qPCR Is a Sensitive and Rapid Method for Detection of Cytomegaloviral DNA in Formalin-fixed, Paraffin-embedded Biopsy Tissue
Summary
This protocol describes qPCR detection of cytomegalovirus in formalin-fixed, paraffin-embedded biopsy tissue, which is rapid, sensitive, specific, and useful for interpreting equivocal hematoxylin and eosin or immunohistochemical staining patterns.
Abstract
It is crucial to identify cytomegalovirus (CMV) infection in the gastrointestinal (GI) tract of immunosuppressed patients, given their greater risk for developing severe infection. Many laboratory methods for the detection of CMV infection have been developed, including serology, viral culture, and molecular methods. Often, these methods reflect systemic involvement with CMV and do not specifically identify local tissue involvement. Therefore, detection of CMV infection in the GI tract is frequently done by traditional histology of biopsy tissue. Hematoxylin and eosin (H&E) staining in conjunction with immunohistochemistry (IHC) have remained the mainstays of examining these biopsies. H&E and IHC sometimes result in atypical (equivocal) staining patterns, making interpretation difficult. It was shown that quantitative polymerase chain reaction (qPCR) for CMV can successfully be performed on formalin-fixed, paraffin-embedded (FFPE) biopsy tissue for very high sensitivity and specificity. The goal of this protocol is to demonstrate how to perform qPCR testing for the detection of CMV in FFPE biopsy tissue in a clinical laboratory setting. This method is likely to be of great benefit for patients in cases of equivocal staining for CMV in GI biopsies.
Introduction
Interpretation of cytomegalovirus (CMV) infection in clinical specimens is critically important. CMV is a member of the betaherpesvirinae subfamily of herpesviruses. Similar to other herpesviruses, CMV has the ability to establish persistent or latent infection characterized by a lack of active viral replication1. Reactivation of virus may occur during times of stress or other immunosuppression, leading to active viral replication characterized by virion release, which may be accompanied by disease manifestations2-4. Gastrointestinal (GI) CMV disease symptoms frequently include abdominal pain and/or bloody stools1.
The diagnosis of CMV gastroenteritis by histology utilizes hematoxylin and eosin (H&E) to identify CMV viral inclusions. In cases where clinical suspicion is high yet no obvious inclusions are observed, immunohistochemistry (IHC) is frequently used as an adjunct test method. However, IHC can also be hampered by rare non-classic appearing cellular staining patterns (equivocal), making interpretation difficult (Figure 2)5. It was sought to use DNA extracted from formalin-fixed, paraffin-embedded (FFPE) GI biopsy tissue and quantitative polymerase chain reaction (qPCR) to detect CMV in GI biopsies. This technique has been shown to be valuable in the detection of CMV in GI biopsies, and is particularly helpful in equivocal IHC staining cases5,6. Also, qPCR has also been shown to correlate well with additional clinical CMV test data when it is available6. Use of qPCR in detecting CMV may lead to earlier diagnosis and treatment in cases where clinical suspicion for CMV is high, but H&E and CMV IHC are negative.
Protocol
With the approval of the Institutional Review Board, a search of the electronic laboratory database was performed to identify formalin-fixed, paraffin embedded (FFPE) blocks representing cases of cytomegalovirus (CMV) infection in gastrointestinal (GI) biopsies diagnosed by histopathology (hematoxylin and eosin (H&E) and/or immunohistochemistry (IHC)).
1. Tissue Processing from FFPE GI Biopsy Tissue
- Collect FFPE GI biopsy tissue blocks for cytomegalovirus (CMV) quantitative polymerase chain reaction (qPCR). These blocks are stored at room temperature.
- Pre-label 1.7 ml microfuge tubes with the block’s unique identification number.
- Lock the microtome handwheel.
- Seat the tissue block into the cassette clamp.
- Using a fresh, unused blade, align the block to the knife angle on the microtome.
- Adjust the microtome to cut 10 μm thick sections.
- Unlock the handwheel and advance the block toward the blade.
- Cut 5 sections of the block at 10 μm thick intervals, allowing each cut to roll into a scroll of tissue. Ensure that each scroll contains an adequate representation of the desired biopsy tissues.
- Lock the microtome handwheel.
- With forceps, place all 5 scrolls of tissue into the pre-labeled centrifuge tube, being careful to only manipulate the scroll by the paraffin in order to reduce potential for cross contamination. Snap the cap on the tube and place it back on the rack.
- Remove the tissue block from the microtome.
- Remove and discard the blade.
- Decontaminate any surfaces or instruments that may have come in contact with the previous tissue.
- Place a fresh, unused blade into the knife holder.
- Repeat steps 1.4-1.14 for each additional block to be processed. The microfuge tubes containing the tissue scrolls may be stored at room temperature until the time of DNA extraction.
2. DNA Extraction from Scrolls of FFPE GI Biopsy Tissue
NOTE: This protocol is adapted from the package insert of the FFPE tissue DNA extraction kit(CAUTION: refer to Material Safety Data Sheet (MSDS)).
- Add 1 ml xylene (CAUTION: Refer to MSDS) to each sample microfuge tube. Close the lid and vortex vigorously for 10 sec.
- Proceed with the DNA extraction as specifically outlined in the manufacturer’s product insert with the following modifications.
- First add 6 μl of the internal control (IC) to each microfuge tube.
- Following the ATL/proteinase K lysis incubation at 56 °C for 1 hr, incubate at 99 °C for 30 min, rather than 90 °C for 1 hr.
- During the final DNA elution step, carefully open the lid of the DNA purification column and apply 50 μl distilled DNAse/RNAse-free water at the center of the membrane, rather than buffer ATE.
NOTE: Refer to the package insert for the appropriate storage and preparation of reagents. DNA should be extracted from all 5 FFPE scrolls. Perform all centrifugation steps at room temperature (15-25 °C).
3. qPCR for CMV
- Polymerase Chain Reaction (PCR) Procedure
- Generate an Excel PCR worksheet for the CMV assay run. This Excel worksheet calculates the total amount of each reagent needed to make up the master mix with Mg. It multiplies the amount of each reagent needed per sample (listed below) times the number of samples and controls plus one (to compensate for loss due to pipetting).
- Include the following reagents in each PCR reaction in the amount indicated:
Reagent Amount per Sample
CMV LC master mix vial 12.5 μl
Mg Solution yellow vial 2.5 μl
Total volume 15.0 μl
- Include the following reagents in each PCR reaction in the amount indicated:
- Include in every assay, run a water or no target control (NTC), negative control, a low positive control, a high positive control, and either QS3 or QS4, and all patient samples. The capillaries should be set up in this order.
- Obtain the cooling block specific for the real-time PCR instrument that is stored at 4 °C. Place the appropriate capillary tubes in the cooling block. Do not touch the surface of the capillary tubes. Always use gloves when handling the capillaries.
- Set up all polymerase chain reactions in a clean hood not contaminated with amplification products.
- Remove the correct number of master vials from the PCR kit to test all patient samples, plus 5 controls. Let the master vials, Mg solution, and PCR grade water thaw in the cooling block. Each master vial contains enough reagents for 12 tests.
- Gently vortex the master vials and quick spin them in the centrifuge at the maximum speed.
- Combine the contents of each master vial into one vial, and gently vortex again.
- Combine the master mix and Mg solution in the amounts indicated on the PCR worksheet into a single sterile microfuge tube.
- Mix gently and aliquot 15 μl of master mix with Mg into each capillary tube, except the capillary in position 5 which is for the quantitation standard (QS).
- Move the cooling block containing the capillary tubes and master mix with Mg from the clean hood to a processing hood.
- Add 2 μl of internal control (IC) to the remaining 30 μl of master mix with Mg. Mix gently and aliquot 15 μl into the capillary in position 5 (position for the QS).
- Pipette 10 μl of water, control DNA, or sample DNA into the appropriate capillary tube. Cap the capillary tubes using the capping tool.
- Take cooling block/samples to the real-time PCR instrument.
- Generate an Excel PCR worksheet for the CMV assay run. This Excel worksheet calculates the total amount of each reagent needed to make up the master mix with Mg. It multiplies the amount of each reagent needed per sample (listed below) times the number of samples and controls plus one (to compensate for loss due to pipetting).
- Program the real-time PCR Instrument (see Table 1)
- Amplification and Detection performed on the real-time PCR instrument
- Log onto the computer
- Double click on the instrument icon and log into the software.
- Turn on the real-time PCR instrument
- Click on “CMV/EBV” from the front screen.
- Follow the prompts of the macro wizard.
- Select the box to run a self-test when prompted by the software. A self-test must be performed once each day the instrument is in use.
- Name the run.
- Adjust the sample count located in the upper left corner of the page to the sum of controls and patients included in the run and enter the identification number of each patient sample.
- Click on the “Abs Quant” tab.
- Under “Sample type”, assign the appropriate sample type for the standard.
- Type in the expected concentrations for the quantitation standard (QS).
- Remove the capillary tubes from the cooler block and place them into the corresponding position in the carousel (cooling block sample #1 should be placed in position #1 on the carousel, etc.).
- Place the carousel into the instrument’s centrifuge and press start.
- Open centrifuge and remove the carousel.
- Place the carousel into the real-time PCR instrument. Close the lid.
- Press “Start Run”.
- The real-time PCR instrument will check each position for a sample. Do not walk away from the instrument until this check has been completed, especially if one of the capillaries is not in the carousel. The instrument will note this and ask if the run is to be continued. Answer yes before leaving.
- Once the run is complete, close the report that was generated.
- Click on “Absolute Quantitation”.
- Click on the arrow next to Color Compensation.
- Click “Select Color Compensation” from the drop-down box.
- Choose the most current color compensation file to apply to the run.
- Click OK when the box pops up.
- Click on the arrow by Standard Curve.
- Select “use external” from the drop-down menu.
- Select the most current external standard curve file, then click OK.
- Look at each curve under the Absolute Quantification menu to ensure that curves are present and not flat line, if a concentration is assigned. The Absolute Quantification for the 705/Back 530 is automatically analyzed.
- Quality control
- Generate a new standard curve whenever a new lot number of reagents is received, or every six months, whichever is more frequent.
- Add 10 μl of each standard to the master mix with Mg. Consider the standards included with the kit as previously extracted DNA.
- Add 1 μl of internal control for each standard to the master mix with Mg used in generating the standard curve in order to assure accurate quantitation. Do not add IC to the master mix with Mg for the No Template Control (NTC).
- Prepare a NTC and three replicates of Quantitation Standards QS 4, 3, 2, 1, and a 1:10 dilution of QS4 for the run to generate the standard curve, totaling in 16 data points.
- Select “replicate of” in the drop down menu for repeated standards
- Choose channel 530/back none and turn on color compensation when analyzing the run,
- Under “Standard Curve (In Run)”, select “save as external” and name the curve. Enter the comment “applied as standard curve” when saving.
- Graph the results. The resulting linearity coefficient must be ≥0.81.
- For daily runs, include one copy of QS3 or QS4 as a control in the run. The current external standard curve may be imported into the run during the analysis phase.
- Run Controls: no DNA control, negative control, low, and high positive controls. Expected viral load is determined and adjusted for each vendor and lot number.
- Add an IC to each sample and control during the DNA extraction.
- If any of the controls fall out of range, record the unacceptable results and refer the assay for troubleshooting.
- Frequency of controls
- Include in every run: a NTC, negative, low positive, high positive, and standard control.
NOTE: Acceptable limits for controls. No peaks should be observed for the no template (NTC) and negative controls. Peaks should be present in the 530 channel and expected viral loads are determined and adjusted for each vendor and lot number for the low and high positive controls.
Representative Results
A total of 228 tissue blocks were tested by quantitative polymerase chain reaction (qPCR), which was comprised of 91 cytomegalovirus (CMV) positive cases based on histology and positive immunohistochemistry (IHC), 18 with equivocal CMV IHC, and 79 negative controls. As illustrated in Figure 2, CMV positive cases would have demonstrated typical CMV viral inclusions (A) and/or positive IHC staining (B). Equivocal cases would have demonstrated rare, non-classic appearing staining patterns (C), whereas negative controls would have shown no suspicious histology or IHC staining5.
Of 91 biopsies positive for CMV by histopathology, 88 were positive by qPCR (Table 2). With a true positive definition based on traditional histology, i.e. typical CMV viral inclusions and/or typical CMV IHC, these data resulted in a sensitivity of 96.7%.
Of 79 negative control biopsies (35 normal colon, 25 colitis, and 19 duodenitis), i.e. negative for viral inclusions and negative by CMV IHC, 78 tested negative and 1 tested positive by qPCR, resulting in a specificity of 98.7%.
Fourteen of 18 (78%) biopsies with equivocal CMV IHC staining tested positive for CMV by qPCR5. Among these 18 equivocal cases, eleven had additional biopsies taken the same day that were positive for CMV by histopathology. Among these 14 qPCR positive samples, 10 had biopsies taken the same day that were positive for CMV on histopathology. Of the 18 equivocal biopsies tested, 11 had additional clinical information regarding CMV testing on other specimens submitted at or within 7 days of the time the biopsy was taken. Among these 11 cases, 8 (73%) tested positive for CMV by qPCR. Of these 11 biopsies, 5 (45%) tested positive by qPCR with additional positive CMV specimens; 3 (27%) tested positive by qPCR with no positive CMV specimens; 3 (27%) tested negative by qPCR with additional negative CMV specimens.
In addition to those studies mentioned above, it was of interest to determine whether obtaining additional hematoxylin and eosin (H&E) levels would be beneficial in cases where no viral inclusions are seen on H&E initially and CMV IHC is equivocal. Therefore, H&E levels times 5 for each of the 18 blocks with equivocal CMV IHC were performed. Two pathologists carefully examined these cases and could not identify discernible viral inclusions on any of the additional slides (data not shown).
| Suggested real-time polymerase chain reaction parameters | ||||||
| Setup Readings: | ||||||
| Default Channel | Seek Temperature | Max Seek Pos | Instrument type | Capillary Size | ||
| 530 | 30 °C | 12 | 6 channel | 20 μl | ||
| Program name: Activation of enzyme | ||||||
| Number of cycles: 1 | ||||||
| Analysis mode: None | ||||||
| Target °C | Hold hh:mm:ss | Ramp Rate | Sec Target | Step Size | Step Delay | Acquisition Mode |
| 95 | 0:10:00 | 20 | 0 | 0 | 0 | None |
| Program name: Touch down step | ||||||
| Number of cycles: 10 | ||||||
| Analysis mode: None | ||||||
| Target °C | Hold hh:mm:ss | Ramp Rate | Sec Target | Step Size | Step Delay | Acquisition Mode |
| 95 | 0:00:05 | 20 | 0 | 0 | 0 | None |
| 65 | 0:00:20 | 20 | 55 | 1 | 1 | None |
| 72 | 0:00:15 | 20 | 0 | 0 | 0 | None |
| Program name: Amplification of the DNA | ||||||
| Number of cycles: 40 | ||||||
| Analysis mode: Quantitation | ||||||
| Target °C | Hold hh:mm:ss | Ramp Rate | Sec Target | Step Size | Step Delay | Acquisition Mode |
| 95 | 0:00:05 | 20 | 0 | 0 | 0 | None |
| 55 | 0:00:20 | 20 | 0 | 0 | 0 | Single |
| 72 | 0:00:15 | 10 | 0 | 0 | 0 | None |
| Program name: Cooling | ||||||
| Number of cycles: 1 | ||||||
| Analysis mode: None | ||||||
| Target °C | Hold hh:mm:ss | Ramp Rate | Sec Target | Step Size | Step Delay | Acquisition Mode |
| 40 | 0:00:30 | 20 | 0 | 0 | 0 | None |
Table 1. The parameters listed should be used as a guideline for target temperatures, cycle numbers, and cycle lengths in the set-up of real-time polymerase chain reaction for detecting cytomegalovirus in formalin-fixed, paraffin-embedded gastrointestinal biopsies. These parameters may require optimization depending on the instrument model and manufacturer.
| Histology Positive Cases | PCR Positive Cases | Sensitivity | Histology Negative Cases | PCR Negative Cases | Specitivity |
| 91 | 88 | 96.7% | 79 | 78 | 98.7% |
Table 2. Sensitivity (96.7%) and specificity (98.7%) of real-time polymerase chain reaction (qPCR) at detecting cytomegalovirus in formalin-fixed, paraffin-embedded gastrointestinal biopsies. This table has been modified from McCoy et al5.
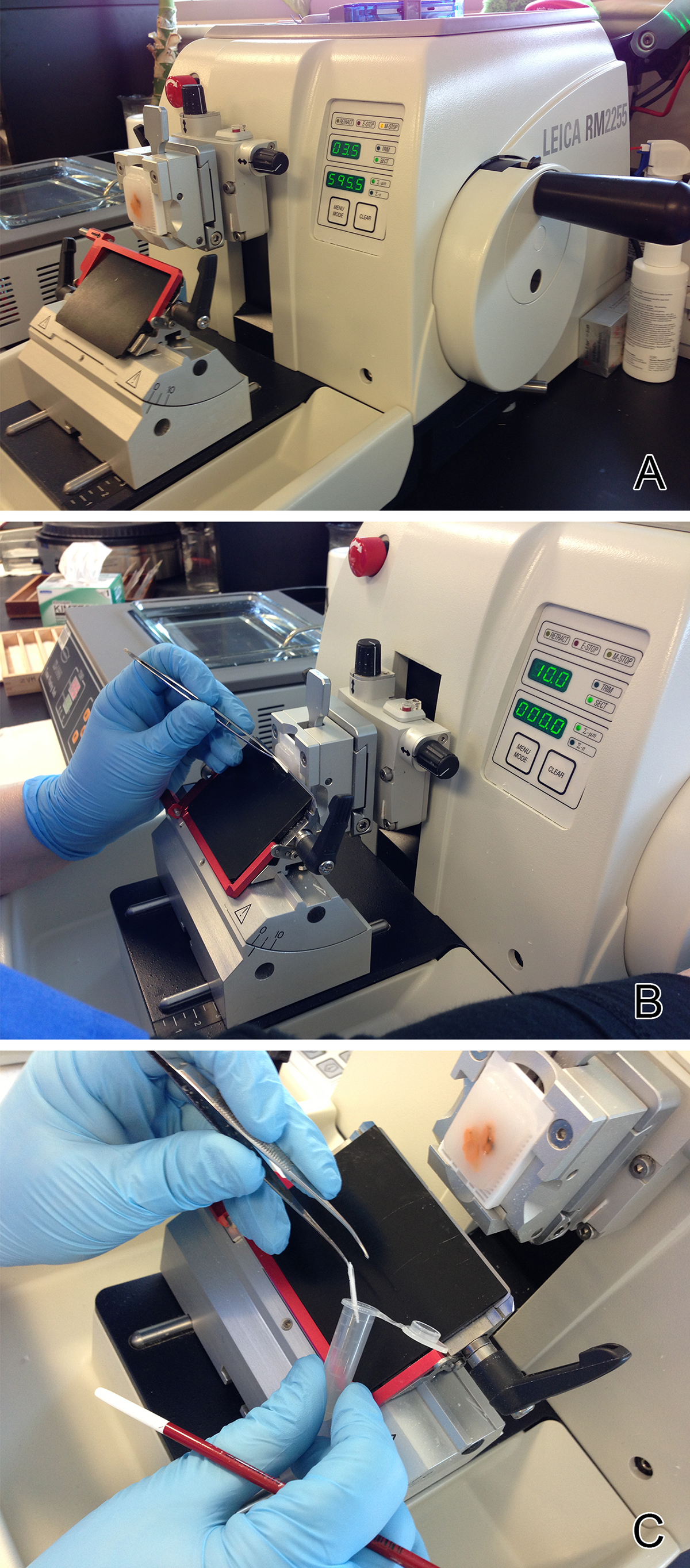

Figure 1. A typical microtome with the tissue block seated correctly in the cassette clamp A) As a 10 micron thick section of biopsy tissue is cut it is allowed to roll into a scroll B), which is transferred to a microfuge tube C). All five scrolls of biopsy tissue are placed in a single microfuge tube for DNA extraction.
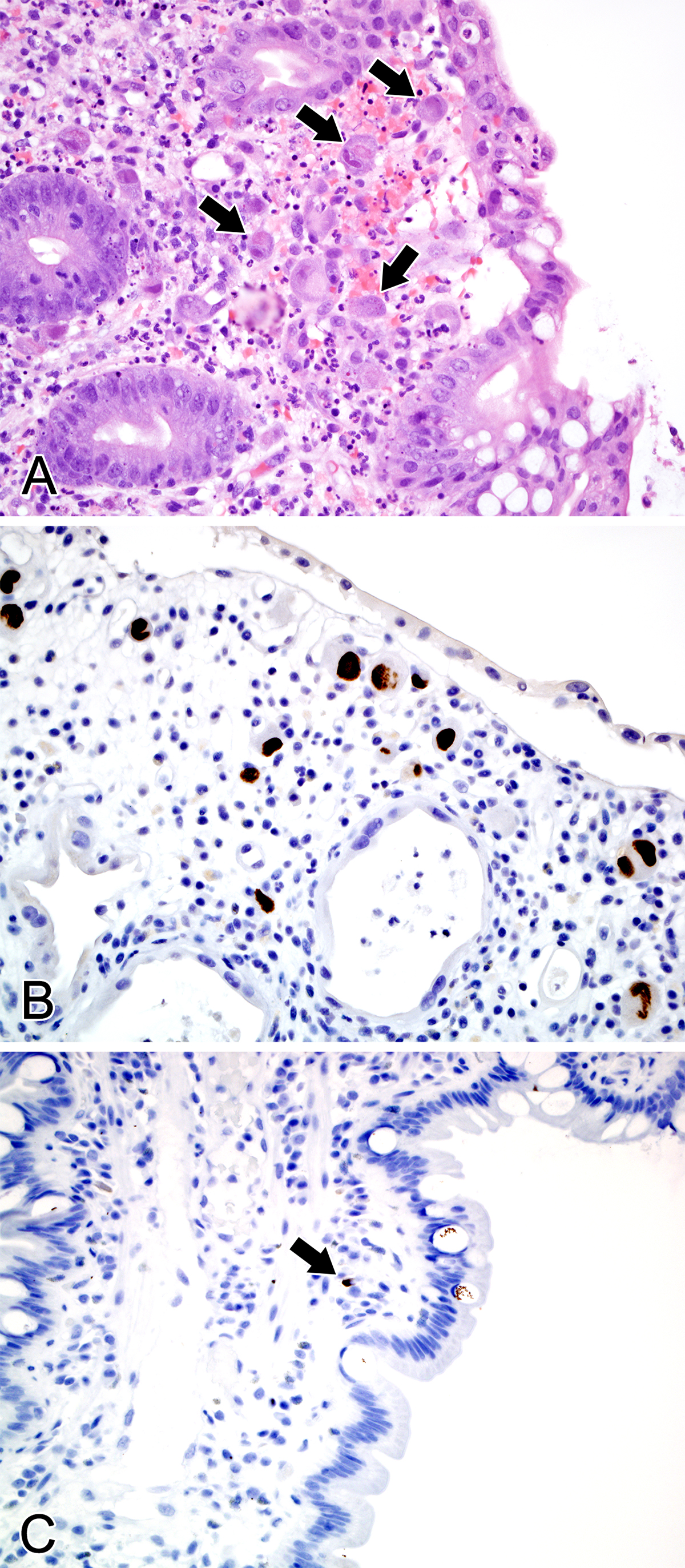
Figure 2. A) Hematoxylin and eosin stained section of a colon biopsy showing abundant, typical CMV inclusions as indicated by arrows (original magnification 400X). B) A colon biopsy stained with a monoclonal antibody against CMV showing the expected IHC staining pattern (original magnification 400X). C) A colon biopsy showing equivocal IHC staining, with a single, small weakly stained cell indicated by the arrow (original magnification 400X). This figure is being reprinted with permission from McCoy et al5.
Discussion
Many laboratory methods are available for the diagnosis of cytomegalovirus (CMV) infection, including serology, viral culture, molecular viremia assays, histology of biopsy material, and, as recently demonstrated, molecular assays of formalin-fixed, paraffin-embedded (FFPE) biopsy tissue5,6. Serology, once a mainstay of diagnosis, only reliably identifies exposure and does not correlate well with acute CMV infection7. Viral culture, both conventional and early antigen shell vial, is hampered by lengthy assay times, at best 2-3 days in the case of the latter8. Molecular viremia assays have been developed, most having the advantage of being quantitative with the potential for monitoring treatment efficacy9-11. Unfortunately, when there is clinical concern for local involvement of the gastrointestinal (GI) tract, viremia assays reflect systemic involvement and do not specifically identify localized infection. Therefore, histology has been the gold standard for identifying CMV infection of the GI tract.
It is crucial to identify active GI tract CMV infections in immunosuppressed patients, such as those with HIV/AIDS, on chemotherapy, or organ/stem cell transplants, given their greater risk for developing severe infection. It was reported quantitative polymerase chain reaction (qPCR) is a highly sensitive and specific method to detect CMV in GI biopsies5. Similar results have been supported by other studies6.
Although all steps throughout the procedure are important, there are a few key critical steps that deserve special consideration. During tissue processing on the microtome, it is imperative the blade be switched between each sample to reduce the possibility of cross-contamination between specimens. Similarly, decontamination of areas on the microtome the tissue may have come in contact with using an appropriate cleaner recommended by the microtome manufacturer will also reduce this risk.
An additional key step during the protocol is the addition of the internal control IC (step 2.3) to the sample during DNA extraction, which acts as a control for both the extraction process and amplification. The amplification and detection of the IC does not interfere with the amplification of the CMV, because detection of the CMV virus occurs in channel 530, whereas the detection of the IC occurs in channel 705. Each laboratory must establish an acceptable range for the IC. After extraction, the DNA quality does not need to be assessed. In other words, spectrophotometric determination of DNA concentration is not routinely performed following DNA isolation. If added prior to DNA extraction, the IC acts as a control for the extraction process. Alternatively, 1 μl of IC may be added to each master mix with Mg vial. If added to the master mix with Mg, the IC acts only as an amplification control.
It is also imperative that during the set-up of all qPCR reactions that a clean hood (including pipettes and tips) be used for reagent set up. This hood should not have any amplification products, i.e. post real-time or conventional PCR, manipulated within it. These amplification products could become aerosolized and contaminate subsequently prepared qPCR reactions leading to false positive results. Many laboratories even go as far as designating hoods and equipment for pipetting non-nucleic acid reagents and a second hood and equipment for adding sample and control nucleic acid. Individual laboratories must decide what works best given their individual space constraints.
Troubleshooting unexpected results may be necessary during the procedure. If there is no evidence of amplification of the IC (i.e. flat line) of a sample, it is recommended to first repeat the qPCR reaction from the original DNA extraction to ensure a pipetting error did not occur during sample reaction preparation. If the IC remains at a flat line on the repeat reaction and there is sufficient biopsy tissue remaining, a second DNA extraction may performed, being mindful to add the IC during the extraction process. If this re-extracted specimen also demonstrates a flat line IC, it is likely there is a reaction inhibitor present within the original specimen. In this case, the test result would be indeterminate for the detection of CMV since the qPCR reaction control did not perform appropriately.
If the water, no target control (NTC) demonstrates amplification of the target, it is first advised to analyze the positivity rate of the run. If the run has a higher than normal positivity rate, it would be recommended to perform swipe tests of areas where any reagent manipulation is performed to identify any potential environmental contamination. If environmental contamination is ruled out, use a newly opened water control along with the usual positive controls. If the newly opened water control does not show an amplification product, it is likely the previous water control was contaminated with target sequence. However, if this new water control is again contaminated, it raises the possibility of any or all of the reagents or hoods being contaminated with target sequence. In this case, it would be best to decontaminate all hoods used during the process and then use all new reagents in running a new set of controls (positive and water). If the problem persists, a reevaluation of the workflow in the processing areas and hoods, as well as contacting the product manufacturers should be done.
If the quality standards (QS) included in the run do not demonstrate the appropriate copy number during amplification, it is advised to first run all the QS (4, 3, 2, 1, and a 1:10 dilution of QS4). If these standards fall within the appropriate ranges, then repeat the samples as previously specified with the appropriate controls. If the QS do not fall within the appropriate ranges, then generate a new standard curve as described in the quality control section of the protocol (3.4). If the problem persists, generating a new standard curve with newly opened QS of the current or a new lot number may be attempted. If these steps are unsuccessful, contacting the manufacturer would be warranted.
Although the sensitivity and specificity of qPCR for detecting CMV in GI biopsies based on calculations using conventional methods (H&E and IHC) as the gold standard do not immediately demonstrate a diagnostic advantage, using the more analytically sensitive qPCR method as the gold standard greatly improves these calculations5. Additionally, 14 (78%) of the 18 equivocal biopsies tested positive for CMV by qPCR in this study. It is felt these points support the notion that qPCR is much more sensitive than either H&E alone, or in conjunction with IHC at detecting CMV in GI biopsy material. Additionally, in cases with equivocal IHC staining patterns, obtaining additional levels is unlikely to be of any added benefit compared to current standard methods.
Taken together, qPCR is highly sensitive and specific to detect CMV infection in FFPE GI biopsy tissue. Particularly useful is qPCR’s performance on GI biopsies with equivocal CMV IHC staining patterns or clinically highly suspicious for CMV. Compared to traditional histology, qPCR testing of GI biopsy material may facilitate an accurate diagnosis in a short time period, thus leading to earlier and more effective patient management.
Disclosures
The authors have nothing to disclose.
Acknowledgements
We acknowledge Amy Thomasson for her assistance with preparing this manuscript, and Fredrik Skarstedt and Ryan Christy for their efforts in the preparation of figures.
Materials
| 1.7 mL microfuge tubes | Costar | 3620 | |
| serrated forceps | Electron Microscopy Services | 62086-1S | Micro Forceps MF-1 |
| Tabletop centrifuges (microfuge) | Eppendorf | 5424 | |
| Xylene | Fisher | Fisher Scientific: X3P | 1 gallon |
| Ethanol | Fisher | Fisher Scientific: ET108 | 96-100% |
| microtome | Leica | RM2255 | |
| QIAamp DNA FFPE Tissue Kit | Qiagen | 56404 | |
| artus CMV LightCycler PCR ASR | Qiagen | 4500025 | |
| CMV LC PCR Supplement Kit | Qiagen | 1031873 | |
| LightCycler centrifuge | Roche | 75005087 | |
| LightCycler Cooling Block | Roche | 10800058001 | |
| LightCycler Capping Tool | Roche | 3357317 | |
| Color Compensation Kit | Roche | 2158850 | |
| LightCycler 20 μL Capillaries | Roche | 1 909 339 | |
| blades | Sakura | Fisher Scientific: 4689 | Accu-Edge low profile |
References
- Hodinka, R. L. . Manual of Clnical Microbiology. 2, 1558-1574 (2011).
- Lasry, S., et al. Interstrain variations in the cytomegalovirus (CMV) glycoprotein B gene sequence among CMV-infected children attending six day care centers. The Journal of infectious diseases. 174, 606-609 (1996).
- Murph, J. R., Baron, J. C., Brown, C. K., Ebelhack, C. L., Bale Jr, J. F. The occupational risk of cytomegalovirus infection among day-care providers. JAMA : the journal of the American Medical Association. 265, 603-608 (1991).
- Pass, R. F., et al. Vaccine prevention of maternal cytomegalovirus infection. The New England journal of medicine. 360, 1191-1199 (2009).
- McCoy, M. H., et al. qPCR increases sensitivity to detect cytomegalovirus in paraffin-embedded, formalin-fixed tissue of gastrointestinal biopsies. Human Pathology. 45, 48-53 (2014).
- Mills, A. M., Guo, F. P., Copland, A. P., Pai, R. K., Pinsky, B. A. A comparison of CMV detection in gastrointestinal mucosal biopsies using immunohistochemistry and PCR performed on formalin-fixed, paraffin-embedded tissue. The American journal of surgical pathology. 37, 995-1000 (2013).
- Krech, U. Complement-fixing antibodies against cytomegalovirus in different parts of the world. Bulletin of the World Health Organization. 49, 103-106 (1973).
- Chou, S. Newer methods for diagnosis of cytomegalovirus infection. Reviews of infectious diseases. 12 Suppl 7, (1990).
- Brytting, M., Xu, W., Wahren, B., Sundqvist, V. A. Cytomegalovirus DNA detection in sera from patients with active cytomegalovirus infections. Journal of clinical microbiology. 30, 1937-1941 (1992).
- Revello, M. G., et al. Human cytomegalovirus in blood of immunocompetent persons during primary infection: prognostic implications for pregnancy. The Journal of infectious diseases. 177, 1170-1175 (1998).
- Shinkai, M., Bozzette, S. A., Powderly, W., Frame, P., Spector, S. A. Utility of urine and leukocyte cultures and plasma DNA polymerase chain reaction for identification of AIDS patients at risk for developing human cytomegalovirus disease. The Journal of infectious diseases. 175, 302-308 (1997).

