Long-Term Culture of Individual Caenorhabditis elegans on Solid Media for Longitudinal Fluorescence Monitoring and Aversive Interventions
Summary
Here, we present a protocol to culture isolated individual nematodes on solid media for lifelong physiological parameter tracking and fluorescence quantification. This culture system includes a palmitic acid barrier around single-worm wells to prevent animals from fleeing, allowing the use of aversive interventions, including pathogenic bacteria and chemical stressors.
Abstract
Caenorhabditis elegans are widely used to study aging biology. The standard practice in C. elegans aging studies is to culture groups of worms on solid nematode growth media (NGM), allowing the efficient collection of population-level data for survival and other physiological phenotypes, and periodic sampling of subpopulations for fluorescent biomarker quantification. Limitations to this approach are the inability to (1) follow individual worms over time to develop age trajectories for phenotypes of interest and (2) monitor fluorescent biomarkers directly in the context of the culture environment. Alternative culture approaches use liquid culture or microfluidics to monitor individual animals over time, in some cases including fluorescence quantification, with the tradeoff that the culture environment is contextually distinct from solid NGM. The WorMotel is a previously described microfabricated multi-well device for culturing isolated worms on solid NGM. Each worm is maintained in a well containing solid NGM surrounded by a moat filled with copper sulfate, a contact repellent for C. elegans, allowing longitudinal monitoring of individual animals. We find copper sulfate insufficient to prevent worms from fleeing when subjected to aversive interventions common in aging research, including dietary restriction, pathogenic bacteria, and chemical agents that induce cellular stress. The multi-well devices are also molded from polydimethylsiloxane, which produces high background artifacts in fluorescence imaging. This protocol describes a new approach for culturing isolated roundworms on solid NGM using commercially available polystyrene microtrays, originally designed for human leukocyte antigen (HLA) typing, allowing the measurement of survival, physiological phenotypes, and fluorescence across the lifespan. A palmitic acid barrier prevents worms from fleeing, even in the presence of aversive conditions. Each plate can culture up to 96 animals and easily adapts to a variety of conditions, including dietary restriction, RNAi, and chemical additives, and is compatible with automated systems for collecting lifespan and activity data.
Introduction
C. elegans are a powerful model organism for research in genetics, cellular biology, and molecular biology, because they are easily cultured in the laboratory, have a short generation time and lifespan, share a high degree of protein homology with mammals, and have a transparent body structure that allows in vivo visualization of fluorescent proteins and dyes1. As a result of the long-standing use of C. elegans as a major model system in a range of fields, including developmental biology and aging, their growth and development are well-understood, their genome has been fully sequenced, and a host of powerful genetic tools have been created, including genome-wide RNAi feeding libraries and thousands of mutant and transgenic strains. Historically, C. elegans are cultivated as populations on solid agar nematode growth media (NGM), and phenotypes are manually evaluated either by direct observation or by imaging and downstream analysis. Fluorescent microscopy is used to capture a variety of molecular phenotypes using dyes or transgenically expressed fluorescent tags in individual C. elegans. Fluorescent imaging typically involves fixing or paralyzing an animal on slides containing thin agarose pads, which is invasive and often lethal. It also involves the use of chemicals, such as levamisole or sodium azide, which can potentially interfere with the molecular process of interest2,3. Together, these approaches allow cross-sectional, population-level data to be collected across a broad range of phenotypes, but do not allow the tracking of individual animals over time.
In recent years, several approaches have emerged to cultivate isolated C. elegans, allowing researchers to capture dynamic changes in physiological and molecular phenotypes of animals over time utilizing new imaging technologies. One category of C. elegans culture approach is microfluidics devices, including WormFarm4, the Nemalife chip5, and the 'behavior' chip by Chronis et al.6, among various others7,8,9. Related to these are liquid-based culture methods, that use multi-well plates to characterize individual worms or small populations over time10,11. Microfluidics and microplate systems provide excellent quantitative measurements of phenotypic responses in C. elegans down to a single animal, but the culture environment presents a key limitation. The vast majority of past research in C. elegans, particularly in the field of aging, has been completed on solid agar-based media. Liquid culture causes C. elegans to swim continuously and represents a distinct environmental context that can alter the underlying biology. For example, animals cultured in liquid media have drastically altered fat content and gene expression — particularly for genes involved in the stress response — relative to animals cultured on agar-based solid NGM12,13. An alternative category of single-animal imaging methods involves polydimethylsiloxane (PDMS) devices that isolate individual animals on solid media, in an effort to more closely mimic the standard environment experienced by worms cultured on solid NGM in group culture on Petri plates. The WorMotel is a 240-well PDMS device designed to culture individual animals on solid media. Each well is filled with a modified NGM using low-melt agarose in place of agar and seeded with bacterial food, creating a solid media environment similar to the most common culture system using Petri plates. The well walls are round, allowing each animal to be imaged regardless of location in the well (avoiding the visual obscuring caused by an animal near a wall in a multi-well plate). Copper sulfate in a narrow moat surrounding each well is used as a deterrent to keep animals in their wells14,15. A limitation of this approach is that the copper sulfate is ineffective at preventing worms from fleeing when aversive environmental conditions are present, including dietary restriction, pathogenic bacteria, or chemicals that induce cellular stress (e.g., paraquat).
A second system that uses solid media is the Worm Corral, which employs a hydrogel to create a small sealed environment for each worm on a slide, allowing long-term monitoring of individually isolated animals16. A key limitation is that animals must be sealed into the environment as eggs, requiring the use of sterile animals to prevent reproduction, and limiting drug treatments to a single application. Multi-dose drug trials can be accomplished in the WorMotel either by conducting multiple rounds of exposure prior to transferring worms to the device or by topically adding additional drugs to the wells during the experiment; however, in the latter case, the actual exposure dose after adding an additional drug to an existing well is difficult to precisely quantify and depends on how rapidly the drug degrades. Both the WorMotel and the Worm Corral are excellent for brightfield or darkfield imaging to capture information related to activity and animal physiology (e.g., growth and development). While these systems can be used to monitor fluorescence, in our experience, the PDMS used to create the other single-worm imaging technologies is prone to forming microbubbles, capturing particulate, and other small abnormalities that generate irregular fluorescent artifacts that interfere with consistent fluorescence visualization and quantification, especially in the emission range for GFP, the most common fluorophore used in C. elegans research. To date, live fluorescence imaging of C. elegans individual animals in a longitudinal manner primarily relies on microfluidics devices17.
Here, we describe a novel method for culturing individual C. elegans on solid media that is compatible with both aversive interventions and direct fluorescent imaging. This approach is similar in concept to other single-worm imaging technologies, except that the custom-molded PDMS chip is replaced with commercially available polystyrene microtrays originally developed for micro cytotoxicity assays (also commonly called Terasaki trays)18. These microtrays feature wells that can be filled with solid media and seeded with bacterial food, closely mimicking the environment experienced by animals under standard solid NGM culture methodology. Each well is surrounded by an aversive barrier of palmitic acid rather than copper sulfate. Palmitic acid is commonly used to prevent worms from fleeing solid media, using standard group culture on Petri plates in experiments where worms are challenged with an aversive environment like dietary restriction or exposure to a chemical stressor. The microtrays also produce minimal and consistent fluorescent background, allowing fluorescent imaging of animals directly in their culture environment. This new single-animal solid agar-based culture system not only allows for tracking individual animals throughout life and monitoring growth, development, activity, and lifespan, but is also compatible with direct fluorescent microscopy. Because the worms can be imaged without paralysis or fixation, in vivo fluorescence biomarkers can be quantified longitudinally in individual animals remaining on their culture media, allowing the observation of dynamic changes over the lifetime of each animal. This culture system is also compatible with current generation automated systems for tracking lifespan and other health metrics14,19. We provide a detailed protocol for culturing individual C. elegans in this microtray-based system, discuss potential pitfalls and troubleshooting, and discuss the advantages and limitations relative to other systems, and in particular, an updated and optimized WorMotel protocol15.
Each single-worm culture environment consists of a microtray mounted inside a standard single-well tray using a custom 3D-printed adapter (Figure 1A). The wells are filled with low-melt agarose nematode growth media (lmNGM), seeded with concentrated bacteria as a food source, and surrounded by a palmitic acid coating to prevent worms from fleeing (Figure 1B). The space between the microtray and the walls of the single-well plate is filled with saturated water crystals to maintain humidity (Figure 1B). A detergent coating is applied to the tray lid to prevent condensation. A single worm is added to each well, and the single-well tray is sealed with Parafilm to maintain moisture and allow oxygen exchange. Up to six microtrays can reasonably be prepared in parallel by a single practiced researcher.
Protocol
1. Recipes
NOTE: Prepare stock solutions before starting microtray plate preparation.
- Stock solutions for low-melt agarose nematode growth media (lmNGM)
- Prepare 1 M K2HPO4 by dissolving 174.18 g of K2HPO4 in 1 L of sterile deionized water in a 1 L bottle. Autoclave the solution at 121 °C, 15 psi, for 30 min and store it at room temperature (RT).
- Prepare 1 M KPi (pH 6.0) by dissolving 136.09 g of KH2HPO4 in 1 L of sterile deionized water in a 1 L bottle. Titrate the solution with 1 M K2HPO4 to achieve pH 6.0. Autoclave the solution at 121 °C, 15 psig, for 30 min and store it at RT.
- Prepare 1 M CaCl2 by dissolving 73.5 g of CaCl2 in 500 mL of sterile deionized water in a 500 mL bottle. Autoclave the solution at 121 °C, 15 psig, for 30 min and store it at RT
- Prepare 1 M MgSO4 by dissolving 123.25 g of MgSO4 in 500 mL of sterile deionized water in a 500 mL bottle. Autoclave the solution at 121 °C, 15 psig, for 30 min and store it at RT.
- Prepare 3 M NaCl for lmNGM: In clean 100 mL bottle, combine 100 mL of ultrapure water and 18.20 g of NaCl. Autoclave at 121 °C, 15 psig, for 30 min.
- Prepare 5 mg/mL cholesterol by combining 2.5 g of cholesterol, 275 mL of 100% ethanol, and 25 mL of sterile deionized water in a 500 mL amber bottle. Autoclave the solution at 121 °C, 15 psig, for 30 min and store it at RT.
- Prepare 50 mM fluorodeoxyuridine (FUdR) by dissolving 0.1231 g of FUdR in 10 mL of sterile deionized water in a 15 mL conical tube. Use a 10 mL syringe and 0.22 µm filter to sterilize the solution. Aliquot 1 mL of the solution each into 10 microcentrifuge tubes and store them at -20 °C.
- Prepare 50 mg/mL carbenicillin by dissolving 500 mg of carbenicillin in 10 mL of sterile deionized water in a 15 mL conical tube. Use a 10 mL syringe and 0.22 µm filter to sterilize the solution. Aliquot 1 mL of the solution each into 10 microcentrifuge tubes and store them at -20 °C.
- Prepare 1 mM isopropyl ß-D-1-thiogalactopyranoside (IPTG) by dissolving 2.38 g of IPTG in 10 mL of sterile deionized water in a 15 mL conical tube. Use a 10 mL syringe and 0.22 µm filter to sterilize the solution. Aliquot 1 mL of the solution each into 10 microcentrifuge tubes and store them at -20 °C.
- Prepare 100 mg/mL ampicillin by dissolving 1 g of ampicillin in 10 mL of sterile deionized water in a 15 mL conical tube. Use a 10 mL syringe and 0.22 µm filter to sterilize the solution. Aliquot 1 mL of the 100 mg/mL ampicillin solution each into 10 microcentrifuge tubes and store the tubes at -20 °C.
- Preparing nematode growth media (NGM) for general C. elegans maintenance
- To make 50 plates, dissolve 10 g of Bacto agar, 1.5 g of NaCl, and 1.25 g of Bacto peptone in 486 mL of ultrapure water in a 1 L flask. Sterilize the medium by autoclaving it on a liquid cycle at 121 °C, 15 psig, for at least 30 min.
- Post-autoclave, allow the medium to cool to 55 °C. Then, add 12.5 mL of 1 M KPi, 500 µL of 1 M MgSO4, 500 µL of 1 M CaCl2, and 500 µL of 5 mg/mL cholesterol to the solution.
- Using a sterile technique, pour 10 mL of the medium prepared in step 1.2.2 into each 60 mm plate (50 total). Leave the plates with media for at least 30 min to solidify. Pipette 300 µL of overnight grown bacterial culture onto the plate's center and leave the plate on the bench. Allow the culture to dry and grow thicker for 1-2 days. Store these NGM plates with bacteria at 4 °C.
- Preparing lmNGM premixture
- In a 250 mL Erlenmeyer flask, combine 2 g of low-melt agarose, 0.25 g of peptone, 1 mL of 3 M NaCl, and 99 mL of ultrapure water. Autoclave the solution at 121 °C, 15 psig, for 30 min.
- Aliquot 10 mL of lmNGM into test tubes and cover with Parafilm to prevent loss of liquid. Cap the tubes and allow them to cool and solidify.
- Prepare 85 mM NaCl for bacterial food: In a 100 mL bottle, combine 515.61 mg of NaCl and fill up to 100 mL with ultrapure water. Autoclave at 121 °C, 15 psig, for 30 min.
- Prepare water-absorbing polyacrylamide crystals by dissolving 1 g of Type S super absorbent polymer in 150 mL of ultrapure water in an autoclavable squeeze bottle. Cut the tip to ensure an adequate dispense size. Autoclave at 121 °C, 15 psig, for 30 min, cap with tinfoil, and shake to mix up the crystals.
- Prepare a detergent solution by combining 3 mL of 100% Tween-20 with 7 mL of ultrapure water in a 15 mL conical vial.
- Prepare 5 mg/mL cholesterol by combining 2.5 g of cholesterol, 275 mL of 100% EtOH, and 25 mL of ultrapure water in a 500 mL bottle.
- Prepare palmitic acid working solution
- Grind 500 mg of solid palmitic acid in a mortar and pestle and combine it with 15 mL of Tween-20 in a 50 mL conical vial. Mix by vortexing, dissolving as many of the crystals as possible.
- Add 17.5 mL of 100% ethanol and vortex the solution to dissolve as much palmitic acid as possible.
- Add 17.5 mL of ultrapure water and vortex rigorously. Gently heat the solution in a bead bath at 80 °C until the palmitic acid completely dissolves. Some palmitic acid may precipitate during cooling.
- Prepare lysogeny broth (LB) by mixing 20 g of LB powder in 1 L of deionized water in a 1 L or larger beaker. Aliquot 500 mL of the broth into two 500 mL beakers. Autoclave the broth at 121 °C, 15 psig, for 30 min. Store the prepared broth at RT.
- Preparing lysogeny broth (LB) agar plates
- Mix 10 g of LB powder and 7.5 g of Bacto agar in 500 µL of deionized water in a 1 L flask. Autoclave the solution on a liquid cycle at 121 °C, 15 psig, for 30 min.
- If desired, add optional antibiotics (500 µL of 100 µg/mL ampicillin) after the autoclaved media is cooled to 55 °C. Using a sterile technique, make LB agar plates by pouring 20 mL of media into each 100 mm plate (25 total). Store the plates at 4 °C.
2. Preparing a population of age-synchronized worms
- Prepare a population of age-synchronized worms that are ready to add to the microtray at larval stage 4 (L4). Alternatively, worms at any life stage can be added (FUdR should be excluded from the microtray media if worms are added at a developmental stage earlier than L4 to prevent developmental arrest).
NOTE: This step should be completed 2 days before adding the worms to the microtray if the target life stage at the start of the experiment is L4. The synchronization timing can be adjusted to allow the worms to reach the desired life stage. - Temperature can impact lifespan. For consistent results, use standard NGM plates (see section 1) to maintain C. elegans at 20 °C, and transfer worms to fresh plates if they run low on bacterial food.
- From the stock plates with many young adult worms, use a standard approach to generate an age-synchronized population, most commonly bleaching20 or timed egg laying21.
- Isolate the eggs and add them to a bacteria-spotted NGM plate. If using wild-type C. elegans, the eggs will hatch and form worms, reaching the L4 larval stage in 2 days.
3. Preparing bacteria culture
- The day before the beginning of the experiment, inoculate the desired bacterial food source. Prepare ~5 mL of bacterial culture per plate.
NOTE: The bacterial streak can be prepared up to 1 month in advance and stored at 4 °C for E. coli food source.- Streak bacteria for single colonies onto the LB agar plate from a frozen glycerol stock.
- Grow the culture at 37 °C overnight.
- Inoculate liquid LB broth with a single colony per inoculation.
- Shake at ~250 rpm at 37 °C for 12-16 h.
4. Applying palmitic acid coating to microtray
NOTE: Palmitic acid serves as an aversive barrier to prevent the fleeing of animals from the individual wells. The coating is applied to the entire bottom surface of the microtray, with the exception of the inside surfaces of the wells. Nystatin is added to the palmitic acid to mitigate fungal contamination. This step can be completed 1 day prior to the experiment start if desired.
- Set the bead bath to 80 °C to allow time to preheat.
- Immediately prior to use, aliquot 1 mL of the palmitic acid working solution per microtray into a separate sterile container (typically a 15 mL conical vial) and add 4 µL of 10,000 units/mL nystatin per 1 mL of palmitic acid solution before heating.
- Spray inside and outside of the microtray with ethanol (70% or greater) to saturation and shake off the excess ethanol.
NOTE: Residual ethanol should evaporate during treatment in UV crosslinking (step 4.4). - Use a UV crosslinker to sterilize the microtray (the following instructions are for the UV crosslinker used in this study):
- Place the microtray and lid in the UV crosslinker separately (i.e., do not place the lids on the plates, as UV does not penetrate the plastic).
- Close the door and turn on the UV crosslinker.
- Press Time, enter 20 min, and then press Start.
- After 20 min, flip over the plate and lid and repeat steps 4.2.2-4.2.3.
- While wearing gloves, place the lids on the microtray to prevent contamination after sterilization.
- Mix the palmitic acid working solution thoroughly by vortexing and inverting.
- Place the 15 mL conical vial with the palmitic acid working solution in the bead bath and allow any visible crystals to dissolve completely before use.
- Place the microtray on the ledge surrounding the bead bath with the lids on for ~2 min to warm up.
- Remove the microtray lid. Slowly add 200 µL of palmitic acid working solution across the back wall (long side) of the microtray. Replace the lids and let the microtray to sit on the bead bath for 2-3 min to allow the palmitic acid solution to settle across the bottom of the microtray.
NOTE: While the full bottom of the microtray will not be completely coated until step 4.10, the solution should spread evenly over more than half of the bottom surface. If the palmitic acid solution is not spreading evenly, turn on a vortex or vibrating motor near the microtray. This will provide a small amount of vibration that can help allow the palmitic acid solution to spread more easily. Keep the microtray lids in place to mitigate contamination. - Repeat step 4.8.
NOTE: Roughly half, or more, of the bottom of the microtray should be visibly covered with the liquid palmitic acid. - Rotate the microtray 180° and repeat steps 4.8 and 4.9 on the other side of the tray.
NOTE: The bottom of the Terasaki tray will only be covered with a thin layer of palmitic acid solution after the full combined volume has been added to both sides (steps 4.9 and 4.10). More or less palmitic acid mixture may be required depending on the mixture preparation, temperature, and other factors. Generally, between 400 to 800 µL of the palmitic acid mixture will be sufficient to coat the entire bottom of the microtray. As soon as the bottom of the microtray is visibly covered, no additional palmitic acid should be added, as this can contaminate the wells. - Dry the tray in a sterile drying box or laminar flow hood for at least 1 h uncovered. Alternatively, complete this step the day before loading plates (section 5), drying the microtray with the lid on at RT for at least 16 h.
5. Loading the microtray with low-melt nematode growth media (lmNGM)
NOTE: This method uses standard NGM mixed with low-melt agarose in place of the usual agar. Agar can begin to gel at 45 °C. When working with the small volumes needed to fill the microtray wells, the agar-based NGM often clogs the pipette tip and/or produces uneven well surfaces due to rapid gelling. NGM using low-melt agarose begins to gel at ~28 °C, allowing the molten media to be easily pipetted and consistently forming flat well surfaces. Prepare lmNGM on the same day that the worms are to be placed on the microtray. If preparing the microtray with lmNGM, seeding with bacteria, and loading the C. elegans all within the same day, ensure the animals are at or near the desired age prior to beginning this set of steps.
- Place a pipette basin in an 80 °C bead bath to warm up.
- Melt one 10 mL lmNGM premixture per microtray being prepared (step 1.1) in the microwave for ~30-45 s.
- Place lmNGM premixture test tubes in a 200 mL beaker, uncovered so they do not tip over.
- After ~10-15 s, or when the lmNGM premixture just starts to bubble, pause the microwave, pour the lmNGM premixture into a sterile 200 mL beaker, and resume microwaving.
- Pause as needed to prevent it from boiling over, and swirl to mix.
- Stop microwaving once all the lmNGM has turned to liquid with no chunks.
NOTE: A single 10 mL aliquot is sufficient to fill a total of 192 wells (two microtrays).
- Prepare lmNGM by adding the following components in the listed order to the warm low melt agarose (volumes are per 10 mL of low melt agarose): 250 µL of 25 mM KPi (pH 6), 10 µL of 1 M CaCl2, 10 µL of 1 M MgSO4, and 10 µL of 5 mg/mL cholesterol
- If the researcher plans to examine adult animals over time and needs to prevent reproduction, also add 10 µL of 50 mM FUdR.
- If conducting an RNAi feeding experiment using the common HT115 RNAi E. coli strain and associated RNAi plasmids, also add 5 µL of 50 mg/mL carbenicillin (to select the RNAi plasmid) and 12 µL of 1 mM IPTG (to induce expression of the double-stranded RNA from the RNAi plasmid).
NOTE: Additional additives can be included depending on the selective antibiotics and other chemicals needed for the desired food source. Drugs or chemical stressors can also be added to lmNGM at this stage. NGM and lmNGM contain the same final concentration of NaCl but differ in how the NaCl is added to the media. For NGM, all of the NaCl is added during the media preparation (step 1.2). For lmNGM, a portion of the NaCl is added during media preparation (step 5.3) and a portion with the bacterial food (section 6). The reason for this change stems from the small media volume in each microtray well. Adding 5 µL of concentrated bacteria suspended in LB to a well containing 20 µL of media will substantially alter the nutrient composition of the media (by adding the components of LB at a comparable volume). To avoid this, the bacteria is instead resuspended in a solution of NaCl to remove the LB components while preventing osmotic stress on the bacteria. The NaCl added to the media is reduced accordingly to account for the salt added to the bacteria.
- Pour lmNGM into the warmed pipette basin.
- With the microtray on a benchtop, pipette 20 µL of lmNGM into each microtray. Cover the microtray when refilling the pipette to minimize contamination.
NOTE: This step is easier with an electronic repeating pipette.
6. Seeding the microtray wells with bacterial food
NOTE: 5 µL of 10x concentrated food from an overnight inoculated culture is sufficient to feed a single C. elegans for the entirety of its lifespan.
- Transfer the bacteria from the overnight culture to a 50 mL conical vial.
- Centrifuge at ~3,500 x g for 20 min.
- Remove the supernatant. Resuspend the bacteria culture at a 10x concentration (1/10 volume of the original culture) with 85 mM NaCl solution.
- Before adding bacteria to the wells, visually inspect the microtray to ensure the solid lmNGM is completely contained within each well. If any lmNGM is hanging off the side of a well, scrape the excess lmNGM off with a pipette tip. This excess media can cause the food to run into palmitic acid if not cleared.
- Carefully pipette 5 µL of 10x concentrated bacterial culture onto the surface of the lmNGM in each well. Try to avoid allowing the bacteria culture to run over the side of the well and into the palmitic acid, as this will attract worms to leave the well.
- Allow the microtrays to dry in a sterile drying box or laminar flow hood for ~35 min. Check every ~5 min and take care not to over-dry. Remove when a few wells are visibly slightly wet, as the wells will dry further while the nematodes are being added.
7. Enclosing each microtray inside a single-well plate
NOTE: In this section, the microtray is mounted inside a standard single-well plate using a custom 3D-printed insert and surrounded by water-absorbing crystals. The single-well plate is then closed and sealed with Parafilm. This allows oxygen exchange while preventing contamination and maintaining sufficient humidity to prevent the lmNGM from drying out over the course of multiple-week experiments (worm lifespan experiments can last 6-8 weeks if examining lifespan-extending mutations or environmental conditions). During preparation, make sure to cover the plate whenever something is not actively added to minimize bacterial or fungal contamination.
- Sterilize a single-well plate and 3D printed adapter and lid by dipping each into a 10% bleach solution. Rinse thoroughly with sterile DI water. Wipe dry with a task wipe.
- Spray the single-well plate and 3D printed adapter and the lid with 70% or greater ethanol solution and wipe clean with a task wipe. Let any residual ethanol air dry.
NOTE: UV radiation can also be used after bleach and ethanol with the same parameters as the microtray, as described above in step 4.4. - Place the sterile 3D-printed insert into a sterile single-well plate.
- Sterilize the outside of the prepared microtray by spraying it with 70% ethanol.
- Place the microtray into the 3D-printed adapter in the single-well plate.
- Discard the microtray lid.
- Liberally dispense water crystals on top of the 3D-printed insert between the outer wall of the microtray and the inner wall of the single-well plate.
- Immediately add the worms (section 8) or seal the microtray to use the next day. Close the tray lid and use a single piece of Parafilm to wrap all four sides to seal the microtray. Repeat the sealing step with Parafilm two more times for a total of three layers.
- After sealing the microtray, leave it on the bench at RT for up to 4 days before adding the worms (section 8).
8. Adding worms to the microtray
NOTE: One worm can be manually added to each well by transferring animals from the age-synchronized worm populations (section 2) using a platinum pick. Only transfer animals at the desired life stage. If the transfer process takes more than 1 h, the lmNGM in the microtray wells can desiccate. Transferring groups of 10 to 20 worms at a time can hasten this step, but it takes some practice to consistently release only a single animal per well.
- Add one nematode per well once they reach the desired life stage.
- Place the lid back over the microtray when picking worms from the source plate to minimize contamination.
9. Finishing preparing the culture environment for long-term use
NOTE: The steps below are performed to ensure that the media and worms in the microtray remain hydrated for the experiment duration.
- Add ~40 µL of the detergent solution to the lid of a single-well plate and use a task wipe to evenly coat the lid. This coating will prevent condensation over time once the plate is sealed. Use additional task wipes to spread the detergent solution until vision through the lid is not distorted (this usually takes about two task wipes).
- Examine the water crystals to ensure they are both level with the top edge of the microtray and not overflowing. If necessary, add or remove water crystals to the plate.
- Using the following approach, wrap the tray with Parafilm (three pieces in total) to ensure that the Parafilm seal will last long-term.
- Stretch the first piece of Parafilm slightly to cover two sides of the tray.
- Stretch the second piece of Parafilm slightly and cover the remaining sides of the tray.
- Stretch the final piece of Parafilm and fully wrap all four sides of the tray, covering the first and second pieces of Parafilm. A properly sealed microtray can remain hydrated for ~2 months.
NOTE: Monitor Parafilm integrity every 1-2 weeks throughout the experiment, and replace as needed. - Wipe the top of the tray with a task wipe, damp with 70% ethanol, to remove any fingerprints.
- Label the single-well plate using tape. The microtray is now ready for imaging and long-term storage to monitor worms over time. Between imaging sessions, keep the tray at 20 °C. The microtray can be opened and resealed multiple times for drug dosing or other procedures, but should be minimized to prevent contamination and moisture loss.
10. Imaging individual worms in microtray wells
NOTE: The purpose of this protocol is to provide a detailed description of how to prepare the microtray culture environment. Once prepared and populated with worms, the microtray single-worm culture environments can be used to longitudinally monitor many phenotypes using techniques established for standard culture on Petri plates. The following section provides basic instructions for measuring some common phenotypes. Fluorescent imaging must be performed in an upright microscope. Refraction of light through the Terasaki tray is minimized in a dark room.
- Lifespan: Measure the lifespan by gently tapping the side or top of the plate or shining a bright blue light on the plate and observing each animal. Score an animal "dead" if it does not move within a few seconds. Repeat this task every 1-3 days until all worms have died.
NOTE: The microtray culture environment is compatible with systems that automate image collection and analysis for lifespan estimation14,19. - Standard microscopy: Perform brightfield (or darkfield) imaging on individual wells or on the entire tray with a high-resolution camera or scanner. This imaging data can be used for a variety of phenotypes, including animal size22,23,24, development25, activity14,19, and lifespan.
- Fluorescent microscopy: Perform fluorescent imaging on single wells either manually or with a motorized platform system. The single-well trays are compatible with standard multi-well plate adapters. Signal-to-noise is minimized if the apparatus is placed in a black-walled box.
NOTE: Because the worms are imaged while remaining free to crawl, this culture system is well-suited to low-resolution imaging to capture whole-body fluorescence and rough tissue localization of fluorophores. High-resolution and high-magnification imaging typically requires that the animals be immobilized to prevent movement. While it should be possible to topically apply a paralyzing agent (e.g., sodium azide or levamisole) to each well and proceed with higher-resolution imaging, this would preclude repeated measurements of the same worm at later time points. The culture system, as described, is also not intended to be inverted, which prevents the use of inverted microscope systems.- If brightfield is not being utilized to measure lifespan and only fluorescence is being captured, measure the lifespan manually. Leaving the blue light (used for GFP excitation) on the worm for 5 s will prompt the animal to move. If the animal has not moved within 5 s, consider the animal as dead.
Representative Results
The microtray-based single-worm culture environment described here can be used to monitor a variety of phenotypes, including lifespan and health span, activity and movement, body shape and crawling geometry, and the expression of transgenically expressed fluorescent biomarkers in individual animals over time. The microtray culture system is compatible with lifespan analysis through either manual scoring or image collection and downstream imaging analysis. As with standard culture on Petri plates21, worms can be manually scored as dead or alive based on movement following a stimulus (e.g., tapping the plate or exposure to blue light). For image-based lifespan analysis, a comparison of frame-to-frame differences between images taken after the stimulus has been applied is used to determine when each worm stops moving. This is compatible with automated image collection and analysis systems and provides additional data in the form of the activity level of individual animals at each time point. This information can then be used not just to estimate lifespan (cessation of movement), but also health span (a threshold of reduced activity; multiple definitions have been proposed). Additional parameters (body size, shape, and posture) can also be measured from these images.
The key features of this microtray culture system are (1) the capacity to track individual animals over time, (2) the compatibility between the culture system and aversive interventions like dietary restriction or chemical stress agents, and (3) the capability to measure fluorescence directly in the culture environment. To demonstrate these features, we exposed worms to 250 µM copper sulfate (CuSO4) to induce heavy metal stress or 10 mM dithiothreitol (DTT) to induce protein misfolding stress and measured daily activity, lifespan, and health span. Both CuSO4and DTT significantly reduced worm lifespan (Figure 2A). Monitoring daily activity allowed us to estimate the individual health span of each worm, here defined as the age at which a worm can no longer move a full body length in response to the stimulus. CuSO4 and DTT reduced health span relative to untreated controls (Figure 2B). In contrast, CuSO4 decreased the proportion of life spent in good health, while DTT did not (Figure 2C). CuSO4 also reduced average activity across individuals within the population to a greater extent than DTT at all ages (Figure 2D), as well as cumulative activity for each animal across the lifespan (calculated as the area under the activity curve [AUC]; Figure 2E). An important advantage of this microtray setup is its ability to correlate phenotypes measured at different time points across individual animals within a population. In this case, we find that cumulative activity for individual animals up to day 10 correlates with lifespan for DTT-challenged animals but not for untreated controls or CuSO4-challenged animals (Figure 2F).
To demonstrate the capability to monitor fluorescent biomarkers across the lifespan, we used the microtray culture system to monitor the expression of mScarlet transgene fused to the C-terminus of the kynu-1 gene, encoding the enzyme kynureninase in the kynurenine metabolic pathway, at its endogenous locus. We previously found that expression of human kynureninase, encoded by Kynu, increases with age in blood26, and that knockdown of kynu-1 di C. elegans is sufficient to extend lifespan27. KYNU-1 displays age-dependent expression; it is nearly undetectable at the L4 larval stage and peaks at day 9 of adulthood. A drastic increase in expression is observed between days 3 and 8 of adulthood, and there is a progressive decline with age thereafter (Figure 3A, Supplementary Figure 1). To assess the relationship between KYNU-1 and C. elegans aging, we quantified the cumulative abundance of KYNU-1 up to day 14 of adulthood (an age at which most animals were still alive) and found a significant correlation with lifespan (Figure 3B). Since KYNU1 expression is dynamic over the course of a lifespan, we compared KYNU-1 expression at different ages to overall survival across individual animals. At no single time point was KYNU-1 expression significantly correlated with survival (Figure 3C-F), demonstrating that lifelong expression is a better indication of lifespan impact than single time point assessments.
Combined, these experiments provide representative data that highlight the capabilities of the microtray-based single-worm culture system to enable dynamic changes in worm activity, health, lifespan, and fluorescent biomarkers to be captured and compared across ages in individual animals. In particular, the ability to monitor individuals at multiple time points allows dynamic changes in both physiological and molecular phenotypes to be tracked in vivo, enabling the detection of patterns that would not be apparent using cross-sectional measurements across populations.
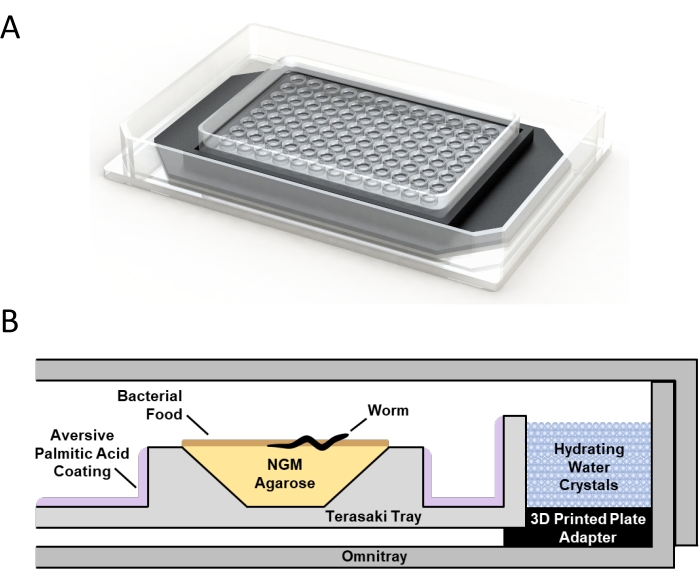
Figure 1: Microtray single-worm culture environment. (A) 3D rendering of a microtray mounted within a single-well plate using a custom 3D-printed adapter. (B) Schematic of the single-worm culture system showing the relative position of a microtray well with low-melt agarose nematode growth media (NGM), bacterial food, isolated C. elegans, aversive palmitic acid coating, the 3D printed adapter, water crystals, and the single-well plate. Please click here to view a larger version of this figure.
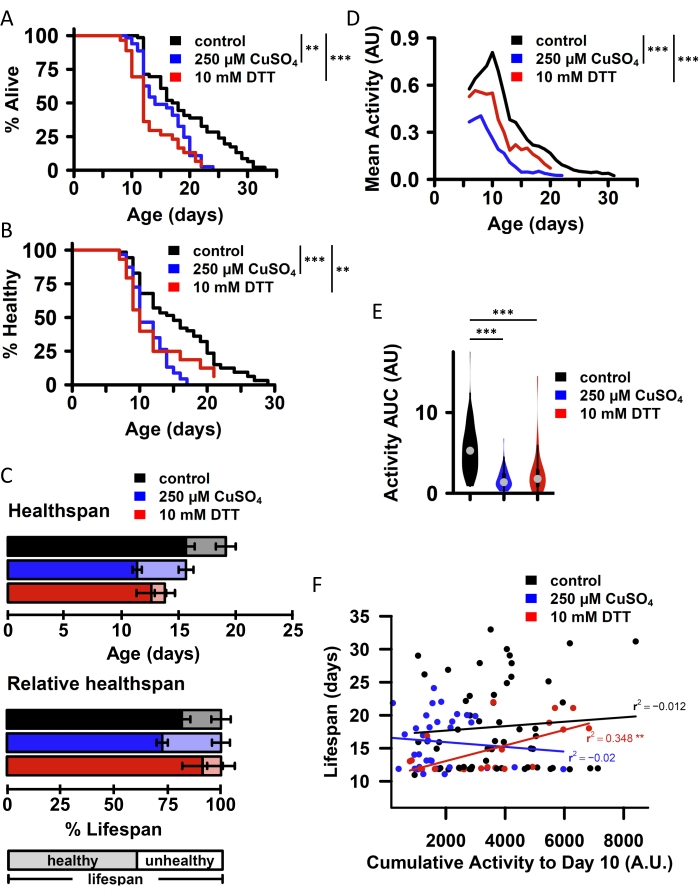
Figure 2: Correlation of phenotypes in individual animals across populations using Microtray single-worm environments. All panels provide data from an experiment comparing three group of wild-type (N2) C. elegans exposed to no additives (control; N = 55), 250 µM copper sulfate (CuS04; N = 38), or 10 mM dithiothreitol (DTT; N = 34) starting at day 2 of adulthood. Lifespan (A) and health span (B) are both moderately reduced by chronic exposure to either CuSO4 or DTT. Health span is defined as the last day an animal can move at least one full body length on the plate. Pairwise significance is calculated using the log-rank test (R survdiff function). (C) Health span and lifespan averages within each population (top: absolute values; bottom: normalized to each group's mean lifespan). (D) Average worm activity (3 day rolling mean of daily activity) within each group across the lifespan and (E) the population average of the individual animal area under the lifetime activity curve (AUC) are both substantially impaired by CuSO4 or DTT. Significance determined by the Mann-Whitney U test to compare AUC for individual animals between groups. (F) Cumulative activity for individual animals to day 10 (bottom) correlates with a lifespan for animals exposed to DTT, but not for control animals or animals exposed to CuSO4, as determined by linear regression (lm function in R). n.s. = not significant, * p < 0.05, ** p < 0.01, *** p < 0.001. The Bonferroni method was used to adjust all p-values for multiple comparisons. Please click here to view a larger version of this figure.
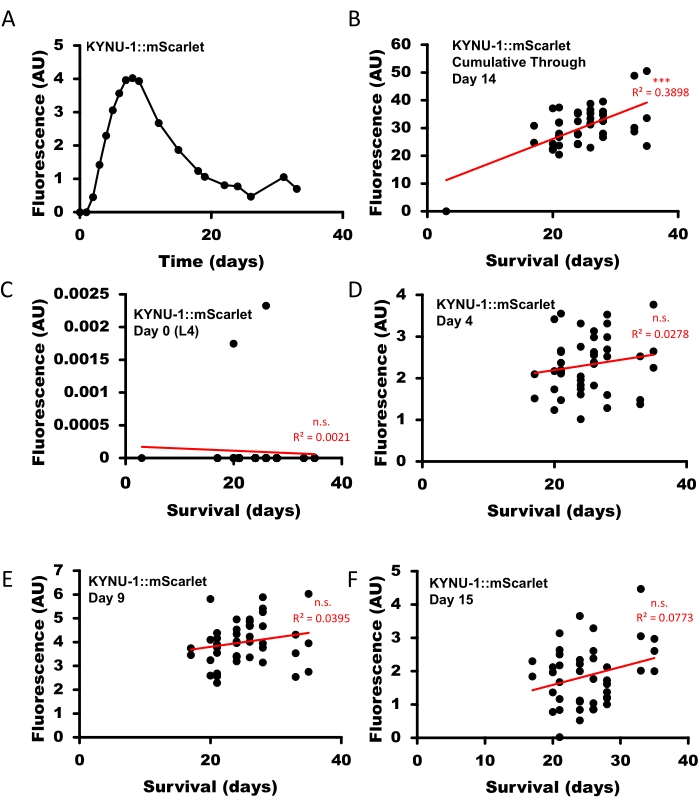
Figure 3: Dynamic quantification of KYNU-1 in individual C. elegans throughout the lifespan. C. elegans with mScarlet transgenically fused in-frame to the C-terminus of the kynu-1 gene (KYNU-1::mScarlet) were imaged using a fluorescent stereo microscope equipped with a monochrome camera every 1-4 days for 36 days until all animals had died. (A) KYNU-1, as measured by daily quantitation of KYNU-1::mScarlet fluorescence, shows a distinct age-dependent expression pattern across the lifespan (average value across worms shown at each age). (B) Cumulative KYNU-1::mScarlet expression through day 14 of adulthood correlates with lifespan in individual animals. KYNU1::mScarlet expression at the (C) L4 larval stage (day 0 of adulthood), (D) day 4 of adulthood, (E) day 9 of adulthood, and (F) day 15 of adulthood did not correlate with lifespan in individual animals. Correlations calculated by linear regression (Data Analysis Regression function in Microsoft Excel). n.s. = not significant, * p < 0.05, ** p < 0.01, *** p < 0.001. All p-values were adjusted for multiple comparisons using the Bonferroni method. Please click here to view a larger version of this figure.
Supplementary Figure 1: Single KYNU-1::mScarlet C. elegans imaged throughout lifespan under red channel. Single well of a prefabricated microtray containing a single C. elegans with mScarlet tagged KYNU-1 living on concentrated E. coli food source. The animal was first imaged at the L4 larval stage and then imaged at increments of 1-5 days for the remainder of its life. Please click here to download this File.
Supplementary Figure 2: Terasaki tray versus WorMotel background and artifact comparison under fluorescence microscopy. Wells of prefabricated microtrays (top) or molded PDMS WorMotels (bottom) were imaged under GFP (A), RFP (B), and DAPI (C) fluorescence channels using a fluorescent stereo microscope equipped with a monochrome camera. Images are shown in false color, and a heatmap is used to highlight points of saturation. Both culture systems have low-level backgrounds in all channels. The WorMotel plates are prone to sporadic high-signal artifacts, which are likely either microbubbles or dust or other particulate captured inadvertently during the PDMS molding process14,15. Please click here to download this File.
Supplemental File 1: Microtray_adapter_2021-10-20.STL. A stereolithography (STL) file for the 3D-printed insert on which the microtray is placed in the single-well plate. This file contains the full version with a solid bottom and is best suited for use in a system where the light source is above the plate. Please click here to download this File.
Supplemental File 2: Microtray_adapter_bottomless_2021-10-20.STL. A stereolithography (STL) file for the 3D-printed insert on which the microtray is placed in the single-well plate. This file contains the bottomless version and is best suited for use in a system where the light source is below the plate. Please click here to download this File.
Discussion
Here, we describe a novel culture system that adapts microtrays, originally developed for human leukocyte antigen tissue typing assays, to allow the isolation and characterization of single C. elegans over time in a solid media environment that is contextually similar to the agar-based NGM that is the standard in C. elegans research. This system is compatible with a variety of interventions, including dietary restriction, exogenous drug treatment, a challenge with chemical or environmental stressors, and RNAi. It further allows longitudinal monitoring of fluorescent biomarkers in vivo in individual animals. The capacity to follow individual animals within a population allows intra-individual variability to be assessed within a population for phenotypes with a distinct temporal component, including behavior, pathogenesis, response to stressful conditions, and decline in health. It also enables early-life traits that vary within a population to be linked with late-life outcomes, including lifespan and health. Similar to other single-worm imaging technologies, this microtray-based culture system enables individual animals to be monitored at multiple time points throughout life while avoiding potential confounding factors, such as the activation of stress response pathways that occur when worms are cultured in liquid environments (as is the case in many microfluidics devices). This allows data collected in this culture system to be more directly compared with the majority of past C. elegans work conducted on solid media. Recent technological advancements allow the automation of many aspects of C. elegans characterization, including growth, development, and aging28. This culture system, when combined with automated microscopy and associated software, is compatible with an automated, high-throughput collection of lifespan, health span, activity, and other physiological parameters14,19.
We routinely use WorMotels14,15 to monitor physiological phenotypes (e.g., lifetime activity), health span, and longevity for individual C. elegans under non-stressful conditions such as drug treatments and RNAi. In our experience, other single-worm imaging technologies provide an excellent tool for monitoring individual variation in these phenotypes, but they have two limitations. First, in the context of fluorescence imaging, the PDMS material used to mold the chip tends to form microbubbles and capture small particulate matter, both of which fluoresce near the GFP emission wavelength. This generates visual artifacts when plates are visualized under various fluorescence channels (Supplementary Figure 2). A second limitation is that copper sulfate is an insufficient deterrent to prevent fleeing when worms are subjected to interventions that are themselves aversive, particularly dietary restriction, pathogenic bacteria, or stress-inducing agents. The moat of other single-worm setups is too narrow and hydrophobic to accommodate palmitic acid, which is a stronger aversive agent commonly used to maintain populations under dietary restriction or other aversive conditions on Petri plates.
The microtray single-worm culture technique presented here addresses both limitations. The larger moat size and hydrophilic material of the microtrays, along with the application of heat and the inclusion of Tween 20, allows palmitic acid to be applied as an enhanced aversive barrier around each well without contaminating the lmNGM pads themselves. The polystyrene construction of the microtrays generates only low-level, consistently distributed autofluorescence, avoiding the issue of autofluorescent microbubbles and particulate inclusions of the PDMS (Supplementary Figure 2). Both background noise and artifacts are further decreased by using a black surface below and around the tray to minimize diffraction. We include 3D printing files for both a hollow adapter (allowing brightfield imaging) and a solid black adapter (providing a darkfield that improves both darkfield and fluorescence imaging; see Supplemental File 1 and Supplemental File 2). While the lmNGM itself does generate some autofluorescence, the impact on image quality is minimized by the small well volume. These factors make microtrays optimal for repeated fluorescent imaging of individual animals in vivo without removing them from their culture environment. The main disadvantage of the microtray single-culture system relative to other single-worm imaging technologies is the sample size; each microtray can accommodate up to 96 individually cultured animals, in contrast to the 240-animal capacity of each WorMotel14,15. Both offer capable single-worm culture systems, and the experimental application determines which is most appropriate.
There are several limitations to the microtray culture system. First, the microtray-based culture system is both more difficult and more time-consuming to set up than standard culture methods using Petri plates. Second, C. elegans are stimulated by several common excitation wavelengths in fluorescence microscopy. Because animals are not immobilized during the imaging process, fluorescent biomarkers that require long exposure times can be blurred. We find that capturing multiple images during each time point is generally sufficient to allow for accurate whole-animal quantification of fluorescent signal. However, because of this issue, the microtray culture system is not compatible with high-resolution imaging, such as with confocal microscopy, and is not optimal for imaging low-signal biomarkers. Even in light of these limitations, this new method provides the potential to understand complex physiologic processes with time-dependent dynamics or high individual-to-individual variability in the context of activity and survival.
Divulgazioni
The authors have nothing to disclose.
Acknowledgements
This work was supported by NIH R35GM133588 to G.L.S., an NIHT32GM008659 training grant to L.E., a United States National Academy of Medicine Catalyst Award to G.L.S., and the State of Arizona Technology and Research Initiative Fund administered by the Arizona Board of Regents.
Materials
| 3D-printed terasaki inserts | Custom printing company | Robot_Terasaki_tray_insert_10-20 -2021.STL |
FDM printing, nozzle size 0.6 mm using standard PLA plus filament |
| AirClean systems AC624LF vertical laminar flow fume hood | Fisher Scientific | 36-100-4376 | |
| Bacto peptone | Thermo Scientific | 211677 | |
| CaCl2 | Acros organics | 349615000 | |
| Caenorhabditis elegans N2 | Caenorhabditis Genetics Center (CGC) | N2 | Wildtype strain |
| Carbenicillin | Goldbio | C-103-25 | |
| Cholesterol | ICN Biomedicals Inc | 101380 | |
| Escherichia coli OP50 | Caenorhabditis Genetics Center (CGC) | OP50 | Standard labratory food for C. elegans |
| Ethanol | Millipore | ex0276-4 | |
| Fisher Vortex Genie 2 | Fisher Scientific | G-560 | |
| FUdR | Research Products International | F10705-1.0 | |
| Hydrating water crystals | M2 Polymer Technologies | Type S | Type S super absorbent polymer |
| Isopropyl ß-D-1-thiogalactopyranoside (IPTG) | GoldBio | I2481C100 | |
| K2HPO4 | Fisher Chemical | P288-500 | |
| Kimwipes | KimTech | 34155 | Task wipes |
| LB Broth, Lennox | BD Difco | 240230 | |
| Leica K5 sCMOS monochrome camera | Leica Microsystems | 11547112 | |
| Leica M205 FCA Fluorescent Stereo Microscope | Leica Microsystems | 10450826 | |
| Low-melt agarose | Research Products International | A20070-250.0 | |
| MgSO4 | Fisher Chemical | M-8900 | |
| NaCl | Fisher bioreagents | BP358-1 | |
| Nunc OmniTray Single-Well Plate | Thermo Scientific | 264728 | |
| Nystatin | Sigma | N1538 | |
| Palmitic acid | Acros organics | 129700010 | |
| Paper towels | Coastwide Professional | 365374 | |
| Parafilm M | Parafilm | 16-101 | |
| Stratagene UV Stratalinker 2400 | Stratagene | 400075 | UV crosslinker |
| Terasaki trays (Lambda) | One Lambda | 151431 | |
| Thermolyne Dri-bath | Thermolyne | DB28125 | |
| Tween | Thermo Scientific | J20605-AP |
Riferimenti
- Shaham, S. Methods in Cell Biology. WormBook: The Online Review of C. elegans Biology. , (2006).
- Boulin, T., et al. Eight genes are required for functional reconstitution of the Caenorhabditis elegans levamisole-sensitive acetylcholine receptor. Proceedings of the National Academy of Sciences. 105 (47), 18590-18595 (2008).
- Massie, M. R., Lapoczka, E. M., Boggs, K. D., Stine, K. E., White, G. E. Exposure to the metabolic inhibitor sodium azide induces stress protein expression and thermotolerance in the nematode Caenorhabditis elegans. Cell Stress & Chaperones. 8 (1), 1-7 (2003).
- Xian, B., et al. WormFarm: a quantitative control and measurement device toward automated Caenorhabditis elegans aging analysis. Aging Cell. 12 (3), 398-409 (2013).
- Rahman, M., et al. NemaLife chip: a micropillar-based microfluidic culture device optimized for aging studies in crawling C. elegans. Scientific Reports. 10 (1), 16190 (2020).
- Chronis, N., Zimmer, M., Bargmann, C. I. Microfluidics for in vivo imaging of neuronal and behavioral activity in Caenorhabditis elegans. Nature Methods. 4 (9), 727-731 (2007).
- Clark, A. S., Huayta, J., Arulalan, K. S., San-Miguel, A. Microfluidic Devices for Imaging and Manipulation of C. elegans. Micro and Nano Systems for Biophysical Studies of Cells and Small Organisms. , 295-321 (2021).
- Levine, E., Lee, K. S. Microfluidic approaches for Caenorhabditis elegans research. Animal Cells and Systems. 24 (6), 311-320 (2020).
- Atakan, H. B., et al. Automated platform for long-term culture and high-content phenotyping of single C. elegans worms. Scientific Reports. 9 (1), 14340 (2019).
- Solis, G. M., Petrascheck, M. Measuring Caenorhabditis elegans life span in 96 well microtiter plates. Journal of Visualized Experiments. (49), e2496 (2011).
- Leung, C. K., Deonarine, A., Strange, K., Choe, K. P. High-throughput screening and biosensing with fluorescent C. elegans strains. Journal of Visualized Experiments. (51), e2745 (2011).
- Laranjeiro, R., Harinath, G., Burke, D., Braeckman, B. P., Driscoll, M. Single swim sessions in C. elegans induce key features of mammalian exercise. BMC Biology. 15 (1), 30 (2017).
- Çelen, &. #. 3. 0. 4. ;., Doh, J. H., Sabanayagam, C. R. Effects of liquid cultivation on gene expression and phenotype of C. elegans. BMC Genomics. 19 (1), 562 (2018).
- Churgin, M. A., et al. Longitudinal imaging of Caenorhabditis elegans in a microfabricated device reveals variation in behavioral decline during aging. eLife. 6, 26652 (2017).
- Turner, E., et al. Long-term culture and monitoring of isolated nematodes on solid media in WorMotels. Journal of Visualized Experiments. , (2022).
- Pittman, W. E., et al. A simple apparatus for individual C. elegans culture. Methods in Molecular Biology. 2144, 29-45 (2020).
- Breimann, L., Preusser, F., Preibisch, S. Light-microscopy methods in C. elegans research. Current Opinion in Systems Biology. 13, 82-92 (2019).
- Mittal, K. K., Mickey, M. R., Singal, D. P., Terasaki, P. I. Serotyping for homotransplantation. 18. Refinement of microdroplet lymphocyte cytotoxicity test. Transplantation. 6 (8), 913-927 (1968).
- . Worm paparazzi-a high throughput lifespan and healthspan analysis platform for individual Caenorhabditis elegans Available from: https://repository.arizona.edu/handle/10150/661628 (2021)
- Porta-de-la-Riva, M., Fontrodona, L., Villanueva, A., Cerón, J. Basic Caenorhabditis elegans methods: synchronization and observation. Journal of Visualized Experiments. (64), e4019 (2012).
- Sutphin, G. L., Kaeberlein, M. Measuring Caenorhabditis elegans life span on solid media. Journal of Visualized Experiments. (27), e1152 (2009).
- Moore, B. T., Jordan, J. M., Baugh, L. R. WormSizer: high-throughput analysis of nematode size and shape. PloS One. 8 (2), 57142 (2013).
- Husson, S. J., Costa, W. S., Schmitt, C., Gottschalk, A. Keeping track of worm trackers. WormBook: The Online Review of C. Elegans Biology. , 1-17 (2013).
- Roussel, N., Sprenger, J., Tappan, S. J., Glaser, J. R. Robust tracking and quantification of C. elegans body shape and locomotion through coiling, entanglement, and omega bends. Worm. 3 (4), 982437 (2014).
- Raizen, D. M., et al. Lethargus is a Caenorhabditis elegans sleep-like state. Nature. 451 (7178), 569-572 (2008).
- Peters, M. J., et al. The transcriptional landscape of age in human peripheral blood. Nature Communications. 6 (1), 8570 (2015).
- Sutphin, G. L., et al. Caenorhabditis elegans orthologs of human genes differentially expressed with age are enriched for determinants of longevity. Aging Cell. 16 (4), 672-682 (2017).
- Felker, D. P., Robbins, C. E., McCormick, M. A. Automation of C. elegans lifespan measurement. Translational Medicine of Aging. 4, 1-10 (2020).

