Epithelial Cell Infection Analyses with Shigella
Summary
The present protocol describes infection assays to interrogate Shigella adherence, invasion, and intracellular replication using in vitro epithelial cell lines.
Abstract
The human-adapted enteric bacterial pathogen Shigella causes millions of infections each year, creates long-term growth effects among pediatric patients, and is a leading cause of diarrheal deaths worldwide. Infection induces watery or bloody diarrhea as a result of the pathogen transiting the gastrointestinal tract and infecting the epithelial cells lining the colon. With staggering increases in antibiotic resistance and the current lack of approved vaccines, standardized research protocols are critical to studying this formidable pathogen. Here, methodologies are presented to examine the molecular pathogenesis of Shigella using in vitro analyses of bacterial adherence, invasion, and intracellular replication in colonic epithelial cells. Prior to infection analyses, the virulence phenotype of Shigella colonies was verified by the uptake of the Congo red dye on agar plates. Supplemented laboratory media can also be considered during bacterial culturing to mimic in vivo conditions. Bacterial cells are then used in a standardized protocol to infect colonic epithelial cells in tissue culture plates at an established multiplicity of infection with adaptations to analyze each stage of infection. For adherence assays, Shigella cells are incubated with reduced media levels to promote bacterial contact with epithelial cells. For both invasion and intracellular replication assays, gentamicin is applied for various time intervals to eliminate extracellular bacteria and enable assessment of invasion and/or the quantification of intracellular replication rates. All infection protocols enumerate adherent, invaded, and/or intracellular bacteria by serially diluting infected epithelial cell lysates and plating bacterial colony forming units relative to infecting titers on Congo red agar plates. Together, these protocols enable independent characterization and comparisons for each stage of Shigella infection of epithelial cells to study this pathogen successfully.
Introduction
Diarrheal diseases caused by enteric bacterial pathogens are a significant global health burden. In 2016, diarrheal diseases were responsible for 1.3 million deaths worldwide and were the fourth leading cause of death in children younger than five years of age1,2. The Gram-negative, enteric bacterial pathogen Shigella is the causative agent of shigellosis, a major cause of diarrheal deaths worldwide3. Shigellosis causes significant morbidity and mortality each year in children from lower- and middle-income countries4,5, while infections in high-income countries are linked to daycare center, foodborne, and waterborne outbreaks6,7,8,9. Ineffective vaccine development10 and rising rates of antimicrobial resistance (AMR)11,12 have complicated the management of large-scale Shigella outbreaks. Recent Centers for Disease Control and Prevention data show that nearly 46% of Shigella infections in the United States displayed drug resistance in 202013,14, while the World Health Organization has declared Shigella as an AMR priority pathogen for which new therapies are urgently needed15.
Shigella infections are easily transmitted via the fecal-oral route upon ingestion of contaminated food or water, or through direct human contact. Shigella has evolved to be an efficient, human-adapted pathogen, with an infectious dose of 10-100 bacteria sufficient to cause disease16. During small intestinal transit, Shigella is exposed to environmental signals, such as elevated temperature and bile17. Detection of these signals induces transcriptional changes to express virulence factors that enhance the ability of the bacteria to infect the human colon17,18,19. Shigella does not invade the colonic epithelium from the apical surface, but rather transits across the epithelial layer following uptake into specialized antigen-presenting microfold cells (M cells) within the follicle-associated epithelium20,21,22. Following transcytosis, Shigella cells are phagocytosed by resident macrophages. Shigella rapidly escapes the phagosome and triggers macrophage cell death, resulting in the release of pro-inflammatory cytokines5,23,24. Shigella then invades colonic epithelial cells from the basolateral side, lyses the macropinocytic vacuole, and establishes a replicative niche in the cytoplasm5,25. Pro-inflammatory cytokines, particularly interleukin-8 (IL-8), recruit polymorphonuclear neutrophil leukocytes (PMNs) to the site of infection, which weakens epithelial tight junctions, and enables bacterial infiltration of the epithelial lining to exacerbate basolateral infection5. The PMNs destroy the infected epithelial lining to contain the infection, which results in the characteristic symptoms of bacillary (bloody) dysentery5. Although invasion and intracellular replication mechanisms have been thoroughly characterized, new research is demonstrating important new concepts in Shigella infection, including virulence regulation during gastrointestinal (GI) transit17, adherence19, improved basolateral access through barrier permeability26, and asymptomatic carriage in malnourished children27.
The ability of Shigella spp. to cause diarrheal disease is restricted to humans and non-human primates (NHP)28. Shigella intestinal infection models have been developed for zebrafish29, mice30, guinea pigs31, rabbits21,32,33, and pigs34,35. However, none of these model systems can accurately replicate the disease characteristics observed during human infection36. Although NHP models of shigellosis have been established to study Shigella pathogenesis, these model systems are expensive to implement and require artificially high infectious doses, up to nine orders of magnitude higher than the infectious dose of humans37,38,39,40,41,42. Thus, the remarkable adaptation of Shigella for infection of human hosts necessitates the use of human-derived cell cultures to recreate physiologically relevant models for accurate interrogation of Shigella pathogenesis.
Here, detailed procedures are described to measure the rates of Shigella adherence to, invasion of, and replication within HT-29 colonic epithelial cells. Using these standardized protocols, the molecular mechanisms by which bacterial virulence genes and environmental signals impact each step of Shigella infection can be interrogated to better understand the dynamic host-pathogen interaction relationship.
Protocol
1. Preparation of reagents and materials
NOTE: All volumes are consistent with an assay using two 6-well plates.
- TSB medium: Add 0.5 L of deionized (DI) water to 15 g of Tryptic Soy Broth (TSB, see Table of Materials) medium and autoclave. Store at room temperature.
- Bile salts medium (TSB + BS): To prepare TSB containing 0.4% (w/v) bile salts, resuspend 0.06 g of bile salts (BS, see Table of Materials) in 15 mL of autoclaved TSB. Filter sterilize using a 0.22 µm PES filter.
NOTE: The bile salts consist of a 1:1 mixture of sodium cholate and sodium deoxycholate. Prepare fresh media immediately before use. - DMEM + 10% (v/v) FBS: Add 5 mL of fetal bovine serum (FBS) to 45 mL of Dulbecco's Modified Eagle Medium (DMEM). Store at 4 °C.
- DMEM + gentamicin: To a 50 mL tube, add 50 mL of DMEM and 50 µL of 50 mg/mL gentamicin (see Table of Materials).
NOTE: Make fresh aliquot and warm in a 37 °C water bath prior to each experiment. - PBS + 1% (v/v) Triton X-100: Add 150 µL of Triton X-100 to 15 mL of Phosphate-buffered saline (PBS).
NOTE: Make fresh aliquot and warm in a 37 °C water bath prior to each experiment. - TSB + Congo red indicator plates: Add 15 g of TSB, 7.5 g of select agar, and 0.125 g of Congo red dye (see Table of Materials) to a 1 L bottle. Add 0.5 L of DI water and autoclave. Pour 10-20 mL of media into individual sterile Petri dishes (100 mm x 15 mm) and let solidify.
CAUTION: Congo red is carcinogenic and a reproductive toxin. Ensure that handling of Congo red is performed using the appropriate personal protective equipment. Consult the product safety data sheet for additional information.
NOTE: Approximately 20 plates are made from 0.5 L Congo red media. Plates can be prepared 2-3 days in advance and left inverted at room temperature until use. For long-term storage, place inverted plates in plastic sleeves at 4 °C for up to 3 months. - DMEM + 10% (v/v) FBS and 5% (v/v) dimethyl sulfoxide (DMSO): Add 42.5 mL of DMEM, 5 mL of FBS, and 2.5 mL of DMSO to a 50 mL tube. Store at 4 °C.
2. Preparation of bacteria
NOTE: All Shigella laboratory cultivation and storage protocols are adapted from Payne, S. M.43.
CAUTION: Shigella spp. are Risk Group 2 pathogens44. Perform all laboratory work in a BSL-2 environment, with additional safety measures undertaken to limit accidental exposures due to the low infectious dose of Shigella spp.
- Growth of Shigella from frozen stocks
- Transfer a small amount of frozen culture from the cryogenic vial to a TSB + Congo red agar plate using a sterile applicator.
- Flame sterilize an inoculating loop and allow it to cool. Streak inoculum back and forth across one quadrant of the plate. Flame the loop, allow it to cool, then streak from the first quadrant onto the second quadrant of the plate. Repeat to streak inoculum into the third and fourth quadrants of the plate.
NOTE: Alternatively, streak inoculum using a fresh sterile applicator between each quadrant. - Invert the plate and incubate at 37 °C overnight.
NOTE: Incubation at temperatures ≥37 °C is required for expression of Shigella virulence factors necessary for observation of Congo red-positive (CR+) phenotype45. Avirulent colonies will have a white appearance and will not be invasive. - Seal the plate with paraffin film and store refrigerated at 4 °C.
NOTE: Bacterial colonies will remain viable on agar plates for 1-2 weeks.
- Overnight growth of Shigella in liquid culture
- Aliquot 3 mL of TSB media into sterile 14 mL culture tubes.
- Pick a single, well-isolated red (CR+) colony using a sterile applicator and resuspend in liquid media.
- Incubate cultures overnight (16-18 h) at 37 °C with shaking at 250 rotations per minute (rpm).
3. Preparation of HT-29 eukaryotic cells
NOTE: All volumes are consistent with an assay using two 6-well plates. HT-29 cell lines were acquired from the American Type Culture Collection (ATCC). HT-29 maintenance protocols are adapted from ATCC recommendations46. All media should be pre-warmed in a water bath at 37 °C prior to use. All HT-29 maintenance protocols should be performed in a biosafety cabinet. Refrain from producing bubbles when mixing/working with HT-29 cells in media to avoid dramatic changes in pH.
- Thawing HT-29 cells from frozen stock
- Thaw the vial of HT-29 cells in a 37 °C water bath.
NOTE: Ensure the cap stays fully above the water to avoid contamination. Thawing should take less than 2 min. - Remove the vial from the water immediately after the culture is fully thawed and decontaminate with 70% ethanol. Ensure that all steps from this point are performed using aseptic techniques.
- Add all the contents of the vial to a 15 mL centrifuge tube containing 9 mL of DMEM + 10% FBS. Centrifuge at 125 x g for 5 min at room temperature.
- Decant the supernatant into a waste container and resuspend the pellet in 10 mL of warm DMEM + 10% FBS. Transfer resuspended cells to a 75 cm2 tissue culture flask (T75) containing 10 mL of warm DMEM + 10% FBS (total volume of 20 mL).
- Incubate cells at 37 °C with 5% CO2 until the cells reach 90% confluency (approximately 6-7 days).
NOTE: Confluency is estimated through visual approximation.
- Thaw the vial of HT-29 cells in a 37 °C water bath.
- Seeding HT-29 cells
- Pre-warm 20 mL of PBS and 50 mL of DMEM + 10% FBS in a 37 °C water bath and pre-warm 3 mL of 0.25% (w/v) Trypsin-EDTA to room temperature.
- Once HT-29 cells (from step 3.1) reach 90% confluency, decant HT-29 cell culture media from the T75 flask into a waste container. Pour ~10 mL of warm PBS into the flask and swirl gently to wash. Decant the PBS into a waste container. Wash with warm PBS again and decant.
- Add 2-3 mL of 0.25% (w/v) Trypsin-EDTA and gently swirl across the entire surface area. Incubate at 37 °C with 5% CO2 for 4 min.
- Remove the flask from the incubator and gently swirl the Trypsin-EDTA, visually ensuring that all cells detach from the surface.
- Immediately add 6 mL of warm DMEM + 10% FBS to deactivate the Trypsin. Pipette up and down to thoroughly mix.
- Transfer all the contents to a 15 mL centrifuge tube and spin it at 500 x g for 5 min at room temperature.
- Gently decant supernatant into a waste container and resuspend the pellet in 6 mL of warm DMEM + 10% FBS.
- Immediately after resuspension, transfer 10 µL of suspended HT-29 cells from the middle of the culture to a 0.2 mL PCR tube. Add 10 µL of Trypan blue dye to the PCR tube and mix.
- Add 10 µL of HT-29 cell/Trypan blue mix to a disposable Countess cell counter chamber slide (see Table of Materials). Enumerate the number of live cells and calculate cell viability.
NOTE: When documenting the number of cells in the sample, read the number under the "live" cell count, not the total cell count. Alternatively, cell enumeration can be performed manually using a hemocytometer. - Seed resuspended HT-29 cells into a fresh T75 flask or 6-well plate.
- For T75 flask:
- Pipet gently to mix, then transfer 2.5 x 106 cells to a fresh T75 flask according to the equation below:
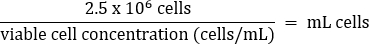
- Add warm DMEM + 10% FBS media to a final volume of 20 mL (final concentration of 1.25 x 105 cells/mL).
- Disperse cells evenly across the flask by gently rocking back and forth.
- Incubate at 37 °C with 5% CO2 until cells reach 80% confluency.
NOTE: For optimal growth, replace the DMEM + 10% FBS media in the T75 flask every ~3 days. Decant media into a waste container and add 10 mL of warm PBS to the flask. Swirl the PBS around gently and decant it into the waste container. Then add 20 mL of fresh, warm DMEM + 10% FBS to the flask and return to the 37 °C, 5% CO2 incubator.
- Pipet gently to mix, then transfer 2.5 x 106 cells to a fresh T75 flask according to the equation below:
- For 6-well plate:
- Pipet gently to mix, then transfer 5.85 x 106 cells to a fresh 50 mL conical tube according to the equation below:
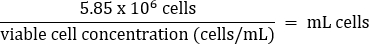
- Add warm DMEM + 10% FBS media to a final volume of 26 mL (final concentration of 2.25 x 105 cells/mL).
- Pipet gently to mix, then dispense 2 mL (4.5 x 105 cells) into individual wells of 6-well plates.
- Disperse cells evenly across the well by gently rocking up/down and left/right 2-3x.
- Incubate at 37 °C with 5% CO2 until cells reach 80%-95% confluency (approximately 3-4 days).
NOTE: 85% confluency is recommended for invasion and intracellular replication assays, while 90%-95% confluency is recommended for adherence assays. Cells should reach ~85% confluency after 48 h incubation with a final concentration of approximately 1 x 106 cells/well. Adjustments to the number of cells seeded and length of incubation may be required.
- Pipet gently to mix, then transfer 5.85 x 106 cells to a fresh 50 mL conical tube according to the equation below:
- For T75 flask:
- Making frozen HT-29 stocks
- Aliquot 1 mL of DMEM + 10% FBS + 5% DMSO media into individual cryogenic vials.
- Add 1 x 106 HT-29 cells from step 3.2.7 to each vial. Calculate the volume of cells according to the formula below:
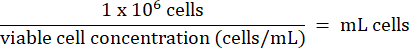
- Store HT-29 cells long-term below -130 °C in liquid nitrogen vapor storage freezer.
4. Adherence assay
NOTE: All volumes are consistent with an assay using two 6-well plates.
- Subculture overnight Shigella cultures via 1:50 dilution into fresh media.
- Vortex, then add 100 µL of each overnight culture to 5 mL of fresh TSB or TSB + BS in a suitably sized culture tube.
NOTE: Limit culture volume to <20% of culture flask or tube volume to ensure proper aeration. - Incubate at 37 °C with shaking at 250 rpm until cells reach an optical density (OD600) of 0.7 (mid-log phase of Shigella growth); about 2-2.5 h.
NOTE: During the subculture, aliquot 50 mL of DMEM and a sufficient volume of PBS for all washing steps and place in a 37 °C water bath. Allow media to reach 37 °C before use.
- Vortex, then add 100 µL of each overnight culture to 5 mL of fresh TSB or TSB + BS in a suitably sized culture tube.
- Transfer 2 x 108 colony forming units (CFUs) subcultured Shigella to individual 2 mL microcentrifuge tubes.
NOTE: 2 x 108 CFUs corresponds to approximately 1 mL of bacterial cells at an OD600 of 0.7. Use OD600 readings to approximate CFU/mL according to the calibration of each individual spectrophotometer. - Wash each Shigella sample 2x with PBS.
- Pellet cells by centrifugation at 17,000 x g for 2 min at room temperature. Aspirate the supernatant, then add 1 mL of warm PBS and resuspend the pellet well, gently pipetting the sample up and down until the mixture is fully homogeneous (8-10x).
- Repeat step 4.3.1 1x additional time.
- Pellet cells by centrifugation at 17,000 x g for 2 min at room temperature, aspirate the supernatant, and resuspend the pellets in 2 mL of warm DMEM.
NOTE: The final concentration of resuspended bacteria will be 1 x 108 CFU/mL.
- Vortex, then add 1 mL (1 x 108 CFUs) of resuspended Shigella to each well of the prepared HT-29 colonic epithelial monolayers in 6-well plates (from step 3.2.10.2).
NOTE: Infections are normally performed at a multiplicity of infection (MOI; ratio of bacterial to epithelial cells) of 100. To test different MOIs, dilute resuspended Shigella in warm DMEM to the desired concentration, then add 1 mL of diluted bacteria to HT-29 monolayers. For example, to test an MOI of 10, dilute bacteria 1:10 by adding 150 µL of 1 x 108 CFU/mL bacteria to 1.35 mL of warm DMEM, then apply 1 mL (1 x 107 CFUs) to HT-29 cells. - Incubate the 6-well plates at 37 °C with 5% CO2 for 3 h.
- During the incubation, determine the bacterial infection titer.
- Prepare 10-fold serial dilutions of resuspended Shigella cells (from step 4.3.3) into PBS.
- Plate 100 µL of the 1 x 10-5 and 1 x 10-6 dilutions onto TSB + Congo red plates and incubate overnight at 37 °C.
NOTE: Plating 100 µL from the 1 x 10-5 and 1 x 10-6 dilutions corresponds to a final dilution factor of 1 x 10-6 and 1 x 10-7, respectively.
- After incubation, wash the monolayers 4-5x with PBS.
- Aspirate media from each well.
NOTE: When aspirating media from 6-well plates, guide the tip of the aspirator along the bottom side of the wells, trying to avoid contact with the HT-29 cells. - Add 1 mL of warm PBS to each well and wash gently.
NOTE: To gently wash 6-well monolayers with PBS, move the plate up and down and side to side on the benchtop. Washing plates in a circular motion and/or removing the plate from the benchtop surface can cause the mechanical removal of cells from plastic. - Repeat steps 4.7.1 and 4.7.2 4x additional times.
- Aspirate media from each well.
- Remove PBS by aspiration and lyse HT-29 cells by adding 1 mL of PBS + 1% Triton X-100 to each well.
- Incubate 6-well plates at 37 °C for 5 min.
- Use a cell scraper or a bent pipette tip to scrape the lysed cells from the bottom of the well and transfer the full 1 mL into a fresh 1.7 mL microcentrifuge tube.
- Determine the number of cell-associated bacteria.
- Vortex each tube (from step 4.10) for at least 30 s to further displace Shigella from the lysed eukaryotic cells.
- Prepare 10-fold serial dilutions of lysates into PBS.
- Plate 100 µL of the 1 x 10-2, 1 x 10-3, and 1 x 10-4 dilutions onto TSB + Congo red plates and incubate overnight at 37 °C.
NOTE: Plating 100 µL from the 1 x 10-2, 1 x 10-3, and 1 x 10-4 dilutions corresponds to a final dilution factor of 1 x 10-3, 1 x 10-4, and 1 x 10-5, respectively.
5. Invasion assay
NOTE: All volumes are consistent with an assay using two 6-well plates.
- Subculture overnight Shigella cultures via 1:50 dilution into fresh media.
- Vortex, then add 100 µL of each overnight culture to 5 mL of fresh TSB or TSB + BS in a suitably sized culture tube.
NOTE: Limit culture volume to <20% of culture flask or tube volume to ensure proper aeration. - Incubate at 37 °C with shaking at 250 rpm until cells reach an OD600 of 0.7 (mid-log phase of Shigella growth); about 2-2.5 h.
NOTE: During the subculture, aliquot 50 mL of DMEM + 50 mg/mL gentamicin and a sufficient volume of PBS for all washing steps and place in a 37 °C water bath. Allow media to reach 37 °C before use.
- Vortex, then add 100 µL of each overnight culture to 5 mL of fresh TSB or TSB + BS in a suitably sized culture tube.
- Transfer 2 x 108 CFUs subcultured Shigella to individual 2 mL microcentrifuge tubes.
NOTE: 2 x 108 CFUs corresponds to approximately 1 mL of bacterial cells at an OD600 of 0.7. Use OD600 readings to approximate CFU/mL according to the calibration of each individual spectrophotometer. - Wash Shigella samples 1x with PBS.
- Pellet cells by centrifugation at 17,000 x g for 2 min at room temperature. Aspirate the supernatant, then add 1 mL of warm PBS and resuspend the pellet well, gently pipetting the sample up and down until the mixture is fully homogeneous (8-10x).
- Repeat step 5.3.1. 1x additional time.
- Pellet cells by centrifugation at 17,000 x g for 2 min at room temperature, aspirate the supernatant, and resuspend the pellets in 2 mL of warm DMEM.
NOTE: The final concentration of resuspended bacteria will be 1 x 108 CFU/mL.
- Vortex, then add 1 mL (1 x 108 CFUs) of resuspended Shigella plus 1 mL of DMEM to each well of the prepared HT-29 colonic epithelial monolayers in 6-well plates (from step 3.2.10.2).
NOTE: Infections are normally performed at a multiplicity of infection (MOI; ratio of bacterial to epithelial cells) of 100. To test different MOIs, dilute resuspended Shigella in DMEM to the desired concentration, then add 1 mL of diluted bacteria to HT-29 monolayers. For example, to test an MOI of 10, dilute bacteria 1:10 by adding 150 µL of 1 x 108 CFU/mL bacteria to 1.35 mL of DMEM, then add 1 mL (1 x 107 CFUs) to HT-29 cells. - To promote bacterial contact with the HT-29 cells, centrifuge the 6-well plates at 2,000 x g for 10 min at room temperature or 37 °C if the temperature setting can be adjusted.
NOTE: Centrifugation promotes bacterial contact with the HT-29 cells, which bypasses the need for adherence factors and allows the bacteria to quickly invade the cells. - Incubate 6-well plates at 37 °C with 5% CO2 for 45 min.
- During the incubation, determine the bacterial infection titer.
- Prepare 10-fold serial dilutions of resuspended Shigella cells (from step 5.3.3) into PBS.
- Plate 100 µL of the 1 x 10-5 and 1 x 10-6 dilutions onto TSB + Congo red plates and incubate overnight at 37 °C.
NOTE: Plating 100 µL from the 1 x 10-5 and 1 x 10-6 dilutions corresponds to a final dilution factor of 1 x 10-6 and 1 x 10-7, respectively.
- Thoroughly wash infected HT-29 cells 3x with 1 mL of PBS.
- Aspirate media from each well.
NOTE: When aspirating media from 6-well plates, guide the tip of the aspirator along the bottom side of the wells, trying to avoid contact with the HT-29 cells. - Add 1 mL of warm PBS to each well and wash gently.
NOTE: To gently wash 6-well monolayers with PBS, move the plate up and down and side to side on the benchtop. Washing plates in a circular motion and/or removing the plate from the benchtop surface can cause the mechanical removal of cells from plastic. - Repeat steps 5.8.1 and 5.8.2 2x additional times.
- Aspirate media from each well.
- Remove PBS by aspiration, then add 2 mL of warm DMEM supplemented with 50 µg/mL gentamicin to each well and incubate for 30 min at 37 °C with 5% CO2.
- Thoroughly wash infected HT-29 cells 3x with 1 mL of PBS.
- Repeat washing step 5.8.
- Remove PBS by aspiration, then add 2 mL of warm DMEM supplemented with 50 µg/mL gentamicin to each well and incubate for 60 min at 37 °C with 5% CO2.
- Thoroughly wash infected HT-29 cells 3x with 1 mL of PBS.
- Repeat washing step 5.8.
- Remove PBS by aspiration and lyse HT-29 cells by adding 1 mL of PBS + 1% Triton X-100 to each well.
- Incubate 6-well plates at 37 °C for 5 min.
- Use a cell scraper or a bent pipette tip to scrape the lysed cells from the bottom of the well and transfer the full 1 mL into a fresh 1.7 mL microcentrifuge tube.
- Determine the number of intracellular bacteria.
- Vortex each tube (from step 5.15) for at least 30 s to further displace Shigella from the lysed eukaryotic cells.
- Prepare 10-fold serial dilutions of lysates into PBS.
- Plate 100 µL of the 1 x 10-2 and 1 x 10-3 dilutions onto TSB + Congo red plates and incubate overnight at 37 °C.
NOTE: Plating 100 µL from the 1 x 10-2 and 1 x 10-3 dilutions corresponds to a final dilution factor of 1 x 10-3 and 1 x 10-4, respectively.
6. Intracellular replication assay
NOTE: All volumes are consistent with an assay using two 6-well plates.
- Subculture overnight Shigella cultures via 1:50 dilution into fresh media.
- Vortex, then add 100 µL of each overnight culture to 5 mL of fresh TSB or TSB + BS in a suitably sized culture tube.
NOTE: Limit culture volume to <20% of culture flask or tube volume to ensure proper aeration. - Incubate at 37 °C with shaking at 250 rpm until cells reach an OD600 of 0.7 (mid-log phase of Shigella growth); about 2-2.5 h.
NOTE: During the subculture, aliquot 50 mL of DMEM + 50 mg/mL gentamicin and a sufficient volume of PBS for all washing steps and place in a 37 °C water bath. Allow media to reach 37 °C before use.
- Vortex, then add 100 µL of each overnight culture to 5 mL of fresh TSB or TSB + BS in a suitably sized culture tube.
- Transfer 2 x 108 CFUs subcultured Shigella to individual 2 mL microcentrifuge tubes.
NOTE: 2 x 108 CFUs corresponds to approximately 1 mL of bacterial cells at an OD600 of 0.7. Use OD600 readings to approximate CFU/mL according to the calibration of each individual spectrophotometer. - Wash Shigella samples 1x with PBS.
- Pellet cells by centrifugation at 17,000 x g for 2 min at room temperature. Aspirate the supernatant, then add 1 mL of warm PBS and resuspend the pellet well, gently pipetting the sample up and down until the mixture is fully homogeneous (8-10x).
- Repeat step 6.3.1. 1x additional time.
- Pellet cells by centrifugation at 17,000 x g for 2 min at room temperature, aspirate the supernatant, and resuspend the pellets in 2 mL of warm DMEM.
NOTE: The final concentration of resuspended bacteria will be 1 x 108 CFU/mL.
- Vortex, then add 1 mL (1 x 108 CFUs) of resuspended Shigella plus 1 mL of DMEM to each well of prepared HT-29 colonic epithelial monolayers in 6-well plates (from step 3.2.10.2).
NOTE: Infections are normally performed at a multiplicity of infection (MOI; ratio of bacterial to epithelial cells) of 100. To test different MOIs, dilute resuspended Shigella in DMEM to the desired concentration, then add 1 mL of diluted bacteria to HT-29 monolayers. For example, to test an MOI of 10, dilute bacteria 1:10 by adding 150 µL of 1 x 108 CFU/mL bacteria to 1.35 mL of DMEM, then apply 1 mL (1 x 107 CFUs) to HT-29 cells. - To promote bacterial contact with the HT-29 cells, centrifuge the 6-well plates at 2,000 x g for 10 min at room temperature or 37 °C if the temperature setting can be adjusted.
NOTE: Centrifugation promotes bacterial contact with the HT-29 cells, which bypasses the need for adherence factors and allows the bacteria to quickly invade the cells. - Incubate 6-well plates at 37 °C with 5% CO2 for 45 min.
- During the incubation, determine the bacterial infection titer.
- Prepare 10-fold serial dilutions of resuspended Shigella cells (from step 6.3.3) into PBS.
- Plate 100 µL of the 1 x 10-5 and 1 x 10-6 dilutions onto TSB + Congo red plates and incubate overnight at 37 °C.
NOTE: Plating 100 µL from the 1 x 10-5 and 1 x 10-6 dilutions corresponds to a final dilution factor of 1 x 10-6 and 1 x 10-7, respectively.
- Thoroughly wash infected HT-29 cells 3x with 1 mL of PBS.
- Aspirate media from each well.
NOTE: When aspirating media from 6-well plates, guide the tip of the aspirator along the bottom side of the wells, trying to avoid contact with the HT-29 cells. - Add 1 mL of warm PBS to each well and wash gently.
NOTE: To gently wash 6-well monolayers with PBS, move the plate up and down and side to side on the benchtop. Washing plates in a circular motion and/or removing the plate from the benchtop surface can cause the mechanical removal of cells from plastic. - Repeat steps 6.8.1 and 6.8.2 2x additional times.
- Aspirate media from each well.
- Remove PBS by aspiration, then add 2 mL of warm DMEM supplemented with 50 µg/mL gentamicin to each well and incubate for 30 min at 37 °C with 5% CO2.
- Thoroughly wash infected HT-29 cells 3x with 1 mL of PBS.
- Repeat washing step 6.8.
- Remove PBS by aspiration, then add 2 mL of warm DMEM with 50 µg/mL gentamicin to each well of the 6-well plates and incubate at 37 °C with 5% CO2 for the desired length of time to allow for intracellular replication (up to 24 h).
- Thoroughly wash cells 2x with 1 mL of PBS.
- Repeat washing step 6.8.
- Remove PBS by aspiration and lyse HT-29 cells by adding 1 mL of PBS + 1% Triton X-100 to each well.
- Incubate 6-well plates at 37 °C for 5 min.
- Use a cell scraper or bent pipette tip to scrape the lysed cells from the bottom of the well and transfer the full 1 mL into a fresh 1.7 mL microcentrifuge tube.
- Determine the number of intracellular bacteria.
- Vortex each tube (from step 6.15) for at least 30 s to further displace Shigella from the lysed eukaryotic cells.
- Prepare 10-fold serial dilutions of lysates into PBS.
- Plate 100 µL of the 1 x 10-2, 1 x 10-3, and 1 x 10-4 dilutions onto TSB + Congo Red plates and incubate overnight at 37 °C.
NOTE: Plating 100 µL from the 1 x 10-2, 1 x 10-3, and 1 x 10-4 dilutions corresponds to a final dilution factor of 1 x 10-3, 1 x 10-4, and 1 x 10-5, respectively.
Representative Results
Adherence, invasion, and intracellular replication assays were performed comparing S. flexneri 2457T wild type (WT) to S. flexneri ΔVF (ΔVF), a mutant hypothesized to negatively regulate Shigella virulence. Since Shigella uses bile salts as a signal to regulate virulence17,18,47, experiments were performed after bacterial subculture in TSB media as well as TSB supplemented with 0.4% (w/v) bile salts18. Bile salts exposure during the subculture step acts as a pre-treatment to replicate small intestinal transit prior to colonic infection17,18,47. Figure 1 analyzes the effect of the ΔVF mutation on the ability of S. flexneri to adhere to HT-29 colonic epithelial cells. Percent adherence is plotted on the y-axis and represents the ratio of recovered bacteria following HT-29 lysis standardized to the number of input bacteria. As expected, both S. flexneri WT and ΔVF strains had a significant increase in adherence when subcultured with bile salts supplementation in comparison to TSB without bile salts supplementation18. However, there was no difference in adherence to HT-29 cells between WT and ΔVF mutant strains within each subculture condition. These data indicate that the ΔVF mutation has no effect on the ability of S. flexneri to adhere to HT-29 epithelial cells with or without the bile salts pre-treatment.
In Figure 2, the effect of the ΔVF mutation on the ability of S. flexneri to invade (Figure 2A) and replicate (Figure 2B) inside of HT-29 colonic epithelial cells with or without bile salts pre-treatment was analyzed. Percent recovery is plotted on the y-axis and represents the ratio of recovered bacterial cells following HT-29 cell lysis standardized to the number of input bacteria. In Figure 2A, there was an expected significant increase in the ability of WT S. flexneri 2457T to invade HT-29 cells after pre-exposure to bile salts48, while the S. flexneri ΔVF mutant displayed a smaller increase in invasion following bile salts pre-exposure compared to the WT strain. The ΔVF mutant had increased invasion rates of HT-29 cells compared to the WT subcultured in TSB, but had similar invasion rates as WT when subcultured in TSB supplemented with bile salts (Figure 2A). These results suggest that the ΔVF mutation enhances the ability of S. flexneri to invade HT-29 cells, which lessens the effect of the bile salts pre-exposure even though the invasion ability of the ΔVF mutant did increase further following bile salts subculture.
Overall, 10-fold more bacteria were recovered following overnight incubation (Figure 2B) compared to the 90 min incubation (Figure 2A), which demonstrates the differences in monitoring intracellular growth versus invasion, respectively. When infected HT-29 cells were incubated for 18 h to allow for intracellular replication of the bacteria (Figure 2B), the impact of the bile salts pre-treatment decreased for both the WT and ΔVF strains. However, the reduced effect of bile salts pre-treatment during intracellular replication was more dramatic for the ΔVF mutant. Since the increase in intracellular replication of both strains when pre-exposed to bile salts was smaller than the increase in invasion rates in the same conditions, we hypothesize that bile salts have a greater impact on the early steps in S. flexneri pathogenesis. The ΔVF mutant displayed an increase in percent recovery from overnight replication inside HT-29 cells compared to WT (Figure 2B) following both subculture conditions. However, the percent recoveries of the ΔVF mutant were similar regardless of bile salts pre-exposure. These data trends suggest that the ΔVF mutant replicates more efficiently inside HT-29 cells compared to WT, and that bile salts pre-exposure does not impact the ability of the ΔVF mutant to replicate intracellularly, as observed for WT (Figure 2B). Since the difference between the mutant and WT strains in the bile salts pre-exposure condition was not observed during the 90 min invasion assay, we hypothesize that the product encoded by the deleted VF gene may also regulate S. flexneri replication inside HT-29 cells. Combined, both analyses demonstrate that the ΔVF mutant is more virulent relative to WT, which suggests that the VF gene product is a negative regulator of S. flexneri virulence.
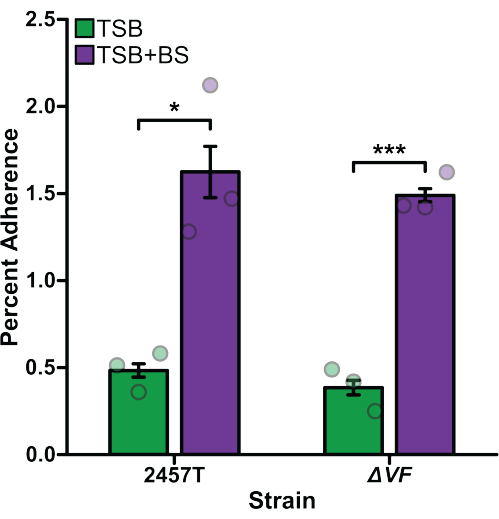
Figure 1: Bile salts pre-exposure induced adherence of S. flexneri to HT-29 cells. S. flexneri 2457T WT and ΔVF mutant cells were subcultured in either TSB or TSB supplemented with 0.4% (w/v) bile salts (TSB+BS) media. The bacteria were then applied to HT-29 cells at a multiplicity of infection (MOI) of 100 and incubated for 3 h to examine adherence. After incubation, infected HT-29 cells were washed and lysed, and serial dilutions of recovered bacteria were plated to enumerate colony-forming units per mL (CFU/mL). The number of adherent bacteria is plotted relative to the input bacteria titers to establish the percent adherence. Data are representative of one biological replicate with three technical replicates (individual dots). Error bars indicate the standard error of the mean (SEM). Statistical significance was determined by a Student's t-test (*p < 0.05; ***p < 0.001). Please click here to view a larger version of this figure.
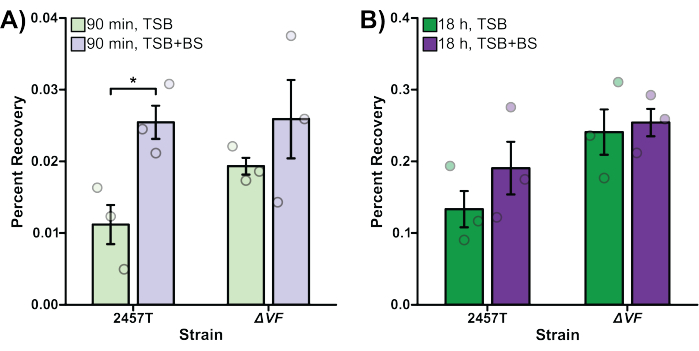
Figure 2: Bile salts pre-exposure increased WT S. flexneri invasion and intracellular replication. S. flexneri 2457T WT and ΔVF mutant cells were subcultured in either TSB or TSB supplemented with bile salts (TSB+BS) media. The bacteria were then applied to HT-29 cells at an MOI of 100, centrifuged onto the cells, and incubated at 37 °C with 5% CO2 for 45 min. Cells were washed with PBS, and extracellular bacteria were lysed by the addition of gentamicin to the DMEM to exclusively recover intracellular bacteria. After 90 min (A, bacterial invasion) or 18 h (B, intracellular replication) incubations, infected HT-29 cells were washed and lysed, and serial dilutions of recovered bacteria were plated to enumerate colony-forming units per mL (CFU/mL). The number of intracellular bacteria is plotted relative to the input bacterial titers to establish the percent recovery. Data are representative of one biological replicate, each with three technical replicates (individual dots). Error bars indicate SEM. Statistical significance was determined by a Student's t-test (*p < 0.05). Please note the differences in the y-axis scales between panels (A) and (B). Please click here to view a larger version of this figure.
Discussion
This protocol describes a set of three standardized assays to study Shigella adherence, invasion, and intracellular replication of intestinal epithelial cells. Although these methods are merely modified versions of classical gentamicin assays used to study the invasion and intracellular replication of various bacterial pathogens within host cells49,50,51, special considerations must be applied when studying Shigella.
Shigella are facultative anaerobes, which grow optimally at 37 °C in rich medium with aeration43. When performing these assays to interrogate the effects of different environmental signals or metabolites on Shigella infections, it is recommended to grow Shigella in rich media overnight, then subculture into defined or supplemented media to mid-log phase prior to infection. Maximal virulence gene expression occurs during the logarithmic phase of growth52,53. Thus, subculturing overnight Shigella cultures and allowing bacterial growth to exponential phase (OD600 ~ 0.7) are necessary steps to properly assess Shigella virulence. Further, virulence gene expression is temperature-dependent. To ensure specificity for expression only during host infection, virulence genes are induced at host physiological temperatures (37 °C) and strictly repressed at lower temperatures (e.g., 30 °C)54. To ensure expression of virulence genes, bacterial subculturing and infections must be carried out at 37 °C, with proper care to ensure the temperature does not drop below 37 °C. A large, 220 kilobase virulence plasmid, which encodes the molecular machinery required for invasion and epithelial cell infection, is an essential virulence determinant for all Shigella spp5. Repeated passaging and prolonged growth at 37 °C promote virulence plasmid instability, resulting in plasmid loss and rendering Shigella cells avirulent55. To ensure virulence plasmid maintenance and expression of crucial virulence genes, it is recommended to restreak bacteria directly from freezer stocks every two weeks onto fresh TSB + Congo red indicator plates. Congo red agar can be used to differentiate between wild-type virulent (red, CR+) colonies and avirulent (white, CR–) colonies that have lost the virulence plasmid45. If virulence plasmid instability is repeatedly observed, overnight cultures of Shigella can be incubated at 30 °C to prevent virulence plasmid loss, followed by subculturing at 37 °C to promote virulence gene expression43. Finally, if analyses with mutants and/or complemented strains are performed, in which antibiotic markers are present in these bacterial strains, it is recommended to use the appropriate antibiotic selection during the bacterial overnight and subculturing steps.
The use of HT-29 epithelial monolayers has many advantages for studying the molecular mechanisms of Shigella pathogenesis in vitro. Since Shigella is a human-adapted pathogen, small animal models do not accurately reflect the characteristic pathogenesis of shigellosis observed in humans36. Thus, the use of standardized protocols for infection of human-derived cell lines enables the quantitative interrogation of individual steps of bacterial pathogenesis to characterize the molecular interplay between this pathogen and its native host. Historically, HeLa cells were predominantly used to study host-pathogen interactions in vitro25,56,57. However, HeLa cells are an immortalized cervical cancer cell line, which are non-native host cells for enteric bacterial pathogens. Thus, in vitro studies of enteric pathogenesis have shifted to the use of colonic adenocarcinoma cell lines (e.g., HT-29, Caco-2, and T84), which more faithfully recapitulate intestinal epithelial morphology, and can be grown on transwells as epithelial monolayers with differentiated apical and basolateral surfaces58,59,60. Although each cell line has its individual strengths and weaknesses, HT-29 cells are used in the assays described here due to phenotypic similarities to intestinal enterocytes and robust expression of cell surface receptors and pro-inflammatory cytokines58,59,61. However, each of these discussed models are cancer-derived cell lines with aberrant metabolic and growth phenotypes that do not accurately represent the natural physiological state of the colonic epithelium in vivo58. Recent advances in human tissue culture techniques have enabled the cultivation of human intestinal stem cells, obtained from tissue biopsies, into organoids or single two-dimensional cell monolayers that recapitulate the various cell types present in the human gastrointestinal (GI) tract62,63,64. Intestinal organoids can be differentiated to include enterocytes, mucus-producing goblet cells, M cells, and other tissue-specific cell types. Recent studies have validated the use of these organoid models for studying enteric pathogens, including various Shigella spp., Salmonella enterica, and Escherichia coli pathovars62,63,64. Although organoids are more accurate models of the human GI epithelium and can even be cultured in varied nutrient conditions to represent the global population65, their relative complexity requires significant training and technical expertise, resulting in higher costs and longer culturing times in comparison with traditional cancer cell lines58.
This protocol describes medium-throughput methods, whereby two 6-well plates are used for a total of 12 monolayers available for testing. However, experiments can easily be scaled to increase the number of monolayers seeded in order to accommodate additional technical replicates, or testing additional Shigella mutants and environmental signals. Tissue culture plates with more wells (e.g., 12-well or 24-well plates) can also be used with appropriate adjustments to account for wells with smaller diameters. Under the HT-29 growth conditions outlined in step 3.2, HT-29 titers are sufficient for seeding about six 6-well plates following cultivation from one T75 flask, with enough remaining cells to maintain the HT-29 cell culture. Further, the seeding of HT-29 monolayers and preparation of inoculating bacteria steps are identical between bacterial adherence, invasion, and intracellular replication assays. Thus, assays testing the same Shigella strains or subculturing conditions can easily be performed in parallel to simultaneously examine the role of each experimental condition in various steps of Shigella infection.
Recovery titers for adherent (step 4), invaded (step 5), and intracellular bacteria (step 6) are determined by plating serial dilutions of infected HT-29 cells following lysis, which enables quantitative assessment of the efficiency of each stage of Shigella infection. The centrifugation steps in the invasion and intracellular replication assays, based on classical assays49,50,51, promote bacterial contact with the host cells for immediate invasion and enhance invasion rates. Thus, centrifugation "skips" the adherence step, which was later discovered to be an important aspect of Shigella infection17,18,19,66. It is essential to note that the centrifuge step cannot be performed with polarized epithelial cells seeded on transwells. For the adherence assay, however, adherence is not forced by centrifugation, and no gentamicin is added to the infected HT-29 cells prior to lysis, thus allowing the enumeration of adherent bacterial cells. The volume of tissue culture media is reduced to 1 mL (as opposed to 2 mL for the invasion and intracellular replication assays) to enable more efficient contact of the bacteria with the HT-29 cells. Furthermore, depending on the rate of adherence, some invasion and intracellular replication can occur during the 3 h incubation, but the amount is typically negligible (e.g., only 0.05% of the recovered bacterial population following host cell lysis). Nevertheless, to properly account for adherent versus intracellular bacteria during the 3 h incubation, it is recommended to perform parallel assays, where, in addition to the stated protocol, a second plate is incubated with 50 µg/mL gentamicin in 2 mL of DMEM per well for 45 min total (15 min incubation, wash, fresh DMEM + gentamycin for 30 min) to thoroughly lyse extracellular bacteria. Following HT-29 cell lysis as noted in the protocol, the intracellular bacteria can be enumerated as noted above, and will represent the number of bacteria that invaded. This value can then be subtracted from the bacteria enumerated from the plate without gentamicin treatment to determine adherent and invaded bacteria appropriately. Parallel analyses can also be performed to give more insight into the intracellular replication of Shigella inside HT-29 cells after invasion, e.g., using multiple time points to perform intracellular growth curves. Finally, along with enumerating intracellular bacteria after an 18 h gentamicin incubation, the culture supernatants can be collected and analyzed for cytokine secretion of infected HT-29 cells. For instance, IL-8 is a chemokine secreted by epithelial cells that largely functions to recruit PMNs to the site of infection. The amount of IL-8 secreted into the culture media of infected HT-29 cells can be analyzed with an IL-8 ELISA assay67.
Bile salts have proved to be an important virulence signal for Shigella during transit through the human GI system, and can be used to supplement bacterial growth media in these protocols to replicate typical GI conditions. Naturally, bile salts are introduced in the duodenum or upper portion of the small intestine to aid in digestion; and in the terminal ileum or end of the small intestine, 95% of bile salts are removed and recycled back into circulation for ultimate deposit in the gallbladder68. Bile salts usually have a concentration between 0.2%-2.0% (w/v) in the small intestine and are naturally bactericidal. However, Shigella, along with most enteric bacteria, resist bile salts and use the signals to enhance infection47. Several studies have documented how bile salts exposure, at times in conjunction with other small intestinal signals like glucose, affects Shigella survival and virulence regulation prior to infection. Shigella has been shown to resist bile salts, alter gene expression, and form and disperse a biofilm in conditions that reproduce small intestinal transit17,69. The studies have demonstrated that these changes result in a hypervirulent phenotype, in which adherence and invasion are induced upon subsequent colonic infection by Shigella17,18,19,48,66. Thus, the conditions outlined above document how to subculture Shigella in bile salts to mimic small intestinal transit prior to performing the adherence, invasion, or intracellular replication assays. The specified TSB formulation contains added glucose relative to typical Luria broth (LB)17. Thus, if LB is used, it is important to also supplement the media with glucose (0.5%-2.0% [w/v]) to appropriately consider the glucose signals in the small intestine17. Furthermore, as mentioned above, all Shigella subcultures are washed to remove bile salts and mimic the transition to the colon for infection analyses.
Traditionally, and as highlighted in the results above (Figure 1 and Figure 2), infection assays are valuable to determine the role of a particular gene in Shigella infection. The phenotype of various mutants, however, may not be truly appreciated without proper supplementation of the bacterial culture media. As prior research has demonstrated, bile salts significantly alter S. flexneri gene expression, including genes for central metabolism, transcription factors, sugar transporters, drug resistance, and virulence genes either encoded on the chromosome or virulence plasmid17. These genes provide insight into how Shigella uses bile salts as a signal to alter gene expression and prepare for eventual infection of the colon, and subsequent mutational analyses thus need to be performed in the appropriate supplemented bacterial growth media prior to examining effects on virulence in the adherence, invasion, and intracellular replication assays. As seen above in Figure 1 and Figure 2, the ΔVF mutation did not affect adherence but did affect invasion and intracellular replication. Since the mutation enhanced invasion and intracellular replication, experiments are currently underway to determine how the gene product regulates infection. The mutant analyses serve as an example of how new understandings in Shigella pathogenesis can be studied, especially in conditions that better replicate the human GI tract. Proper complementation analyses are recommended to validate the phenotypes of various mutants.
In combination, these procedures describe quantitative experiments that will provide important insight into the molecular mechanisms of Shigella adherence to, invasion of, and replication inside HT-29 colonic epithelial cells by best mimicking the natural environment of the human GI tract during infection. Future studies can expand on the mutants and experimental conditions to gain a better understanding of how Shigella prepares for and effectively infects human hosts. Despite decades of research, there is still so much more to discover about Shigella infection.
Divulgations
The authors have nothing to disclose.
Acknowledgements
Support for the authors includes Massachusetts General Hospital's Department of Pediatrics, the Executive Committee on Research Interim Support Funding (ISF) award 2022A009041, the National Institute of Allergy and Infectious Diseases grant R21AI146405, and the National Institute of Diabetes and Digestive and Kidney Diseases grant Nutrition Obesity Research Center at Harvard (NORCH) 2P30DK040561-26. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Materials
| 0.22 μm PES filter | Millipore-Sigma | SCGP00525 | Sterile, polyethersulfone filter for sterilizing up to 50 mL media |
| 14 mL culture tubes | Corning | 352059 | 17 mm x 100 mm polypropylene test tubes with cap |
| 50 mL conical tubes | Corning | 430829 | 50 mL clear polypropylene conical bottom centrifuge tubes with leak-proof cap |
| 6-well tissue culture plates | Corning | 3516 | Plates are treated for optimal cell attachment |
| Bile salts | Sigma-Aldrich | B8756 | 1:1 ratio of cholate to deoxycholate |
| Congo red dye | Sigma-Aldrich | C6277 | A benzidine-based anionic diazo dye, >85% purity |
| Countess cell counting chamber slide | Invitrogen | C10283 | To be used with the Countess Automated Cell Counter |
| Dimethyl sulfoxide (DMSO) | Sigma-Aldrich | D8418 | A a highly polar organic reagent |
| Dulbecco’s Modified Eagle Medium (DMEM) | Gibco | 10569-010 | DMEM is supplemented with high glucose, sodium pyruvate, GlutaMAX, and Phenol Red |
| Fetal Bovine Serum (FBS) | Sigma-Aldrich | F4135 | Heat-inactivated, sterile |
| Gentamicin | Sigma-Aldrich | G3632 | Stock concentration is 50 mg/mL |
| HT-29 cell line | ATCC | HTB-38 | Adenocarcinoma cell line; colorectal in origin |
| Paraffin film | Bemis | PM999 | Laboratory sealing film |
| Petri dishes | Thermo Fisher Scientific | FB0875713 | 100 mm x 15 mm Petri dishes for solid media |
| Phosphate-buffered saline (PBS) | Thermo Fisher Scientific | 10010049 | 1x concentration; pH 7.4 |
| Select agar | Invitrogen | 30391023 | A mixture of polysaccharides extracted from red seaweed cell walls to make bacterial plating media |
| T75 flasks | Corning | 430641U | Tissue culture flasks |
| Triton X-100 | Sigma-Aldrich | T8787 | A common non-ionic surfactant and emulsifier |
| Trypan blue stain | Invitrogen | T10282 | A dye to detect dead tissue culture cells; only live cells can exclude the dye |
| Trypsin-EDTA | Gibco | 25200-056 | Reagent for cell dissociation for cell line maintenance and passaging |
| Tryptic Soy Broth (TSB) | Sigma-Aldrich | T8907 | Bacterial growth media |
References
- Karambizi, N. U., McMahan, C. S., Blue, C. N., Temesvari, L. A. Global estimated Disability-Adjusted Life-Years (DALYs) of diarrheal diseases: A systematic analysis of data from 28 years of the global burden of disease study. PloS one. 16 (10), e0259077 (2021).
- WHO. WHO methods and data sources for country-level causes of death 2000-2016. World Health Organization. , (2018).
- Kotloff, K. L. Shigella infection in children and adults: a formidable foe. Lancet Glob Health. 5 (12), e1166-e1167 (2017).
- Kotloff, K. L., et al. Burden and aetiology of diarrhoeal disease in infants and young children in developing countries (the Global Enteric Multicenter Study, GEMS): A prospective, case-control study. Lancet. 382 (9888), 209-222 (2013).
- Schroeder, G. N., Hilbi, H. Molecular pathogenesis of Shigella spp.: Controlling host cell signaling, invasion, and death by type III secretion. Clin Microbiol Rev. 21 (1), 134-156 (2008).
- Arvelo, W., et al. Transmission risk factors and treatment of pediatric shigellosis during a large daycare center-associated outbreak of multidrug resistant shigella sonnei: Implications for the management of shigellosis outbreaks among children. Pediatr Infect Dis J. 28 (11), 976-980 (2009).
- Kozyreva, V. K., et al. Recent outbreaks of Shigellosis in California caused by two distinct populations of Shigella sonnei with either increased virulence or fluoroquinolone resistance. mSphere. 1 (6), 1-18 (2016).
- Bowen, A., et al. Importation and domestic transmission of Shigella sonnei resistant to ciprofloxacin – United States, May 2014-February 2015. MMWR Morb Mortal Wkly Rep. 64 (12), 318-320 (2015).
- Tansarli, G. S., et al. Genomic reconstruction and directed interventions in a multidrug-resistant Shigellosis outbreak in Seattle, WA, USA: a genomic surveillance study. Lancet. 3099 (22), 1-11 (2023).
- Barry, E. M., et al. Progress and pitfalls in Shigella vaccine research. Nat Rev Gastroenterol Hepatol. 10 (4), 245-255 (2013).
- Increase in Extensively Drug-Resistant Shigellosis in the United States. CDC Health Alert Network. Centers for Disease Control and Prevention Available from: https://emergency.cdc.gov/han/2023/han00486.asp?ACSTrackingID=USCDC_511-DM100260&ACSTrackingLabel=HAN%20486%20-%20General%20Public&deliveryName=USCDC_511-DM100260 (2023)
- Shiferaw, B., et al. Antimicrobial susceptibility patterns of Shigella isolates in Foodborne Diseases Active Surveillance Network (FoodNet) sites, 2000-2010. Clin Infect Dis. 54, S458-S463 (2012).
- Centers for Disease Control and Prevention. COVID-19: U.S. Impact on Antimicrobial Resistance, Special Report 2022. Atlanta, GA: U.S. Department of Health and Human Services. CDC. , (2022).
- Centers for Disease Control and Prevention. Antibiotic resistance threats in the United States, 2019. CDC. 10 (1), (2019).
- WHO. Prioritization of pathogens to guide discovery, research and development of new antibiotics for drug-resistant bacterial infections, including tuberculosis. WHO. , (2017).
- DuPont, H. L., Levine, M. M., Hornick, R. B., Formal, S. B. Inoculum size in shigellosis and implications for expected mode of transmission. J Infect Dis. 159 (6), 1126-1128 (1989).
- Nickerson, K. P., et al. Analysis of Shigella flexneri resistance, biofilm formation, and transcriptional profile in response to bile salts. Infect Immun. 85 (6), 1-18 (2017).
- Faherty, C. S., Redman, J. C., Rasko, D. A. Shigella flexneri effectors OspE1 and OspE2 mediate induced adherence to the colonic epithelium following bile salts exposure. Mol Microbiol. 85 (1), 107-121 (2012).
- Chanin, R. B., et al. Shigella flexneri adherence factor expression in in vivo-like conditions. mSphere. 4 (6), e00751 (2019).
- Baranov, V., Hammarström, S. Carcinoembryonic antigen (CEA) and CEA-related cell adhesion molecule 1 (CEACAM1), apically expressed on human colonic M cells, are potential receptors for microbial adhesion. Histochem Cell Biol. 121 (2), 83-89 (2004).
- Wassef, J. S., Keren, D. F., Mailloux, J. L. Role of M cells in initial antigen uptake and in ulcer formation in the rabbit intestinal loop model of shigellosis. Infect Immun. 57 (3), 858-863 (1989).
- Sansonetti, P. J., Arondel, J., Cantey, J. R., Prévost, M. C., Huerre, M. Infection of rabbit Peyer’s patches by Shigella flexneri: Effect of adhesive or invasive bacterial phenotypes on follicle-associated epithelium. Infect Immun. 64 (7), 2752-2764 (1996).
- Sansonetti, P. J., et al. Caspase-1 activation of IL-1beta and IL-18 are essential for Shigella flexneri-induced inflammation. Immunity. 12 (5), 581-590 (2000).
- Zychlinsky, A., Fitting, C., Cavaillon, J. M., Sansonetti, P. J. Interleukin 1 is released by murine macrophages during apoptosis induced by Shigella flexneri. J Clin Invest. 94 (3), 1328-1332 (1994).
- Sansonetti, P. J., Ryter, A., Clerc, P., Maurelli, A. T., Mounier, J. Multiplication of Shigella flexneri within HeLa cells: lysis of the phagocytic vacuole and plasmid-mediated contact hemolysis. Infect Immun. 51 (2), 461-469 (1986).
- Maldonado-Contreras, A., et al. Shigella depends on SepA to destabilize the intestinal epithelial integrity via cofilin activation. Gut Microbes. 8 (6), 544-560 (2017).
- Collard, J. -. M., et al. High prevalence of small intestine bacteria overgrowth and asymptomatic carriage of enteric pathogens in stunted children in Antananarivo, Madagascar. PLoS Negl Trop Dis. 16 (5), e0009849 (2022).
- Mattock, E., Blocker, A. J. How do the virulence factors of shigella work together to cause disease. Front Cell Infect Microbiol. 7, 1-24 (2017).
- Mostowy, S., et al. The zebrafish as a new model for the in vivo study of Shigella flexneri interaction with phagocytes and bacterial autophagy. PLoS Pathog. 9 (9), e1003588 (2013).
- Martinez-Becerra, F. J., et al. Parenteral immunization with IpaB/IpaD protects mice against lethal pulmonary infection by Shigella. Vaccine. 31 (24), 2667-2672 (2013).
- Shim, D. -. H., et al. New animal model of shigellosis in the Guinea pig: its usefulness for protective efficacy studies. J Immunol. 178 (4), 2476-2482 (2007).
- Marteyn, B., et al. Modulation of Shigella virulence in response to available oxygen in vivo. Nature. 465 (7296), 355-358 (2010).
- West, N. P., et al. Optimization of virulence functions through glucosylation of Shigella LPS. Science. 307 (5713), 1313-1317 (2005).
- Maurelli, A. T., et al. Shigella infection as observed in the experimentally inoculated domestic pig, Sus scrofa domestica. Microbial Pathog. 25 (4), 189-196 (1998).
- Jeong, K. -. I., Zhang, Q., Nunnari, J., Tzipori, S. A piglet model of acute gastroenteritis induced by Shigella dysenteriae Type 1. J Infect Dis. 201 (6), 903-911 (2010).
- Kim, Y. -. J., Yeo, S. -. G., Park, J. -. H., Ko, H. -. J. Shigella vaccine development: prospective animal models and current status. Curr Pharm Biotechnol. 14 (10), 903-912 (2013).
- Kent, T. H., Formal, S. B., LaBrec, E. H., Sprinz, H., Maenza, R. M. Gastric shigellosis in rhesus monkeys. Am J Pathol. 51 (2), 259-267 (1967).
- Shipley, S. T., et al. A challenge model for Shigella dysenteriae 1 in cynomolgus monkeys (Macaca fascicularis). Comp Med. 60 (1), 54-61 (2010).
- Higgins, R., Sauvageau, R., Bonin, P. Shigella flexneri Type 2 Infection in captive nonhuman primates. Can Vet J. 26 (12), 402-403 (1985).
- Oaks, E. V., Hale, T. L., Formal, S. B. Serum immune response to Shigella protein antigens in rhesus monkeys and humans infected with Shigella spp. Infect Immun. 53 (1), 57-63 (1986).
- Formal, S. B., et al. Protection of monkeys against experimental shigellosis with a living attenuated oral polyvalent dysentery vaccine. J Bacteriol. 92 (1), 17-22 (1966).
- Levine, M. M., Kotloff, K. L., Barry, E. M., Pasetti, M. F., Sztein, M. B. Clinical trials of Shigella vaccines: two steps forward and one step back on a long, hard road. Nat Rev Microbiol. 5 (7), 540-553 (2007).
- Payne, S. M. Laboratory cultivation and storage of Shigella. Curr Protoc Microbiol. 55 (1), 93 (2019).
- NIH Guidelines. NIH guidelines for research involving recombinant or synthetic nucleic acid molecules. NIH Guidelines. 2, 142 (2019).
- Maurelli, A. T., Blackmon, B., Curtiss, R. Loss of pigmentation in Shigella flexneri 2a is correlated with loss of virulence and virulence-associated plasmid. Infect Immun. 43 (1), 397-401 (1984).
- HT-29 cell line product sheet. ATCC Available from: https://www.atcc.org/products/htb-38 (2023)
- Sistrunk, J. R., Nickerson, K. P., Chanin, R. B., Rasko, D. A., Faherty, C. S. Survival of the fittest: How bacterial pathogens utilize bile to enhance infection. Clin Microbiol Rev. 29 (4), 819-836 (2016).
- Stensrud, K. F., et al. Deoxycholate interacts with IpaD of Shigella flexneri in inducing the recruitment of IpaB to the type III secretion apparatus needle tip. J Biol Chem. 283 (27), 18646-18654 (2008).
- Mandell, G. L. Interaction of intraleukocytic bacteria and antibiotics. J Clin Invest. 52 (7), 1673-1679 (1973).
- Elsinghorst, E. A. Measurement of invasion by gentamicin resistance. Methods Enzymo. 236 (1979), 405-420 (1994).
- Elsinghorst, E. A., Weitz, J. A. Epithelial cell invasion and adherence directed by the enterotoxigenic Escherichia coli tib locus is associated with a 104-kilodalton outer membrane protein. Infect Immun. 62 (8), 3463-3471 (1994).
- Dorman, C. J., McKenna, S., Beloin, C. Regulation of virulence gene expression in Shigella flexneri, a facultative intracellular pathogen. Int J Med Microbiol. 291 (2), 89-96 (2001).
- Porter, M. E., Dorman, C. J. Positive regulation of Shigella flexneri virulence genes by integration host factor. J Bacteriol. 179 (21), 6537-6550 (1997).
- Maurelli, A. T., Blackmon, B., Curtiss, R. Temperature-dependent expression of virulence genes in Shigella species. Infect Immun. 43 (1), 195-201 (1984).
- Schuch, R., Maurelli, A. T. Virulence plasmid instability in Shigella flexneri 2a is induced by virulence gene expression. Infect Immun. 65 (9), 3686-3692 (1997).
- Formal, S. B., Hale, T. L., Sansonetti, P. J. Invasive enteric pathogens. Rev Infect Dis. 5, S702-S707 (1983).
- Pál, T., Hale, T. L. Plasmid-associated adherence of Shigella flexneri in a HeLa cell model. Infect Immun. 57 (8), 2580-2582 (1989).
- Noben, M., et al. Human intestinal epithelium in a dish: Current models for research into gastrointestinal pathophysiology. United European Gastroenterol J. 5 (8), 1073-1081 (2017).
- Liévin-Le Moal, V., Servin, A. L. Pathogenesis of human enterovirulent bacteria: lessons from cultured, fully differentiated human colon cancer cell lines. Microbiol Mol Biol Rev R. 77 (3), 380-439 (2013).
- Mitchell, D. M., Ball, J. M. Characterization of a spontaneously polarizing HT-29 cell line, HT-29/cl.f8. In Vitro Cell Dev Biol – Anim. 40 (10), 297-302 (2004).
- Gagnon, M., Zihler Berner, A., Chervet, N., Chassard, C., Lacroix, C. Comparison of the Caco-2, HT-29 and the mucus-secreting HT29-MTX intestinal cell models to investigate Salmonella adhesion and invasion. J Microbiol Methods. 94 (3), 274-279 (2013).
- Koestler, B. J., et al. Human intestinal enteroids as a model system of Shigella pathogenesis. Infect Immun. 87 (4), 00733 (2019).
- Ranganathan, S., et al. Evaluating Shigella flexneri pathogenesis in the human enteroid model. Infect Immun. 87 (4), (2019).
- Nickerson, K. P., et al. A versatile human intestinal organoid-derived epithelial monolayer model for the study of enteric pathogens. Microbiol Spectr. 9 (1), 1-17 (2021).
- Perlman, M., Senger, S., Verma, S., Carey, J., Faherty, C. S. A foundational approach to culture and analyze malnourished organoids. Gut Microbes. 15 (2), 2248713 (2023).
- Pope, L. M., Reed, K. E., Payne, S. M. Increased protein secretion and adherence to HeLa cells by Shigella spp. following growth in the presence of bile salts. Infect Immun. 63 (9), 3642-3648 (1995).
- Faherty, C. S., et al. The synthesis of OspD3 (ShET2) in Shigella flexneri is independent of OspC1. Gut Microbes. 7 (6), 486-502 (2016).
- Ridlon, J. M., Kang, D. -. J., Hylemon, P. B. Bile salt biotransformations by human intestinal bacteria. J Lipid Res. 47 (2), 241-259 (2006).
- Köseoğlu, V. K., Hall, C. P., Rodríguez-López, E. M., Agaisse, H. The Autotransporter IcsA promotes Shigella flexneri biofilm formation in the presence of bile salts. Infect Immun. 87 (7), 1-14 (2019).

