Quartz-Crystal Microbalance Biosensor-Based Biopanning to Study Protein-Drug Interactions
Abstract
Source: Takakusagi, Y. Biosensor-based High Throughput Biopanning and Bioinformatics Analysis Strategy for the Global Validation of Drug-protein Interactions. J. Vis. Exp. (2020)
This video demonstrates a method to study protein-drug interactions using QCM biosensor-based biopanning of the T7 phage-displayed peptide library. The biopanning of phages displaying a specific peptide on the QCM biosensor allows specific isolation of high-affinity peptides, followed by amino acid sequencing, which helps to identify specific interaction sites on the target protein.
Protocol
NOTE: The following are the steps for screening drug-recognizing T7 phages using a QCM biosensor and recovering the screened phages via E. coli (BLT5615) infection. The protocols for the synthesis of a derivative of a small molecule that forms a self-assembled monolayer (SAM) and for the construction of the T7 phage-displayed 15-mer random peptide library can be found elsewhere.
1. Preparation of the QCM sensor chip
- Attach a ceramic sensor chip on the oscillator of a 27-MHz QCM apparatus, and record the intrinsic frequency (Hz) in the air phase before small molecule immobilization.
- Detach the chip and drop 20 μL of a solution (1 mM in 70% ethanol) of a small molecule derivative that forms SAM onto the gold electrode of the sensor chip using a pipette.
CAUTION: The sensor chip crystal where the gold electrode (Au, 0.1 mm thick, 2.5 mm i.d., 4.9 mm2) is located is extremely thin and may crack easily (SiO2, 0.06 mm thick, 9 mm i.d.). Hence, pipette carefully. - Leave for 1 h at room temperature (around 20 °C) in a Petri dish with moistened tissue and shielded from room lights.
- Wash the electrode surface gently with ultrapure water; then, remove the water drops by blowing air with a syringe or air duster.
- Set up the sensor chip for the QCM apparatus and record the reduction in frequency in the air phase to measure the amount of the small molecule that has been immobilized.
NOTE: At least, 100 Hz of intrinsic frequency is necessary for successful small molecule immobilization (1 Hz immobilizes 30 pg).
2. Biopanning of the T7 phage library using a QCM biosensor (Figure 1)
- Set a cuvette with a dedicated magnetic stirrer on the QCM biosensor and pour 8 mL of the reaction buffer (10 mM Tris-HCl, 200 mM NaCl, pH 7-8) into the cuvette.
- Attach the QCM sensor chip to the oscillator and pull down the arm of the oscillator to immerse the chip into the buffer being stirred at 1000 rpm.
- Start monitoring the QCM frequency on the personal computer (PC) and wait until the sensorgram equilibrates to around 3 Hz/min of the frequency drift.
- Inject 8 μL of a T7 phage library (1—2 × 1010 pfu/mL) into the cuvette (final concentration: 1—2 × 107 pfu/mL) and mark the injection point on the sensor.
- Monitor the frequency reduction caused by the binding of T7 phages to the small molecule immobilized on the gold electrode surface for 10 min.
- Stop the QCM frequency monitor, dislodge the sensor chip from the oscillator, and remove the buffer by blowing air and/or wicking away with wipes.
- Put the sensor chip into a humid Petri dish and drop the 20 μL of the suspension of E. coli (BLT5615) (OD600 = 0.5—1.0 after shaking at 37 °C by adding IPTG to 1 mM) host cells in the log phase onto the gold electrode.
- Close the lid of the dish and cover it with aluminum foil to block out light.
- Incubate the dish at 37 °C for 30 min on a 96-well microplate mixer (1000-1500 rpm) for enhancing the recovery of the bound T7 phages.
- Recover the 20 μL of the solution and suspend it into 200 μL of LB medium.
NOTE: The samples obtained in this step can be preserved at 4 °C for one week. - Prepare a dilution series of the phage solution for plaque isolation and DNA sequencing according to the general procedure described in the manufacturer's instructions.
Representative Results
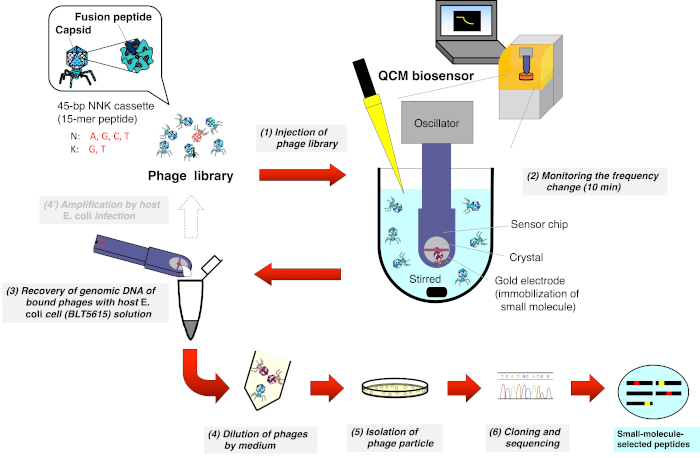
Figure 1: Schematic representation of the QCM biosensor-based biopanning of the T7 phage-displayed peptide library. A T7 phage library that displays random peptides is injected into the cuvette containing the buffer (under stirring) where the QCM biosensor chip is immersed and the frequency is stabilized. After monitoring the frequency reduction due to the binding of T7 phages to small molecules immobilized on the gold electrode surface of the sensor chip, the sensor chip is detached from the oscillator. DNA from the bound T7 phage is then directly recovered after host E. coli (BLT5615) infection. The resulting T7 phages are isolated via plaque formation and, finally, the amino acid sequence of the drug affinity-selected peptide displayed on the T7 phage capsid is determined according to the general phage display method.
Divulgations
The authors have nothing to disclose.
Materials
| AFFINIXQN | ULVAC, Inc. (Tokyo, Japan) | QCM2008-STKIT | Contains Glass cuvette, stir magnet, operation and analysis software with a Windows PC |
| Ceramic Sensor Chip | ULVAC, Inc. (Tokyo, Japan) | QCMST27C | 4 sensor chips/package |
| Dimethyl sulfoxide | Sigma-Aldrich (St. Louis, MO, USA) |
D8418 | |
| Ethanol | Merck (Kenilworth, NJ, USA) | 09-0850 | |
| Immobilization kit for AFFINIX | ULVAC, Inc. (Tokyo, Japan) | QCMIMKT | SAM reagent and amine-coupling reagent |
| Liquid LB medium | Sigma-Aldrich (St. Louis, MO, USA) |
L3522 | Autoclave for 20 min |
| NaCl | Merck (Kenilworth, NJ, USA) | S3014 | |
| Tris | Merck (Kenilworth, NJ, USA) | 252859 |

