The Colon-26 Carcinoma Tumor-bearing Mouse as a Model for the Study of Cancer Cachexia
Summary
Mice bearing the Colon-26 (C26) carcinoma represent a classical model of cancer cachexia. Progressive muscle wasting occurs in association with tumor growth, over-expression of muscle-specific ubiquitin ligases, and reductions in muscle cross-sectional area. Fat loss is also observed. Cachexia is studied in a time-dependent manner with increasing severity of wasting.
Abstract
Cancer cachexia is the progressive loss of skeletal muscle mass and adipose tissue, negative nitrogen balance, anorexia, fatigue, inflammation, and activation of lipolysis and proteolysis systems. Cancer patients with cachexia benefit less from anti-neoplastic therapies and show increased mortality1. Several animal models have been established in order to investigate the molecular causes responsible for body and muscle wasting as a result of tumor growth. Here, we describe methodologies pertaining to a well-characterized model of cancer cachexia: mice bearing the C26 carcinoma2-4. Although this model is heavily used in cachexia research, different approaches make reproducibility a potential issue. The growth of the C26 tumor causes a marked and progressive loss of body and skeletal muscle mass, accompanied by reduced muscle cross-sectional area and muscle strength3-5. Adipose tissue is also lost. Wasting is coincident with elevated circulating levels of pro-inflammatory cytokines, particularly Interleukin-6 (IL-6)3, which is directly, although not entirely, responsible for C26 cachexia. It is well-accepted that a primary mechanism by which the C26 tumor induces muscle tissue depletion is the activation of skeletal muscle proteolytic systems. Thus, expression of muscle-specific ubiquitin ligases, such as atrogin-1/MAFbx and MuRF-1, represent an accepted method for the evaluation of the ongoing muscle catabolism2. Here, we present how to execute this model in a reproducible manner and how to excise several tissues and organs (the liver, spleen, and heart), as well as fat and skeletal muscles (the gastrocnemius, tibialis anterior, and quadriceps). We also provide useful protocols that describe how to perform muscle freezing, sectioning, and fiber size quantification.
Introduction
Muscle wasting is a serious complication of various clinical conditions such as cancer, sepsis, liver, cirrhosis, heart and kidney failure, chronic obstructive pulmonary disease, and AIDS. In particular, muscle wasting is evident in at least 50% of patients with cancer1. Loss of skeletal muscle in cancer results from increased protein degradation due to the over-activation of the skeletal muscle proteolytic systems and/or from decreased protein synthesis6. Lipolysis is also evident, leading to the depletion of adipose tissue. Clinically, cachexia is associated with reduced quality and length of life and is estimated to be the cause of death in 20 – 30% of cancer patients7. Use of experimental models that resemble the human disease as closely as possible would be beneficial. An optimal animal model is characterized by high reproducibility, as well as by limited interference from different therapies and the unpredictable factors of diet, sex, and genetic background that are usually associated with the clinical condition8. So far, cancer cachexia has been studied mainly in animal models characterized by transplantation of cancer cells or injection of carcinogens, although a new method is to use genetically modified mice susceptible to the development of cancer.
Mice bearing the C26 carcinoma (also referred to as colon-26 and adenocarcinoma) represent a well-characterized and extensively used model of cancer cachexia2,5. The growth of the C26 tumor results in body and muscle weight loss, mainly through enhanced fat and protein catabolism9. Generally, a 10% tumor weight versus total body weight is associated with a reduction of 20-25% in skeletal muscle weight and a greater depletion of fat3,10. Hepatomegaly and splenomegaly are also observed with tumor growth, along with the activation of the acute phase response and the elevation of pro-inflammatory cytokine levels3,11. Among these, it is well known that IL-6 plays a pivotal role in mediating muscle wasting in the C26 model, even though this cytokine is probably not the only inducer of cachexia12. Elevated IL-6 causes muscle atrophy through activation of the JAK/STAT3 pathway, and inhibiting this transcription factor can prevent muscle wasting3,4.
During C26-induced muscle wasting, as in many conditions of muscle atrophy, muscle mass is lost largely through reductions in muscle protein content across muscle fibers, not through cell death or loss of fibers13. In C26 cachexia, a shift towards smaller cross-sectional areas is observed in both glycolytic and oxidative fibers2. This is also consistent with reduced muscle strength5. Many groups worldwide have taken advantage of the C26 model in order to discover new mediators of muscle wasting or clinically relevant drugs for cancer cachexia. However, many different procedures for the use of this model have been reported, raising concerns about the consistency of the obtained data and posing barriers to reproducibility in different experimental conditions. Here we report a typical use of this model for the study of cancer cachexia that yields standardized and reproducible data.
Protocol
Ethics Statement: All studies described were approved by the Institutional Animal Care and Use Committees of the Thomas Jefferson University and Indiana University School of Medicine.
1. C26 Cell Growth and Preparation
- Obtain C26 colorectal cancer cells (Ohio State University Medical Center (OSUMC)) and prepare complete growth medium (i.e., high-glucose Dulbecco's Modified Eagle's Medium (DMEM) containing 10% Fetal Bovine Serum (FBS), 1 mM sodium pyruvate, 1% glutamine, and 1% streptomycin/penicillin).
NOTE: In order to allow better inter-experimental reproducibility, it is preferable to use C26 cells at a passage as low as possible. - Use a commercially available cell counter to plate 50,000 cells/cm2 in flasks with complete growth medium. Incubate them in a humidified atmosphere at 37 °C and 5% CO2 until sub-confluent. Avoid culturing them at high density to prevent differentiation and drifting of cell phenotypes.
- After the first passage, plate enough cells to be implanted in the planned number of animals. On the day of cell implantation, dissociate the cells by trypsinization with 0.02 ml of 0.25% trypsin-EDTA solution per cm2, neutralize the trypsin by adding at least 3 ml of fresh medium per ml of trypsin, determine cell concentration in cells/ml, and resuspend the number of cells needed for the experiment (1 x 106 cells per mouse) in sterile PBS.
NOTE: C26 cells reach confluency when the density is about 500,000 cells/cm2.
2. Mice
- Randomize adult CD2F1 mice (at least 8 weeks of age) into groups (usually controls and tumor-bearers) with at least n = 6 per group.
NOTE: C26 tumors grow in Balb/c mice as well. However, based on our experience, CD2F1 mice represent a more versatile and economic host for this particular tumor model. Moreover, similar to the clinical settings and to findings reported in other experimental models of cancer-associated muscle wasting14-16, sex differences in the response to C26 cachexia may also be observed17 (see also Table 1). - Anesthetize the mouse using isoflurane by inhalation (3% in oxygen). Maintain the animal under anesthesia exclusively for the time necessary to perform the tumor inoculation. Make sure the mouse lies on a heated pad in order to prevent the loss of body heat. Before starting the tumor injection procedure, confirm proper anesthetization by gently pinching the rear toes.
NOTE: Due to the duration of this procedure (generally a few seconds), the use of veterinary eye ointment is not required. For the same reason, control animals undergoing the sham operation are not expected to lose weight. - Using a sterile 1-ml insulin syringe with a 26-gauge needle, quickly inject 250 μl of the solution containing the tumor cells subcutaneously and intrascapularly in the fat pad of the mouse.
NOTE: The control animals will receive 250 µl of sterile saline solution intrascapularly in the fat pad. - Place the animal back in its cage. Directly observe the mouse at least once every 15 min until it can respond to gentle manipulation and has regained a righting reflex.
NOTE: Do not leave the mouse unattended, and keep it in a warm recovery cage that contains a solid substrate. Furthermore, do not return the mouse to the animal room until it is fully recovered from anesthesia. Generally, mice do not need to be individually housed unless the investigator aims to assess individual food intake. If this is the case and the use of automated metabolic cages is not available, manual recording of food consumption must be performed, possibly daily and at the same time each day. - Check the mice daily and record their body weights.
NOTE: Recording the body condition score and home cage behaviors can differentiate degrees of cachexia and sickness behavior, which is useful for treatment studies18.
3. Euthanasia and Blood Collection
- When body weight loss in the tumor hosts versus controls reaches 5%, 10%, or 15% (mild, moderate, and severe cachexia, respectively) compared to their initial body weights, euthanize the mice.
NOTE: A typical experiment will be carried out until one group reaches 10% weight loss, upon which time all groups are euthanized. Body weight loss is assessed by measuring the body weight inclusive of the tumor. - Place the mouse under isoflurane general anesthesia (3% in oxygen). Draw blood by means of a cardiac puncture (approximately 1 – 1.5 ml).
- Collect the blood in K2-EDTA (18 mg) empty tubes and place them on ice. Centrifuge the blood (2,000 x g for 15 min at 4 °C) and collect the plasma/serum.
- Euthanize the mouse by means of cervical dislocation. Proceed to tissue excision and organ collection.
4. Tissue and Organ Excision
NOTE: For use of tissues in biochemical or molecular biology assays, plan to weigh each organ and tissue and place a fragment immediately into pre-labeled cryotubes. Snap freeze in liquid nitrogen and store at -80 °C.
- In order to avoid contamination of tissue samples with fur, spray 70% ethanol over the entire body.
- Place the mouse on a dissection bed in a supine position and extend a limb vertically by attaching the foot to an elevated buret clamp.
- Grasp the skin of the lower limb with Dumont forceps and use the curved fine scissors to gently remove the skin and fascia, exposing the underlying muscle bellies of the lower limb.
- Identify the gastrocnemius muscle on the posterior lower limb by its origin at the lateral and medial condyles on the posterior femur and its insertion at the calcaneus via the calcaneal tendon.
- Proceed to cut the gastrocnemius calcaneal tendon. Grasp the distal end of the gastrocnemius with Dumont forceps and pull the muscle belly toward its origin. Using scissors, cut the muscle at the origin as close as possible to the femur. Place the muscle in a dish. Weigh it immediately and freeze it in a tube in liquid nitrogen.
NOTE: Be sure not to excise the soleus muscle along with the gastrocnemius. If that occurs, carefully remove soleus by gently pulling it off with the forceps.
- Proceed to cut the gastrocnemius calcaneal tendon. Grasp the distal end of the gastrocnemius with Dumont forceps and pull the muscle belly toward its origin. Using scissors, cut the muscle at the origin as close as possible to the femur. Place the muscle in a dish. Weigh it immediately and freeze it in a tube in liquid nitrogen.
- Proceed to identify and excise the tibialis anterior muscle from its origin at the anterolateral surface of the tibia and its insertion at the medial cuneiform via its distal tendon.
- In order to have leverage, hold the foot by grasping the digits with the index finger and thumb. Insert the tip of Dumont forceps immediately under the superficial distal tendon of the tibialis and move the forceps such that the blunt side can be used to detach the tibialis muscle belly from the underlying connective tissue.
- Cut the distal tendon using the fine curved scissors, and then cut the muscle at the origin, as close as possible to the tibia.
- Open the abdominal cavity by performing a sagittal incision starting in the hypogastric region and moving superiorly to the epigastric region, stopping the incision just above the diaphragm.
NOTE: The depth of the incision should only be enough to breach the abdominal muscle wall and the underlying peritoneal fascia. - Carefully remove the liver by grasping it with the blunt-tipped forceps, and use the fine curved scissors to dissect away any blood vessels or connective tissue that adhere the liver to the cavity. Take care not to leave any lobes behind. Weigh and snap freeze the liver in a tube in liquid nitrogen.
- Using the blunt-tipped forceps, move away the intestine, and then expose and delicately remove the spleen. Grasp the spleen gently with the blunt-tipped forceps and use the back or blunt side of the fine curved scissors to dissect away any connective tissue or blood vessels adhering the spleen to the cavity. Weigh and snap freeze it in a tube in liquid nitrogen.
- Identify the gonadal fat pads, adjacent to the epididymis (in males) or uterus (in females). Gently grasp the epididymal fat pad with the blunt-tipped forceps and pull the tissue out of the cavity. Using the fine curved scissors, cut away any gonadal tissue that may be attached to the fat pad. Weigh the fat pad and snap freeze it in a tube in liquid nitrogen.
NOTE: Generally, this adipose tissue is considered representative of the overall body fat mass in mice and declines in weight with cachexia. Other fat pads can be similarly identified, dissected, removed, and weighed. - Use the curved fine scissors to open the chest of the animal. Cut away the ribs attached to both lateral borders of the sternum. Remove the sternum. Using two pairs of forceps, grasp each side of the rib cage and pull each side laterally to open the thoracic cavity.
- Delicately pull the heart with the forceps. Carefully excise the heart with the curved fine scissors by severing the aorta at the left atrium. Make sure to empty the organ of any residual blood by delicately blotting it against some absorbent paper. Weigh and snap freeze it in a tube in liquid nitrogen.
5. Muscle Freezing and Mounting
- Immerse the bottom half of a small plastic beaker (50 ml) containing isopentane into liquid nitrogen. The isopentane is ready to use when it becomes slightly viscous and forms a solid white laminate lining the inside of the beaker (temperature: -160 °C).
NOTE: Always use nonsparking bronze or aluminum hand tools. Avoid breathing product vapor. Use with adequate ventilation. If dealing with a spill and ventilation is impossible or impractical, wear a suitable respirator. - Freeze some embedding medium on a chuck (cork) by dipping it briefly (10 sec) into isopentane.
- Handle the muscle (e.g., the tibialis or gastrocnemius) by holding it by the tendon, vertically relative to the cork, in order to maintain the orientation of the fibers and to allow for cross sections.
- Carefully position the end portion of the fresh muscle on top of the frozen embedding medium.
NOTE: The muscle will stick. It is important NOT to completely surround the muscle with embedding medium. It is also important to maintain the vertical orientation for subsequent determination of the cross-sectional area. - Dip the chuck with the attached muscle into the isopentane bath (the usual freezing time is 7 – 15 sec, depending on the specimen size and composition).
NOTE: Immersion in the freezing solution should not last more than is needed to completely freeze the specimen. Freezing too long will fracture the tissue block, while too short a time will cause ice crystal formation. A well-frozen specimen will be chalky white. - After freezing the specimen, place it into a small plastic bag or specimen tube (50 ml) and immediately store it in a deep freezer at -80 °C or in liquid nitrogen.
6. Analysis of the Raw Data
- In order to control for variations in size of the mice, normalize the final body weight (FBW); tumor-free body weight; and carcass, organ, and tissue weight (expressed in grams) according to the initial body weight (IBW), and express them as "weight/100 mg IBW".
NOTE: Alternatively, some investigators normalize the muscle mass to the tibia length. - Use a software for data analysis. Find the mean and SD of the normalized weights. Perform statistical analysis with unpaired Student's t-tests between controls and C26-bearing mice.
NOTE: Results can be presented relative to body (IBW and tumor-free FBW), tumor, organ, and tissue weights.
7. Muscle Sectioning
- Set the working temperature of the cryostat inner chamber to around -23 °C.
- Allow the specimen (previously stored at -80 °C) to acclimatize at the working temperature (a couple of hours should be enough).
- Cut multiple 8-μm thick sections of the specimen (preferably the tibialis anterior or gastrocnemius muscles) and collect them on glass slides.
NOTE: Cut the section at the mid-belly region of the muscle. Also, for the sake of accuracy, cut the sections perpendicularly to the mounting axis. - Keep the glass slides inside the cryostat chamber. Do not let them thaw if the aim is to perform IF/IHC studies or enzymatic staining.
- Store the slides at -80 °C for further analyses.
8. Evaluation of Myofiber Size by Laminin Immunofluorescence (Alternative 1)
NOTE: For the evaluation of muscle morphology and cross-sectional area (CSA), hematoxylin and eosin (H&E)- as well as immunofluorescence (IF)-based staining methods are accepted systems to determine fiber size19. Although the H&E staining of muscle sections represents a valuable and convenient method for the analysis of morphology, the use of an IF approach14 is significantly faster and modestly more accurate than the H&E-based method. H&E methods are described below.
- Remove tissue sections from either the cryostat or the -80 °C storage and allow them to equilibrate to room temperature (approximately 5 – 10 min).
NOTE: In order to avoid non-specific interactions associated with primary antibodies raised in mice, it is advisable to either use antibodies raised in another species or to make use of commercially available "mouse-on-mouse" immunodetection kits. - Immerse the sections in pre-cooled methanol (-20 °C) for 10 min.
- Wash the sections in room temperature 1x PBS for 5 min.
- Remove the PBS and use a pen for immunohistochemical applications to encircle the sections on the slide. Do NOT let sections completely dry at any point.
- Apply a suitable blocking buffer (5 – 8% FBS in PBS or 5% goat serum in PBS) onto the sections for 1 hr at RT.
- Remove the blocking buffer.
- Apply either a rabbit anti-laminin (1:30) or a mouse anti-dystrophin (1:30) primary antibody in blocking buffer to the sections for 2 hr at RT (or overnight at 4 °C) in a humid chamber.
- Wash the sections with 1x PBS for 5 min.
- Remove the wash buffer and wash sections again in 1x PBS for 5 min.
- Remove the wash buffer and apply a proper fluorescent secondary antibody (1:1,000 in blocking buffer) to the sections in a humid chamber for 1 hr at RT.
- Remove the secondary antibody and wash the sections with 1x PBS for 5 min.
- Remove the wash buffer and repeat the wash with 1x PBS for 5 min. Remove the wash buffer and cover the sections with a cover glass using an aqueous-based medium.
- Identify the morphological features of the muscle by using digital images of the stained section. In order to do so, use an inverted fluorescent light microscope for IF sections (a 10X or 20X magnification objective is appropriate), along with a digital camera and image capture software. Place a scale bar in each digital image (20 – 30 random fields are recommended for each muscle section) to facilitate the quantification of the CSA. Images should be saved in .TIF format to be compatible with the image analysis software using the ImageJ program.
- Use an ImageJ macro to distinguish connective tissue from contractile tissue (myofiber size) through the presence of the fluorescent secondary antibody in the endomysium20.
NOTE: The macro and instructions are available from the authors, Richard Lieber and Shannon Bremner. - Open the ImageJ program by double-clicking the icon on the desktop.
- Load the CSA macro by clicking once the "Plugins" tab. Move the cursor down to select the "Install" option by clicking it once.
- Load the cross-sectional macro by selecting it from the folder it has been saved in, and then click "Open".
- Once the macro has been opened, open the histological image containing the scale bar by clicking "File" and then "Open" from the ImageJ toolbar.
- Once the image is opened, select the "straight line tool" in the ImageJ program by clicking on the icon with the straight line.
- Once the "Straight line tool" is selected, click once on the left-most end of the scale bar and then click once on the right-most end of the scale bar. When done correctly, there should be a yellow line overlaid on the scale bar.
- Making sure that the yellow line is still overlaid on the scale bar, select the "Analyze" tab from the ImageJ toolbar by clicking it once.
- In the "Analyze" tab, move the cursor down and select "set scale" by clicking it once.
- In the "Set scale" window, DO NOT change the value in the "Distance in pixels" box. Enter the known distance of the scale bar in the "Known distance" box. Change the units in the "Unit of length" box to correspond with the scale bar units (i.e., micrometers = µm). Finally, select "global" by clicking the box to the left of "Global" once.
- Close the "Set scale" window.
- Open the first image to be measured by pressing the "O" button on the keyboard. Observe a window pop-up that allows the user to find and select the image to be measured from the folder it has been saved in. Select the image by clicking on it once and then clicking "Open."
- Once opened, a window should appear for the selection of the acceptable range for CSA measurements and for circularity.
NOTE: These settings are usually left to their default values. However, if one wishes to change these settings, be sure to keep the values the same for each consecutive image to be measured in the experiment. - After setting the CSA range and circularity, observe the image covered in red pixels with a "Threshold" window. Adjust the threshold of the image using the bars in the "Threshold" window until the muscle fibers are accurately filled in with red pixels.
NOTE: Attempt to obtain the greatest number of fibers that are not touching one another; if fibers are touching, they will be measured as one large fiber, which will need to be edited downstream. - After setting the threshold, press the "1" button on the keyboard to measure the muscle fibers.
- Circle the individual muscle fibers with a yellow line. Move through the image and remove artifacts or double fibers that have been selected by the program. To do this, select the structures that are not fibers by clicking them once, and then press the "D" button on the keyboard. This will delete them from the measurements.
- Select and copy the measurements from the "Measurement value window," and paste them into a spreadsheet for further analysis.
NOTE: The values obtained can also be used to assess the fiber distribution analysis, which generally represents a useful parameter to determine whether tumor growth promotes a shift towards smaller muscle fibers. - Close all of the open windows in ImageJ by clicking the "X" in the top-left corner of the windows.
- Open a new image to be measured by pressing "O" and repeating steps 8.24 – 8.29.
9. Evaluation of Myofiber Size from H&E Stained Sections (Alternative 2)
- Place the glass slides into Coplin jars containing filtered Harris hematoxylin for 8 min.
NOTE: Alternatively, Mayer hematoxylin can be used if a less intense stain is desired. - Rinse the slides in deionized water.
- Dip the slides in tap water for up to 5 min.
- Quickly rinse them in deionized water, and then incubate them in filtered 1% eosin solution for less than 2 min.
NOTE: If a more intense stain is desired, add 1 – 2 drops of acetic acid (0.01% or less) to the eosin solution. - Begin the dehydration steps. Rinse the slides in 70% ethanol for less than 1 min.
- Dip the slides in 90% ethanol for 1 min.
- Dip the slides in 95% ethanol for 1 min.
- Dip the slides in 100% ethanol for 2 min.
- Dip the slides in xylene for 2 min. Move the slides to another Copley jar containing fresh xylene for 2 more min.
NOTE: Keep the slides in Xylene (no longer than 1 hr) until they are coverslipped. - Place the coverslips on the muscle sections using Permount or a Xylene-based mounting medium of choice.
- Observe the H&E-stained glass slides under a microscope with a 20X magnification, and acquire pictures of the entire muscle section area. Obtain images by using a bright-light microscope, digital camera, and image capture software.
NOTE: The presence of a scale bar or image of a micrometer taken at the same magnification will be required to determine the CSA. As above, record at least 20 – 30 random fields for each muscle section. - Evaluate the myofiber size using ImageJ software. Open every digital image. Set the scale as above. Measure at least 1,500-2,000 muscle fibers per muscle using the freehand selection tool, tracing the fiber circumference with a mouse or, for faster input, with a tablet input device and pen.
10. Data Collection and Analysis
- In order to establish muscle fiber size, assess the average fiber "area." Calculate means and SD for the measurements obtained for each muscle, including both the area and Feret's diameter. In addition, in order to evaluate whether the experimental setting is associated with a shift towards either smaller or larger muscle fibers, analyze the "fiber distribution."
- Evaluate the significance of the result by performing an unpaired T-test for two groups. Determine the ratios between the values obtained for the C26 mice group and for the control group. Plot the values on a graph reporting the means SD and the frequency distribution/histogram aimed at demonstrating the shift in fiber sizes.
Representative Results
C26 tumor growth kinetics show a lag phase for the first 7 – 8 d after injection, followed by exponential cell growth (4 – 5 d). The tumor mass eventually reaches ~10% of the body weight (about 2 g; Figure 1A-B). During the first phase, the tumor can be located by palpation only and appears as a small protrusion of the skin. In the second phase, the tumor is observed as a mass under the skin. Rarely, the tumor becomes ulcerated, resulting in an open wound; in this case, the mouse is excluded from the experimental group and is humanely euthanized.
Body weight is unchanged in the first phase, but it is significantly reduced in the second phase, when it reaches 10 – 15% of the initial body weight (30% in the case of tumor-free weight; Figure 1A). Tumor-bearing mice appear wasted and showed disheveled fur at the end of the experimental period, with a body condition (BC) score equal to 118 (Figure 1C). BC1 represents a severely emaciated mouse, where skeletal structures are evident and the vertebrae are distinctly segmented. Body weight loss is mainly accounted for by both skeletal muscle and fat tissue wasting (Figure 1B). Body weight loss is consistent with a reduction of about 20 – 30% in skeletal muscle weight, in particular in the gastrocnemius, tibialis anterior, and quadriceps (Figure 2). The cardiac muscle is also significantly reduced in weight, although to a lesser extent when compared to the other skeletal muscles (Figure 2). Interestingly, hepatomegaly (+16%, p < 0.01) and splenomegaly (+110%, p < 0.01) are generally detected in tumor hosts, while fat mass, similar to skeletal muscle, is severely depleted (-70%, p < 0.001; Figure 3).
Skeletal muscle weight loss is also consistent with and proportional to the reduction in muscle fiber size, as observed after morphometric evaluation of muscle fiber CSA by means of an IF method (Figure 4A-B). In particular, the frequency distribution analysis showed a shift towards smaller-size fibers in C26-bearing mice, thus suggesting that the whole muscle undergoes atrophy in the presence of a C26 tumor (Figure 4C). Similar results can be observed by taking advantage of a traditional H&E-based methodology for the quantification of fiber size, although the magnitude of the change in muscle CSA associated with cancer growth is slightly different (38% versus control, p < 0.01 for the IF-based method; -18% versus control, p < 0.01 for the H&E-based method; Figure 5).
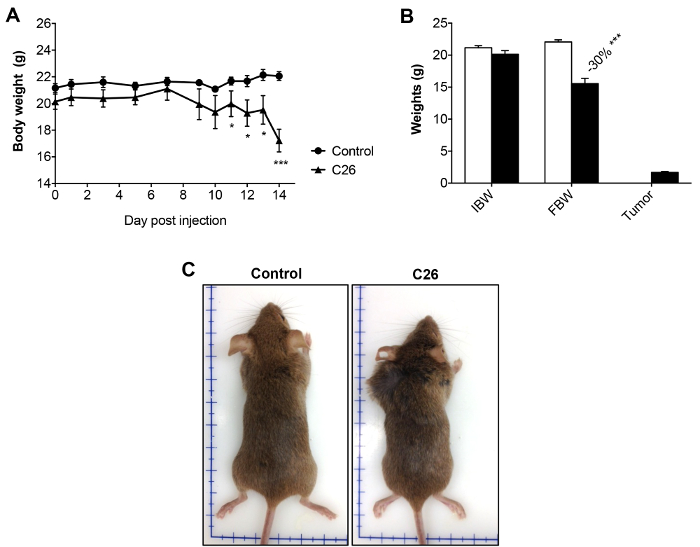
Figure 1: Body Weights in Control and C26-bearing Mice. A) Body weight curve in control and C26 hosts over the entire experimental period (14 days after tumor injection). B) Initial body weight (IBW), final tumor-free body weight (FBW), and tumor weight in control and C26-bearing animals. C) Representative pictures of controls and C26-bearers at the time of sacrifice (day 14 after tumor inoculation). Data are expressed as the mean SD. n = 6. Significance of the differences: *p < 0.05, **p < 0.01, ***p < 0.001 versus the control group using Student's t-test. Please click here to view a larger version of this figure.
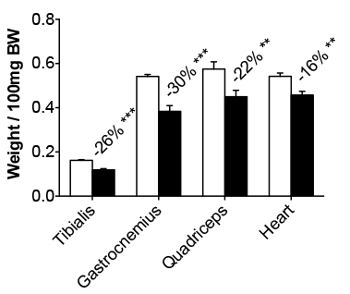
Figure 2: Muscle Weights in Control and C26-bearing Mice. The tibialis, gastrocnemius, quadriceps, and heart weights in control and C26 tumor-bearers are reported as the weight/100 mg IBW. The data are expressed as the mean SD. n = 6. Significance of the differences: **p < 0.01 and ***p < 0.001 versus the control group using Student's t-test.
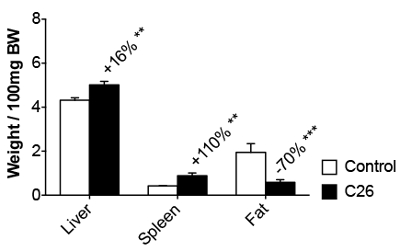
Figure 3: Organ Weights in Control and C26-bearing Mice. The liver, spleen, and epididymal fat pad weights in control and C26 tumor-bearers are reported as the weight/100 mg IBW. The data are expressed as the mean SD. n = 6. Significance of the differences: **p < 0.01 versus the control group.
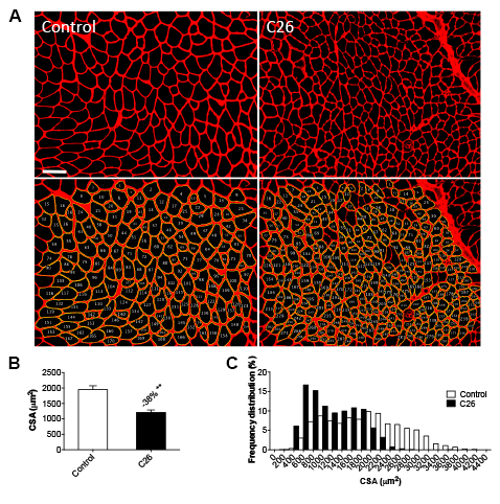
Figure 4: Morphometry for Myofiber Size by Immunofluorescence. A) Fluorescent immunohistochemical reaction of tibialis sections using an anti-dystrophin antibody (Vector Laboratories) from control and C26 hosts. Magnification: 10X. Size bar: 100 µm. B) Quantification of CSA in the tibialis muscle measured with the ImageJ macro. C) Frequency distribution of CSA in the tibialis muscle of control and C26-bearing mice. The data are expressed as the mean ± SD. n = 6 for each group. Significance of the differences: **p < 0.01 versus the control group using Student's t-test.
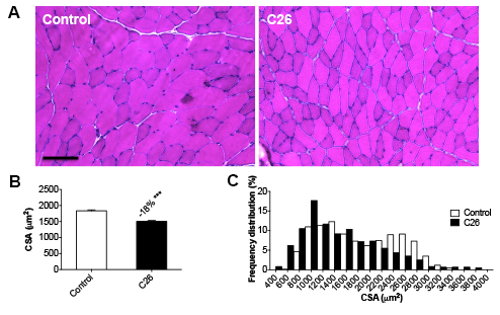
Figure 5: Morphometry for Myofiber Size by H&E. A) Hematoxylin & eosin staining of tibialis sections from control and C26 hosts. Magnification: 20X. Size bar: 100 µm. B) Quantification and frequency distribution of CSA in the tibialis muscle. The data are expressed as the mean SD. n = 6. Significance of the differences: ***p < 0.001 versus the control group using Student's t-test.
| Autori | Mouse strain | Mouse gender | Cell number | Origin | Implanted as solid tumor? | Site of injection | Days |
| Lazarus et al., 1999 | CD2F1 | M | 5 x 105 | NCI | NO | Dorsal region | 17 |
| Al-Majid and McCarthy, 2001 | CD2F1 | F | 2.5 x 106 | OSUMC | NO | Dorsal region | 17 |
| Samuels et al., 2001 | Balb/c | M | 0.5 g/ml of tumor suspension | ns | YES | Dorsal region | Up to 11 |
| Acharyya et al., 2004 | CD2F1 | M | 1 x 107 | OSUMC | NO | Right flank | Up to 24 |
| Aulino et al., 2010 | Balb/c | ns | 0.5 mm3 | NCI | YES | Dorsal region | 21 |
| Benny-Klimek et al., 2010 | CD2F1 | F | 1 x 106 | OSUMC | NO | Dorsal region | 14 |
| Bonetto et al., 2011; 2012 | CD2F1 | F | 1 x 106 | OSUMC | NO | Dorsal region | 14 |
| Cosper and Leinwand, 2010 | CD2F1 | M+F | 5 x 105 | ns | NO | Right flank | Up to 27 |
| Murphy et al., 2012 | CD2F1 | ns | 5 x 105 | NCI/OSUMC | NO | Dorsal region | 14 |
| Cornwell et al., 2014 | CD2F1 | M | 5 x 105 | NCI | NO | Right&Left flank | Up to 26 |
| Assi et al., 2016 | Balb/c | ns | 1 x 106 | Cell Line Service | NO | Dorsal region | Up to 22 |
Table 1: Comparison of the Different Protocols used for C26 Implantation. Different protocols for C26 implantation are provided, with particular reference to the mouse strain used, the cell number and type of tumor inoculated, the cell origin, the site of injection, and the length of the experimental period. OSUMC: Ohio State University Medical Center. NCI: National Cancer Institute. Commercial cell line service NS: not specified.
Discussion
Especially in its latest stages, colorectal cancer is associated with the development of cachexia, which is responsible for poorer outcomes and reductions in patient quality of life. Many studies have focused on the treatment of conditions secondary to cancer; however, despite many efforts in this direction, there is still no approved therapy for cancer cachexia21. Thus, it is imperative that animal models resemble the human pathology as closely as possible in order to maximize the translation of findings.
C26 tumor-bearing mice are a commonly used experimental model of cancer cachexia22-24. This model closely resembles the human disease, showing reductions in body, muscle, and fat mass, as well as muscle fiber atrophy and increased expression of inflammatory genes and ubiquitin ligases consistent with marked protein hypercatabolism2,5. Despite this, a few disparities with data obtained in cancer patients have been reported25,26. Indeed, some features of the model, including the non-physiologic growth environment (e.g., an ectopic tumor grown subcutaneously instead of orthotopically implanted in the GI tract), the relatively short experimental period compared to other models (e.g., genetic or orthotopic injection of tumors), the dependency on IL-6 action, and the necessary use of CD2F1 or Balb/c mice may represent severe limitations and could constrain the interpretation of results.
The current protocol has proven to be highly reproducible across many experiments performed in our laboratory, maintaining the same characteristics and allowing comparisons between results obtained at different times, in different geographic locations, and by different researchers3,10. However, in order to promote the reproducibility of data, it is important to establish clear and common guidelines.
As gathered from the literature, several caveats might prevent the reproduction of the same data in two different laboratories. For the same reasons, the use of different protocols may generate different phenotypes and outcomes, as well as lead to conflicting and questionable results. Indeed, differences in performing this animal model have been reported, mainly resulting from the strain (Balb/c or CD2F1)2,5,27-29 or sex of mice used17, the type of tumor that was implanted (a cell suspension3,29 rather than a solid tumor in a graft2,30), the tumor source (NCI, ATCC, or OSUMC), the number of C26 cells that were injected, and the site of injection or implantation (the flanks6,17,31 or the dorsal region3,10,32). No direct comparison of the effects associated with the implantation of C26 cell lines obtained from different sources (mainly OSUMC versus NCI) has ever been performed, thus preventing us from drawing any definitive conclusions. However, it is likely that the choice of the source of cells may also significantly influence the expected outcomes. When considering the choice of males versus females, investigators should be reminded of significant differences in the outcomes. Indeed, as reported by Cosper and Leinwand17, male tumor-bearing mice, due to the absence of estrogens, may show a more severe phenotype than females, including greater cardiac mass loss and mortality, a more robust pro-inflammatory response to the tumor, and greater cardiac autophagy.
Especially for those investigators that are not familiar with this model and may initially face problems, based on both a power analysis and our previous experience, the use of at least 6 animals per experimental condition (n = 6) is advisable in order to detect statistically significant differences. It is also critical that randomization is performed carefully, so that initial body weights in the experimental animals do not differ significantly among groups. Similarly, it is advised that, in order to avoid inter-operator variability, the same investigator perform the tissue and organ collection on every animal. Further, it is imperative that tissues (particularly muscle) are frozen as fast as possible in order to preserve the RNAs and enzymatic structure and activity, especially if the goal of the study is to assess gene expression or to evaluate enzymatic activities. Moreover, in the attempt to determine muscle CSA, we showed that reporting the fiber area is generally accepted and representative of muscle fiber size. Here, we present two distinct methods to assess muscle CSA. As shown in Figures 4-5, both methods were able to detect a significant reduction in myofiber size between controls and tumor hosts, although the degree of wasting appeared different.
This discrepancy may result from the fact that, although both methods are acceptable ways to assess muscle fiber size, quantification of muscle characteristics from H&E-stained slides is still a manual or semi-automatic process, most often labor-intensive, time-consuming, and affected by limited accuracy. Based on our experience, we believe that the IF-based method is a more accurate technique to report muscle CSA. Indeed, the number of fibers that can be measured automatically by taking advantage of this technique is significantly larger, thus increasing the accuracy of the measurement. Of note, for both techniques, an alternative method of reporting muscle size is assessing the Feret's diameter. Interestingly, this is considered a very reliable tool due to the fact that this parameter is largely independent of the angle of sectioning. Other parameters, such as the "minimal inner diameter" and the "minimal outer diameter" are also insensitive to the plane of sectioning and can be used instead of the Feret's diameter as alternative indications of the fiber size.
In conclusion, although the cachexia community clearly needs to establish more physiological models for the study of tumor-associated muscle wasting, we believe that mice bearing the C26 colon carcinoma represent a well-standardized and easy-to-use model to investigate molecular alterations and physiological abnormalities usually detected after the occurrence of a tumor. Future applications will involve the investigation of whether orthotopic implantation of C26 cells into the colon might represent a proper and more physiological model of colorectal cancer cachexia.
Divulgazioni
The authors have nothing to disclose.
Acknowledgements
We thank Richard Lieber and Shannon Bremner for their ImageJ macro and instructions. While at Thomas Jefferson University, this work was supported by the Pennsylvania Department of Health CURE Grant TJU No. 080-37038-AI0801. Subsequently, this study was supported by a grant to AB from the National Institutes of Health (R21CA190028), and by grants to TAZ from the National Institutes of Health (R01CA122596, R01CA194593), the IU Simon Cancer Center, the Lustgarten Foundation, the Lilly Foundation, Inc., and the IUPUI Pancreas Signature Center.
Materials
| Cell culture Flasks | Falcon – Becton Dickinson | 35-5001 | |
| DMEM | Cellgro | 10-017-CV | |
| FBS | Gibco | 26140 | |
| Streptomycin-Penicillin | Cellgro | 30-002-CI | |
| CD2F1 mice | Harlan | 060 | |
| Anesthesia apparatus | EZ-Anesthesia | EZ-7000 | |
| 2-Methyl Butane | Sigma-Aldrich | M32631 | |
| OCT | Tissue-Tek | 4583 | |
| Cryostat | Leica | CM1850 | |
| Cork disks | Electron Microscopy Sciences | 63305 | |
| Superfrost plus glass slides | VWR | 48311-703 | |
| Anti-Laminin Rabbit polyclonal antibody | Sigma-Aldrich | L9393 | |
| Anti-Dystrophin Mouse Monoclonal antibody | Vector Laboratories | VP-D508 | |
| Alexa Flour 594 anti-mouse IgG | Life Technologies | A11062 | |
| Alexa Flour 594 anti-rabbit IgG | Life Technologies | A21211 | |
| Hematoxylin | Sigma-Aldrich | GHS216 | |
| Eosin | Sigma-Aldrich | HT110332 | |
| Xylene | Acros Organics | 422680025 | |
| Cytoseal-XYL | Thermo | 8312-4 | |
| Microscope | Zeiss | Observer.Z1 | |
| Bamboo Tablet | Wacom | CTH-661 | |
| Prism 7.0 for Mac OS X | GraphPad Software, Inc. | ||
| Excel for Mac 2011 | Microsoft Corp. | ||
| Image J | US National Institutes of Health | IJ1.46 | http://rsbweb.nih.gov/ij/download.html |
| Microtainer | BD | 365873 |
Riferimenti
- Tan, B., Fearon, K. Cachexia: prevalence and impact in medicine. Curr Opin Clin Nutr Metab Care. 11, 400-407 (2008).
- Aulino, P., et al. Molecular cellular and physiological characterization of the cancer cachexia-inducing C26 colon carcinoma in mouse. BMC cancer. 10, 363 (2010).
- Bonetto, A., et al. STAT3 activation in skeletal muscle links muscle wasting and the acute phase response in cancer cachexia. PloS One. 6, e22538 (2011).
- Bonetto, A., et al. JAK/STAT3 pathway inhibition blocks skeletal muscle wasting downstream of IL-6 and in experimental cancer cachexia. Am J Physiol Endocrinol Metab. 303, E410-E421 (2012).
- Bonetto, A., et al. Deacetylase inhibitors modulate the myostatin/follistatin axis without improving cachexia in tumor-bearing mice. Current Cancer Drug Targets. 9, 608-616 (2009).
- Acharyya, S., et al. Cancer cachexia is regulated by selective targeting of skeletal muscle gene products. The Journal of Clinical Investigation. 114, 370-378 (2004).
- Fearon, K., et al. Definition and classification of cancer cachexia: an international consensus. The Lancet Oncology. 12, 489-495 (2011).
- Holecek, M. Muscle wasting in animal models of severe illness. Int J Exp Pathol. 93, 157-171 (2012).
- Acharyya, S., et al. Dystrophin glycoprotein complex dysfunction: a regulatory link between muscular dystrophy and cancer cachexia. Cancer Cell. 8, 421-432 (2005).
- Benny Klimek, ., E, M., et al. Acute inhibition of myostatin-family proteins preserves skeletal muscle in mouse models of cancer cachexia. Biochemical and Biophysical Research Communications. 391, 1548-1554 (2010).
- Pedroso, F. E., et al. Inflammation, organomegaly, and muscle wasting despite hyperphagia in a mouse model of burn cachexia. Journal of Cachexia, Sarcopenia and Muscle. 3, 199-211 (2012).
- Soda, K., Kawakami, M., Kashii, A., Miyata, M. Manifestations of cancer cachexia induced by colon 26 adenocarcinoma are not fully ascribable to interleukin-6. International journal of cancer. 62, 332-336 (1995).
- Costelli, P., et al. IGF-1 is downregulated in experimental cancer cachexia. American journal of physiology. Regulatory, Integrative and Comparative Physiology. 291, R674-R683 (2006).
- Palus, S., Akashi, Y., von Haehling, S., Anker, S. D., Springer, J. The influence of age and sex on disease development in a novel animal model of cardiac cachexia. International Journal of Cardiology. 133, 388-393 (2009).
- Norman, K., et al. Effect of sexual dimorphism on muscle strength in cachexia. Journal of Cachexia, Sarcopenia and Muscle. 3, 111-116 (2012).
- Stephens, N. A., et al. Sexual dimorphism modulates the impact of cancer cachexia on lower limb muscle mass and function. Clinical Nutrition. 31, 499-505 (2012).
- Cosper, P. F., Leinwand, L. A. Cancer causes cardiac atrophy and autophagy in a sexually dimorphic manner. Ricerca sul cancro. 71, 1710-1720 (2011).
- Ullman-Cullere, M. H., Foltz, C. J. Body condition scoring: a rapid and accurate method for assessing health status in mice. Laboratory Animal Science. 49, 319-323 (1999).
- Bonetto, A., Andersson, D. C., Waning, D. L. Assessment of muscle mass and strength in mice. Bonekey Rep. 4, 732 (2015).
- Minamoto, V. B., et al. Increased efficacy and decreased systemic-effects of botulinum toxin A injection after active or passive muscle manipulation. Dev Med Child Neurol. 49, 907-914 (2007).
- Murphy, K., Lynch, G. Update on emerging drugs for cancer cachexia. Expert Opin Emerg Drugs. 14, 619-632 (2009).
- Seto, D. N., Kandarian, S. C., Jackman, R. W. A Key Role for Leukemia Inhibitory Factor in C26 Cancer Cachexia. The Journal of Biological Chemistry. 290, 19976-19986 (2015).
- Judge, S. M., et al. Genome-wide identification of FoxO-dependent gene networks in skeletal muscle during C26 cancer cachexia. BMC Cancer. 14, 997 (2014).
- Kliewer, K. L., et al. Adipose tissue lipolysis and energy metabolism in early cancer cachexia in mice. Cancer Biol Ther. 16, 886-897 (2015).
- Aversa, Z., et al. Changes in myostatin signaling in non-weight-losing cancer patients. Ann Surg Oncol. 19, 1350-1356 (2012).
- Bonetto, A., et al. Early changes of muscle insulin-like growth factor-1 and myostatin gene expression in gastric cancer patients. Muscle Nerve. 48, 387-392 (2013).
- Lazarus, D. D., et al. A new model of cancer cachexia: contribution of the ubiquitin-proteasome pathway. The American Journal of Physiology. 277, E332-E341 (1999).
- al-Majid, S., McCarthy, D. O. Resistance exercise training attenuates wasting of the extensor digitorum longus muscle in mice bearing the colon-26 adenocarcinoma. Biol Res Nurs. 2, 155-166 (2001).
- Bonetto, A., et al. JAK/STAT3 pathway inhibition blocks skeletal muscle wasting downstream of IL-6 and in experimental cancer cachexia. American journal of physiology. Endocrinology and metabolism 303. 303, E410-E421 (2012).
- Samuels, S. E., et al. Liver protein synthesis stays elevated after chemotherapy in tumour-bearing mice. Cancer Lett. 239, 78-83 (2006).
- Cornwell, E. W., Mirbod, A., Wu, C. L., Kandarian, S. C., Jackman, R. W. C26 cancer-induced muscle wasting is IKKbeta-dependent and NF-kappaB-independent. PloS One. 9, e87776 (2014).
- Penna, F., et al. Muscle wasting and impaired myogenesis in tumor bearing mice are prevented by ERK inhibition. PloS One. 5, e13604 (2010).

