Direct Synthesis of EM-Visible Gold Nanoparticles in Cells for Protein Localization Analysis with Well-Preserved Ultrastructure
Summary
The present protocol describes a clonable electron microscopy labeling technology for detecting metallothionein-tagged proteins in cells using a novel autonucleation suppression mechanism-based gold nanoparticle synthesis technique.
Abstract
Analyzing the precise localization of protein molecules in cells with ultrastructural resolution is of great significance for the study of various physiological or pathological processes in all living organisms. Therefore, the development of clonable tags that can be used as electron microscopy probes is of great value, just as fluorescent proteins have played a crucial role in the field of optical imaging. The autonucleation suppression mechanism (ANSM) was recently uncovered, which allows for the specific synthesis of gold nanoparticles (AuNPs) on cysteine-rich tags, such as metallothionein (MT) and antifreeze protein (AFP).
Based on the ANSM, an electron microscopy labeling technology was developed, which enables the specific detection of tagged proteins in prokaryotic and eukaryotic cells with an unprecedented labeling efficiency. This study illustrates a protocol for the detection of MTn (an engineered MT variant lacking aldehyde-reactive residues) fusion proteins in mammalian cells with well-preserved ultrastructure. In this protocol, high-pressure freezing and freeze-substitution fixation were performed using non-aldehyde fixatives (such as tannic acid, uranyl acetate) to preserve near-native ultrastructure and avoid damage to the tag activity caused by aldehyde crosslinking.
A simple one-step rehydration was used prior to the ANSM-based AuNP synthesis. The results showed that the tagged proteins targeted various organelles, including the membranes and the lumen of the endoplasmic reticulum (ER), and mitochondrial matrices were detected with high efficiency and specificity. This research provides biologists with a robust protocol to address an enormous range of biological questions at the single-molecule level in cellular ultrastructural contexts.
Introduction
In the postgenomic era, the development of green fluorescent protein (GFP) as a single-molecule reporter for light microscopy has revolutionized the field of modern life science research1,2. For decades, electron microscopy (EM) has been a powerful tool for intuitively observing the cellular ultrastructure with nanoscale resolution3; however, the precise identification and localization of protein molecules remain challenging.
The most commonly used EM labeling technique is the immunoelectron microscopy (IEM) labeling technique, which is based on the antigen-antibody reaction. However, although many techniques have been developed in the field of IEM labeling, including pre-embedding IEM and post-embedding IEM (on resin sections or hydrated cryosections), it still suffers from low labeling efficiency (<10%)4,5, which is related to the sample preparation and antibody quality. To overcome these limitations, developing genetically encoded tags has great application potential.
Two main types of EM tags have been thoroughly explored in recent years. One type is the DAB staining method, which utilizes tags such as APEX2 to oxidize 3,3'-diaminobenzidine (DAB) to osmiophilic polymers for EM visualization6,7,8,9,10,11,12. It enables the labeling of high-abundance proteins in subcellular regions but is not suitable for single-molecule counting. The other type utilizes metal-binding proteins, such as ferritin13 and metallothionein (MT)14,15,16,17,18,19,20,21,22,23,24,25,26, to generate electron-dense metal deposits in situ for EM visualization. Only the latter has real potential for single-molecule visualization and counting. The molecular size of ferritin is too large (~450 kD) for it to be used as a promising tag, whereas the small size (~5 kD) of MT and its ability to bind various ions through its 20 cysteines have attracted great attention. Several labs have tried to label purified MT-fusion proteins or MT-expressing cells by incubating directly with Au+. These attempts have initially proved that MT tags can bind gold ions to form high-contrast signals, but none have really achieved the effective identification of individual proteins in cells, and they are not widely applicable14,15,16,17,18,19,20,21,22,23.
The ANSM-based AuNP synthesis technique, which involves synthesizing 2-6 nm-sized AuNPs directly on cysteine-rich tags (e.g., MT, the MT variants MTn and MTα, AFP) as electron-dense labels for EM visualization, is the first reliable and applicable approach for protein labeling and single molecule detection in cells24,25,26. It allows for the specific synthesis of AuNPs on isolated tag fusion proteins and has achieved an unprecedented labeling efficiency in nonfixed or chemically fixed prokaryotic (E. coli) and eukaryotic (S. pombe) cells. However, implementing the same protocol in more advanced systems such as mammalian cells or even tissues involves additional challenges, such as the more complex intracellular redox homeostasis and more fragile cellular structure.
This study presents a clonable EM labeling technology, which combines the novel ANSM-based AuNP synthesis technique for labeling genetically encoded cysteine-rich tags (MT) with the HPF/FSF-rehydration-HPF/FSF sample preparation method, enables the unambiguous single-molecule identification of tagged proteins in the ER membrane, ER lumen, and mitochondrial matrix in HeLa cells. The current method combines the characteristics of high labeling efficiency, a high signal-to-noise ratio, single-molecule labeling, and strong universality, and this method has broad application prospects in life science research.
Protocol
All the supplies used in this experiment are listed in the Table of Materials. The step-by-step workflow of the current protocol is shown in Figure 1.
1. Cell culture on sapphire discs
- Transfer 3 mm x 0.16 mm sapphire discs to a 2 mL centrifuge tube containing 1 mL of ethanol, and sonicate in an ultrasonic cleaner for 10 min.
- Burn each sapphire disc in an alcohol lamp flame until there are no visible deposits on the surface.
NOTE: Through the above method, sapphire discs can be recycled many times. - Label one side of each sapphire disc with a number using a solvent-resistant pen (Figure 1A).
NOTE: The marker pen used needs to be tested to determine whether it is resistant to acetone and methanol solvents in advance to prevent the loss of marks. - Place the labeled sapphire discs on the bottom of a 35 mm culture dish with the labeled side facing up.
- Sterilize the cell culture dish with sapphire discs under UV light for 30 min.
- Flip the sapphire discs with tweezers (the labeled side facing down), and sterilize under UV light for an additional 30 min. Now, the culture dish containing the sapphire discs is ready for cell culture.
NOTE: The sapphire discs can also be sterilized by autoclaving. - Seed the cells on the sapphire discs prepared above, and grow to 80%-90% confluency in a cell incubator at 37 °C, 95% humidity, and 5% CO2 (Figure 1B).
NOTE: Stable HeLa cell lines expressing MTn tags were generated as described in Jiang et al.26.
2. High-pressure freezing (HPF)
CAUTION: Liquid nitrogen is used in this experiment. When working with liquid nitrogen, use proper safety procedures and personal protective equipment to prevent frostbite and asphyxiation.
- Prepare the high-pressure freezing machine at least 2 h before the experiment according to the manufacturer's instructions.
- Transfer type A HPF aluminum carriers (recess: 0.025/0.275 mm) to a 2 mL centrifuge tube containing 1 mL of ethanol, and sonicate in an ultrasonic cleaner for 10 min.
- Dry the carriers in a Petri dish covered with qualitative filter paper (medium speed).
- Soak the precleaned carriers in 1-hexadecene.
NOTE: BSA is not recommended as a cryoprotectant in the current experiment as it will interfere with the subsequent gold nanoparticle synthesis. - Use fine tweezers to pick up a sapphire disc with cells from the culture dish; touch the labeled surface with qualitative filter paper (medium speed) to remove the excess medium.
NOTE: The thermal conductivity of water is very low, and too much residual water will affect the freezing effect. Retain minimal water to prevent the cells from drying out. - Mount the sapphire disc into the HPF specimen holder, and quickly cap the disc with a 0.025 mm deep aluminum carrier containing 1-hexadecene (Figure 1C).
- Aspirate excess solution with qualitative filter paper (medium speed).
- Load the specimen holder for high-pressure freezing (Figure 1D).
NOTE: The specimen loading process needs to be as fast as possible (within 1 min) to minimize physiological alterations due to changes in the temperature, humidity, pH, and oxygen content. - Unload the sapphire disc-carrier assembly under liquid nitrogen in a foam cryobox, and store the assembly in a cryovial in liquid nitrogen before use.
NOTE: The protocol may be paused here. The specimens in cryovials can be stored in a liquid nitrogen dewar for years.
3. Freeze-substitution fixation (FSF) and rehydration (Figure 1E)
- Fill the automated freeze-substitution machine (AFS) with liquid nitrogen, and cool the chamber to −90 °C.
CAUTION: Liquid nitrogen is used in this experiment. When working with liquid nitrogen, use proper safety procedures and personal protective equipment to prevent frostbite and asphyxiation. - Prepare 2 mL of 0.01% tannic acid (w/v) in acetone (FSF solution 1) in a 20 mm round polypropylene container (Figure 1E) in a fume hood and freeze it in liquid nitrogen.
- Load the specimens (the sapphire disc-carrier assemblies) into the container with frozen FSF solution 1 under LN2 using a pair of precooled tweezers.
- Transfer the container to the precooled AFS chamber for FSF processing at −90 °C.
- Keep the specimens at −90 °C for 1 h; then, separate the sapphire discs from the carriers with precooled tweezers, and make sure the marked sides of the sapphire discs are facing down.
NOTE: The tweezers must be sufficiently precooled to prevent temperature changes. The sapphire discs should be easily separated. - Keep at −90 °C for a further 8-10 h; then, warm to −60 °C within 3 h.
- Replace the FSF solution 1 in the container with precooled acetone, and incubate for 1 h. Repeat this acetone wash 2x.
- Warm to −30 °C within 3 h. During this period, prepare and precool 2 mL of 0.01% uranyl acetate in acetone (FSF solution 2, diluted from 10% uranyl acetate in methanol) in a 2 mL centrifuge tube.
CAUTION: Uranyl acetate, used in the current experiment, is radioactive and highly toxic; it must be handled according to proper safety procedures and disposed of as hazardous chemical waste. - Replace the acetone with FSF solution 2, and incubate at −30 °C for 3 h.
- Replace the FSF solution 2 in the container with precooled acetone, and incubate for 30 min at −30 °C. Repeat this acetone wash 2x.
- Warm from −30 °C to 4 °C within 2 h.
- Replace the acetone with 2 mL of 0.2 M HEPES buffer containing 1 mM CaCl2 and 1 mM MgCl2 at pH 5.5.
- Change the buffer once, and incubate at room temperature for 1 h or at 4 °C overnight.
NOTE: The protocol may be paused here for one night or continued for at least 6 h until the specimens are frozen again in step 5.
4. ANSM-based AuNP synthesis for mammalian cells (Figure 1F)
- Wash with PBS-A buffer 3x for 5 min each time.
- Transfer the sapphire discs to a 35 mm culture dish containing 1 mL of PBS-A buffer at room temperature.
NOTE: Always keep the marked sides of the sapphire discs facing down, and avoid overlapping the sapphire discs and rubbing off the cells. - Prepare the reducing solution by adding 4.28 µL of 2-mercaptoethanol into a 2 mL centrifuge tube containing 1 mL of PBS-A buffer and mixing well in a fume hood.
CAUTION: 2-Mercaptoethanol is toxic and has a pungent odor; it must be handled according to proper safety procedures. - Replace the PBS-A buffer in the culture dish with 1 mL of reducing solution, and incubate for 1 h at room temperature.
- Prepare the gold precursor before use.
- Add 80 µL of 10 mM HAuCl4 in ddH2O into a 2 mL centrifuge tube containing 1 mL of reducing solution, and immediately vortex.
NOTE: The solution may become cloudy. - Add 80 µL of 500 mM D-penicillamine in ddH2O into the above solution, and immediately vortex.
NOTE: The solution may become clear again.
- Add 80 µL of 10 mM HAuCl4 in ddH2O into a 2 mL centrifuge tube containing 1 mL of reducing solution, and immediately vortex.
- Replace the solution in the culture dish with the 1 mL of gold precursor solution prepared in step 4.5, and incubate for 2 h at 4 °C.
NOTE: One milliliter of the gold precursor is sufficient to react with approximately 1 × 106 cells (confluent on a 35 mm dish). Larger volumes of the precursor are needed when the AuNPs become significantly smaller. - Weigh 0.0038 g of NaBH4 into a 1.5 mL centrifuge tube, add 1 mL of ice-cold ddH2O, and vortex to make fresh 100 mM NaBH4 just prior to use.
- Add 20-100 µL of 100 mM NaBH4 to the solution from step 4.6, immediately shake to mix well, and incubate for 5 min at room temperature.
NOTE: Prepare 100 mM NaBH4 freshly for immediate use. After adding the NaBH4, the color of the solution will gradually turn red and then deepen. If no color change is observed, it largely indicates that the experiment has failed. - Replace the solution with 2 mL of PBS-A buffer to stop the reaction.
NOTE: Stopping the reaction within 30 min is critical, as a prolonged reaction will gradually break down the AuNPs.
5. High-pressure freezing and freeze-substitution fixation (Figure 1C-F)
NOTE: The specimens are HPF and FSF again, and the procedure is almost the same as that previously described in section 2 and section 3 with a few modifications.
- Prepare 1% osmium tetroxide and 0.1% uranyl acetate in acetone (FSF solution 3, diluted from 4% osmium tetroxide in acetone and 10% uranyl acetate in methanol).
CAUTION: Osmium tetroxide and uranyl acetate are highly toxic chemicals; they must be handled according to proper safety procedures and disposed of as hazardous chemical waste. - Load the specimens (the sapphire disc-carrier assemblies) into the container with frozen FSF solution 3 under LN2 using a pair of precooled tweezers.
- Transfer the container to the precooled AFS chamber for FSF processing at −90 °C, and run the FSF program as follows: keep at −90 °C for 8-10 h; then, warm to −60 °C within 3 h; keep at −60 °C for 3 h; then, warm to −30 °C within 3 h; keep at −30 °C for 3 h; then, warm to 4 °C within 2 h.
- When the temperature reaches 4 °C, transfer the specimens to the fume hood, and maintain at room temperature for 15 min.
- Wash with acetone 3x for 15 min each time.
- Maintain in acetone at room temperature for at least 2 h.
6. Resin infiltration, embedding, and polymerization
- Prepare the epoxy resin mixture according to the manufacturer's instructions before use.
CAUTION: The epoxy resin used in the current experiment is toxic prior to polymerization; it must be handled according to proper safety procedures. Unpolymerized resin should be disposed of as hazardous chemical waste or polymerized prior to disposal. - Transfer the sapphire discs to flat-bottom embedding capsules, and infiltrate with resin for at least 30 min at room temperature.
NOTE: Keep the marked sides of the sapphire discs facing down. - Polymerize the specimen at 60 °C in an oven for at least 18 h.
NOTE: The protocol can be paused here after polymerization. Polymerized specimen blocks can be stored in a dry container for years.
7. Ultrathin sectioning
- Trim the polymerized specimen blocks carefully using a razor to expose the sapphire discs.
- Dip the block tips with sapphire discs into liquid nitrogen for several seconds, and thaw at room temperature. Repeat the freeze-thaw action several times to detach the discs from the blocks.
NOTE: Avoid prolonged freezing, as this may crack the specimen blocks. Gently detach the sapphire discs with a blade, avoiding excessive force, which may damage the samples. - Trim the block face to a trapezoidal shape for ultrathin-sectioning (Figure 1G).
- Obtain 70-100 nm ultrathin sections with an ultramicrotome (Figure 1H).
- Pick the sections on 200-mesh hexagonal copper grids.
NOTE: The protocol can be paused here after sectioning. The sections on girds can be stored in a dry container for years. If desired, the sections on grids can be stained with 2% uranyl acetone to obtain better membrane contrast. Lead staining should be avoided as it will mask the nanogold particle signals.
8. TEM imaging (Figure 1I)
- Examine the ultrathin sections on copper grids using a transmission electron microscope. Typically, a magnification range from 11 kX to 30 kX is suitable for visualizing AuNPs in cells, and an appropriate defocus value is needed to ensure 2-3 nm AuNPs are visible.
Representative Results
The ANSM-based AuNP synthesis technique is an extremely useful tool for labeling and detecting MT-tagged proteins with TEM26. To validate its robustness in mammalian cells, three stable cell lines expressing EGFP-MTn-KDEL, Ost4-EGFP-MTn, or Mito-acGFP-MTn in Hela cells were generated. KDEL is a canonical C-terminal endoplasmic reticulum (ER) retention/retrieval sequence, which maintains the fusion protein EGFP-MTn-KDEL within the ER lumen or the perinuclear space of the nuclear envelope (NE). Ost4 is a subunit of the oligosaccharyltransferase complex, which is a membrane protein complex localized in the ER and NE that catalyzes the N-glycosylation of nascent polypeptides. The C terminus of the Ost4 fusion protein Ost4-EGFP-MTn faces the cytosol. Mito is a mitochondrial targeting sequence that targets the fusion protein Mito-acGFP-MTn in the mitochondrial matrix.
The HPF/FSF sample preparation, combined with the use of tannic acid and uranyl acetate instead of aldehyde fixatives, preserved excellent ultrastructure with good membrane contrast (Figure 2, Figure 3, and Figure 4). The overall structure of the sample was dense, without obvious cytoplasm and lipid extraction. The membrane structure was smooth, without obvious deformation, and the phospholipid bilayer structure was clearly revealed.
In addition to the well-preserved ultrastructure, efficient labeling was observed in all three cases representing distinct organelle specificities. The EGFP-MTn-KDEL protein appeared as 2-5 nm sized gold nanoparticles exclusively distributed in the peripheral ER lumen and in the perinuclear space of the NE (Figure 2A–C). The well-preserved ultrastructure not only enabled the single-molecule identification of tagged proteins but also facilitated the analysis of organelle interactions, such as ER-mitochondria interactions (Figure 2D,E). Nanoparticles of the Ost4-EGFP-MTn protein delineated the ER membrane (Figure 3A–D) and the outer membrane of the NE (Figure 3E). Nanoparticles were also distributed on the inner membrane of the NE, but the number of nanoparticles there was lower than on the outer membrane, indicating that the protein composition of the inner and outer membranes was different (Figure 3E). Likewise, the Mito-acGFP-MTn-expressing cells exhibited specific labeling in the mitochondrial matrix (Figure 4A–E). No particles were observed in the vesicles or ER (Figure 4A–D), and few particles were shown in the MVBs (Figure 4D).
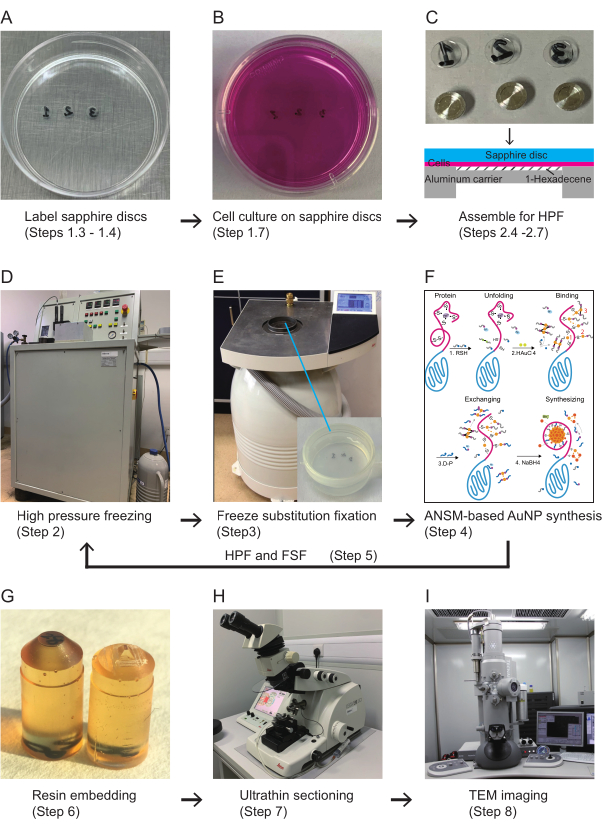

Figure 1: Scheme for the workflow of the clonable electron microscopy labeling technology. (A) Sapphire discs are labeled and sterilized for cell culture. (B) Cells are grown on sapphire discs in a 35 mm culture dish. (C) The sapphire discs with cells are capped with 0.025 mm deep aluminum carriers for high-pressure freezing, and the extra space is filled with 1-hexadecene. (D) The cells are cryofixed by HPF. (E) Freeze-substitution fixation. (F) The schematic steps of the ANSM-based AuNP synthesis. (G) The sapphire discs with cells are embedded in flat-bottom embedding capsules. The resin blocks are trimmed for ultrathin sectioning. (H) The trimmed resin blocks are sectioned with an ultramicrotome. (I) The ultrathin sections are imaged with TEM. Abbreviations: ANSM = autonucleation suppression mechanism; AuNP = gold nanoparticle; HPF = high-pressure freezing; FSF = freeze-substitution fixation; TEM = transmission electron microscopy. Please click here to view a larger version of this figure.
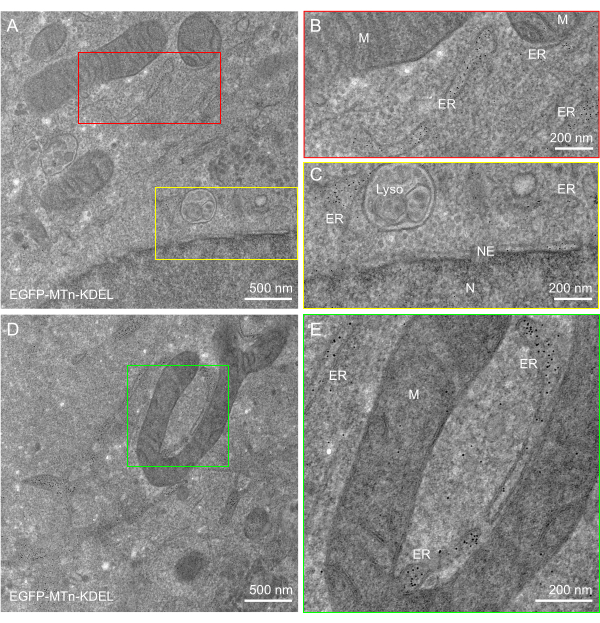

Figure 2: ANSM-based AuNP synthesis on EGFP-MTn-KDEL expressed in HeLa cells. (A,D) EM images of a 90 nm thick section of HeLa cells expressing EGFP-MTn-KDEL show AuNPs specifically accumulated in the lumen of the endoplasmic reticulum and in the perinuclear space of the nuclear envelope. Few particles are observed in the mitochondria, lysosome, nucleus, or cytosol. (B,C) Zoomed-in images of the red and yellow rectangle areas, respectively, in (A). (E) Zoomed-in image of the green rectangle area in (D). This figure has been modified from Jiang et al.26. Scale bars = (B,C,E) 200 nm; (A,D) 500 nm. Abbreviations: ANSM = autonucleation suppression mechanism; AuNP = gold nanoparticle; EGFP = enhanced green fluorescent protein; ER = endoplasmic reticulum; NE = nuclear envelope; M = mitochondria; Lyso = lysosome; N = nucleus. Please click here to view a larger version of this figure.
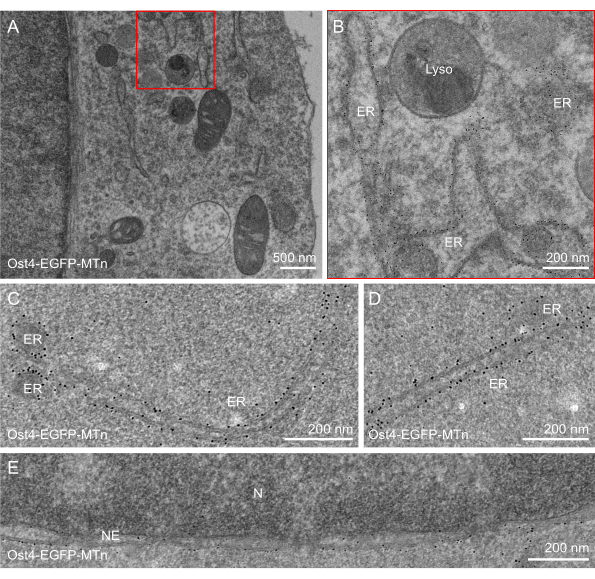
Figure 3: ANSM-based AuNP synthesis on Ost4-EGFP-MTn expressed in HeLa cells. (A–D) EM images of a 90 nm thick section of HeLa cells expressing Ost4-EGFP-MTn show AuNPs specifically accumulated on the membrane of the endoplasmic reticulum. (E) An EM image of a 90 nm thick section of HeLa cells expressing Ost4-EGFP-MTn shows AuNPs specifically accumulated on the membrane of the NE (nuclear envelope). Few particles are observed in the mitochondria, lysosome, nucleus, or cytosol. (C) Zoomed-in image of the red rectangle area in (A). This figure has been modified from Jiang et al.26. Scale bars = (B–E) 200 nm; (A) 500 nm. Abbreviations: ANSM = autonucleation suppression mechanism; AuNP = gold nanoparticle; EGFP = enhanced green fluorescent protein; ER = endoplasmic reticulum; NE = nuclear envelop; M = mitochondria; Lyso = lysosome; N = nucleus; EM = electron microscopy. Please click here to view a larger version of this figure.
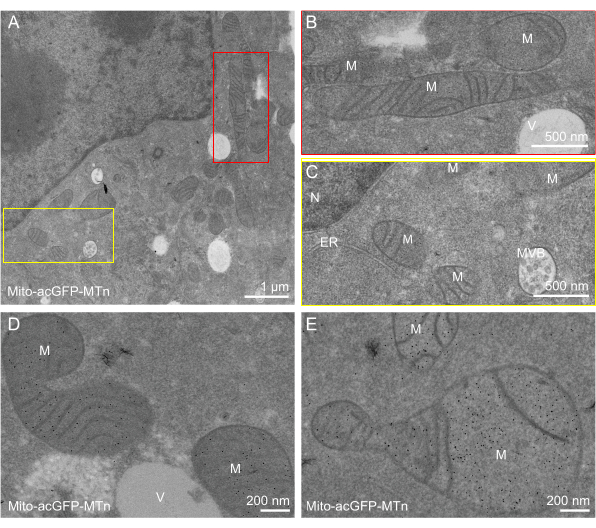
Figure 4: ANSM-based AuNP synthesis on Mito-acGFP-MTn expressed in HeLa cells. (A,D,E) EM images of a 90 nm thick section of HeLa cells expressing Mito-acGFP-MTn show AuNPs specifically accumulated in the matrix of the mitochondria (M). Few particles are observed in the endoplasmic reticulum, nucleus, vesicle, multi-vesicular body, or cytosol. (B,C) Zoomed-in image of the red and yellow rectangle areas, respectively, in (A). This figure has been modified from Jiang et al.26. Scale bars = (D,E) 200 nm; (B,C) 500 nm; (A) 1 µm. Abbreviations: ANSM = autonucleation suppression mechanism; AuNP = gold nanoparticle; EGFP = enhanced green fluorescent protein; ER = endoplasmic reticulum; NE = nuclear envelope; M = mitochondria; Lyso = lysosome; N = nucleus; V= vesicle; MVB = multi-vesicular body. Please click here to view a larger version of this figure.
Discussion
The study presents here a robust clonable EM labeling technology for the single-molecule visualization of protein molecules within the cellular environment with ultrastructural resolution. The AuNPs directly synthesized on genetically encoded cysteine-rich tags provide unambiguous and precise localization of the target proteins. High-pressure freezing and freeze-substitution technique excellently preserve the ultrastructure of biological samples. Taken together, the clonable electron microscopy labeling technology presented here provides a powerful tool for the efficient localization and recognition of a single molecule in cellular ultrastructural contexts in situ with EM.
The ANSM mechanism suppresses the autonucleation process of thiolate-Au(I) polymers26, which is a typical reaction in the classic Brust-Schiffrin method (BSM)27 that is widely used for synthesizing thiolate-capped gold nanoclusters. Therefore, the ANSM-based AuNP synthesis technique can specifically synthesize 2-6 nm-sized AuNPs directly on cysteine-rich tags as electron-dense labels for EM visualization with a high signal-to-noise ratio and high efficiency. Unlike APEX2 or HPR labeling techniques that rely on DAB polymer deposition, which is uncountable and not suitable for soluble proteins in the cytosol, the ANSM-based AuNP synthesis technique is the only technique that enables single-molecule detection thus far.
The drawback of the clonable electron microscopy labeling technology is that the ANSM-based AuNP synthesis technique relies on the activity of the thiol group of cysteine, which is sensitive to aldehyde fixatives. The oxidative protection of thiols by 3,3'-dithiodipropionic acid (DTDPA) has been introduced prior to aldehyde fixation in E. coli and the fission yeast S. pombe, resulting in good morphology and excellent labeling efficiency24,25,26. In this work, the oxidation/fixation method worked properly for those tags expressed in a relatively oxidative compartment, such as the ER lumen and mitochondria matrix (in EGFP-MTn-KDEL and Mito-acGFP-MTn cells). However, it was suboptimal for cytosolic tags (Ost4-GFP-MTn) in mammalian cells. The reason may be that the tags in the reducing compartments are not well folded, and the reducing substances such as GSH that are present in abundance in the cytoplasm can resist oxidation.
HPF-FSF technique is the gold standard in the ultrastructural preservation of biological specimens for electron microscopy. In this work, the method of HPF/FSF-rehydration-HPF/FSF, which completely avoids the conventional chemical fixation and permeabilization, achieved efficient labeling with Ost4-GFP-MTn and yielded excellent cell structure. However, this method still has certain limitations. First, a second HPF may lead to significant ice crystal damage. Second, different batches of uranyl acetate have problems of inconsistent quality and solubility. Acicular precipitates occasionally appeared in the rehydrated samples when visualized by electron microscopy. Washing with HEPES buffer at pH 5.5 may alleviate this problem. Third, the sample preparation for tissue samples needs further optimization.
In conclusion, the clonable EM labeling technology, combining the ANSM-based AuNP synthesis technique with the sample preparation method of HPF/FSF-rehydration-HPF/FSF, enables the unambiguous single-molecule identification of genetically tagged proteins in cells by electron microscopy. The current protocol should allow an enormous range of biological questions to be addressed.
Disclosures
The authors have nothing to disclose.
Acknowledgements
The protocol described here was derived from the article published by Jiang et al. (2020). This work was supported by grants from the MOST (973 Programs nos. 2011CB812502 and 2014CB849902) and by funding support from the Beijing Municipal Government.
Materials
| 0.025 mm/0.275 mm Aluminum carrier | Beijing Wulundes Biotech Ltd., or Engineering Office of M. Wohlwend | ||
| 0.2 M HEPES buffer | Dissolve HEPES (0.2 M) in 980 mL of ddH2O, then add 10 mL of 100 mM MgCl2 and 10 mL of 100 mM CaCl2 (final concentration 1 mM), respectively, adjust pH to 5.5 | ||
| 1.5 mL MaxyClear snaplock microtube | Axygen Scientific | MCT150C | |
| 2 mL polypropylene screw cap microtubes | Biologix | 81-0204 | |
| 200 mesh hexagonal copper grid | Tedpella inc | G200HEX | |
| 2-mercaptoethanol | Amresco | 0482-250ML | |
| 35 mm cell culture dishes | Corning | 430165 | |
| 50 mL polypropylene centrifuge tubes | Corning | 430928 | |
| Acetone | Beiijng Tong Guang Fine Chemicals Company | 31025 | |
| Automated freeze substitution machine | Leica | AFS2 | |
| Customized 3.05 mm x 0.66 mm specimen holders for HPF | Beijing Wulundes Biotech Ltd. | ||
| D-penicillamine | TCI | P0147 | |
| Dulbecco’s modified Eagle medium | GBICO | C11965500BT | |
| Fetal bovine serum | GBICO | 10099-141C | |
| Flat bottom embedding capsule | Tedpella inc | ||
| Foam cryobox | |||
| Formvar 15/95 resin | Electron Microscopy Sciences | 15800 | |
| HAuCl4 | Sigma | 4022-1G | |
| HEPES | sigma | H3375-500G | |
| HPF machine | Wohlwend | HPF compact01 | |
| Methonal | Beiijng Tong Guang Fine Chemicals Company | 12397 | |
| NaBH4 | Sigma | 480886-25G | |
| OsO4 | Electron Microscopy Sciences | 19110 | |
| PBS-A buffer | Dissolve NaH2PO4 (1.125 mM), Na2HPO4 (3.867 mM), NaCl (100 mM) in 1 L of ddH2O, adjust to pH 7.4 | ||
| Qualitative filter paper (medium speed) | Beyotime Biotechnology | FFT08 | |
| Sapphire discs | Beijing Wulundes Biotech Ltd., or Engineering Office of M. Wohlwend | ||
| Solvent resistant pen | Electron Microscopy Sciences | 62053-B | |
| SPI-Pon 812 resin | SPI Inc | 02659-AB | |
| Transmission electron microscopy | FEI | Tecnai G2 spirit | |
| Trypsin-EDTA | GBICO | C25200-056 | |
| Tweezers | Dumont | ||
| Ultramictotome | Leica | FC7 | |
| Uranyl acetate | Electron Microscopy Sciences | 22400 |
References
- Tsien, R. Y. The green fluorescent protein. Annual Review of Biochemistry. 67, 509-544 (1998).
- Shaner, N. C., Patterson, G. H., Davidson, M. W. Advances in fluorescent protein technology. Journal of Cell Science. 120, 4247-4260 (2007).
- Masters, B. R. History of the electron microscope in cell biology. Encylopedia of Life Sciences. , (2009).
- Griffiths, G., Hoppeler, H. Quantitation in immunocytochemistry: Correlation of immunogold labeling to absolute number of membrane antigens. Journal of Histochemistry & Cytochemistry. 34 (11), 1389-1398 (1986).
- Tokuyasu, K. T. A technique for ultracryotomy of cell suspensions and tissues. Journal of Cell Biology. 57 (2), 551-565 (1973).
- Connolly, C. N., Futter, C. E., Gibson, A., Hopkins, C. R., Cutler, D. F. Transport into and out of the Golgi complex studied by transfecting cells with cDNAs encoding horseradish peroxidase. Journal of Cell Biology. 127 (3), 641-652 (1994).
- Grabenbauer, M., et al. Correlative microscopy and electron tomography of GFP through photooxidation. Nature Methods. 2 (11), 857-862 (2005).
- Gaietta, G., et al. Multicolor and electron microscopic imaging of connexin trafficking. Science. 296 (5567), 503-507 (2002).
- Shu, X., et al. A genetically encoded tag for correlated light and electron microscopy of intact cells, tissues, and organisms. PLoS Biology. 9 (4), e1001041 (2011).
- Martell, J. D., et al. Engineered ascorbate peroxidase as a genetically encoded reporter for electron microscopy. Nature Biotechnology. 30 (11), 1143-1148 (2012).
- Lam, S. S., et al. Directed evolution of APEX2 for electron microscopy and proximity labeling. Nature Methods. 12 (1), 51-54 (2015).
- Mavlyutov, T. A., et al. APEX2-enhanced electron microscopy distinguishes sigma-1 receptor localization in the nucleoplasmic reticulum. Oncotarget. 8 (31), 51317-51330 (2017).
- Wang, Q., Mercogliano, C. P., Lowe, J. A ferritin-based label for cellular electron cryotomography. Structure. 19 (2), 147-154 (2011).
- Sano, T., Glazer, A. N., Cantor, C. R. A streptavidin-metallothionein chimera that allows specific labeling of biological materials with many different heavy metal ions. Proceedings of the National Academy of Sciences of the United States of America. 89 (5), 1534-1538 (1992).
- Mercogliano, C. P., DeRosier, D. J. Gold nanocluster formation using metallothionein: Mass spectrometry and electron microscopy. Journal of Molecular Biology. 355 (2), 211-223 (2006).
- Mercogliano, C. P., DeRosier, D. J. Concatenated metallothionein as a clonable gold label for electron microscopy. Journal of Structural Biology. 160 (1), 70-82 (2007).
- Morphew, M. K., et al. Metallothionein as a clonable tag for protein localization by electron microscopy of cells. Journal of Microscopy. 260 (1), 20-29 (2015).
- Nishino, Y., Yasunaga, T., Miyazawa, A. A genetically encoded metallothionein tag enabling efficient protein detection by electron microscopy. Journal of Electron Microscopy. 56 (3), 93-101 (2007).
- Bouchet-Marquis, C., Pagratis, M., Kirmse, R., Hoenger, A. Metallothionein as a clonable high-density marker for cryo-electron microscopy. Journal of Structural Biology. 177 (1), 119-127 (2012).
- Fukunaga, Y., et al. Electron microscopic analysis of a fusion protein of postsynaptic density-95 and metallothionein in cultured hippocampal neurons. Journal of Electron Microscopy. 56 (4), 119-129 (2007).
- Diestra, E., Fontana, J., Guichard, P., Marco, S., Risco, C. Visualization of proteins in intact cells with a clonable tag for electron microscopy. Journal of Structural Biology. 165 (3), 157-168 (2009).
- Risco, C., et al. sensitive, high-resolution detection of protein molecules in eukaryotic cells using metal-tagging transmission electron microscopy. Structure. 20 (5), 759-766 (2012).
- Ding, S. -. W., et al. Noncanonical role for the host Vps4 AAA+ ATPase ESCRT protein in the formation of tomato bushy stunt virus replicase. PLoS Pathogens. 10 (4), e1004087 (2014).
- Jiang, Z., et al. Direct synthesis of gold nanoparticles on cysteine-rich tags in yeast cells. Protocol Exchange. , (2020).
- Jiang, Z., et al. Direct synthesis of gold nanoparticles on cysteine-rich tags in mammalian cells. Protocol Exchange. , (2020).
- Jiang, Z., et al. Genetically encoded tags for direct synthesis of EM-visible gold nanoparticles in cells. Nature Methods. 17 (9), 937-946 (2020).
- Brust, M., Walker, M., Bethell, D., Schiffrin, D. J., Whyman, R. Synthesis of thiol-derivatised gold nanoparticles in a two-phase liquid-liquid system. Journal of the Chemical Society, Chemical Communications. 7, 801-802 (1994).

