Stem Cell-Derived Viral Ag-Specific T Lymphocytes Suppress HBV Replication in Mice
Summary
Presented here is a protocol for the effective suppression of hepatitis B virus (HBV) replication in mice by utilizing adoptive cell transfer (ACT) of stem cell-derived viral antigen (Ag)-specific T lymphocytes. This procedure may be adapted for potential ACT-based immunotherapy of HBV infection.
Abstract
Hepatitis B virus (HBV) infection is a global health issue. With over 350 million people affected worldwide, HBV infection remains the leading cause of liver cancer. This is a major concern, especially in developing countries. Failure of the immune system to mount an effective response against HBV leads to chronic infection. Although HBV vaccine is present and novel antiviral medicines are being created, eradication of virus-reservoir cells remains a major health topic. Described here is a method for the generation of viral antigen (Ag) -specific CD8+ cytotoxic T lymphocytes (CTLs) derived from induced pluripotent stem cells (iPSCs) (i.e., iPSC-CTLs), which have the ability to suppress HBV replication. HBV replication is efficiently induced in mice through hydrodynamic injection of an HBV expression plasmid, pAAV/HBV1.2, into the liver. Then, HBV surface Ag-specific mouse iPSC-CTLs are adoptively transferred, which greatly suppresses HBV replication in the liver and blood as well as prevents HBV surface Ag expression in hepatocytes. This method demonstrates HBV replication in mice after hydrodynamic injection and that stem cell-derived viral Ag-specific CTLs can suppress HBV replication. This protocol provides a useful method for HBV immunotherapy.
Introduction
Following acute infection, the adaptive immune system (i.e., humoral and cellular immunity) controls the bulk of acute HBV-related hepatitis. Still, a number of people in the HBV-endemic regions cannot eliminate the viruses and subsequently convert as chronic individuals. More than 25% of chronic patients (>250 million people) worldwide develop progressive liver disease, resulting in liver cirrhosis and/or hepatocellular carcinoma (HCC)1. As a result, eradication of insistently infected cells remains a general healthiness problem, even though there is an available vaccine2 and numerous antiviral medicines are under development. Standard treatment for HBV infection includes IFN-α, nucleoside, and nucleotide analogues. These agents have direct antiviral activity and immune modulatory capacities. Nevertheless, seroconversion of HBe antigen (Ag)+ carriers with anti-HBe antibody (Ab) and loss of serum HBV deoxyribonucleic acid (DNA) appear individually in approximately 20% of treated patients, and whole immunological control of the virus verified by the deprivation of the HBsAg is no more than 5%3. Moreover, the response to treatment is often not durable. Prophylactic vaccination with recombinant HBs Ag is highly effective in preventing infection, but therapeutic HBs Ag vaccination is not effective. Clearly, T cell-mediated immune responses play a critical role in controlling HBV infection and liver impairment; however, in chronic hepatitis patients, HBV-reactive T cells are often deleted, dysfunctional, or convert exhausted4,5,6. Consequently, in individuals with persistent HBV infection, no attempts to reinstate HBV-specific immunity (i.e., T cell-based immunity) by means of anti-viral remedy, immuno-modulatory cytokines, or curative immunization have achieved success.
Adoptive cell transfer (ACT) of HBV Ag-specific T cells is an efficient treatment directed to eventually eradicate remaining hepatocytes wih HBV7,8. ACT of HBV-specific CTLs into HBV-infected mice has been shown to cause transient, mild hepatitis, and a dramatic drop in HBV ribonucleic acid (RNA) transcripts in hepatocytes. In these studies, CTLs did not inhibit transcription of HBV genes but enhanced the degradation of HBV transcripts9. HBV-specific CTLs are important to prevent viral infection and mediate the clearance of HBV10,11. For ACT-based remedies, in vitro expansion of HBV-specific T cells with a high reactivity for in vivo resettlement has been suggested to be an ideal method12,13,14; nevertheless, the present approaches are restricted regarding their abilities to generate, separate, and grow appropriate quantities and qualities of HBV-specific T cells from patients for the potential therapies.
Although clinical trials present safety, practicability, and prospective therapeutic activity of cell-based treatments by means of engineered T cells that are specific to HBV virus-infected hepatocytes, there are worries about the unfavorable effects occurring from autoimmune responses because of cross-reactivity from mispairing T cell receptor (TCR)15,16, off-target Ag recognition by non-specific TCR17 and on-target off-toxicity by a chimeric Ag receptor (CAR)18,19 with healthy tissues. Currently, the genetically modified T cells, which only have short-term persistence in vivo, are usually intermediate or later effector T cells. To date, pluripotent stem cells (PSCs) are the only source available to generate high numbers of naive single-type Ag-specific T cells20,21,22,23. Induced PSCs (iPSCs) are simply converted from a patient’s somatic cells through the use of gene transduction of several transcription factors. As a result, the iPSCs have similar characteristics as those of embryonic stem cells (ESCs)24. Owing to the flexibility and possibility for the infinite ability to self-renew, in addition to tissue replacement, iPSC-based treatments may be widely applied in regenerative medicine. Furthermore, the regiments underlying iPSCs may substantially improve current cell-based therapies.
The overall goal of this method is to generate a large amount of HBV-specific CTLs from iPSCs (i.e., iPSC-CTLs) for ACT-based immunotherapy. The advantages over alternative techniques are that HBV-specific iPSC-CTLs have a single-type TCR and naive phenotype, which results in more memory T cell development after the ACT. It is demonstrated that the ACT of HBV-specific iPSC-CTLs increases the migration of functional CD8+ T cells in the liver and reduces HBV replication in both the livers and blood of administered mice. This method reveals a potential use of viral Ag-specific iPSC-CTLs for HBV immunotherapy and may be adapted to generate other viral Ag-specific iPSC-T cells for viral immunotherapy.
Protocol
All animal experiments are approved by The Texas A&M University Animal Care Committee (IACUC; #2018-0006) and are conducted in compliance with the guidelines of the Association for the Assessment and Accreditation of Laboratory Animal Care. Mice are used during 6–9 weeks of age.
1. Generation of viral Ag-specific CD8+ T cells from iPSCs (iPSC-CD8+ T cells)
- Creation of the retroviral constructs
NOTE: TCR α and β genes are linked with 2A self-cleaving sequence. The retroviral vector MSCV-IRES-DsRed (MiDR) is DsRed+ 23.- Sub-clone HBs183-191 (FLLTRILTI)-specific A2-restricted human-murine hybrid TCR (s183 TCR) genes (Vα34 and Vβ28) into the MiDR to create the s183 MiDR construct (Figure 1A)25.
- Retroviral transduction
NOTE: The Platinum-E (Plat-E) cells are used for packaging retroviruses (carrying s183 TCR genes), which will be used for retroviral transduction. The Plat-E cells are an effective retrovirus packaging cells underlying the 293T cells, which were developed by means of unique packaging constructs via the EF1α promoter to express retroviral structure protein, including gag, pol, and ecotropic env.- On a 100 mm culture dish, seed 3 x 106 Plat-E cells in 8 mL of DMEM culture medium containing 10% fetal calf serum (FCS) in an incubator at 37 °C with 5% CO2, 1 day before transfection.
- On day 0, transfect the s183 MiDR construct into the Plat-E cells using a DNA transfection reagent25.
- On day 1, seed 1 x 106 iPSCs (GFP+) into a gelatin pre-coated culture plate.
- On days 2–3, collect retroviruses-containing supernatant from Plat-E cell culture to transduce iPSCs with the s183 TCR in the presence of 1,6-dibromohexane solution25.
- On day 4, trypsinize the s183 TCR gene-transduced iPSCs, centrifuge at 400 x g for 5 min and seed 3 × 105 iPSCs in a 100 mm culture dish pre-coated with 3 x 106 irradiated SNL76/7 (irSNL76/7) feeder cells25.
- On day 5 or 6 of confluence, trypsinize the cells, centrifuge at 400 x g for 5 min and process for cell sorting. Gating on live cells, sort GFP and DsRED double-positive cells (the s183 TCR gene-transduced iPSCs) using a high-speed cell sorter. Similar to step 1.2.5, co-culture the sorted cells on irSNL76/7 feeder cells for future use25.
- Differentiation of HBV-specific iPSC-CD8+ T cells
NOTE: The OP9-DL1/DL4 stromal cells overexpress both Notch ligands DL1 and DL4, and co-culturing iPSCs with the iPSCs can promote Notch signaling-mediated T cell differentiation26.- Grow s183 TCR gene-transduced iPSCs (s183/iPSCs) in the OP9-DL1-DL4 cell monolayer in α-minimum essential medium (MEM) media containing 20% fetal bovine serum (FBS)27. Include murine Flt3 ligand (mFlt-3L; final concentration = 5 ng/mL) in the culture.
- On day 0, seed 0.5–1.0 x 105 S183/iPSCs in a 10 cm culture dish previously grown with OP9-DL1-DL4 cells. Validate that the OP9-DL1-DL4 cells are in a condition of 80%–90% confluency.
- On day 5, rinse the iPSCs with 10 mL of phosphate-buffered saline (PBS), aspirate off the PBS, add this to 4 mL of 0.25% trypsin, and incubate in a 37 °C incubator for 10 min. Afterward, add a supplementary 8 mL of iPSC media to end trypsin digestion. Accumulate all the digestive solutions containing the cells and centrifuge at 400 x g for 5 min at 15–30 °C.
- Aspirate the supernatant and resuspend cells in 10 mL of iPSC media. Transfer the cell suspension to a new 10 cm Petri plate and incubate in an incubator for 30 min at 37 °C.
- After 30 min, collect the iPSC media containing the floating cells. Pass the cell suspension through a 70 μm cell strainer and calculate the cell number.
- Seed 5 x 105 cells in the culture dish previously grown OP9-DL1-DL4 cells with a condition of 80%–90% confluent as described in step 1.3.1.
NOTE: For T cell differentiation, each 2–3 days, the iPSC-derived cells need to re-seed with a fresh layer of the OP9-DL1-DL4 cells.
- Evaluation
- Morphological changes of differentiating iPSCs
- Observe cells under a microscope on various days.
NOTE: By day 5, colonies have mesoderm-like characteristics, exhibiting a classic spindle-shape morphology resembling human dermal fibroblasts and sustained growth in vitro. By day 8, small round clusters of cells begin to appear.
- Observe cells under a microscope on various days.
- Analysis of differentiating iPSCs by flow cytometry
- On various days of co-culture, analyze iPSC-derived cells as described previously25 (Figure 1B,C).
- Functional analysis of differentiating iPSCs
- On day 28 of co-culture, collect iPSC-CD8+ T cells from cultures through harvesting the floating cells, trypsinize the leftover cells with 0.25% trypsin, re-suspend in 8 mL of iPSC media, centrifuge for 5 min at 400 x g at 15–30 °C, remove the media, then resuspend the cells in 10 mL of media.
- Keep the re-suspended cells in a fresh 10 cm dish in a 37 °C incubator for 30 min and assemble the floating cells. Then rinse the cells one times with a cold PBS.
- Incubate 3 x 106 T cell-depleted splenocytes (CD4–CD8–) from spleens of H-2 class I knockout, HLA-A2.1-transgenic (HHD) mice with 5 µM s183 peptide (FLLTRILTI) in 200 μL of media at 4 °C for 30 min.
- Produce a mixture of iPSC-CD8+ T cells with splenocytes pulsed with s183 peptide (T cells: splenocytes = 1:4; use 0.75 x 106 T cells). Incubate the mixture of cells at 37 °C in a CO2 incubator for 40 h. During the last 7 h, add 4 µL of diluted brefeldin A into the culture (final concentration of 1,000x, which will be diluted in 1x culture media) to block transport processes during cell activation.
- Stain the cells and perform flow cytometric analysis of intracellular IFN-γ as described previously (Figure 2).
- Morphological changes of differentiating iPSCs
2. Induction of HBV replication through hydrodynamic delivery of HBV plasmid
NOTE: pAAV/HBV1.2 construct was generated as described previously9. The HBV 1.2 complete DNA is incorporated in the vector pAAV.
- Hydrodynamic deliveries of HBV plasmid through the tail vein
- Heat HHD mice using a heat lamp for 5 min in the cage in order to dilate the tail vein.
- Detain the animal by means of a restrainer, and clean mouse tails with 70% ethanol spray.
- Delivery of plasmid into the liver.
- Measure the mouse body weight by using a measuring scale.
- Dilute 10 μg of HBV plasmid in the 8% equivalent of body mass PBS (e.g., 1.6 mL for a 20 g mouse). Load diluted plasmid in a 3 mL syringe with a 26 G × 1⁄2″ (0.45 × 12 mm) hypodermic needle.
- Locate one of the two lateral tail veins in the middle third of the tail and position the needle into either lateral vein. Administer the injection containing HBV plasmid through the tail vein within 3 – 5 s.
- Quantification of viremia from blood serum of infected mice
NOTE: HBV replication occurs from day 3–35 in the mouse serum. The DNA replication peaks on day 7 and reduces gradually. The HBV DNA is not cleared from the serum until day 35.- Collection of blood serum from infected mice
- Collect approximately 0.1 mL of blood in a microcentrifuge tube from each mouse on 3, 5, 7, and 10 days post-infection by retro-orbital bleeding in a 1.5 mL microcentrifuge tube, then incubate at room temperature (RT) for 20 min.
- Centrifuge the sample at 6,000 x g for 15 min at 4 °C and collect the serum supernatant after centrifugation.
- Purify the DNA from the blood serum using a commercially available DNA extraction kit following the manufacturer’s recommendations. Briefly, lyse the cells in a silica-based column, perform the washes, and extract DNA by adding 100% ethanol to the elution column and centrifuging at 6000 x g for 1 min. Elute the DNA in 50 µL of RNase-free water.
- Use 100 ng of HBV DNA from the elution for real-time PCR analysis. Use the following primers and probes: forward 5' TAGGAGGCTGTAGGCATAAATTGG 3'; reverse 5' GCACAGCTTGGAGGCTTGT 3'; probe 5' TCACCTCTGCCTAATC 3'.
- Use HBV genome containing plasmid (pAAV/HBV1.2) for standard curve and perform real-time PCR in a total volume of 10 μL (Figure 3).
- Set up the PCR reaction in a total volume of 10 μL as shown in Table 1.
- Set up the PCR program in the thermocycler as shown in Table 2. The programmed temperature transition rate is 20 °C/s for denaturation/annealing and 5 °C/s for extension. Measure the fluorescence at the end of the annealing phase for each cycle for real-time PCR monitoring.
- Collection of blood serum from infected mice
3. Reduction of HBV replication by ACT of viral Ag-specific iPSC-CD8+ T cells
- Adoptive cell transfer (ACT)
- Differentiate s183/iPSCs (1.3.7) upon the OP9-DL1-DL4 stromal cells in the presence of mFlt-3L and mIL-7 for 8 days as described in section 1.3.
- On day 22, collect iPSC-CD8+ T cells from the 10 cm plate with trypsin, then wash and resuspend each 10 cm plate in 10 mL of fresh media. Add the cells to a fresh 10 cm plate and return to the incubator for 30 min as done in section 1.3. After 30 min, collect floating cells.
- Use a 70 μm nylon strainer to pass cells to eliminate cell clusters and count the cell number. Adapt the cells to a concentration of 1.5 x 107 cells/mL in cold PBS solution and use a 70 μm nylon strainer to pass cells to eliminate cell clusters again if needed. Keep cells on ice until the ACT.
- Inject 200 μL cell suspension (3 x 106 cells) into 4–6 week-old HHD mice through the tail vein.
- Induction of HBV replication
- On day 14 after cell transfer, perform the hydrodynamic delivery of HBV plasmid through the tail vein as described in section 2.1.
- Virus protein detection from infected liver
- Sacrifice mice on days 3, 5, 7, 14, and 21 post-infection. For euthanasia, in each cage, use 1–2 L of carbon dioxide (CO2) in the first stage. When the animal develops the loss of consciousness, gain the CO2 flow rate around 4–5 L/min. Perform mouse euthanasia using CO2 inhalation.
- Separate the liver by cutting the surface skin of the peritoneum utilizing scissors and forceps and slightly dragging the liver back to uncover the internal skin lining the peritoneal cavity. Collect and cut liver samples (length x width x height = 0.5 cm x 0.5 cm X 0.3 cm) of the infected mice to fit easily into the embedding cassette and block in 10% neutral buffer formalin for 4–24 h.
- Decalcify the liver samplesusing 2.5 M formic acid; rinse in xylene for 3 min, rinse 2x in 100% ethanol, rinse 2x in 95% ethanol, rinse 2x in deionized (DI) water for 2 min, then decalcify the liver samples in 1 mM ethylenediaminetetraacetic acid (EDTA) correspondingly. Handle at a sub-boiling temperature (90 °C) for 20 min.
- Cool the fixed liver tissues for 30 min. Rinse the tissues with 1x PBS for 4 min and embed the tissues in paraffin. Dehydrate the tissues in a series of increasing concentrations of ethanol to replace the water, and then immerse in xylene immersion. Embed the infiltrated tissues into wax blocks. Make evenly vertical and horizontal sectioning for staining. Prepare 4 μm sections with a sliding microtome.
- Pass the sections through the deparaffinization and rehydration using xylene and ethanol, then perform immunofluorescent staining of the sections.
- Stain the sections with 200 μL of HBV surface Ag-specific antibody (1:100 dilution in the blocking solution). Incubate the sections with the antibody for 2 h at RT in a 75%–100% humidified chamber, and wash 5x in 1x PBS for 5 min.
- Use an anti-fade reagent containing 4',6-diamidino-2-phenylindole (DAPI) for nuclear staining to counterstain the slides. Add the coverslip with about 300 µL of the diluted DAPI staining solution (300 nM in 1x PBS) and validate the entire coverslip covered. Keep the slides in the dark at 4 °C until inspection under a fluorescent microscope.
- Inflammatory cell infiltration in infected mice
- Sacrifice mice as described in step 3.3.1. Collect the liver and make sections as described in step 3.3.4.
- Stain the section with hematoxylin and eosin (H&E) to evaluate infiltration of inflammatory cells into the liver.
- Store slides in the dark at 4 °C until further analysis under a fluorescent microscope.
- Visualize slides under a fluorescence microscope to detect the infiltration of inflammatory cells into the liver (Figure 4).
- HBV DNA detection of infected liver
- Isolation of viral DNA.
- Euthanize mice and collect the liver samples according to section 3.3.
- Lyse the liver tissues in Nonindet P-40 (NP-40) lysis buffer (50 mM Tris-HCL, 1 mM EDTA, 1% NP-40) containing a protease inhibitor cocktail.
- Briefly centrifuge at 16,000 x g to remove the nuclei and cell debris.
- Incubate the cytoplasmic lysate with micrococcal nuclease (nuclease S7; 150 units/mL) and CaCl2 (5 mM) at 37 °C for 90 min to degrade the nucleic acid outside nucleocapsids (NCs).
- Inactivate the nuclease S7 by the addition of 10 mM EDTA.
- Precipitate the nucleocapsids with polyethylene glycol (PEG), disrupt by 0.5% sodium dodecyl sulfate (SDS), and digest with 0.6 mg/mL proteinase K (PK) at 37 °C for 1 h.
- Recover the viral nucleic acids by phenol chloroform extraction and ethanol precipitation.
- Resolve the extracted viral DNA on a 1.2% agarose gel and detect by standard Southern blot analysis using a32 P-labeled HBV DNA probe.
Representative Results
As shown here, HBV viral Ag-specific iPSC-CD8+ T cells are generated by an in vitro culture system. After ACT of these viral Ag-specific iPSC-CD8+ T cells substantially suppress HBV replication in a murine model (Supplemental File 1). Mouse iPSC are transduced with the MIDR retroviral construct encoding a human-mouse hybrid HBV TCR gene (HBs183-191-specific, s183), then the gene-transduced iPSCs are co-cultured with OP9-DL1/DL4 cells expressing Notch ligands (both DL1 and DL4) molecules in the presence of rFlt3L and rIL-7. On day 28 of in vitro co-culture, the iPSC-derived cells substantially express CD3 and Ag-specific TCR (T cell markers). Flow cytometric analysis of the CD3+CD8+ population shows that the HBV TCR transduction dramatically increases the generation of viral s183-specific CD8+ T cells (Figure 1).
On day 28 of in vitro co-culture, the CD4–CD8+ single-positive (SP) iPSC-CD8+ T cells are isolated and stimulated by T-depleted splenocytes pulsed with s183 peptide, and cytokine production is assessed. The iPSC-CTLs produce large amounts of IL-2 and IFN-ϒ as detected by intracellular staining or ELISA (Figure 2). HBV infection is induced in HLA-A2.1 transgenic (HHD) mice by hydrodynamic injection of 10 µg of pAAV/HBV1.2 DNA plasmid through the tail veins of mice. HBV replication is detected from day 3–21 in the serum of HHD mice. The DNA replication peaks on day 6 and reduces gradually using real-time PCR analysis (Figure 3).
Two weeks following the ACT, mice are challenged with a hydrodynamic injection of pAAV/HBV1.2 DNA plasmid. The viral titer is measured by real-time PCR from serum at various timepoints after the injection. The results demonstrate that viral replication is significantly reduced at all timepoints in the mice receiving HBV viral Ag-specific T cells compared to the mice receiving control cells (Figure 4A). HBV surface protein expression is also examined in the liver in the above setting of treatment. Mice are euthanized at various days after the HBV injection and the liver samples are isolated for histologic examination. Samples are stained for HBV surface protein and examined under a fluorescence microscope. HBV surface protein is observed as substantially decreased in the mice receiving HBV viral Ag-specific pre-iPSC-CTLs, as compared with the mice receiving control cells (Figure 4B). This experiment clearly demonstrates that viral Ag-specific iPSC-CD8+ T cells have the ability to reduce HBV replication in the murine model.
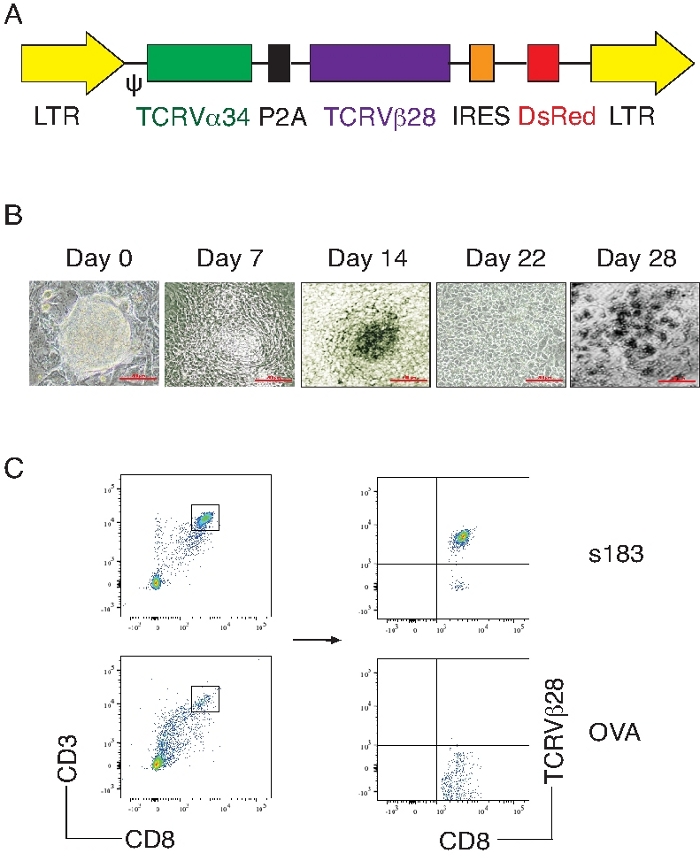
Figure 1: Generation of HBV viral Ag-specific iPSC-CD8+ T cells.
Mouse iPSCs are transduced with the following retroviral constructs: HBs183-91 TCR (MiDR-s183 TCR) or OVA257–264 TCR (MiDR-OVA TCR), and the transduced iPSCs are co-cultured with OP9-DL1/DL4 stromal cells for T lineage differentiation. (A) Schematic representation of the retroviral construct MiDR-s183 TCR expressing s183-specific TCR. Ψ = packaging signal; 2A = picornavirus self-cleaving 2A sequence; LTR = long terminal repeats. (B) Morphology of T cell differentiation on days 0, 7, 14, and 22. (C) Flow cytometric analysis for the iPSC-derived cells on day 28. CD3+CD8+ cells (left) are gated as indicated and analyzed for the expression of CD8 and TCRVβ28 (right). Data shown are representative of three individual experiments. The values represent mean ± SD (**p < 0.01; paired t-tests). Please click here to view a larger version of this figure.
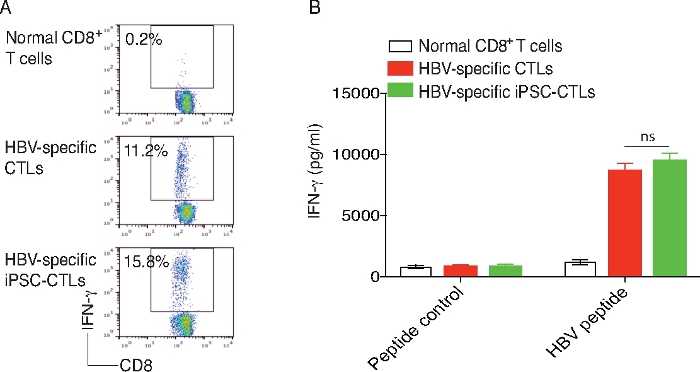
Figure 2: Functional analysis of the HBV viral Ag-specific iPSC-CD8+ T cells.
On day 28 of in vitro co-culture, the SP CD8+s183 TCR pentamer+ iPSC-T cells are sorted. The iPSC-T cells and CD8+ T cells transduced with MiDR-s183 TCR are stimulated by T-depleted splenocytes (APCs) from HHD mice and pulsed with s183 peptide (FLLTRILTI). (A) Intracellular staining of IFN-ϒ after 7 h (gated on CD8+ cells) (T/APCs = 1:4). (B) ELISA of IFN-γ after 40 h. Data shown are representative of three individual experiments. The values represent mean ± SD (n.s., p > 0.05; paired t-tests). Please click here to view a larger version of this figure.
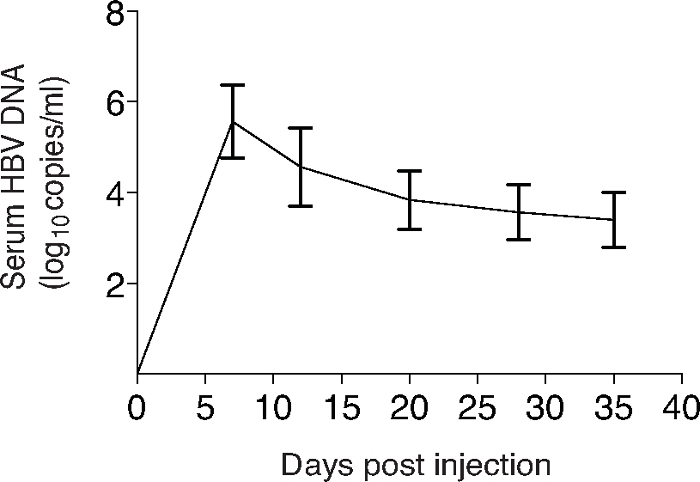
Figure 3: Induction of HBV replication in HHD mice by hydrodynamic injection.
HHD mice are i.v. administrated with HBV plasmid via hydrodynamic tail vein injection. 10 μg of the plasmid is injected with 8% of total body mass PBS. On indicated time points after injection, the serum is isolated from the blood and DNA is extracted for real-time PCR. Data shown are representative of three individual experiments. The values represent mean ± SD. Data are representative of five mice per group of three independent experiments. Please click here to view a larger version of this figure.
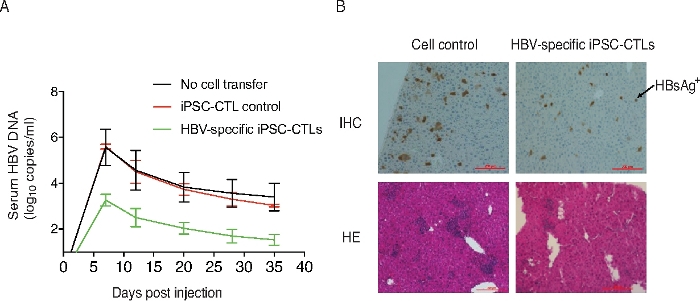
Figure 4: Reduction of HBV replication by ACT of viral Ag-specific iPSC-CD8+ T cells.
HHD mice are i.v. adoptively transferred with viral Ag-specific iPSC-CD8+ T cell progenitors (on day 22 of the in vitro culture) and administrated with HBV plasmid at week 2 after the cell transfer. (A) Serum HBV copies. At the indicated timepoints after injection, serum is isolated from blood, and DNA is extracted for real-time PCR analysis. Data shown are representative of three individual experiments. The values represent mean ± SD. (B) Liver tissue histology. Mice are euthanized at day 8 post-HBV infection. Liver samples are isolated and stained for histologic examination. Upper panel shows the HBsAg protein expression in infected mice (IHC staining) and lower panel shows the inflammatory cell infiltration (HE-staining). Data are representative of five mice per group of three independent experiments. Please click here to view a larger version of this figure.
| DNA template | 2 ml |
| DNA master hybridization mixture ((Taq DNA polymerase, PCR reaction buffer, 10 mM MgCl2, and dNTP mixture) | 1 ml |
| 25 mM MgCl2 | 0.8 ml |
| 0.3 μM the probe | 3 ml |
| 5 μM of each primer | 3.2 ml |
| Total | 10 ml |
Table 1: PCR reaction volume
| Temperature | Time | |
| Denaturation | 95 °C | 5 s |
| Annealing | 53 °C | 10 s |
| Extension | 72 °C | 20 s |
| 5 °C |
Table 2: PCR program
Supplemental File 1: Please click here to download this file.
Discussion
This protocol presents a method to generate the viral Ag-specific iPSC-CTLs for use as ACT to suppress HBV replication in a murine model. In chronic HBV infection, the viral genome forms a stable mini chromosome, the covalently closed circular DNA (cccDNA) that can persist throughout the lifespan of the hepatocyte. Targeting the clearance of the viral mini chromosome may result in a cure of chronic HBV infection. Current antiviral therapy targets the virus reverse transcriptase but rarely establishes immunological control over HBV replication driven by cccDNA. HBV-specific CD8+ CTLs can mediate the killing of the infected hepatocytes and accelerate the clearance of cccDNA. Nevertheless, the HBV-specific CTLs are deleted, dysfunctional, or succumb to exhaustion in patients with chronic HBV infection. ACT with the HBV-specific CTLs is a favorable treatment for chronic HBV infection28,29. Naive or central memory T cell-derived T lymphocytes (i.e., highly reactive immune cells) are ideal effectors for ACT-based therapies owing to their great proliferation, lower tendency for death compared to terminally differentiated cells, and excellent ability to respond to homeostatic cytokines. However, such ACT has been often not feasible due to difficulties in obtaining sufficient numbers of CTLs from patients. It has been previously showed that reprogramming of Ag-specific CTLs or Tregs from iPSCs can be used for cell-based therapies20,23,27,30. This report demonstrates a method to generate viral Ag-specific iPSC-CTLs and use these cells for ACT-based immunotherapy in a murine model of HBV replication.
Although there are transgenic mouse models of HBV replication, these models are challenging because the central tolerance induced by the transgenic gene products causes mice to be immune tolerant to HBV Ags. Additionally, transgenic mice are not suitable for monitoring viral clearance as the integrated HBV genome persists in each mouse cell31,32. Also, despite that successful vaccines have been developed for preventing the infection, the treatment or immunotherapy after HBV infection has not been developed. Furthermore, the experimental approaches to HBV pathogenesis have been hampered because the host range of HBV infection is limited to men and chimpanzees, and in vitro culture system for the propagation of HBV is not sufficient. CD8+ T cells are promising effector cells against various types of viral infection; however, T cell response against HBV is not abundant. Specificity, functioning, and lack of sufficient numbers to mount an immune response may be the cause. This report reveals the method of hydrodynamic injection that efficiently induces HBV replication in mice. The method allows delivery of a large amount of HBV plasmid directly into the liver. The model exhibits continuous viremia for more than 8 weeks with detection of HBV mRNA, protein, and DNA at different days after injection. This method for HBV replication in mice is useful for HBV immunotherapy.
In summary, HBV replication was successfully induced in mice through hydrodynamic injection procedure, and viral Ag-specific iPSC-CTLs were used as immunotherapy for HBV replication. However, there are two limitations of the method: (1) more than three weeks of in vitro T cell differentiation may reduce the translational applications of the generated T cells for ACT; and (2) HBV hydrodynamic injection can efficiently induce HBV replication in the liver; however, it does not form the cccDNA, which is the main reason for the persistent HBV infection. Nevertheless, the method provides an alternative approach for HBV replication and treatment. A combination regimen using anti-HBV drugs and the ACT of viral Ag-specific iPSC-CTLs is likely to reduce HIV reservoirs, thereby resulting in a therapy for chronic HIV infection.
Divulgazioni
The authors have nothing to disclose.
Acknowledgements
The authors thank Dr. Adam J Gehring from Toronto General Hospital Research Institute for providing cDNA for HBs183-91 (s183) (FLLTRILTI)- specific A2-restricted human-murine hybrid TCR genes, and Dr. Pei-Jer Chen from National Taiwan University for providing pAAV/HBV 1.2 construct. This work is supported by the National Institute of Health Grant R01AI121180, R01CA221867 and R21AI109239 to J. S.
Materials
| HHD mice | Institut Pasteur, Paris, France | H-2 class I knockout, HLA-A2.1-transgenic (HHD) mice | |
| iPS-MEF-Ng-20D-17 | RIKEN Cell Bank | APS0001 | |
| SNL76/7 | ATCC | SCRC-1049 | |
| OP9 | ATCC | CRL-2749 | |
| pAAV/HBV1.2 plasmid | Dr. Dr. Pei-Jer Chen (National Taiwan University Hospital, Taiwan) | HBV DNA construct | |
| HBs183-91(s183) (FLLTRILTI)-specific TCR genes | Dr. Adam J Gehring (Toronto General Hospital Research Institute, Toronto, Canada) | FLLTRILTI-specific A2-restricted human-murine hybrid TCR genes (Vα34 and Vβ28) | |
| OVA257–264-specific TCR genes | Dr. Dario A. Vignali (University of Pittsburgh, PA) | SIINFEKL-specific H-2Kb-restricted TCR genes | |
| Anti-CD3 (17A2) antibody | Biolegend | 100236 | |
| Anti-CD44 (IM7) antibody | BD Pharmingen | 103012 | |
| Anti-CD4 (GK1.5) antibody | Biolegend | 100408 | |
| Anti-CD8 (53-6.7) antibody | Biolegend | 100732 | |
| Anti-IFN-γ (XMG1.2) antibody | Biolegend | 505810 | |
| Anti-TNF-a (MP6-XT22) antibody | Biolegend | 506306 | |
| α-MEM | Invitrogen | A10490-01 | |
| Anti-HBs antibody | Thermo Fisher | MA5-13059 | |
| ACK Lysis buffer | Lonza | 10-548E | |
| Brefeldin A | Sigma | B7651 | |
| DMEM | Invitrogen | ABCD1234 | |
| FBS | Hyclone | SH3007.01 | |
| FACSAria Fusion cell sorter | BD | 656700 | |
| Gelatin | MilliporeSigma | G9391 | |
| GeneJammer | Agilent | 204130 | |
| HLA-A201-HBs183-91-PE pentamer | Proimmune | F027-4A – 27 | |
| HRP Anti-Mouse Secondary Antibody | Invitrogen | A27025 | |
| mFlt-3L | Peprotech | 250-31L | |
| mIL-7 | Peprotech | 217-17 | |
| Nuclease S7 | Roche | 10107921001 | |
| Paraformaldehyde | MilliporeSigma | P6148-500G | Caution: Allergenic, Carcenogenic, Toxic |
| Permeabilization buffer | Biolegend | 421002 | |
| Polybrene | MilliporeSigma | 107689 | |
| ProLong™ Gold Antifade Mountant with DAPI | Invitrogen | P36931 | |
| QIAamp MinElute Virus Spin Kit | Qiagen | 57704 |
Riferimenti
- Scaglione, S. J., Lok, A. S. Effectiveness of hepatitis B treatment in clinical practice. Gastroenterology. 142 (6), 1360-1368 (2012).
- Osiowy, C. From infancy and beyond… ensuring a lifetime of hepatitis B virus (HBV) vaccine-induced immunity. Human Vaccines & Immunotherapeutics. 14 (8), 2093-2097 (2018).
- Gish, R. G., et al. Loss of HBsAg antigen during treatment with entecavir or lamivudine in nucleoside-naive HBeAg-positive patients with chronic hepatitis B. Journal of Viral Hepatitis. 17 (1), 16-22 (2010).
- Kurktschiev, P. D., et al. Dysfunctional CD8+ T cells in hepatitis B and C are characterized by a lack of antigen-specific T-bet induction. Journal of Experimental Medicine. 211 (10), 2047-2059 (2014).
- Fisicaro, P., et al. Antiviral intrahepatic T-cell responses can be restored by blocking programmed death-1 pathway in chronic hepatitis B. Gastroenterology. 138 (2), 682-693 (2010).
- Schurich, A., et al. The third signal cytokine IL-12 rescues the anti-viral function of exhausted HBV-specific CD8 T cells. PLoS Pathogens. 9 (3), 1003208 (2013).
- Gehring, A. J., et al. Engineering virus-specific T cells that target HBV infected hepatocytes and hepatocellular carcinoma cell lines. Journal of Hepatology. 55 (1), 103-110 (2011).
- Xia, Y., et al. Interferon-gamma and Tumor Necrosis Factor-alpha Produced by T Cells Reduce the HBV Persistence Form, cccDNA, Without Cytolysis. Gastroenterology. 150 (1), 194-205 (2016).
- Huang, L. R., Wu, H. L., Chen, P. J., Chen, D. S. An immunocompetent mouse model for the tolerance of human chronic hepatitis B virus infection. Proceedings of the National Academy of Sciences U.S.A. 103 (47), 17862-17867 (2006).
- Wong, P., Pamer, E. G. CD8 T cell responses to infectious pathogens. Annual Review of Immunology. 21, 29-70 (2003).
- Murray, J. M., Wieland, S. F., Purcell, R. H., Chisari, F. V. Dynamics of hepatitis B virus clearance in chimpanzees. Proceedings of the National Academy of Sciences USA. 102 (49), 17780-17785 (2005).
- Hinrichs, C. S., et al. Adoptively transferred effector cells derived from naive rather than central memory CD8+ T cells mediate superior antitumor immunity. Proceedings of the National Academy of Sciences U.S.A. 106 (41), 17469-17474 (2009).
- Hinrichs, C. S., et al. Human effector CD8+ T cells derived from naive rather than memory subsets possess superior traits for adoptive immunotherapy. Blood. 117 (3), 808-814 (2011).
- Kerkar, S. P., et al. Genetic engineering of murine CD8+ and CD4+ T cells for preclinical adoptive immunotherapy studies. Journal of Immunotherapy. 34 (4), 343-352 (2011).
- Kuball, J., et al. Facilitating matched pairing and expression of TCR chains introduced into human T cells. Blood. 109 (6), 2331-2338 (2007).
- van Loenen, M. M., et al. Mixed T cell receptor dimers harbor potentially harmful neoreactivity. Proceedings of the National Academy of Sciences USA. 107 (24), 10972-10977 (2010).
- Cameron, B. J., et al. Identification of a Titin-derived HLA-A1-presented peptide as a cross-reactive target for engineered MAGE A3-directed T cells. Science Translational Medicine. 5 (197), (2013).
- Fedorov, V. D., Themeli, M., Sadelain, M. PD-1- and CTLA-4-based inhibitory chimeric antigen receptors (iCARs) divert off-target immunotherapy responses. Science Translational Medicine. 5 (215), (2013).
- Maus, M. V., et al. T cells expressing chimeric antigen receptors can cause anaphylaxis in humans. Cancer Immunolology Research. 1 (1), 26-31 (2013).
- Haque, R., et al. Programming of regulatory T cells from pluripotent stem cells and prevention of autoimmunity. Journal of Immunology. 189 (3), 1228-1236 (2012).
- Vizcardo, R., et al. Regeneration of human tumor antigen-specific T cells from iPSCs derived from mature CD8(+) T cells. Cell Stem Cell. 12 (1), 31-36 (2013).
- Nishimura, T., et al. Generation of rejuvenated antigen-specific T cells by reprogramming to pluripotency and redifferentiation. Cell Stem Cell. 12 (1), 114-126 (2013).
- Lei, F., et al. In vivo programming of tumor antigen-specific T lymphocytes from pluripotent stem cells to promote cancer immunosurveillance. Ricerca sul cancro. 71 (14), 4742-4747 (2011).
- Kim, J. B., et al. Oct4-induced pluripotency in adult neural stem cells. Cell. 136 (3), 411-419 (2009).
- Lei, F., Haque, R., Xiong, X., Song, J. Directed differentiation of induced pluripotent stem cells towards T lymphocytes. Journal of Visualized Experiments. (63), e3986 (2012).
- Lei, F., Haque, M., Sandhu, P., Ravi, S., Ni, Y., Zheng, S., Fang, D., Jia, H., Yang, J. M., Song, J. Development and characterization of naive single-type tumor antigen-specific CD8+ T lymphocytes from murine pluripotent stem cells. OncoImmunology. 6, (2017).
- Haque, M., et al. Melanoma Immunotherapy in Mice Using Genetically Engineered Pluripotent Stem Cells. Cell Transplantation. 25 (5), 811-827 (2016).
- Tan, A. T., et al. Use of Expression Profiles of HBV DNA Integrated Into Genomes of Hepatocellular Carcinoma Cells to Select T Cells for Immunotherapy. Gastroenterology. , (2019).
- Wu, L. L., et al. Ly6C(+) Monocytes and Kupffer Cells Orchestrate Liver Immune Responses Against Hepatitis B Virus in Mice. Hepatology. , (2019).
- Haque, M., et al. Stem cell-derived tissue-associated regulatory T cells suppress the activity of pathogenic cells in autoimmune diabetes. Journal of Clinical Investigation Insights. , (2019).
- Chisari, F. V., et al. Structural and pathological effects of synthesis of hepatitis B virus large envelope polypeptide in transgenic mice. Proceedings of the National Academy of Sciences USA. 84 (19), 6909-6913 (1987).
- Wirth, S., Guidotti, L. G., Ando, K., Schlicht, H. J., Chisari, F. V. Breaking tolerance leads to autoantibody production but not autoimmune liver disease in hepatitis B virus envelope transgenic mice. Journal of Immunology. 154 (5), 2504-2515 (1995).

