Fabrication of a Crystalline Nanocellulose Embedded Agarose Biomaterial Ink for Bone Marrow-Derived Mast Cell Culture
Summary
This protocol highlights a method to rapidly assess the biocompatibility of a crystalline nanocellulose (CNC)/agarose composite hydrogel biomaterial ink with mouse bone marrow-derived mast cells in terms of cell viability and phenotypic expression of the cell surface receptors, Kit (CD117) and high-affinity IgE receptor (FcεRI).
Abstract
Three-dimensional (3D) bioprinting utilizes hydrogel-based composites (or biomaterial inks) that are deposited in a pattern, forming a substrate onto which cells are deposited. Because many biomaterial inks can be potentially cytotoxic to primary cells, it is necessary to determine the biocompatibility of these hydrogel composites prior to their utilization in costly 3D tissue engineering processes. Some 3D culture methods, including bioprinting, require that cells be embedded into a 3D matrix, making it difficult to extract and analyze the cells for changes in viability and biomarker expression without eliciting mechanical damage. This protocol describes as proof of concept, a method to assess the biocompatibility of a crystalline nanocellulose (CNC) embedded agarose composite, fabricated into a 24-well culture system, with mouse bone marrow-derived mast cells (BMMCs) using flow cytometric assays for cell viability and biomarker expression.
After 18 h of exposure to the CNC/agarose/D-mannitol matrix, BMMC viability was unaltered as measured by propidium iodide (PI) permeability. However, BMMCs cultured on the CNC/agarose/D-mannitol substrate appeared to slightly increase their expression of the high-affinity IgE receptor (FcεRI) and the stem cell factor receptor (Kit; CD117), although this does not appear to be dependent on the amount of CNC in the bioink composite. The viability of BMMCs was also assessed following a time course exposure to hydrogel scaffolds that were fabricated from a commercial biomaterial ink composed of fibrillar nanocellulose (FNC) and sodium alginate using a 3D extrusion bioprinter. Over a period of 6-48 h, the FNC/alginate substrates did not adversely affect the viability of the BMMCs as determined by flow cytometry and microtiter assays (XTT and lactate dehydrogenase). This protocol describes an efficient method to rapidly screen the biochemical compatibility of candidate biomaterial inks for their utility as 3D scaffolds for post-print seeding with mast cells.
Introduction
The recent interest in 3D culture systems and 3D bioprinting has focused attention on hydrogels and hydrogel composites. These composites serve as viscous yet porous biomimetics and can be composed of up to 99% water content by weight, which is comparable to biological tissues1,2,3. These features of hydrogel composites thereby permit the growth of cells without affecting their viability and function. One such composite is crystalline nanocellulose (CNC), which has been used as a reinforcing material in hydrogel composites, cell scaffolds in the development of biomaterial implants, and in two-dimensional (2D) and 3D in vitro cell culture4,5. For the most part, matrices composed of CNC are not overtly cytotoxic to human corneal epithelial cells6, intestinal epithelial cells7, human bone marrow-derived mesenchymal stem cells8, or neuron-like cells9. However, metabolic activity and proliferation of human bone marrow-derived mesenchymal stem cells decreases in correlation with the increased viscosity of wood-based nanocellulose composites, suggesting that the composition of the matrix must be carefully tested for its deleterious effects on cell functions8.
Similarly, CNC can induce inflammatory responses in macrophages upon internalization, which could have serious consequences in 3D immune cell culture systems10,11. In fact, there is very little data available on how CNC may influence other immune cell responses, particularly allergic inflammatory responses that are initiated by mast cells. Mast cells are granulated leukocytes that express the high-affinity IgE receptor, FceRI, responsible for activating inflammatory responses to allergens. Their proliferation and differentiation are dependent-upon stem cell factor (SCF), which binds the tyrosine receptor, Kit. Mast cells are derived from bone marrow progenitor cells that enter the circulation and subsequently migrate peripherally to disperse ubiquitously in all human tissues12. As mast cells function in a 3D tissue environment, they are an ideal immune cell candidate for studying immunological processes in in vitro 3D tissue models. However, to date, there is no viable in vitro 3D tissue model containing mast cells.
Due to the highly sensitive nature of mast cells and their propensity to elicit pro-inflammatory responses to external stimuli, careful consideration of the 3D matrix constituents and the bioprinting method of introducing mast cells into the 3D scaffold is required, as discussed further. Tissue constructs can be biofabricated from two broad categories of biomaterials, i.e., bioinks and biomaterial inks. The distinction lies in the fact that bioinks are cell-laden hydrogel composites, whereas biomaterial inks are hydrogel composites that are devoid of cells, as defined by Groll et al.13,14. Hence, 3D constructs printed with bioinks contain cells pre-embedded within the hydrogel matrix, whereas 3D constructs printed with biomaterial inks need to be seeded with cells post-printing. The biofabrication of culture scaffolds from hydrogel-based bioinks/biomaterial inks is most commonly performed using extrusion 3D bioprinters, which extrude the bioink/biomaterial ink through a microscale nozzle under pressure via either a pneumatically or mechanically driven piston14. Extrusion bioprinters fabricate 3D scaffolds by depositing the bioink in 2D cross-sectional patterns that are sequentially stacked upon each other in a 'bottom-up' approach.
To be compatible with extrusion bioprinting, the hydrogel-based bioink/biomaterial ink must possess thixotropic (shear-thinning) properties, whereby the constituent hydrogel polymers of the bioink/biomaterial ink flow like a fluid through a microchannel nozzle when subjected to shear stress, but revert to a viscous, gel-like state upon removal of the shear stress15. Due to their high water content, the polymers of hydrogel-based bioinks/biomaterial inks must be crosslinked, either physically or covalently, to maintain the architecture and structural integrity of the 3D bioprinted structure. In the case of cell-laden bioinks, the cells are directly subjected to chemical stresses during the crosslinking process. The process of extruding cells encapsulated within the bioink hydrogel matrix also subjects the cells to shear stress, which can lead to reduced viability and/or cell death. Once the 3D tissue model has been bioprinted, it is difficult to discriminate between the levels of cytotoxicity elicited by the hydrogel matrix itself and the extrusion and crosslinking processes, respectively. This is particularly challenging in the context of 3D scaffolds where the cells are pre-embedded within the hydrogel matrix, thus making it difficult to remove the cells for subsequent analyses, which would be detrimental to the viability of mast cells.
A gentler approach to generating 3D tissue constructs containing mast cells involves seeding the cells into pre-printed, porous biomaterial ink 3D scaffolds from a cell culture suspension, which leverages the innate ability of mast cells to migrate from the circulation into peripheral tissues. The benefits of this cell seeding approach are two-fold: (i) the mast cells are not subjected to shear and chemical stresses from the extrusion and crosslinking processes, respectively, and (ii) the cells can be easily removed from the 3D scaffold after exposure by gentle washing for analysis without adversely affecting their viability. The additional benefit of seeding and analyzing the cell viability of mast cells on 3D bioprinted, porous hydrogel scaffolds as opposed to 2D hydrogel discs is that the 3D bioprinted hydrogel scaffolds recapitulate microscale topographical features of in vivo tissues, which are not present in bulk, 2D planar hydrogel discs. This approach is a suitable, rapid, and cost-effective approach to determine the potentially catastrophic cytotoxic effects of candidate bioink hydrogel matrices on mast cells, as well as other immunological cells, prior to investment in costly 3D tissue engineering experiments.
Protocol
NOTE: This protocol is composed of five sections: (1) isolation of mouse bone marrow and differentiation of mouse bone marrow-derived mast cells (BMMCs), (2) fabrication of CNC/agarose/D-mannitol hydrogel substrates in a 24-well system and culture of BMMCs on the substrates, (3) removal of BMMCs from the CNC/agarose/D-mannitol hydrogel substrates and analysis of viability and biomarker expression using flow cytometry, (4) 3D bioprinting of hydrogel scaffolds from a commercially available fibrillar nanocellulose (FNC)/sodium alginate composite biomaterial ink, and (5) culture of BMMCs on FNC/sodium alginate hydrogel scaffolds and analysis of viability using flow cytometry, XTT, and lactate dehydrogenase (LDH) microtiter assays.
1. Generation of the BMMC culture
NOTE: Mice were euthanized by CO2 asphyxiation following isoflurane anesthesia. The tibia and femur were isolated, and whole bone marrow was harvested. All animal studies were conducted in accordance with the Canadian Council on Animal Care Guidelines and Policies with approval from the Health Science Animal Care and Use Committee for the University of Alberta.
- Prepare complete RPMI-1640 culture medium by adding the supplements listed in Table 1 to the indicated final concentrations. Adjust the pH of the medium to 7.4-7.6 using NaOH, filter-sterilize using a single-use, bottle-top filter (0.2 µm pore size), and store at 4 °C in the dark for up to 2 months.
- Obtain femurs from mice in accordance with the local University Animal Care Committee protocols and procedures.
NOTE: In this study, femurs were isolated from C57BL/6 mice, but any strain can be used if it is relevant to the study. - Using scissors, remove skin and muscle from the hip joint, and then gently pull the femur by twisting slightly out of the hip socket (Figure 1). Be careful not to break off the femoral head. Cut the tibia away from the femur at the medial fabella using scissors. Remove the paw by cutting just above the calcaneum and discard.
- Using a scalpel, remove skin and cartilage from femur and tibia pieces. Using scissors, make a clean cut on each end of the femur and tibia, exposing the red bone marrow within. If required, remove the fibula at this point, but note that it may aid in the manipulation of the tibia during bone marrow flushing.
- Prepare a 26 G needle with a 5 mL luer-lock syringe, and fill with approximately 5 mL of complete media prepared as described above.
NOTE: At this point, incomplete RPMI without the additives can also be used to save on reagents. - Holding the femur or tibia with forceps, insert the needle into one end of the bone and gently press the syringe. Hold the bone over a 50 mL sterile conical tube, and gently inject approximately 5 mL of medium into the bone, applying some pressure. Observe pieces of bone marrow fall out the other end of the bone as thin red ribbons and into the conical tube.
- Spin down the aspirates and resuspend in complete medium, or transfer directly into a T175 cm2 sterile tissue culture flask containing 50 mL of complete RPMI-1640 medium. Maintain the aspirates in complete RPMI-1640 medium with 30 ng/mL mouse recombinant interleukin (IL)-3.
- After 1 day, observe the cultures under a light microscope. The culture will be composed of a heterogeneous population of cells of varying shapes and sizes and even larger pieces of bone marrow. Feed cell cultures by adding 15 mL of fresh medium every 3-5 days using the complete RPMI-1640 medium as described above, and ensure the cell density never exceeds 1 × 106 cells/mL. Once per week, spin down cells at 200 × g for 5 min at room temperature, resuspend in fresh complete RPMI-1640 medium, and split 1:4 into 50 mL of medium in fresh T175 flasks.
- After 4 weeks, determine cell purity by measuring the surface expression of CD117 (Kit) and FceRI receptors by flow cytometry as described below (section 3.2).
NOTE: After 4 weeks, 99% of cells should be double-positive for CD117 (Kit) and FcεRI. - At 4 weeks and onward, maintain the BMMCs with 20 ng/mL of IL-3 in complete RPMI-1640 medium. At approximately 6 weeks, the cells will stop dividing and will no longer require splitting. Feed the cultures every 3–5 days, pellet the cells by centrifugation once per week, and resuspend in fresh complete RPMI-1640 medium.
2. Fabrication of the CNC/agarose/D-mannitol hydrogel substrates and BMMC culture
- In a glass bottle, add 0.2 g of agarose powder to phosphate-buffered saline (PBS) (2% w/v) to a final volume of 10 mL, and boil for 1 min at 100 °C to dissolve.
- Dissolve 2.5 g of CNC powder in PBS to a final volume of 10 mL to prepare a 25% w/v mixture.
- Dissolve 2 g of CNC powder in PBS to a final volume of 10 mL to prepare a 20% w/v mixture, and serially dilute the 20% w/v mixture to prepare 10, 5, and 2% w/v CNC mixtures.
- Heat the 25, 20, 10, 5 and 2% w/v CNC mixtures to 37 °C for 10 min (Figure 2A).
- Add 0.46 g of D-mannitol to the hot agarose solution (4.6% w/v) and dissolve.
- Mix the pre-warmed 25, 20, 10, 5 and 2% w/v CNC-PBS mixtures with the hot agarose/D-mannitol solution in separate conical tubes at a 1:1 ratio to yield final CNC concentrations of 12.5, 10, 5, 2.5, and 1% w/v in the hydrogel mixtures.
- Working quickly, add 500 µL/well of the above solutions prepared in step 2.6 in quadruplicate to a 24-well culture plate, and let stand for 30 min at room temperature to facilitate polymerization (Figure 2B).
- Carefully layer 1000 µL of the BMMC suspension at a density of 1.1 × 106 cells/mL on top of the hydrogel substrates with a micropipettor, and avoid touching the gel with the tip of the pipette as this will damage the gel surface and generate particles.
- Incubate the BMMCs on the hydrogel substrates in a sterile incubator (5% CO2, humidified atmosphere) at 37 °C for 18 h.
3. Flow cytometric analyses
- Flow cytometric analysis of BMMC viability via PI exclusion
- Carefully remove the culture medium containing BMMCs from the top of the CNC/agarose/D-mannitol, avoiding the gel, and transfer the cells into a 1.5 mL microfuge tube.
- Wash the BMMCs twice with PBS (pH 7.4) containing 0.5% w/v bovine serum albumin at 300 × g for 5 min at room temperature, and resuspend in 180 µL of PBS-0.5% w/v BSA to a final density of 1.5 × 106 cells/mL. Transfer the cells to a 96 well round-bottom plate.
NOTE: Filter-sterilize all PBS-0.5% w/v BSA solutions twice using a 0.2 µm pore-size syringe filter, and store at 4 °C. - Add 20 µL of PBS-0.5% w/v BSA, or 10x PI (100 µg/mL) prepared in PBS-0.5% w/v BSA, to the cells to a final concentration of 10 µg/mL, and incubate for either 1 h at 4 °C or 15 min at room temperature in the dark. Acquire fluorescence data using a flow cytometer equipped with an Argon ion laser (488-514 nm) and bandpass filter to enable detection of fluorescence emission at 578 nm. Acquire 20,000 events per sample at a flow rate of 30 µL/min at room temperature. Analyze the data as described below (sections 3.2.7-3.2.10).
- Flow cytometric analysis of BMMCs for Kit (CD117) and FcεRI surface receptor expression
- Carefully remove the culture medium containing BMMCs from the top of the CNC/agarose/D-mannitol, avoiding the gel, and transfer the cells into a 1.5 mL microfuge tube.
- Wash BMMCs twice with filtered PBS-0.5% BSA at 300 × g for 5 min at room temperature, and resuspend in 180 µL of PBS containing 0.5% w/v BSA (1.5 × 106 cells/mL). Transfer the resuspended cells to a 96-well round-bottom plate.
- Add 20 µL of 10x antibody working solutions to the cells to achieve a final concentration of 0.06 µg/mL of CD117 (c-Kit)-phycoerythrin (PE) and 0.06 µg/mL of FcεRIα-allophycocyanin (APC), respectively.
NOTE: The 10x antibody working solutions must be prepared in filter-sterilized PBS-0.5% w/v BSA. - Add 20 µL of 10x isotype control working solutions containing either rat IgG2b κ isotype control-PE or Armenian hamster IgG isotype control-APC to separate wells containing cells that have not been stained with either of the antibody-fluorophore conjugates in step 3.2.3.
NOTE: The isotype controls, rat IgG2b κ isotype control-PE and Armenian hamster IgG isotype control-APC serve to control for non-specific attachment of antibodies to the BMMC. As BMMCs do not express high levels of FcR, there is rarely high background fluorescence detected with isotype control antibodies. Prepare the 10x isotype control working solutions in filter-sterilized PBS-0.5% w/v BSA. - Incubate the 96-well round-bottom plate for 1 h at 4 °C in the dark.
- Wash the cells by adding 200 µL of fresh PBS-0.5% w/v BSA buffer to each well, and centrifuge at 300 × g for 5 min at room temperature. Repeat the wash step once more. Carefully remove the supernatant without touching the cell pellet; leave behind some supernatant to ensure that the cell pellet remains undisturbed.
- After washing, resuspend cells in 100 µL of 0.5% w/v BSA, 0.05% w/v sodium azide in PBS (pH 7.4) that has been sterile-filtered twice with a 0.2 µm pore-size syringe filter. Acquire fluorescence data in the PE and APC detector channels using a flow cytometer, as outlined above in step 3.1.3.
- Analyze data using software capable of viewing and analyzing flow cytometric data files with the extension ".fcs".
- Begin analysis by plotting forward scatter (FSC) along the x-axis and side scatter (SSC) along the y-axis, and gate for the conserved cell population with a FSC range of log10 1.5-5.0 and an SSC range of log10 0.75-3.25 in the untreated and unstained cells as shown in Figure 3A(i),(ii).
NOTE: This serves to ensure that cell debris with low FSC and SSC values are eliminated from the analysis. Mast cells are normally very granular (high SSC) cells and large (high FSC) when compared to other immunological cell types, such as monocytes and lymphocytes. - Next, generate FSC vs. SSC dot plots and histogram profiles of untreated samples that have been stained with the dye, antibodies, or isotype controls, and obtain mean fluorescence intensity (MFI) in the PE or APC channels according to the emission spectrum of the specific antibody-fluorophore conjugate or dye used. Apply the same set of gates and parameters to all BMMC samples incubated with different CNC/agarose/D-mannitol substrates and stained with the corresponding isotype controls, antibodies, or dye; obtain MFIs.
NOTE: Essentially, the FSC vs SSC dot plots of the untreated stained samples should be indistinguishable from untreated/unstained samples, obtained in step 3.2.9, to ensure that the staining procedure does not alter the cell size (measured by FSC) or granularity (measured by SSC) (Figure 3A(i),(ii)). - Calculate average MFIs and standard error of mean (SEM) for each sample from 4 independent experiments followed by generation of graphs using a statistical analysis software package.
- Prepare histogram overlays in the flow cytometric data analysis software (Figure 3B, 4A(ii),B(ii)).
4. 3D Bioprinting of FNC/sodium alginate hydrogel substrates
NOTE: The 3D bioprinter used in this study is a pneumatic-extrusion system equipped with two independent, temperature-controlled printheads. The biomaterial ink used to 3D bioprint the hydrogel scaffolds is formulated of (a) highly hydrated fibrillar nanocellulose (FNC), which is morphologically similar to collagen, (b) sodium alginate, and (c) D-mannitol. It is supplied as a sterile hydrogel suspension in 3 mL cartridges to which sterile luer-lock conical bioprinting nozzles (22, 25, or 27 G) can be attached.
- Use this protocol with the printheads and printbed at room temperature, where the room temperature is 20-25 °C.
- Store the biomaterial ink cartridges at 4 °C to maintain stability of the hydrogel composite. Prior to commencing 3D bioprinting, remove a cartridge (3 mL) from the refrigerator and allow it to equilibrate to room temperature.
- Installation of a Bioink cartridge onto the INKREDIBLE+ 3D Bioprinter
- Remove the blue end caps from the 3 mL biomaterial ink cartridge (Figure 5B(i)), and affix a sterile 22 G (blue) conical bioprinting nozzle to the luer-lock end of the Bioink cartridge.
NOTE: It is essential to affix the desired conical bioprinting nozzle to the bioink cartridge prior to installing the cartridge and performing the subsequent calibration steps, as failure to do so will result in improper calibration of the z-axis of the 3D bioprinter. - Connect the air pressure supply tubing for printhead 1 (PH1, left) to the opposite end of the cartridge, and insert the cartridge with its attached conical bioprinting nozzle into the vertical slot of PH1 (Figure 5A(ii)).
- Firmly seat the cartridge in PH1 with the conical bioprinting nozzle extending below the printhead. Tighten the screw on PH1 clockwise until finger-tight to lock the Bioink cartridge in place. Note that 3D bioprinting can be performed with PH2 left empty.
- Remove the blue end caps from the 3 mL biomaterial ink cartridge (Figure 5B(i)), and affix a sterile 22 G (blue) conical bioprinting nozzle to the luer-lock end of the Bioink cartridge.
- Calibrating the x-y-z axes of the 3D bioprinter
- Power on the 3D bioprinter, and launch the bioprinter software on a PC connected to the 3D bioprinter via a USB 2.0 cable.
- In the software, click the CONNECT button to sync the software with the 3D bioprinter.
NOTE: The sync is successful when the printhead ultraviolet light-emitting diode lights cycle on and off, and the HEPA filter fan cycles off and on again. The software must be synced to the 3D bioprinter prior to homing the bioprinter axes and the starting point calibration, as outlined in the subsequent steps. - Observe four options in the main control menu of the 3D bioprinter: (i) PREPARE BIOPRINT, (ii) BIOPRINT, (iii) UTILITIES MENU, and (iv) STATUS SCREEN. Select the PREPARE BIOPRINT option, then scroll down to and select the HOME AXES option.
NOTE: The 3D bioprinter will then move the printhead assembly backward and leftward to calibrate the y– and x-axes, respectively. Thereafter, with the printheads paused in the far left position, the bioprinter will raise the printbed until the z-axis calibration switch (located on the printbed) makes contact with the conical bioprinting nozzle installed on printhead 1 - After successful calibration of the x-y-z axes, observe the slight lowering of the printbed and the relocation of the printheads to above the center point of the printbed (Figure 5A(i)).
- Starting point calibration of the bioprinter for bioprinting in 24-well plates
- Remove a sterile 24-well culture plate from its sealed plastic wrapping, and mark a dot of at the center point of well D1 on the underside of the plate with a permanent marker. Remove the plate cover, and place the 24-well culture plate on the printbed with well D1 located at the front left corner of the printbed.
- On the main control menu, select UTILITIES MENU and then select MOVE AXIS. Move the printheads in 1 mm increments along the x– and y-axes until the conical bioprinting nozzle of PH1 is directly over the dot marked under well D1. If necessary, fine-tune the position of the conical bioprinting nozzle over the dot by moving the printheads in 0.1 mm increments.
- Record the x– and y-coordinates of the conical bioprinting nozzle when directly over the center of well D1, as indicated on the control panel screen of the 3D bioprinter.
NOTE: In the case of the INKREDIBLE+ 3D bioprinter used here, the x– and y-coordinates are as follows: x : -46.5 and y : -27.5. These coordinates serve as the starting point at the center of well D1 for bioprinting in a 24-well plate. - Next, raise the printbed in 1 mm increments until the bottom of well D1 is almost touching the conical bioprinting nozzle installed in printhead 1. Fine-tune the movement of the printbed in 0.1 mm increments if necessary (Figure 5A(ii)). Then, from the UTILITIES MENU, select the Z-AXIS CALIBRATION option, and further select and confirm the STORE Z CALIBRATION option.
- Return to the main menu, and select the PREPARE BIOPRINT option. Scroll down to and select the CALIBRATE Z option.
NOTE: The 3D bioprinter will now lower the printbed upon successfully executing the z-axis calibration (Figure 5A(iii)).
- Updating 24-well plate G-code file with correct starting point coordinates
- Open the provided 24-well plate geometric code (G-code) file in the bioprinter software (Supplemental File 1).
NOTE: This 24-well G-code file encodes the printing of 2-layer 5 x 5 x 1 mm rectilinear constructs in each well (Figure 5C(i),(iii)). The provided G-code files can be used with any 3D bioprinting software. - Note that Line 1 of the G-code file reads G0 X-50.0 Y-33.5 ;Center Position of D1 well. Update the x- and y-coordinates on Line 1 with the values obtained in step 4.5.3, i.e., Line 1 should now read G0 X-46.5 Y-27.5 ;Center Position of D1 well. Save the file under a new name.
NOTE: This procedure serves to calibrate the G-code file so that printing will commence in the center of well D1 based on the x,y coordinates of the specific 3D bioprinter being used. This approach can be used to calibrate the 24-well plate G-code file for printing with any extrusion 3D bioprinter. For the purpose of this study, only the left half of the 24-well plate, i.e., wells A1-3, B1-3, C1-3, and D1-3 were printed. A separate G-code file that encodes the printing of 2-layer 5 x 5 x 1 mm rectilinear constructs in a 3 x 4 grid is also provided (Supplemental File 2, Figure 5C(ii)).
- Open the provided 24-well plate geometric code (G-code) file in the bioprinter software (Supplemental File 1).
- Adjustment of extrusion pressure for Printhead 1 (PH1)
- Ensure that the pneumatic pump is tightly connected to the rear air intake port of the INKREDIBLE+ 3D bioprinter, and turn on the pneumatic pump.
- Pull out the forward control knob located on the right side of the INKREDIBLE+ 3D bioprinter.
NOTE: The forward control knob adjusts the pressure for PH1, while the rear control knob adjusts the pressure for PH2. - Observe the digital pressure gauges for PH1 and PH2 located on the front of the bioprinter each read close to 0 kPa. Slowly rotate the forward control knob clockwise until the pressure indicated on the left gauge for PH1 reaches 12 kPa (Figure 5A(iii)).
- Place a folded tissue paper or piece of waterproof, sealing film under the print nozzle of the installed cartridge, being careful not to touch the print nozzle.
- From the main control menu on the 3D bioprinter, select PREPARE BIOPRINT.
- Navigate to and select TURN ON PH1. Note that the biomaterial ink starts to extrude from the print nozzle. If necessary, increase the extrusion pressure by rotating the control knob clockwise until the biomaterial ink is extruded in a continuous filament, and record the new pressure setting. Work quickly to avoid wasting biomaterial ink.
- Select TURN OFF PH1 to stop extrusion of the biomaterial ink, remove the tissue paper or film containing the extruded biomaterial ink from the printbed, and close the bioprinter door.
NOTE: The 3D bioprinter will automatically turn on the air pressure supply to PH1 during printing as instructed by the G-code file.
- 3D bioprinting of rectilinear hydrogel substrates in a 24-well plate format
- From the main control menu on the 3D bioprinter, select UTILITIES MENU. Navigate to and select DISABLE SD PRINT, which will allow the bioprinter software to transmit G-code files to the 3D bioprinter for printing.
- Click on the LOAD button in the bioprinter software, and select the updated 24-well plate G-code file saved in step 4.6.2.
- In the right-hand control panel in the software, select the PRINT PREVIEW tab, and click on the PRINT button to commence bioprinting in well D1.
NOTE: If using the 3 x 4 grid well plate G-code file, the bioprinting will terminate in well A3 of the 24-well plate (Figure 5C(ii),D), as opposed to well A6 if a full 24-well plate is printed. - Upon completion of bioprinting of the rectilinear constructs, cover the 24-well plate with its lid and move it to a class II biosafety cabinet.
- Immerse each rectilinear construct in two drops of sterile 50 mM CaCl2 solution (Figure 5B(ii)), and incubate at room temperature for 5 min.
NOTE: This serves to ionically crosslink the alginate polymer chains with the Ca2+ ions and allow the rectilinear constructs to maintain their structural integrity. - Carefully aspirate the CaCl2 solution from each construct, and rinse once in 1 mL of 1x PBS (pH 7.4) to remove excess CaCl2 (Figure 5D).
- To prevent dehydration of the bioprinted hydrogel constructs, maintain the 3D bioprinted rectilinear constructs in fresh 1x PBS until the BMMCs are ready to be seeded onto the constructs (Figure 5D).
5. Incubation of BMMCs on 3D bioprinted rectilinear scaffolds and viability testing
- Incubation of BMMCs on 3D bioprinted rectilinear hydrogel scaffolds
- Seed aliquots of BMMCs onto the 3D bioprinted rectilinear scaffolds in triplicate for four different durations, e.g., 6, 18, 24, and 48 h, so that all treatments terminate simultaneously. Use BMMCs seeded into empty wells in triplicate for the same durations as untreated controls.
- Aseptically transfer BMMCs from a T175 cm2 culture flask to a sterile 50 mL conical tube, and pellet the BMMCs at 200 × g for 5 min at room temperature.
- Resuspend the pellet in 50 mL of fresh complete RPMI medium containing 20 ng/mL of IL-3, and calculate the live cell density by counting an aliquot of trypan blue-stained BMMCs using a hemacytometer.
- Based on the cell density calculated in step 5.1.3, prepare a 6 mL aliquot of the BMMC suspension at a final density of 1 × 106 cells/mL.
- For the 48 h incubation setup, aspirate PBS from wells A1-3 containing the 3D bioprinted rectilinear hydrogel scaffolds, and pipette 1 mL of BMMC (1 × 106 cells/mL) suspension into each well. Also, pipette 1 mL of BMMC (1 × 106 cells/mL) suspension into wells A4-6 without bioprinted constructs to serve as the untreated control. Incubate the remaining wells containing bioprinted scaffolds (B1-3, C1-3, D1-3) in RPMI without additives to prevent dehydration until the appropriate time is reached for seeding with BMMC for the 24, 18, and 6 h treatment durations. Fill the wells without bioprinted scaffolds (B4-6, C4-6, D4-6) with 1 mL of PBS until the appropriate time is reached for seeding with the BMMCs for the 24, 18, and 6 h time points.
- Incubate the 24-well plate at 37 °C in a 5% CO2 humidified atmosphere.
- For the 24 h incubation setup, aspirate pre-existing culture medium/buffer from wells B1-6, and seed 1 mL of BMMC (1 × 106 cells/mL) suspension into these wells. Return the plate to the incubator.
- For the 18 h incubation setup, aspirate pre-existing culture medium/buffer from wells C1-6, and seed 1 mL of BMMC (1 × 106 cells/mL) suspension into these wells. Return the plate to the incubator.
- For the 6 h incubation setup, aspirate pre-existing culture medium/buffer from wells D1-6, and seed 1 mL of BMMC (1 × 106 cells/mL) suspension into these wells. Return the plate to the incubator.
- For incubation termination, dispense samples from each well of the 24-well plate for (i) PI staining and analysis by flow cytometry, (ii) XTT cell viability assay, and (iii) LDH release assay. Therefore, ensure that a round-bottom 96-well plate and the necessary 1.5 mL microfuge tubes are labeled well in advance of the endpoint of the incubation.
- Cell sampling for XTT metabolic assay
- Thoroughly resuspend the BMMCs in the culture medium within each well, being careful not to damage and fragment the 3D bioprinted hydrogel scaffolds.
- Aseptically transfer 40 µL of the homogeneous BMMC suspension from each well to sterile 1.5 mL microfuge tubes containing 760 µL of fresh complete RPMI medium (final volume = 800 µL), and vortex briefly to mix.
- Dispense 100 µL of the diluted BMMC suspensions in triplicate into a flat-bottom 96-well microtiter plate that is suitable for recording absorbance measurements (see the Table of Materials) on a microplate spectrophotometer.
- Dispense 50 µL of XTT working solution to each well containing the BMMC cell suspension using a multichannel micropipette.
NOTE: Prepare XTT working solution by mixing 5 mL of XTT reagent with 100 µL of electron-coupling reagent. Thaw these components from -20 °C in a 37 °C water bath immediately before use. - Incubate the 96-well plate at 37 °C in a 5% CO2 humidified atmosphere for 24 h.
- At the end of the incubation, remove the 96-well plate from the incubator and allow to cool to room temperature. Record absorbance values on a microplate spectrophotometer at 450 nm with background subtraction at 650 nm.
- Supernatant sampling for lactate dehydrogenase (LDH) assay
- Transfer the remaining BMMC suspensions (960 µL) from each well of the 24-well plate to microfuge tubes, and pellet the cells at 200 × g for 5 min at room temperature.
- Pipette three 100 µL aliquots of the cell culture supernatant from each microfuge tube into a 96-well microtiter flat-bottom plate. Discard the leftover supernatants from each microfuge tube, and retain the BMMC pellets for further staining with PI in sections 5.4.1-5.4.8.
- Pipette 50 µL of the LDH working solution into each 100 µL aliquot of supernatant.
NOTE: Refer to Supplemental File 3 for the preparation of LDH assay reagents. - Incubate for 1 h in the dark at room temperature.
- Pipette 50 µL of 1 M acetic acid to each well to stop the reaction.
- Record absorbance values on a microplate spectrophotometer at 490 nm with background subtraction at 680 nm.
- Flow cytometric analysis of BMMC viability via propidium iodide (PI) exclusion
- Resuspend the BMMC pellets in flow wash buffer (PBS [pH 7.4] containing 0.5% w/v BSA), and pellet the cells at 300 × g for 5 min at room temperature.
- Repeat the wash step with flow wash buffer once more.
- Resuspend each BMMC pellet in 400 µL of flow wash buffer.
NOTE: There will be 24 resuspended BMMC samples in total: 12 BMMC samples incubated on bioprinted hydrogel substrates (4 time points in triplicate) and 12 BMMC samples incubated in wells without bioprinted constructs (4 time points in triplicate). - In a round-bottom 96-well microtiter plate, pipette 20 µL of flow wash buffer into wells A1-12 and B1-12. In the same microtiter plate, pipette 20 µL of 10x PI staining solution (100 µg/mL PI in flow wash buffer) into wells C1-12 and D1-12.
- Dispense 180 µL of the 12 BMMC samples incubated on bioprinted hydrogel substrates into wells A1-12 (one sample per well). Dispense 180 µL of the 12 BMMC samples incubated without bioprinted hydrogel substrates into wells B1-12 (one sample per well). Use the BMMC samples dispensed into wells A1-12 and B1-12 as unstained controls for each treatment condition (24 controls in total).
- Dispense 180 µL of the 12 BMMC samples incubated on bioprinted constructs into wells C1-12 (one sample per well). Dispense 180 µL of the 12 BMMC samples incubated without bioprinted constructs into wells D1-12 (one sample per well).
NOTE: The BMMC samples in wells C1-12 and D1-12 will be stained at a final effective PI concentration of 10 µg/mL (24 stained samples in total). - Incubate the plate either for 1 h at 4 °C or 15 min at room temperature in the dark.
- At the end of the incubation period, analyze the samples directly from the microtiter plate on a flow cytometer as previously described under the section 3.1.
Representative Results
One of the most crucial characteristics of a successful biomaterial ink or culture substrate is that of biocompatibility. Primarily, the substrate must not induce cellular death. There are several microtiter-based and flow cytometric methods of quantifying cell viability and necrosis; however, these methods are not amenable to analyzing cells embedded within a hydrogel matrix. In this protocol, the above mentioned limitation is circumvented by seeding the BMMCs onto the hydrogel substrate or bioprinted scaffold. After a specific incubation period (6-48 h in this study), the BMMCs are easily harvested by micropipetting without the need for mechanical disruption or hydrolysis of the hydrogel substrate or bioprinted scaffold, which would otherwise cause physical damage to the cells. The viability of the BMMCs is then rapidly analyzed using the PI permeability method via flow cytometry. PI is a membrane-impermeant dye that is excluded from viable cells with intact cell membranes and fluoresces only upon intercalating between the bases pairs of DNA16.
As such, only cells with compromised cell membranes are permeable to PI and therefore, will fluoresce. PI permeability analysis demonstrated no changes in BMMC viability when they are cultured on the CNC/agarose/D-mannitol substrate. In fact, none of the CNC concentrations tested (up to 12.5% w/v) elicited any adverse effects on BMMC viability as compared to the BMMCs cultured in the absence of the CNC/agarose/D-mannitol hydrogel substrates (Figure 3B,C). It is also vital that the bioink or culture substrate does not alter the morphology of the cells. Based on the flow cytometric FSC vs. SSC plots, the hydrogel substrate with the highest CNC concentration (12.5% w/v) did not alter the native size (FSC) or granularity (SSC) of the BMMCs as compared to the BMMCs cultured in the absence of the CNC/agarose/D-mannitol hydrogel substrates (Figure 3A(i), (ii)).
Another important characteristic of a biomaterial ink or culture substrate is that it must possess the ability to maintain the differentiation and phenotype of the cells it supports. Two of the most quintessential biomarkers of mast cells are the high-affinity IgE receptor, FcεRI, which facilitates BMMC responses to antigens and the stem cell factor receptor, Kit (CD117), which is required for mast cell survival and differentiation. Mature mast cells are defined by the expression of these two surface receptors, and for a bioink substrate to maintain them in culture, it must not significantly modify the expression of these two receptors. The CNC/agarose/D-mannitol substrate appeared to increase FcεRI and Kit expression at CNC concentrations ≥ 2.5% (Figure 4A(iii),B(iii)). Interestingly, the elevated FcεRI and Kit expression levels remained relatively consistent between 2.5% and 12.5% CNC, which is indicative of a plateau effect and suggests an effect that is not dependent upon the concentration of CNC, but on some other parameter of the hydrogel composite.
The 3D bioprinted hydrogel biomaterial ink scaffolds generated in this study consist of fibrillar nanocellulose (FNC), instead of crystalline nanocellulose, and sodium alginate as the gelator, instead of agarose. Fibrillar nanocellulose exhibits distinct nanoscale topographical features as compared to CNC17 which, in addition to the microscale architecture of the 3D bioprinted FNC hydrogel bioink substrates, could potentially affect the viability of the BMMCs. Following the 3D bioprinting and ionic crosslinking of FNC/alginate/D-mannitol bioink scaffolds (Figure 6), BMMCs were cultured either alone or on the 3D bioprinted hydrogel scaffolds for 6, 18, 24, and 48 h, respectively, in order to assess the dynamic changes in the viability of the BMMCs in response to the FNC/alginate/D-mannitol scaffolds, if any, over extended periods of time. The XTT assay indicated that the metabolic activity of the BMMCs cultured on the 3D bioprinted hydrogel scaffolds remained relatively consistent at ~100% across all tested time points when compared to BMMCs cultured alone (Figure 7A). The lysis of cells results in the release of LDH into the cell culture medium, which the LDH assay detects as a function of its oxido-reductive enzymatic activity.
The BMMCs cultured on the 3D bioprinted hydrogel scaffolds exhibited a gradual time-dependent increase in LDH release when compared to BMMCs cultured alone; however, this trend had a non-significant standard deviation of less than 9% (Figure 7B). PI staining of the BMMCs cultured on the 3D bioprinted hydrogel scaffolds revealed no significant changes in viability compared to BMMCs cultured alone at each time point (Figure 7C). Notably, the MFI of PI-stained BMMCs cultured on the 3D bioprinted hydrogel scaffolds remained relatively consistent (PI-MFI of 7000) across all time points and was similar to the BMMCs cultured on the CNC/agarose/D-mannitol substrates, which exhibited a consistent PI-MFI of 7000 across all CNC concentrations (2.5-12.5%). Collectively, these data demonstrate that the FNC/alginate/D-mannitol hydrogel scaffolds do not adversely affect the viability of the BMMCs.
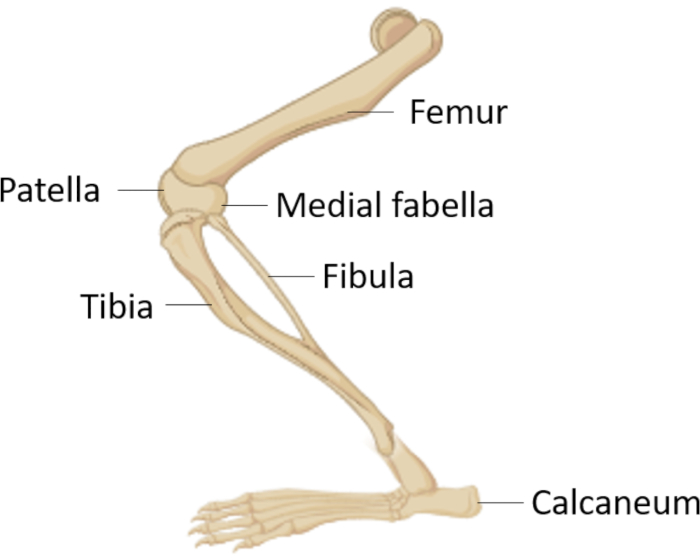
Figure 1: Anatomy of mouse leg, depicting the tibia and femur from which bone marrow is isolated. Please click here to view a larger version of this figure.
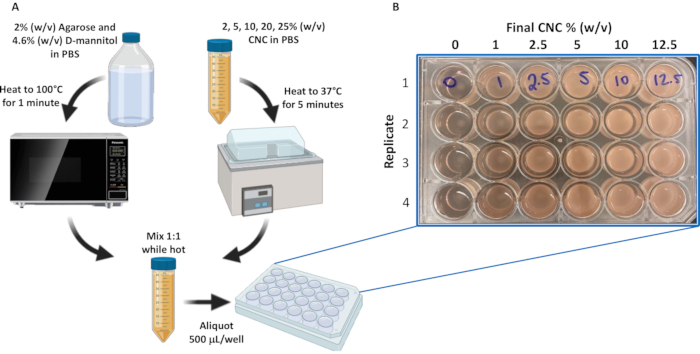
Figure 2: Preparation of the CNC/agarose/D-mannitol hydrogel substrate. (A) Schematic of CNC/agarose/D-mannitol hydrogel substrate preparation protocol. (B) CNC/agarose/D-mannitol preparations (0, 1, 2.5, 5, 10, and 12.5% (w/v) CNC in PBS/agarose/D-mannitol) were loaded onto a 24-well plate in quadruplicate as illustrated. Abbreviations: PBS = phosphate-buffered saline; CNC = crystalline nanocellulose. Please click here to view a larger version of this figure.
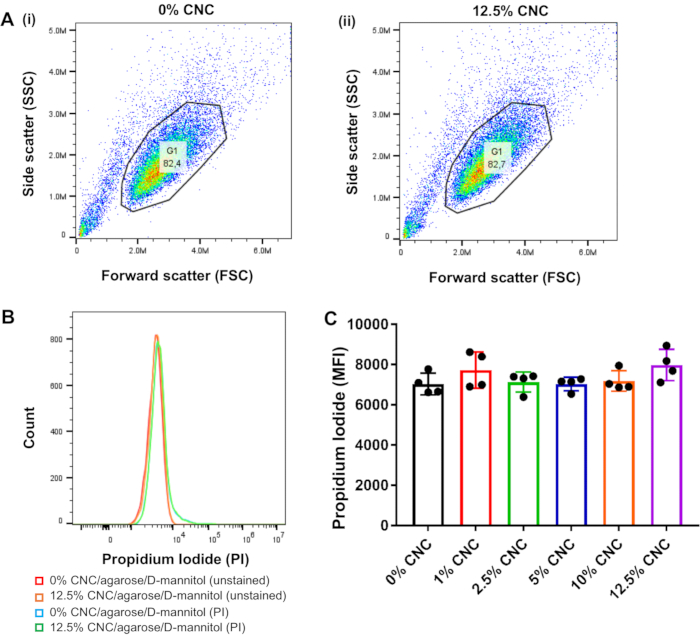
Figure 3: Flow cytometric analysis of BMMC viability via propidium Iodide exclusion following incubation on CNC/agarose/D-mannitol substrates. BMMCs were removed from the CNC/agarose/D-mannitol hydrogel substrates, washed twice, resuspended in PBS-0.5% w/v BSA, stained with PI for 1 h at 4 °C, and analyzed by flow cytometry (n=4). A total of 20,000 cells per sample were acquired including PI fluorescence emission detection in the PE channel. Data analysis was performed using flow cytometry analysis software. (A) Forward scatter (x-axis) versus side scatter (y-axis) dot plot analysis of total cell population from (i) untreated control BMMC (0% CNC/agarose/D-mannitol) and (ii) BMMCs cultured on 12.5% CNC/agarose/D-mannitol, depicting the gated cell population used for data analysis. (B) Histogram overlay profile of gated cells (unstained or stained with PI), from untreated control BMMCs (0% CNC/agarose/D-mannitol) and BMMCs cultured on 12.5% CNC/agarose/D-mannitol. (C) BMMCs were cultured on different CNC/agarose/D-mannitol substrates (1-12.5% (w/v) CNC) for 18 h, and cell viability was determined by flow cytometric analysis of PI-stained cells. Graphical representation of PI MFIs for BMMCs incubated on different CNC/agarose/D-mannitol substrates (1-12.5% (w/v) CNC) relative to BMMCs that were untreated and stained with PI. Abbreviations: BMMCs = bone marrow-derived mast cells; CNC = crystalline nanocellulose; PBS = phosphate-buffered saline; BSA = bovine serum albumin; PI = propidium iodide; PE = phycoerythrin. Please click here to view a larger version of this figure.
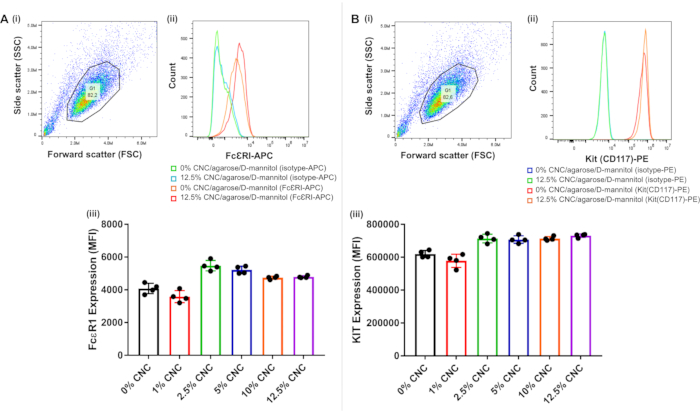
Figure 4: Flow cytometric analysis of FcεRI and Kit (CD117) cell surface receptor expression. BMMCs were cultured either in the absence or presence of CNC/agarose/D-mannitol substrates for 18 h, removed, and analyzed for receptor expression. Data is representative of 4 replicates. Forward scatter (x-axis) versus side scatter (y-axis) dot plot of the total cell population in the untreated BMMC sample (0% CNC/agarose/D-mannitol), stained with (A)(i) the isotype control antibody-APC or (B)(i) the isotype control antibody-PE, and the gated cell population used for data analysis. Histogram overlay profiles of gated BMMCs (stained with isotype control antibody or (A)(ii) anti-FcεRI-APC antibody or (B)(ii) anti-Kit-PE antibody), incubated either alone (0% CNC/agarose/D-mannitol) or on 12.5% CNC/agarose/D-mannitol substrates. BMMCs were cultured for 18 h on different CNC/agarose/D-mannitol substrates (1-12.5% (w/v) CNC) followed by FcεRI and Kit surface receptor expression analysis, respectively, via flow cytometry (n=4). Graphical representation of MFIs of BMMCs stained with (A)(iii) anti-FcεRI-APC or (B)(iii) anti-Kit-PE antibodies, respectively, following culture on different CNC/agarose/D-mannitol substrates (1-12.5% (w/v) CNC) relative to cells that were untreated (0% CNC) and stained similarly. Abbreviations: CNC = crystalline nanocellulose; APC = allophycocyanin; PE = phycoerythrin; BMMCs = bone marrow-derived mast cells; MFI = mean fluorescence intensities. Please click here to view a larger version of this figure.
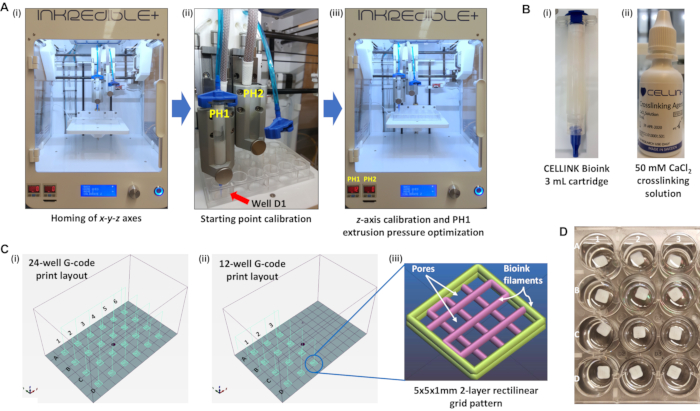
Figure 5: 3D bioprinter equipment and consumables required to bioprint 5 x 5 x 1 mm 2-layer rectilinear bioink hydrogel scaffolds in a 24-well plate format. (A)(i) A pneumatic-extrusion 3D bioprinter with the printhead assembly position at its resting position upon completing the homing of its x-y-z axes. (A)(ii) Starting point calibration of printhead 1 with a bioink cartridge installed and placement of the print nozzle directly over the middle of well D1 in the x-y-z dimensions. (A)(iii) Position of PH1 following z-axis calibration and the extrusion pressure of PH1 set to 12 kPa. (B)(i) Bioink cartridge (3 mL) containing NFC/Alginate/D-mannitol biomaterial ink formulation. (B)(ii) Droplet dispenser of 50 mM CaCl2 crosslinking solution. (C)(i) Schematic representation of a print layout from a G-code file encoding the printing of 5 x 5 x 1 mm 2-layer rectilinear bioink hydrogel scaffolds in all wells of a 24-well plate. (C)(ii) Schematic representation of a print layout from a G-code file encoding the printing of 5 x 5 x 1 mm 2-layer rectilinear bioink hydrogel scaffolds only in wells A1-3, B1-3, C1-3, and D1-3 of a 24-well plate. (C)(iii) Expanded view of the 5 x 5 x 1 mm 2-layer rectilinear bioink hydrogel scaffold pattern in the slicing program, Slic3r. (D)Topview of actual 3D bioprinted 5 x 5 x 1 mm 2-layer rectilinear bioink hydrogel scaffolds in a 24-well plate format immersed in PBS. Abbreviations: 3D = three-dimensional; PH1 = printhead 1; NFC = nanofibrillar cellulose; G-code = geometric code; PBS = phosphate-buffered saline. Please click here to view a larger version of this figure.
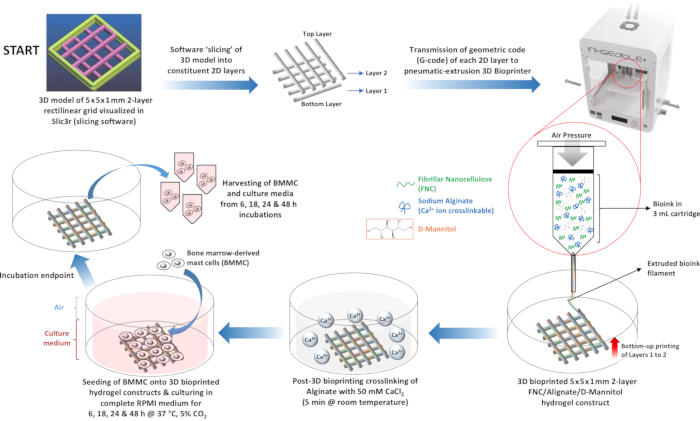
Figure 6: Schematic diagram depicting workflow of 3D bioprinting and BMMC culture on FNC/alginate/D-mannitol hydrogel scaffolds. 3D bioprinting of hydrogel scaffolds with FNC/Alginate/D-mannitol bioink from a G-code file encoding a 5 x 5 x 1 mm 2-layer grid pattern, crosslinking of the hydrogel scaffolds with CaCl2, and culturing of BMMCs on the crosslinked FNC/Alginate/D-mannitol hydrogel constructs. Abbreviations: 3D = three-dimensional; 2D = two-dimensional; G-code = geometric code; FNC = fibrillar nanocellulose; BMMCs = bone-marrow-derived mast cells. Please click here to view a larger version of this figure.
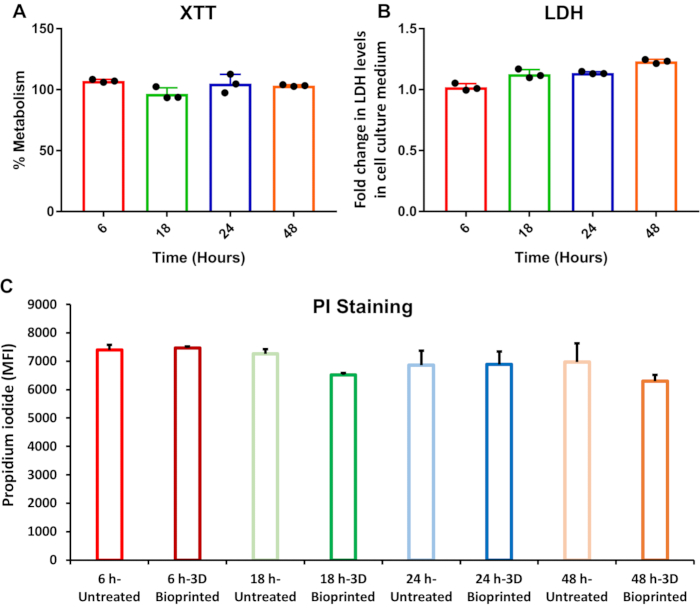
Figure 7: Flow cytometric (PI) and microtiter (XTT and LDH) assays of BMMC viability following incubation on FNC/alginate/D-mannitol scaffolds. (A) Cell proliferation (XTT) metabolic assay data for BMMCs cultured on FNC/Alginate/D-mannitol bioink substrates for 6, 18, 24, and 48 h, respectively. Values are presented as a percentage of the XTT metabolic data for BMMCs cultured alone for 6, 18, 24, and 48 h, respectively. Error bars indicate standard deviation (n=3). (B) LDH enzyme release assay data for BMMCs cultured on FNC/Alginate/D-mannitol bioink scaffolds for 6, 18, 24, and 48 h, respectively. Values are presented as fold-changes relative to the LDH enzyme released by BMMCs cultured alone for 6, 18, 24, and 48 h, respectively. Error bars indicate standard deviation (n=3). (C) Flow cytometric data of PI-stained BMMCs that were cultured either alone or on FNC/Alginate/D-mannitol bioink scaffolds for 6, 18, 24, and 48 h, respectively. Error bars indicate standard deviation (n=3). Abbreviations: LDH = lactate dehydrogenase; BMMCs = bone marrow-derived mast cells; FNC = fibrillar nanocellulose; PI = propidium iodide; MFI = mean fluorescence intensity. Please click here to view a larger version of this figure.
| 1.1.1. 500 mL bottle of RPMI (HyClone, GE Healthcare, USA) |
| 1.1.2. 4 mM L-glutamine (Gibco, Waltham Massachusetts, USA) |
| 1.1.3. 50 mM β-Mercaptoethanol (Fisher Scientific, Hampton, New Hampshire, USA) |
| 1.1.4. 1 mM Sodium Pyruvate (Gibco) |
| 1.1.5. 100 U/mL penicillin (Gibco) |
| 1.1.6. 100 µg/mL streptomycin (Gibco) |
| 1.1.7. 0.1 mM MEM non-essential amino acids (Gibco) |
| 1.1.8. 25 mM HEPES (Fisher) |
| 1.1.9. 10% heat inactivated FBS (Gibco) |
| 1.1.10. 30 ng/mL mouse recombinant IL-3 (Peprotech, Rocky Hill, New Jersey, USA) |
Table 1: Complete RPMI-1640 media supplements.
Supplemental File 1. Please click here to download this File.
Supplemental File 2. Please click here to download this File.
Supplemental File 3. Please click here to download this File.
Discussion
The fabrication of 3D biomimetic tissues requires the successful amalgamation of the bioink, which mimics components of the extracellular matrix, with the cellular component(s) to create physiological analogs of in vivo tissues. This necessitates the use of primary cells, and not transformed cells, when fabricating physiological biomimetic tissues. Primary immunological cells, such as mast cells, however, are particularly susceptible to cytotoxic effects and phenotypic changes that may be elicited by the bioink matrix itself, which is undesirable. Therefore, the ability to rapidly assess the effects of the candidate biomaterial ink on the viability and phenotype (surface receptor expression) of the mast cells is highly advantageous, especially prior to 3D bioprinting of complex tissues containing different cell types embedded within the bioink matrix, which is costly and time-consuming.
The approach of this protocol involves culturing BMMCs, an example of primary mast cells, on the surface of preformed candidate hydrogel bioink substrates (1-12.5% CNC embedded in agarose), which permits the easy retrieval of the BMMCs for subsequent analysis of cell viability and surface receptor expression by flow cytometry. The isolation of bone marrow from mouse femur and tibia requires precision and attention to detail due to the fragility and small size of these bones. After successful isolation and deposition of the bone marrow into cell culture medium, it is essential to maintain the cell culture with 30 ng/mL mouse recombinant IL-3 continuously for 4 weeks, which ensures sustained stimulation to differentiate the hematopoietic progenitor cells into mast cells that are double-positive for the Kit (CD117) and FcεRI cell surface receptors.
Flow cytometric analysis of mast cells for the surface expression of Kit (CD117) and FcεRI receptors should employ isotype antibody controls to assist with differentiating non-specific background signal from the specific signals of the antibodies used to target these receptors, as demonstrated in Figure 4A(ii),B(ii). It is also essential to gate on a well-defined mast cell population to exclude cell debris as shown in Figure 4A(i) and 4B(i), which can also non-specifically bind antibodies. Gentle micropipetting should be used when retrieving the BMMCs from the surface of the bioink hydrogel substrates to reduce the shear forces exerted on the cells, which could damage the cell membrane and result in artificially elevated staining with PI. The PI-stained BMMCs should be analyzed by flow cytometry immediately after the staining incubation period ends as the PI staining medium, which is based on PBS-0.5% w/v BSA, is not intended to sustain cells for extended periods of time outside of cell culture medium. Because PI only permeates cells with compromised cell membranes, it is able to stain necrotic cells with high selectivity and sensitivity. However, PI is unable to detect apoptotic cells, which maintain intact cell membranes.
The PI-exclusion flow cytometric assay described in this protocol may be augmented with Annexin-V-fluorescein isothiocyanate (FITC), which specifically binds to the phospholipid, phosphatidylserine, that is translocated to the outer leaflet of the cell membrane of apoptotic cells. However, compensation is necessary to account for the fluorescence emission bleed-through of FITC into the detector channel used to acquire fluorescence emission from PI and vice versa. In preparing the bioprinter for 3D bioprinting from the provided 24-well plate G-code, it is essential that the starting point calibration be performed accurately, whereby the x– and y-coordinates of the center of well D1 are recorded, and the G-code file is updated with these x– and y-coordinates on line 1. If these initial calibration steps are not performed accurately, the printheads will not move to the correct starting point at the commencement of printing. Correct calibration of the z-axis is equally important as failure to do so can result in the nozzle of the printhead colliding with the printbed or the 24-well plate at the commencement of printing.
The schematic diagram of the 5 x 5 x 1 mm 2-layer rectilinear grid construct (Figure 5C(iii)), illustrates the presence of pores between the extruded bioink filaments that form the substrate. The size of the pores and diameter of the filaments are dependent on three parameters: (i) the travel speed of the print nozzle, (ii) the extrusion pressure applied to the bioink within the cartridge, and (iii) the print nozzle diameter. Bioink substrates with large filament diameters and small pores can be printed using a slower print nozzle speed, higher extrusion pressure (>12 kPa), and larger diameter print nozzle (22 G). Conversely, a faster print nozzle speed, lower extrusion pressure (<12 kPa), and smaller diameter print nozzle (27 G) will result in substrates with finer filaments and larger pores. However, an insufficiently high extrusion pressure will result in inconsistent clumps of bioink being extruded instead of continuous filaments, which will adversely affect the quality and structural integrity of the 3D bioprinted substrate.
The CNC/agarose/D-mannitol candidate bioink presented in this protocol undergoes thermal gelation when agarose cools down to room temperature. In contrast, the commercial bioink used for 3D bioprinting in this protocol undergoes ionic crosslinking with CaCl2 due to the inclusion of sodium alginate in the bioink formulation. It is, therefore, necessary to immerse the bioprinted FNC/alginate/D-mannitol scaffolds in 50 mM CaCl2 solution after bioprinting to facilitate crosslinking (gelation) of the substrate. Omission of the ionic crosslinking step with CaCl2 will result in the dissolution of the FNC/alginate/D-mannitol scaffolds when they are immersed in cell culture medium. The foremost limitation of the CNC/agarose/D-mannitol bioink formulation in its application in actual 3D bioprinting is the exceptionally high melting temperature of agarose (90-95 °C). Even though the printheads of the 3D bioprinter used in this study can reach a maximum temperature of 130 °C, the extruded filaments of CNC/agarose/D-mannitol will likely disperse rapidly as successive layers are printed to construct the substrate. This limitation of the CNC/agarose/D-mannitol bioink formulation can be circumvented by either printing the CNC/agarose/D-mannitol bioink directly onto a cooled printbed to accelerate gelation, or substituting agarose with sodium alginate, which can be extruded at room temperature and undergoes rapid ionic crosslinking with 50 mM CaCl2 within 5 min.
This protocol offers a reliable approach to rapidly screen and exclude bioink formulations that exhibit poor biochemical compatibility with sensitive immunological cells, such as mast cells, that should not be subjected to pneumatic-extrusion 3D bioprinting. Furthermore, this protocol is cost-effective as it requires relatively low quantities of cells to perform a multiplexed screening assay. In contrast, pre-formulated bioink-cell mixtures need to be bioprinted at very high cell densities (>107 cells/mL), which can prove to be costly and time-consuming to perform a biocompatibility screening assay especially when culturing primary cells that are dependent on expensive recombinant growth factors.
Disclosures
The authors have nothing to disclose.
Acknowledgements
We thank Alberta Innovates for providing the CNC and Ken Harris and Jae-Young Cho for their technical advice when preparing the CNC/agarose/D-mannitol matrix. We also thank Ben Hoffman, Heather Winchell and Nicole Diamantides for their technical advice and support with the setup and calibration of the INKREDIBLE+ 3D bioprinter.
Materials
| A | |||
| Acetic Acid (glacial) | Sigma Aldrich | AX0074-6 | |
| Agarose (OmniPur) | EMD Millipore Corporation | 2125-500GM | |
| Armenian Hamster IgG Isotype Control, APC (Clone: eBio299Arm) | Thermo Fisher Scientific | 17-4888-82 | |
| B | |||
| b-Mercaptoethanol | Fisher Scientific | O3446I-100 | |
| b-Nicotinamide adenine dinucleotide sodium salt (NAD) | Sigma Aldrich | N0632-5G | |
| BD 5 mL Syringe (Luer-Lok Tip) | BD | 309646 | |
| BD PrecisionGlide Needle 26G x 1/2 in | BD | 305111 | |
| BioLite 24 Well Multidish | Thermo Fisher Scientific | 930-186 | |
| BioLite 96 Well Multidish | Thermo Fisher Scientific | 130-188 | |
| BioLite 175 cm2 Flask Vented | Thermo Fisher Scientific | 130-191 | |
| Biosafety Cabinet Class II | Microzone Corp., Canada | BK-2-6-B3 | |
| BSA, Fraction V (OmniPur) | EMD Millipore Corporation | 2930-100GM | |
| C | |||
| C57BL/6 mice | The Jackson Laboratory | 000664 | |
| CD117 (c-Kit) Monoclonal Antibody, PE (Clone: 2B8) | Thermo Fisher Scientific | 12-1171-82 | |
| CELLINK BIOINK (3 x 3 mL Cartridge) | CELLINK LLC | IK1020000303 | |
| CELLINK CaCl2 Crosslinking Agent – Sterile Bottle 1 x 60 mL | CELLINK LLC | CL1010006001 | |
| CELLINK Empty Cartridges 3cc with End and Tip Caps | CELLINK LLC | CSC0103000102 | |
| CELLINK HeartWare for PC | CELLINK LLC | Version 2.4.1 | |
| CELLINK INKREDIBLE+ 3D BIOPRINTER | CELLINK LLC | S-10003-001 | |
| CELLINK Sterile Standard Conical Bioprinting Nozzles 22G | CELLINK LLC | NZ4220005001 | |
| CELLINK Sterile Standard Conical Bioprinting Nozzles 25G | CELLINK LLC | NZ4250005001 | |
| CELLINK Sterile Standard Conical Bioprinting Nozzles 27G | CELLINK LLC | NZ4270005001 | |
| Cell Proliferation Kit II (XTT) (Roche) | Sigma Aldrich | 11465015001 | |
| Centrifuge (Benchtop) | Eppendorf | 5804R | |
| Corning Costar 96 Well Clear Flat-Bottom Non-Treated PS Microplate | Sigma Aldrich | CLS3370 | |
| CO2 Incubator | Binder GmbH, Germany | 9040-0113 | |
| CytoFLEX Flow Cytometer | Beckman Coulter | A00-1-1102 | |
| D | |||
| D-mannitol (MilliporeSigma Calbiochem) | Fisher Scientific | 44-390-7100GM | |
| F | |||
| Falcon 15 mL Polystyrene Conical Tubes, Sterile | Corning | 352095 | |
| Falcon 50 mL Polystyrene Conical Tubes, Sterile | Corning | 352070 | |
| FceR1 alpha Monoclonal Antibody, APC (Clone: MAR-1) | Thermo Fisher Scientific | 17-5898-82 | |
| Fetal Bovine Serum (FBS), qualified, heat inactivated | Thermo Fisher Scientific | 12484028 | |
| FlowJo Software | Becton Dickinson & Co. USA | Version 10.6.2 | |
| G | |||
| GraphPad Prism | GraphPad Software, LLC | Version 8.4.3 | |
| H | |||
| Hemacytometer (Improved Neubauer 0.1 mmm deep levy) | VWR | 15170-208 | |
| HEPES Sodium Salt | Fisher Scientific | BP410-500 | |
| I | |||
| Iodonitrotetrazolium chloride (INT) | Sigma Aldrich | I10406-5G | |
| L | |||
| L-Glutamine 200 mM (Gibco) | Thermo Fisher Scientific | 25030-081 | |
| Lithium L-lactate | Sigma Aldrich | L2250-100G | |
| M | |||
| MEM Non-Essential Amino Acids 100 mL 100x (Gibco) | Thermo Fisher Scientific | 11140-050 | |
| 1-Methoxy-5-methylphenazinium methyl sulfate (MPMS) | Sigma Aldrich | M8640 | |
| Microtubes (1.7 mL clear) | Axygen | MCT-175-C | |
| Microtubes (2.0 mL clear) | Axygen | MCT-200-C | |
| MilliQ Academic (for producing MilliQ ultrapure water) | Millipore | ZMQS60001 | |
| N | |||
| Nalgene Rapid-Flow 90 mm Filter Unit (0.2 mm Pore size, 500 mL) | Thermo Fisher Scientific | 566-0020 | |
| Nalgene Syringe filter (0.2 mm PES, 25 mm) | Thermo Fisher Scientific | 725-2520 | |
| P | |||
| Penicillin Streptomycin 100 mL (Gibco) | Thermo Fisher Scientific | 15140-122 | |
| PBS pH 7.4, No Calcium/Magnesium, 500 mL (Gibco) | Thermo Fisher Scientific | 10010-023 | |
| Propidium iodide, 1.0 mg/mL (Invitrogen) | Thermo Fisher Scientific | P3566 | |
| R | |||
| Rat IgG2b kappa Isotype Control, PE (Clone: eB149/10H5) | Thermo Fisher Scientific | 12-4031-82 | |
| Recombinant Murine IL-3 | PeproTech, Inc. | 213-13 | |
| RPMI-1640 Medium 1X + 2.05 mM L-Glutamine (HyClone) | GE Healthcare | SH30027.01 | |
| S | |||
| Sarstedt 96 well round base PS transparent micro test plate (82.1582.001) | Fisher Scientific | NC9913213 | |
| Sodium Azide, 500 g | Fisher Scientific | BP922I-500 | |
| Sodium Pyruvate (100 mM) 100X (Gibco) | Thermo Fisher Scientific | 11360-070 | |
| T | |||
| Tris Base (2-amino-2(hydroxymethyl)-1,3-propanediol) | Sigma Aldrich | 252859 | |
| Trypan Blue solution (0.4%, for microscopy) | Sigma Aldrich | 93595 | |
| V | |||
| VARIOSKAN LUX Microplate Spectrophotometer (Type: 3020) | Thermo Fisher Scientific | VLBL00D0 |
References
- Tibbitt, M. W., Anseth, K. S. Hydrogels as extracellular matrix mimics for 3D cell culture. Biotechnology and Bioengineering. 103 (4), 655-663 (2009).
- Drury, J. L., Mooney, D. J. Hydrogels for tissue engineering: Scaffold design variables and applications. Biomaterials. 24 (24), 4337-4351 (2003).
- Lee, K. Y., Mooney, D. J. Hydrogels for tissue engineering. Chemical Reviews. 101 (7), 1869-1879 (2001).
- Halib, N., Ahmad, I. Nanocellulose: Insight into health and medical applications. Handbook of Ecomaterials. , 1345-1363 (2019).
- Alonso-Lerma, B., et al. High performance crystalline nanocellulose using an ancestral endoglucanase. Communications Materials. 1 (1), 57 (2020).
- Tummala, G. K., Lopes, V. R., Mihranyan, A., Ferraz, N. Biocompatibility of nanocellulose-reinforced PVA hydrogel with human corneal epithelial cells for ophthalmic applications. Journal of Functional Biomaterials. 10 (3), 35 (2019).
- Fey, C., et al. Bacterial nanocellulose as novel carrier for intestinal epithelial cells in drug delivery studies. Materials Science and Engineering: C. 109, 110613 (2020).
- Ojansivu, M., et al. Wood-based nanocellulose and bioactive glass modified gelatin-alginate bioinks for 3D bioprinting of bone cells. Biofabrication. 11 (3), 035010 (2019).
- Jonsson, M., et al. Neuronal networks on nanocellulose scaffolds. Tissue Engineering Part C: Methods. 21 (11), 1162-1170 (2015).
- Samulin Erdem, J., et al. Cellulose nanocrystals modulate alveolar macrophage phenotype and phagocytic function. Biomaterials. 203, 31-42 (2019).
- Menas, A. L., et al. Fibrillar vs crystalline nanocellulose pulmonary epithelial cell responses: Cytotoxicity or inflammation. Chemosphere. 171, 671-680 (2017).
- Halova, I., Draberova, L., Draber, P. Mast cell chemotaxis chemoattractants and signaling pathways. Frontiers in Immunology. 3, 1-19 (2012).
- Groll, J., et al. A definition of bioinks and their distinction from biomaterial inks. Biofabrication. 11 (1), 013001 (2019).
- Schwab, A., et al. Printability and shape fidelity of bioinks in 3D bioprinting. Chemical Reviews. 120 (19), 11028-11055 (2020).
- Jungst, T., Smolan, W., Schacht, K., Scheibel, T., Groll, J. Strategies and molecular design criteria for 3D printable hydrogels. Chemical reviews. 116 (3), 1496-1539 (2016).
- Sasaki, D. T., Dumas, S. E., Engleman, E. G. Discrimination of viable and non-viable cells using propidium iodide in two color immunofluorescence. Cytometry. 8 (4), 413-420 (1987).
- Usov, I., et al. Understanding nanocellulose chirality and structure-properties relationship at the single fibril level. Nature Communications. 6 (1), 7564 (2015).

