Multiplexed Single-molecule Force Proteolysis Measurements Using Magnetic Tweezers
Summary
In this article we describe the use of magnetic tweezers to study the effect of force on enzymatic proteolysis at the single molecule level in a highly parallelizable manner.
Abstract
The generation and detection of mechanical forces is a ubiquitous aspect of cell physiology, with direct relevance to cancer metastasis1, atherogenesis2 and wound healing3. In each of these examples, cells both exert force on their surroundings and simultaneously enzymatically remodel the extracellular matrix (ECM). The effect of forces on ECM has thus become an area of considerable interest due to its likely biological and medical importance4-7.
Single molecule techniques such as optical trapping8, atomic force microscopy9, and magnetic tweezers10,11 allow researchers to probe the function of enzymes at a molecular level by exerting forces on individual proteins. Of these techniques, magnetic tweezers (MT) are notable for their low cost and high throughput. MT exert forces in the range of ~1-100 pN and can provide millisecond temporal resolution, qualities that are well matched to the study of enzyme mechanism at the single-molecule level12. Here we report a highly parallelizable MT assay to study the effect of force on the proteolysis of single protein molecules. We present the specific example of the proteolysis of a trimeric collagen peptide by matrix metalloproteinase 1 (MMP-1); however, this assay can be easily adapted to study other substrates and proteases.
Protocol
1. Flow Cell Preparation
- Coverslips (#1.5, 22×22 mm and 22×40 mm, VWR) are cleaned using sonication.
- Add the coverslips to a small glass container capable of holding coverslips and fitting in the sonicator (see step 2).
- Fill the container with isopropanol and sonicate in a bath sonicator for 20 minutes.
- Discard the isopropanol and rinse the coverslips with copious quantities of deionized water produced by a Barnsted MilliQ apparatus or similar device. Fill the container with water and sonicate for 20 minutes.
- After sonication, dry the coverslips in a stream of filtered, dust-free air. Use only coverslips that appear clean (i.e. no smudges).
- Gently flame the coverslips for a few seconds to clean them of any remaining dust and moisture: pass the coverslips through a gas flame produced with a Bunsen burner, taking care not to warp the coverslip due to excess heat. This step removes residual surface contaminants.
- Using double-sided scotch tape, attach the two coverslips together. Cut the tape ~3 cm in length, and 0.3 cm in width. Attach the tape on 22×40 mm coverslip leaving a 1.6 cm channel in the middle. Use a pipette tip to gently press down on the coverslip to ensure that the tape has properly adhered.
2. Magnetic Tweezers Setup and Calibration
- Magnetic tweezers were setup using permanent rare-earth magnets as described previously10,12. An aluminum L-bracket was constructed with a 1 mm diameter pinhole and mounted on a vertical z-translator (Thorlabs) to modulate the height of the tweezers. Two permanent rare-earth magnets were attached to the bracket on either side of the pinhole to create the magnetic trap (Figure 1).
- DNA preparation: DNA from lambda phage labeled at its 5′ and 3′ ends with biotin and digoxigenin (IDT DNA, San Diego, CA) was used to calibrate the magnetic tweezers.
- Mix 50 μL of 4 nM λ-DNA with 2 μL of 40 nM biotin conjugated oligonucleotide and heat at 60 °C for 10 minutes. Then slowly cool at room temperature over 1 hour.
- Add DNA ligase according to manufacturer’s specifications and ligate at room temperature for 3 hours.
- Add 2 μL of 400 nM digoxigenin conjugated oligonucleotide and incubate at room temperature for 45 minutes.
- Add 1.5 μL of 10 mM ATP and continue ligation of the digoxigenin conjugated oligonucleotide to the λ-DNA for 3 hours.
- Remove the excess oligonucleotides and the DNA ligase using 100 kDa spin filters (Microcon Ultracell YM-100). A total of 1000-fold buffer exchange in TE buffer is adequate to remove the excess oligos. Note: It is important not to use spin speeds >500xG because it will lead to the shearing of the DNA.
- Examine a sample of the DNA via electrophoresis on a 0.7% agarose gel to ensure that the DNA is not denatured or sheared.
- Measure concentration of DNA. The use of a Nanodrop UV-Vis spectrometer (Thermo Scientific) is recommended because the available sample volume may be very small.
- Prepare flow cells as described in section 1.
- For the attachment of the DNA to the flow cell, add anti-digoxigenin antibody (sheep IgG; 1 μg mL-1; Roche Bioscience, Manheim, Germany) and allow it to adhere to the glass surface for 15 minutes.
- Passivate the surface of the flow cell to prevent non-specific sticking of DNA by adding 2 mg mL-1 BSA, 0.1% Tween-20 in 1x PBS. Incubate at room temperature for 45 minutes.
- Repeat step 2.5.
- Add 50 pM of fuctionalized λ-DNA and allow to attach to the coverslip for 15 minutes.
- Wash away excess DNA with 1x PBS.
- Add streptavidin-coated superparamagnetic beads (1 μm beads = 2 μg mL-1; 2.8 μm beads = 20 μg mL-1) and allow them to attach to the immobilized DNA for 15 minutes.
- Wash away excess beads with 1x PBS.
- Magnetic Tweezers Calibration
- Move the permanent magnets to a position that is far away from the sample surface (>15 mm).
- Place the prepared flow cell in the microscope.
- Move the permanent magnets into position above the flow cell.
- Choose the focal plane of λ-DNA functionalized beads by focusing on the beads away from both glass surfaces. The correct focal plane is one in which the desired beads have a bright center.
- Take a video of bead motion at 80 Hz or greater for 500 frames.
- Move the magnet closer to the sample surface. For distances beyond 5 mm, move in 1 mm increments. For distances between 1 and 5 mm, move in 0.5 mm increments. For distances less than 1 mm, move in 0.25 mm increments.
- Repeat steps 2.11.5 and 2.11.6 at each new magnet position.
- For each data set, track the center of the beads using a 2D Gaussian fit.
- Calculate the force via Brownian fluctuations using:
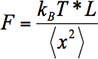
where kBT is the Boltzmann thermal energy, L is the length of λ-DNA, and <x2> is the variance in centroid position.
- Confirm the accuracy of the forces by checking the force calculation via a power spectrum Lorentzian fit in order to find the rolloff frequency. Applied force is related to the rolloff frequency by the relation:
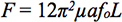
Where ƒ0 is the rolloff frequency, a is the bead radius, and μ is the fluid viscosity12.
3. Collagen Peptide Attachment to Flow Cells
In order to provide a concrete example for the application of our methodology, we describe recent work in our laboratory that characterizes the proteolysis of a trimeric collagen model peptide. We anticipate that this general approach may be broadly applied to other proteins and polynucleotides.
- The collagen model peptide consists of a N-terminal 6x His-tag for purification, followed by a 5x myc tag, (GPP)10 to enforce triple helix formation, the collagen α1 residues 772-786 (GPQGIAGQRGVVGL), which form the MMP-1 recognition site, the foldon sequence (GSGYIPEAPRDGQAYVRKDGEWVLLSTFL), and a C-terminal KKCK to facilitate labeling with biotin-maleimide. Foldon derives from the T4-phage protein fibritin, and stabilizes collagen model trimers when fused at either the N– or C-terminus13,14.
- The collagen peptide and MMP-1 proteins were expressed and purified as described previously4,15.
- Add anti-myc (15 μg mL-1) to the flow cell and incubate at room temperature for 20 minutes to allow the anti-myc to attach to the surface of the flow cell.
- Passivate the flow cell to prevent non-specific attachment of proteins using 5 mg mL-1 BSA in the same way as described in step 2.5.
- Add 150 pM collagen peptide and allow it to attach to the antibody via the myc tag for 45 minutes.
- Wash away excess collagen using PBS buffer.
- Add streptavidin coated superparamagnetic beads (1 μm beads = 2 μg mL-1and 2.8 μm beads = 20 μg mL-1) and allow them to attach to the collagen molecules for 45 minutes. The 1 μm beads should be diluted to the desired concentration in PBS. The 3 μm beads should be separated from the solution using strong magnets, then resolubilized in PBS. This process should be repeated 3 times. We noticed that this step is essential to get the 3 μm beads to bind to surface-immobilized collagen peptide.
4. Force Proteolysis Assay
- Once the flow cell is assembled with collagen peptide and magnetic beads, image the flow cell in the magnetic trap under low, ~1 pN force. In our experience, a subpopulation of the beads is sometimes weakly adhered to the surface in a non-specific fashion. The application of modest force ensures that these beads are liberated.
- Add activated enzyme to the flow cell. In this case MMP-1 was pre-activated by adding 3.5 mM APMA (4-aminophenylmercuric acetate) and incubated at 37 °C for 3 hours. The activation was verified using SDS-PAGE.
- As soon as the flow cell is re-introduced into the magnetic tweezers apparatus, record a video spanning a few fields of view, typically yielding several hundred attached beads. This will be t = 0.
- Repeat the same process of recording several fields of view (while returning to the same field of view) at regular time points, till all the beads detach or no further proteolysis is discernable.
- Kinetic data analysis
- Count the beads in the various fields of view at different time points and average them. Normalize the number of beads by dividing the average number of beads at each time point by the number of beads at t = 0. This ensures that we can compare the rates of proteolysis from different experiments, because the absolute number of beads will differ from experiment to experiment.
- Plot the ratio as a function of time. In this case, we fitted it to an exponential decay plus a constant.
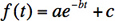
where ƒ(t) is the ratio of the beads attached (or collagen molecules unproteolyzed), t is time, a is the fraction of beads attached by a collagen tether, b is the decay rate constant, and c is the fraction of beads that are attached non-specifically.
- The same experiment is repeated at varying force and MMP-1 concentrations to elucidate the effect of force on collagen proteolysis by MMP-1.
5. Representative Results
The above protocol describes a novel use of magnetic tweezers (Figure 1) for studying the effect of force on enzymatic proteolysis. We calibrated the tweezers for 1 μm and 3 μm beads using both the magnitude of the observed Brownian fluctuations and calculation of the roll-off frequency at varying magnet positions (Figure 2). In the force proteolysis experiments, the setup is similar except that the DNA is replaced with collagen (Figures 3, 4). The normalized number of beads remaining can be plotted as function of time to find the proteolysis rates (Figure 5), and this process can be repeated for varying enzyme concentrations and forces.
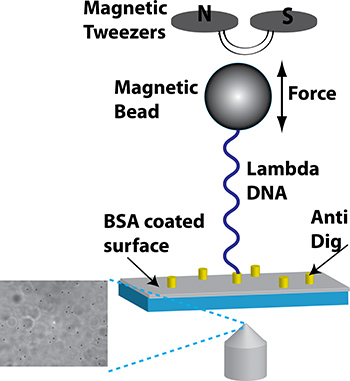
Figure 1. Schematic of the magnetic trap calibration process (not to scale). Two permanent rare earth magnets create a magnetic field that pulls on the superparamagnetic bead. Translating the magnet up and down adjusts the applied force. The beads are imaged using a conventional bright-field microscope with the light passing through a pinhole between the two magnets. Inset: Image taken with a 40x air objective. The sharp, round spots correspond to the beads attached to the coverslip surface. The out-of-focus objects are detached beads.
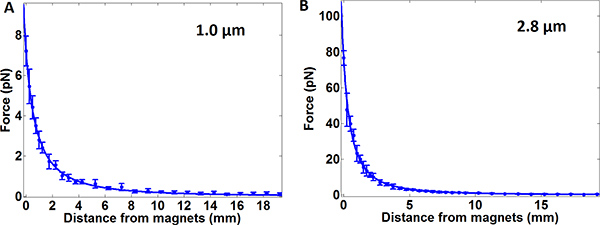
Figure 2. Plots of the calibrated force as a function of magnet distance from the sample surface for 1 μm (left) and 2.8 μm (right) beads. Data were fit to the empirical function 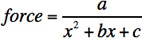 , where x is the distance from the magnet. a = 31.8, b= 5.61, and c = 4.39 for the 1 μm beads and a = 140, b = 3.10 and c = 1.86 for the 3 μm beads. These values are specific to our specific instrument and experimental geometry, and each instrument should be calibrated individually. The error bars at each point represent a ~10% variability in applied force on a bead to bead basis, due to the variability in bead size. The force applied is proportional to the volume of the beads, and the bead volume varies ~9% over the mean (according to manufacturer specifications).
, where x is the distance from the magnet. a = 31.8, b= 5.61, and c = 4.39 for the 1 μm beads and a = 140, b = 3.10 and c = 1.86 for the 3 μm beads. These values are specific to our specific instrument and experimental geometry, and each instrument should be calibrated individually. The error bars at each point represent a ~10% variability in applied force on a bead to bead basis, due to the variability in bead size. The force applied is proportional to the volume of the beads, and the bead volume varies ~9% over the mean (according to manufacturer specifications).
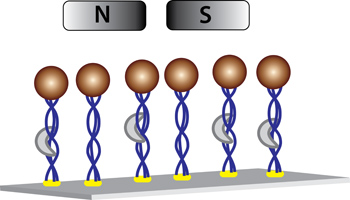
Figure 3. Force proteolysis assay setup (Not to scale). The collagen model trimer is attached to the surface of the coverslip via myc/anti-myc conjugation. The streptavidin-coated superparamagnetic beads are attached to the collagen trimers via a biotin-streptavidin linkage. Activated MMP-1 cuts the collagen over time, causing the beads to detach from the surface and move away from the focal plane.
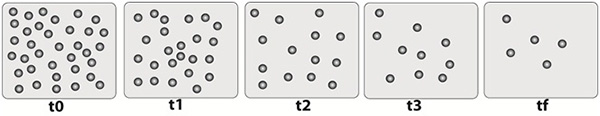
Figure 4. Schematic cartoon of proteolysis as a function of time. The cartoon shows a sample field of view over time as proteolysis occurs. Over time, MMP-1 cuts the collagen and the beads detach and move away from the focal plane under the influence of the magnetic field.
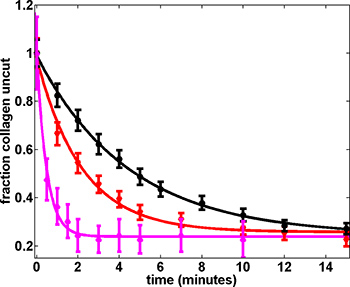
Figure 5. Proteolysis rates depend on applied force16. Shown are data collected at 1.0 pN (3 μM MMP-1; black), 6.2 pN (3 μM MMP-1; red) and 13 pN (0.2 μM MMP-1; magenta). The rates of proteolysis (fit parameters) are: 0.22 ± 0.02 min-1 (1 pN), 0.46 ± 0.09 min-1 (6.2 pN), and 2.08 ± 0.18 min-1 (13 pN). The fraction of beads unproteolyzed at long time points (> 15 minutes) remains approximately constant at ~0.25 across different experiments. Error bars correspond to the Poisson statistics reflecting the number of observations at each time point. The error at each time point for n beads is n1/2. The error in fraction of beads attached was calculated by error propagation.1 μm beads were used for 1.0 pN and 6.2 pN experiments and 2.8 μm beads were used for 13 pN experiments.
Discussion
This protocol describes a new use for a classical single molecule technique. Magnetic tweezers allow medium to high-throughput single molecule assays in a cost-efficient manner. However, like all experimental techniques there are challenges and potential pitfalls.
Limitations of magnetic tweezers
Compared to an optical trap the spatial and temporal resolution of a MT apparatus is low. Moreover, the forces generated by the simple MT described here are 30 pN or less, significantly less than forces routinely accessed in AFM experiments. This limitation can be a virtue: MT are well suited for applying sub-pN forces.
Unlike an optical trap or AFM, conventional MT, such as the apparatus here, do not allow the manipulation of individual particles. Optical traps are usually used to apply force parallel to the coverslip, while MT are best-suited for vertical pulling experiments. Electromagnetic MT address these limitations, although at the cost of increased complexity.
Tweezer Calibration
Both the Brownian fluctuation and power spectrum analysis for force calibration have their limitations. We found that the Brownian fluctuation method is particularly sensitive to camera blur. As the frame rate increased (exposure time decreased) the calculated force decreased. In practice, we increased frame rate until the forces calculated from Brownian fluctuations agreed with the forces calculated using the power spectrum to within 10%. Conversely, the power spectrum analysis is more robust against camera blur, but slightly more difficult to implement. We recommend performing both calculations to check for the robustness of the results.
We note that variations in bead size and magnetic particle content result in variations in applied force of ~10%. This effect was not important in our recent experiment because we average over hundreds of beads in each measurement. However, experiments that extract unique information from each tethered bead could potentially require a more accurate calibration method.
Controls to ensure single point attachments
Achieving specific, oriented, single-point attachments is often a practical challenge in single molecule studies. The following controls confirmed specific attachment of collagen to antibody, and beads to collagen:
- Added only beads to a flow cell passivated with BSA.
- Added anti-myc, passivated with BSA and then added beads.
- Added anti-myc, passivated with BSA, added non-biotinylated collagen and then added beads.
- Passivated with BSA, added non-biotinylated collagen and then added beads.
- Passivated with BSA, added biotinylated collagen and then added beads.
- Added anti-myc, passivated with BSA, added biotinylated collagen and then added beads.
In cases 1-5, where some component of the attachment series was deliberately omitted, anywhere from 0 to 4 beads were seen attached to the flow cell per field of view (80,000 μm2). Only in case 6, where all the attachment moieties are present, did we see ~30-60 2.8 μm beads and 80-200 1 μm beads attached to the surface. The proteolysis kinetics observed are adequately fit by a single exponential plus a constant term, which strongly suggests that there is only one rate limiting step, and hence only a single tether. The constant term likely reflects beads that are non-specifically attached to the coverslip.
We report the concentrations of antibody and protein that worked for our experiment. A wide range of concentrations for each component should be examined when developing a new assay. BSA worked well as a surface passivation agent in our experiment. However, this method may not work in all circumstances. Other protocols for surface passivation have also been used with success. Examples include polyethylene glycol functionalized glass slides17, supported lipid bilayers18, or casein as a blocking agent.
Notes on data analysis
In force proteolysis experiments, there is always some non-specific detachment (especially at higher forces19) due to dissociation of non-covalent (e.g. biotin – streptavidin) interactions. We account for this phenomenon by measuring proteolysis at various MMP-1 concentrations, including no MMP-1, as described in Adhikari et. al. Measuring proteolysis at various MMP-1 concentrations for all forces allows us to generate Michaelis-Menten curves, where are used to calculate the catalytic efficiency kcat/KM of the enzyme. Alternatively, if various concentrations of the enzyme are not used, we recommend that a control measurement be done with no enzyme at the given forces to subtract off the rate of non-specific detachment.
Potential applications
The magnetic tweezers cover a wide force range. The forces that can be exerted depend upon the size of the beads. 1 μm and 2.8 μm beads overlap in accessible forces, and in our experience, using either of the beads at the intermediate forces works well.
We anticipate that similar experimental approaches may be broadly applicable in studying force dependent proteolysis4,20, unfolding21 and binding interactions22. We and others23 have noted that the radius of diffraction rings for out of focus beads provides a sensitive means for tracking the position of the bead relative to the coverslip23. Although generally not used in this way, we believe that MT assays have the capacity to provide nanometer-precision measurements of protein structural dynamics. This additional information may be particularly valuable in the context of force-dependent proteolysis or binding measurements.
Disclosures
The authors have nothing to disclose.
Acknowledgements
This work was supported by the Burroughs Wellcome Career Award at the Scientific Interface (A.R.D.), the National Institutes of Health through the NIH Director’s New Innovator Award Program 1-DP2-OD007078 (A.R.D.), the William Bowes Jr. Stanford Graduate Fellowship (A.S.A.), and the Stanford Cardiovascular Institute Younger Predoctoral Fellowship (J.C.). The authors thank James Spudich for loaning microscopy equipment.
Materials
| Name of Reagent | Company | Catalogue Number |
| Micro Cover Glass #1.5 (22×22) | VWR | 48366-067 |
| Micro Cover Glass #1.5 (22×40) | VWR | 48393-048 |
| Lambda DNA | Invitrogen | 25250-010 |
| T4 DNA Ligase | Invitrogen | 15224-041 |
| Microcon Ultracel YM-100 | Millipore | 42413 |
| Anti-Digoxigenin | Roche Diagnostics | 11-333-089-001 |
| Tween 20 | Sigma | P9416-100ML |
| Anti-myc Antibody | Invitrogen | 46-0603 |
| Bovine Serum Albumin | Sigma | B4287-5G |
| Dynabeads M-280 Streptavidin | Invitrogen | 658.01D |
| Dynabeads MyOne T1 Streptavidin | Invitrogen | 658.01D |
| p-Aminophenylmercuric Acetate | Calbiochem | 164610 |
| Biotin-Maleimide | Sigma Aldrich | B1267 |
| Biotin labeled oligo | IDT DNA | Custom synthesis |
| Digoxigenin labeled oligo | IDT DNA | Custom synthesis |
| Collagen peptide gene | DNA 2.0 | Custom synthesis |
| MMP-1 cDNA | Harvard Plasmid Database | |
| z-translator | Thorlabs | MTS50 |
| Servo controller for translator | Thorlabs | TDC001 |
References
- Ingber, D. E. Can cancer be reversed by engineering the tumor microenvironment. Semin. Cancer Biol. 18, 356-364 (2008).
- Hahn, C., Schwartz, M. A. Mechanotransduction in vascular physiology and atherogenesis. Nat. Rev. Mol. Cell Biol. 10, 53-62 (2009).
- Laurens, N., Koolwijk, P., de Maat, M. P. Fibrin structure and wound healing. J. Thromb. Haemost. 4, 932-939 (2006).
- Adhikari, A. S., Chai, J., Dunn, A. R. Mechanical Load Induces a 100-Fold Increase in the Rate of Collagen Proteolysis by MMP-1. J. Am. Chem. Soc. 133, (2011).
- Zareian, R. Probing collagen/enzyme mechanochemistry in native tissue with dynamic, enzyme-induced creep. Langmuir. 26, (2010).
- Ellsmere, J. C., Khanna, R. A., Lee, J. M. Mechanical loading of bovine pericardium accelerates enzymatic degradation. Biomaterials. 20, 1143-1150 (1999).
- Jesudason, R. Mechanical forces regulate elastase activity and binding site availability in lung elastin. Biophys. J. 99, 3076-3083 (2010).
- Neuman, K. C., Block, S. M. Optical trapping. Rev. Sci. Instrum. 75, 2787-2809 (2004).
- Lee, C. K., Wang, Y. M., Huang, L. S., Lin, S. Atomic force microscopy: determination of unbinding force, off rate and energy barrier for protein-ligand interaction. Micron. 38, 446-461 (2007).
- Smith, S. B., Finzi, L., Bustamante, C. Direct mechanical measurements of the elasticity of single DNA molecules by using magnetic beads. Science. 258, 1122-1126 (1992).
- Tanase, M., Biais, N., Sheetz, M. Magnetic tweezers in cell biology. Methods Cell Biol. 83, 473-493 (2007).
- Neuman, K. C., Nagy, A. Single-molecule force spectroscopy: optical tweezers, magnetic tweezers and atomic force microscopy. Nat. Methods. 5, 491-505 (2008).
- Frank, S. Stabilization of short collagen-like triple helices by protein engineering. J. Mol. Biol. 308, 1081-1089 (2001).
- Stetefeld, J. Collagen stabilization at atomic level: crystal structure of designed (GlyProPro)10foldon. Structure. 11, 339-346 (2003).
- Chung, L. Collagenase unwinds triple-helical collagen prior to peptide bond hydrolysis. EMBO J. 23, 3020-3030 (2004).
- Bell, G. I. Models for the specific adhesion of cells to cells. Science. 200, 618-627 (1978).
- Selvin, P. R., Ha, T. . Single-molecule techniques: a laboratory manual. , (2008).
- Graneli, A., Yeykal, C. C., Prasad, T. K., Greene, E. C. Organized arrays of individual DNA molecules tethered to supported lipid bilayers. Langmuir. 22, 292-299 (2006).
- Danilowicz, C., Greenfield, D., Prentiss, M. Dissociation of ligand-receptor complexes using magnetic tweezers. Anal. Chem. 77, 3023-3028 (2005).
- Zhang, X., Halvorsen, K., Zhang, C. Z., Wong, W. P., Springer, T. A. Mechanoenzymatic cleavage of the ultralarge vascular protein von Willebrand factor. Science. 324, 1330-1334 (2009).
- Woodside, M. T. Nanomechanical measurements of the sequence-dependent folding landscapes of single nucleic acid hairpins. Proc. Natl. Acad. Sci. U.S.A. 103, 6190-6195 (2006).
- del Rio, A. Stretching single talin rod molecules activates vinculin binding. Science. 323, 638-641 (2009).
- Gosse, C., Croquette, V. Magnetic tweezers: micromanipulation and force measurement at the molecular level. Biophys. J. 82, 3314-3329 (2002).

