. Science. 357 (6354), 925-928 (2017).
- Goldstein, B. Embryonic polarity: a role for microtubules. Current Biology. 10 (22), 820-822 (2000).
- Uchida, S., Shumyatsky, G. P. Deceivingly dynamic: Learning-dependent changes in stathmin and microtubules. Neurobiology of Learning and Memory. 124, 52-61 (2015).
- Fife, C. M., McCarroll, J. A., Kavallaris, M. Movers and shakers: cell cytoskeleton in cancer metastasis. British Journal of Pharmacology. 171 (24), 5507-5523 (2014).
- Bouchet, B. P., Akhmanova, A. Microtubules in 3D cell motility. Journal of Cell Science. 130 (1), 39-50 (2017).
- Dent, E. W. Of microtubules and memory: implications for microtubule dynamics in dendrites and spines. Molecular Biology of the Cell. 28 (1), 1-8 (2017).
- Craddock, T. J., Tuszynski, J. A., Hameroff, S. Cytoskeletal signaling: is memory encoded in microtubule lattices by CaMKII phosphorylation. PLOS Computational Biology. 8 (3), (2012).
- Smythies, J. Off the beaten track: the molecular structure of long-term memory: three novel hypotheses-electrical, chemical and anatomical (allosteric). Frontiers in Integrative Neuroscience. 9, 4 (2015).
- Kaganovsky, K., Wang, C. Y. How Do Microtubule Dynamics Relate to the Hallmarks of Learning and Memory. Journal of Neuroscience. 36 (22), 5911-5913 (2016).
- Mitchison, T., Kirschner, M. Dynamic instability of microtubule growth. Nature. 312 (5991), 237-242 (1984).
- Dubey, J., Ratnakaran, N., Koushika, S. P. Neurodegeneration and microtubule dynamics: death by a thousand cuts. Frontiers in Cellular Neuroscience. 9, 343 (2015).
- Parker, A. L., Kavallaris, M., McCarroll, J. A. Microtubules and their role in cellular stress in cancer. Frontiers in Oncology. 4, 153 (2014).
- Honore, S., Pasquier, E., Braguer, D. Understanding microtubule dynamics for improved cancer therapy. Cell and Molecular Life Sciences. 62 (24), 3039-3056 (2005).
- Straube, A. . Methods in Molecular Biology. , (2011).
- Budde, P. P., Desai, A., Heald, R. Analysis of microtubule polymerization in vitro and during the cell cycle in Xenopus egg extracts. Methods. 38 (1), 29-34 (2006).
- Gierke, S., Kumar, P., Wittmann, T. Analysis of microtubule polymerization dynamics in live cells. Methods in Cell Biology. 97, 15-33 (2010).
- Matov, A., et al. Analysis of microtubule dynamic instability using a plus-end growth marker. Nature Methods. 7 (9), 761-768 (2010).
- Bailey, M., Conway, L., Gramlich, M. W., Hawkins, T. L., Ross, J. L. Modern methods to interrogate microtubule dynamics. Integrative Biology (Camb). 5 (11), 1324-1333 (2013).
- Galjart, N. Plus-end-tracking proteins and their interactions at microtubule ends. Current Biology. 20 (12), 528-537 (2010).
- Stepanova, T., et al. Visualization of microtubule growth in cultured neurons via the use of EB3-GFP (end-binding protein 3-green fluorescent protein). Journal of Neuroscience. 23 (7), 2655-2664 (2003).
- Zwetsloot, A. J., Tut, G., Straube, A. Measuring microtubule dynamics. Essays in Biochemistry. 62 (6), 725-735 (2018).
- Bayguinov, P. O., et al. Modern Laser Scanning Confocal Microscopy. Current Protocols in Cytometry. 85 (1), 39 (2018).
- Nakano, A. Spinning-disk confocal microscopy — a cutting-edge tool for imaging of membrane traffic. Cell Structure and Function. 27 (5), 349-355 (2002).
- Nipkow, P. Elektrisches teleskop. Germany patent. , (1884).
- Yin, S., Lu, G., Zhang, J., Yu, F. T., Mait, J. N. Kinoform-based Nipkow disk for a confocal microscope. Applied Optics. 34 (25), 5695-5698 (1995).
- Tanaami, T., Kenta, M. Nipkow disk for confocal optical scanner. European patent application. , (1992).
- Oreopoulos, J., Berman, R., Browne, M. Spinning-disk confocal microscopy: present technology and future trends. Methods in Cell Biology. 123, 153-175 (2014).
- Rusan, N. M., Fagerstrom, C. J., Yvon, A. M., Wadsworth, P. Cell cycle-dependent changes in microtubule dynamics in living cells expressing green fluorescent protein-alpha tubulin. Molecular Biology of the Cell. 12 (4), 971-980 (2001).
- Rusan, N. M., Fagerstrom, C. J., Yvon, A. -. M. C., Wadsworth, P. Cell Cycle-Dependent Changes in Microtubule Dynamics in Living Cells Expressing Green Fluorescent Protein-α Tubulin. Molecular Biology of the Cell. 12 (4), 971-980 (2001).
- Liu, D., Davydenko, O., Lampson, M. A. Polo-like kinase-1 regulates kinetochore-microtubule dynamics and spindle checkpoint silencing. Journal of Cell Biology. 198 (4), 491-499 (2012).
- Maiato, H., Sunkel, C. E. Kinetochore-microtubule interactions during cell division. Chromosome Research. 12 (6), 585-597 (2004).
- Muller, C., et al. Inhibitors of kinesin Eg5: antiproliferative activity of monastrol analogues against human glioblastoma cells. Cancer Chemotherrapy and Pharmacology. 59 (2), 157-164 (2007).
- Mayer, T. U., et al. Small molecule inhibitor of mitotic spindle bipolarity identified in a phenotype-based screen. Science. 286 (5441), 971-974 (1999).
- Kapoor, T. M., Mayer, T. U., Coughlin, M. L., Mitchison, T. J. Probing spindle assembly mechanisms with monastrol, a small molecule inhibitor of the mitotic kinesin Eg5. The Journal of Cell Biology. 150 (5), 975-988 (2000).
- Ertych, N., et al. Increased microtubule assembly rates influence chromosomal instability in colorectal cancer cells. Nature Cell Biology. 16 (8), 779-791 (2014).
- Brito, D. A., Yang, Z., Rieder, C. L. Microtubules do not promote mitotic slippage when the spindle assembly checkpoint cannot be satisfied. The Journal of Cell Biology. 182 (4), 623-629 (2008).
- Shaner, N. C., Patterson, G. H., Davidson, M. W. Advances in fluorescent protein technology. Journal of Cell Science. 120 (24), 4247-4260 (2007).
- Phelan, M. C., Lawler, G. Cell Counting. Current Protocols in Cytometry. 00 (1), 3 (1997).
- Merriam, E. B., et al. Synaptic regulation of microtubule dynamics in dendritic spines by calcium, F-actin, and drebrin. Journal of Neuroscience. 33 (42), 16471-16482 (2013).
- Samora, C. P., et al. MAP4 and CLASP1 operate as a safety mechanism to maintain a stable spindle position in mitosis. Nature Cell Biology. 13 (9), 1040-1050 (2011).
- Applegate, K. T., et al. plusTipTracker: Quantitative image analysis software for the measurement of microtubule dynamics. Journal of Structural Biology. 176 (2), 168-184 (2011).
- Jaqaman, K., et al. Robust single-particle tracking in live-cell time-lapse sequences. Nature Methods. 5 (8), 695-702 (2008).
- Stout, A., D’Amico, S., Enzenbacher, T., Ebbert, P., Lowery, L. A. Using plusTipTracker Software to Measure Microtubule Dynamics in Xenopus laevis Growth Cones. Journal of Visualized Experiments. , e52138 (2014).
- Linkert, M., et al. Metadata matters: access to image data in the real world. The Journal of Cell Biology. 189 (5), 777-782 (2010).
- Brouhard, G. J. Dynamic instability 30 years later: complexities in microtubule growth and catastrophe. Molecular Biology of the Cell. 26 (7), 1207-1210 (2015).
- Burbank, K. S., Mitchison, T. J. Microtubule dynamic instability. Current Biology : CB. 16 (14), 516-517 (2006).
- Caplow, M., Shanks, J., Ruhlen, R. L. Temperature-jump studies of microtubule dynamic instability. Journal of Biological Chemistry. 263 (21), 10344-10352 (1988).
- Prasad, V., Jordan, M. A., Luduena, R. F. Temperature sensitivity of vinblastine-induced tubulin polymerization in the presence of microtubule-associated proteins. Journal of Protein Chemistry. 11 (5), 509-515 (1992).
- Wasteneys, G. O. Microtubules Show their Sensitive Nature. Plant and Cell Physiology. 44 (7), 653-654 (2003).
- Turi, A., Lu, R. C., Lin, P. -. S. Effect of heat on the microtubule disassembly and its relationship to body temperatures. Biochemical and Biophysical Research Communications. 100 (2), 584-590 (1981).
- Safinya, C. R., et al. The effect of multivalent cations and Tau on paclitaxel-stabilized microtubule assembly, disassembly, and structure. Advances in Colloid and Interface Science. 232, 9-16 (2016).
- Sandoval, I. V., Weber, K. Calcium-Induced Inactivation of Microtubule Formation in Brain Extracts. European Journal of Biochemistry. 92 (2), 463-470 (1978).
- Vater, W., Böhm, K. J., Unger, E. Tubulin assembly in the presence of calcium ions and taxol: Microtubule bundling and formation of macrotubule-ring complexes. Cell Motility. 36 (1), 76-83 (1997).
- Yamashita, N., et al. Three-dimensional tracking of plus-tips by lattice light-sheet microscopy permits the quantification of microtubule growth trajectories within the mitotic apparatus. Journal of Biomedical Optics. 20 (10), 1-18 (2015).
- Pamula, M. C., et al. High-resolution imaging reveals how the spindle midzone impacts chromosome movement. Journal of Cell Biology. 218 (8), 2529-2544 (2019).

