Bottom-Up In Vitro Methods to Assay the Ultrastructural Organization, Membrane Reshaping, and Curvature Sensitivity Behavior of Septins
Summary
Septins are cytoskeletal proteins. They interact with lipid membranes and can sense but also generate membrane curvature at the micron scale. We describe in this protocol bottom-up in vitro methodologies for analyzing membrane deformations, curvature-sensitive septin binding, and septin filament ultrastructure.
Abstract
Membrane remodeling occurs constantly at the plasma membrane and within cellular organelles. To fully dissect the role of the environment (ionic conditions, protein and lipid compositions, membrane curvature) and the different partners associated with specific membrane reshaping processes, we undertake in vitro bottom-up approaches. In recent years, there has been keen interest in revealing the role of septin proteins associated with major diseases. Septins are essential and ubiquitous cytoskeletal proteins that interact with the plasma membrane. They are implicated in cell division, cell motility, neuro-morphogenesis, and spermiogenesis, among other functions. It is, therefore, important to understand how septins interact and organize at membranes to subsequently induce membrane deformations and how they can be sensitive to specific membrane curvatures. This article aims to decipher the interplay between the ultra-structure of septins at a molecular level and the membrane remodeling occurring at a micron scale. To this end, budding yeast, and mammalian septin complexes were recombinantly expressed and purified. A combination of in vitro assays was then used to analyze the self-assembly of septins at the membrane. Supported lipid bilayers (SLBs), giant unilamellar vesicles (GUVs), large unilamellar vesicles (LUVs), and wavy substrates were used to study the interplay between septin self-assembly, membrane reshaping, and membrane curvature.
Introduction
Septins are cytoskeletal filament-forming proteins that interact with lipid membranes. Septins are ubiquitous in eukaryotes and essential to numerous cellular functions. They have been identified as the main regulators of cell division in budding yeast and mammals1,2. They are involved in membrane reshaping events, ciliogenesis3, and spermiogenesis4. Within mammalian cells, septins can also interact with actin and microtubules5,6,7 in a binder of Rho GTPases (BORG)-dependent manner8. In various tissues (neurons9, cilia3, spermatozoa10), septins have been identified as regulators of diffusion barriers for membrane-bound components11. Septins have also been shown to regulate membrane blebbing and protrusion formation12. Septins, being multi-tasking proteins, are implicated in the emergence of various prevalent diseases13. Their misregulation is associated with the emergence of cancers14 and neurodegenerative diseases15.
Depending on the organism, several septin subunits (two in Caenorhabditis elegans to 13 in humans) assemble to form complexes whose organization varies in a tissue-dependent fashion16. The basic septin building block gathers two to four subunits, present in two copies and self-assembled in a rod-like palindromic manner. In budding yeast, septins are octameric17,18. In situ, septins are often localized at sites with micrometer curvature; they are found at division constriction sites, at the base of cilia and dendrites, and at the annulus of spermatozoa19,20. At the membrane, the role of septins seems to be dual: they are implicated in reshaping the lipid bilayer and in maintaining membrane integrity21. Hence, investigating the biophysical properties of septin filament-forming proteins and/or subunits at the membrane is crucial for understanding their role. To dissect specific properties of septins in a well-controlled environment, bottom-up in vitro approaches are appropriate. So far, only a few groups have described the biophysical properties of septins in vitro20,22,23. Hence, as compared with other cytoskeletal filaments, the current knowledge on the behavior of septins in vitro remains limited.
This protocol describes how the organization of septin filaments, membrane reshaping, and curvature sensitivity can be analyzed19. To this end, a combination of optical and electron microscopy methods (fluorescence microscopy, cryo-electron microscopy [cryo-EM], and scanning electron microscopy [SEM]) has been used. The membrane reshaping of micrometer-sized giant unilamellar vesicles (GUVs) is visualized using fluorescence optical microscopy. The analysis of the arrangement and ultrastructure of septin filaments bound to lipid vesicles is performed using cryo-EM. Analysis of septin curvature sensitivity is carried out using SEM, by studying the behavior of septin filaments bound to solid-supported lipid bilayers deposited on wavy substrates of variable curvatures, which enables the analysis of curvature sensitivity for both positive and negative curvatures. As compared with previous analysis20,24, here, we propose to use a combination of methods to thoroughly analyze how septins can self-assemble, synergistically deform membrane, and be curvature-sensitive. This protocol is believed to be useful and adaptable to any filamentous protein that displays an affinity for membranes.
Protocol
1. Determination of membrane reshaping using giant unilamellar vesicles (GUVs)
NOTE: In this section, GUVs are generated to mimic the membrane deformations possibly induced by septins in a cellular context. Indeed, in cells, septins are frequently found at sites with micrometer curvatures. GUVs have sizes ranging from a few to tens of micrometers and can be deformed. They are thus appropriate to assay any micrometer-scale septin-induced deformations. Fluorescent lipids, as well as fluorescently labeled septins (using Green Fluorescent Protein [GFP]), are used to follow the behavior of both lipids and proteins via fluorescence microscopy.
- Preparation of buffers and solutions
- Prepare the GUV growth buffer (50 mM NaCl, 50 mM sucrose, and 10 mM Tris [pH = 7.8]) and the observation buffer (75 mM NaCl and 10 mM Tris [pH = 7.8]).
- Measure (using a commercial osmometer) and adjust the osmolarity of the observation and growth buffers (170 mOsmol·L-1, in theory) by adding small amounts of NaCl until their respective osmolarities are equal. Filter the buffers using a 0.2 µm filter. Aliquot the growth buffer and store it at -20 °C for further use. Store the observation buffer at 4 °C.
NOTE: The difference in osmolarity between both buffers should not exceed 5%. - Prepare a 5 mg·mL-1 β-casein solution in the observation buffer. Ensure complete dissolution (after several hours at 4 °C with magnetic stirring). Filter the solution with a 0.2 µm filter, aliquot, and store at -20 °C.
- Prepare the lipid mixtures at a total lipid concentration of 3 mg·mL-1 in chloroform. Use a composition (mol%) of 56.8% Egg L-α-phosphatidylcholine (EggPC), 15% cholesterol, 10% 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine (DOPE), 10% 1,2-dioleoyl-sn-glycero-3-phospho-L-serine (DOPS), 8% brain L-α-phosphatidylinositol-4,5-bisphosphate (PI(4,5)P2), and 0.2% Bodipy-TR-Ceramide to enhance the protein-lipid interactions and favor the incorporation of PI(4,5)P225.
NOTE: Handle chloroform under a fume hood using nitrile gloves and safety glasses. Pipette chloroform solutions with glass syringes and avoid plastic as chloroform dissolves plastic. Before and after use, rinse the syringes by pipetting chloroform 5x-10x. Use separate syringes to pipette specific fluorescent lipids to prevent cross-contamination. Lipids can be stored in chloroform at -20 °C in an amber glass vial capped with Teflon. The vials must be filled with argon before closing and sealed with parafilm to prevent any lipid oxidation. - Electro-formation of GUVs using a platinum wires setup
NOTE: Figure 1 presents the scheme of the experimental steps and a picture of the chamber.- Clean the chamber and platinum wires thoroughly as follows to remove any lipid residues.
- Plunge the wires and the chamber in acetone and sonicate for 10 min. Wipe them carefully with a paper tissue using acetone.
- Assemble the chamber by inserting the wires, plunge again in acetone, and sonicate for 10 min. Wipe once more with acetone, making sure that the wires are fully cleaned. Plunge the chamber in ethanol, sonicate for 10 min, and wipe with ethanol.
- Finally, plunge the chamber in deionized water, sonicate for 10 min, and dry with a stream of nitrogen or air.
NOTE: The Teflon chamber (Figure 1B) was custom-made in the in-house workshop. It accommodates three compartments that can be sealed on both sides using glass coverslips. Platinum wires can be inserted into the chamber through 1.3 mm diameter holes. - After cleaning the chamber, deposit 3-4 drops per compartment (each drop is about 0.1 μL)) of the 3 mg·mL-1 lipid mixture on each platinum wire. Turn the wires by 180° and deposit 3-4 lipid droplets per compartment on the opposite side of each platinum wire. Ensure that the drops do not contact one another. About 5 µL of the lipid mixture is required per whole chamber.
- Place the growth chamber in a vacuum chamber for 30 min to remove any trace of chloroform.
NOTE: Deep vacuum (0.1 mbar) is best. When dried, lipids are vulnerable to oxidation and, therefore, should not be left in air for more than a few minutes. - Deposit high-vacuum grease at the bottom of the chamber (the side closest to the wires) along the periphery of the three compartments using a syringe and press a clean (22 mm x 40 mm) coverslip against the grease to ensure perfect sealing. Seal both extremities of the chamber (i.e., at the entry/exit sites of the wires) using sealing paste (wax plates). Similarly, apply vacuum grease on the other side of the chamber.
- Fill the compartments with growth buffer (~1 mL per chamber) using a pipette. Do not stir the solution too rapidly or strongly to prevent any detachment of the lipid film from the wires. Seal the top of the chamber hermetically using a 22 mm x 40 mm coverslip by pressing it against the grease. To avoid the formation of air bubbles, gently press the glass coverslip from the center to the edges.
- Place the chamber in a 4 °C fridge and connect the wires to a wave function generator (sine function at 500 Hz). Set an effective voltage of 350 mV for a shorter growth period (i.e., 6 h) or 250 mV for a longer growth period (i.e., 12-16 h), as already presented and optimized in a study by Beber et al.25.
- Remove the coverslips, wipe off the sealant and grease, and remove the wires. Wash and scrub the chamber with a paper tissue using water and ethanol (≥70%) alternately.
NOTE: The optimal voltage and time scale for GUV growth depend on many parameters, from the buffer salt concentration to the chamber geometry (i.e., the distance between the wires and chamber size). Use the same chamber every time the experiment is repeated to ensure reproducibility. The wires are in proximity with the bottom of the chamber so that the lipids can be imaged using fluorescent microscopy. Image the lipids at each step (see Figure 1C,D) to make sure that the electro-formation process is successful.
- Septin incubation with the vesicles
- Collect GUVs from the wires using pre-cut pipette tips (~1 mm opening) that are brought in close vicinity to the wires. Then, pipette the solution all along the wire. This procedure prevents the generation of strong lamellar flows that could disrupt the GUVs. Following this step, cutting pipette tips is no longer required. Indeed, lamellar flows do not damage GUVs in the solution.
NOTE: Due to the presence of PI(4,5)P2 in the lipid mixture, GUVs that have been collected must be stored for no more than 2-3 h before the experiment. Indeed, PI(4,5)P2 gets solubilized rapidly and septins no longer bind to the membranes a few hours after their formation. However, once septins are bound to the membrane, they remain bound for a few days. - Dilute the septins stock solution in Tris 10 mM (pH 8) exclusively to reach an osmolarity equal to that of the growth buffer; if needed, dilute the septin solution further in the observation buffer. Add the intended volume of collected GUVs (50-100 µL for a total volume of 200 µL). Perform the incubation directly in the observation chamber after passivation with β-casein (see below). 20-30 min waiting time is required to reach equilibrium.
NOTE: The expression and purification of septin octameric complexes (human or budding yeasts) are extensively described in other articles17. Briefly, septins were expressed in Escherichia coli, purified in the lab using affinity, size-exclusion, and ion-exchange chromatography steps, and stored at -80 °C in an aqueous solution of 50 mM Tris-HCl (pH 8), 300 mM KCl, and 5 mM MgCl2 at ~1 mg·mL-1 (3 µM) concentration. A high salt concentration is used to avoid septin aggregation. Septin complexes should not be concentrated through a filter centrifugation device, which induces aggregation and thus reduces the protein yield.
- Collect GUVs from the wires using pre-cut pipette tips (~1 mm opening) that are brought in close vicinity to the wires. Then, pipette the solution all along the wire. This procedure prevents the generation of strong lamellar flows that could disrupt the GUVs. Following this step, cutting pipette tips is no longer required. Indeed, lamellar flows do not damage GUVs in the solution.
- Imaging with confocal and/or spinning disk microscope
- To prevent the GUVs from sticking to the surface and/or exploding, passivate the observation chamber by incubating it with a 5 mg·mL-1 β-casein solution for 30 min.
- Remove the β-casein solution and transfer the septin-GUV solution (Step 1.4.2.) into the observation chamber using a pipette. Let the GUVs sediment to the bottom of the chamber for 10-15 min.
NOTE: The composition discrepancy between the interior of the GUV and the external buffer creates both a density and refractive index mismatch. Owing to the refractive index mismatch, the GUVs are visible with transmission light optical microscopy. - Using confocal microscopy, visualize the fluorescent signal of the lipids to check for the GUVs quality and membrane lamellarity state. Assess the density of septins bound to GUVs by recording the septins' fluorescent signal after performing a proper calibration25. Perform Z-stack acquisitions at 0.4 µm spatial interval to analyze and visualize the 3D deformations of the vesicles induced by the interaction between the septins and the membrane.
NOTE: Oil immersion objectives with 60- or 100-fold magnifications were used. Standard confocal or spinning disk microscopes (Table of Materials) with pixel sizes of 250 nm and 110 nm, respectively, were used. One must adapt imaging conditions to a given piece of equipment. No specific anti-photo bleaching agents were added to the solution.
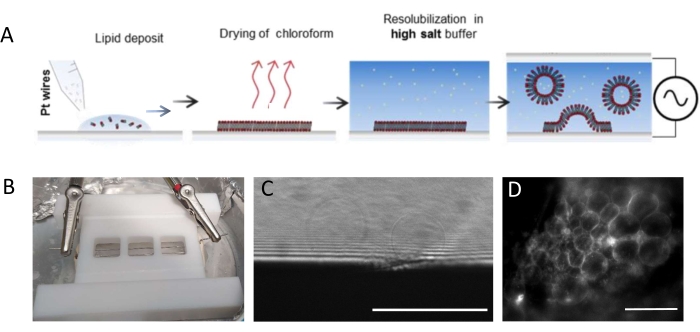
Figure 1: Electro-formation of GUVs. (A) Schematic representation of the electro-formation process using platinum wires. (B) Picture of the Teflon home-made device assembled with platinum wires used for generating GUVs by electro-formation. Wires are 0.5 mm in diameter and 3 mm apart. (C) GUVs (spherical objects) observed by transmission optical microscopy during the growth process. The opaque zone at the bottom of the image is the platinum wire. (D) GUVs (round fluorescent objects) observed with fluorescent microscopy during the growth on the platinum wire. Scale bars = 100 μm. Please click here to view a larger version of this figure.
2. Analysis of ultrastructural organization of septin filaments by cryo-electron microscopy
NOTE: Vesicles are not suitable for imaging with standard electron microscopy methods. Indeed, samples are dried using standard negative stain methods. Upon dehydration, vesicles are likely to undergo unspecific deformations, frequently resulting in lipid protrusions. Cryo-electron microscopy is thus a much better strategy to observe specific deformations of vesicles. Using cryo-EM, the samples are embedded within a thin (~100-200 nm) layer of vitrified ice, which preserves the samples close to the native state. GUVs are, however, too large (several tens of micrometers) to be embedded within thin ice and thus imaged by transmission electron microscopy. Hence, large unilamellar vesicles (LUVs), whose diameters range from ~50-500 nm, are generated to determine how septins can deform vesicles and how they arrange on vesicles.
- Generation of large unilamellar vesicles (LUVs)
- Prepare 50 µg of lipid mix solubilized in chloroform with a molar composition of 57% EggPC, 15% cholesterol, 10% DOPE, 10% DOPS, and 8% brain PI(4,5)P2), which is optimized to enhance septin interaction with the membrane in a glass vial.
- Dry the solution under argon flow to generate a dried lipid film in the vial. Put the vial under vacuum for 30 min to completely dry the lipid.
- Resolubilize the lipid film in 50 µL of aqueous solution (50 mM Tris-HCl [pH 8]; 50 mM KCl; 2 mM MgCl2) to obtain a final concentration of 1 mg·mL-1, vortex for 10 s, and transfer the solution to a tube.
NOTE: LUVs must be used at once for incubation with septins. Otherwise, the protein-lipid interaction will be weak because of PI(4,5)P2 solubilization. This crude re-solubilization process generates a heterogeneous population of vesicles with diameters ranging from 50 nm to 500 nm. Hence, a whole range of diameters and thus curvatures are simultaneously assayed.
- Incubate the septins with the solubilized lipids at final lipid and septin concentrations of 0.1 mg·mL-1 (about 300 nM) and 20 nM, respectively, in a high-salt buffer (50 mM Tris-HCl [pH 8], 300 mM KCl, 2 mM MgCl2). Incubate the sample for 1 h at room temperature.
- Plunge-freezing to vitrify the sample
- Glow-discharge holey carbon grids (300 mesh) on the carbon side for 30 s at 5 mA using plasma generator equipment and insert the grid inside a plunge freezing machine in a humid environment.
- Adsorb 4 µL of the sample (Step 2.2.) on the glow-discharged carbon side of the grid. Immediately before sample adsorption, add 5-10 nm gold beads into the solution, in case the samples will be used to generate tilted series by cryo-tomography.
NOTE: The density of gold beads must be screened and adjusted empirically and depends on the provider. The optimized density is 10-15 gold beads in the field of view. - Blot dry the samples from the bare side to aspirate the sample drop on the opposite side.
NOTE: The blotting time is typically 4 s, and the position of the filter paper and blotting force are adjusted empirically by testing and screening alternative positions of the filter paper to optimize the ice quality (thickness) and the material density. - Transfer the grids into a microscope or store them in a liquid nitrogen container.
NOTE: Using holey carbon grids is essential to accommodate the polydispersity in the size of the vesicles. Blotting the sample from the opposite side favors enhanced adsorption of the biological material on the grid.
- Cryo-electron microscopy imaging
- Insert the grids into an electron microscope (EM) equipped for cryo-EM observation. Screen the whole grid by generating a map of the whole sample at low magnification (typically at 120x magnification) to select areas displaying better ice (i.e., thin and well vitrified).
- For cryo-EM 2D data collection, collect images at a pixel size of about 2 Å per pixel to check for the quality of the sample. Ensure that both the septin filaments and the lipid bilayer are visible.
- For cryo-electron tomography data collection, select areas of interest displaying deformed vesicles. Ensure that enough gold beads (at least 10) are present in the field of view.
- According to the software used for tilted series data collection, select focus and tracking positions to be far enough from the area of interest. Collect tilted series by varying the tilt angle from -60° to +60°, collecting an image every 2°-3° degrees.
NOTE: The total dose should be about, or less than, 100 electrons/Å2. The pixel size varies from 1.3 Å to 2.1 Å, depending on the microscope used for data collection. Ideally, an angular symmetric scheme for data collection is preferred as follows: 0°, -3°, +3°, -6°, +6°, -9°, +9° […] -60°, +60°. However, goniometers of side-entry microscopes do not provide enough mechanical stability to achieve those symmetric schemes. Alternatively, the acquisition can be started at 0° to 34°, followed by a second angular sequence from -2° to -60°, and ended by a final sequence from 36° to 60°. The purpose is to collect the first images (with the lowest radiation damages) at the lowest angles. Moreover, to visualize the ultrastructure of vesicles and bound septin filaments, a standard cryo-EM microscope can be used (200 kV, Lanthanum hexaboride (LaB6) filament, and sample side entry). However, if one aims to pursue further image processing (sub-tomogram averaging, for instance), it is best to use last-generation field emission gun (FEG) microscopes equipped with direct detectors. In this protocol, we restrict our description to the acquisition of 3D reconstructions and leave out sub-tomogram averaging.
- 3D reconstruction from cryo-tomography and segmentation
- Use the IMOD software suite (Table of Materials) for tilted series image alignment and 3D reconstruction26,27. Within IMOD, perform tilt series alignment based on fiducials (gold beads) positioning. In addition, if needed, carry out contrast transfer function (CTF) determination and correction within IMOD26. Finally, achieve 3D reconstruction with IMOD, following each step rigorously.
- Manually segment the lipid bilayers and septin filaments using 3Dmod27 from the IMOD software suite, for display.
3. Analysis of curvature sensitivity of septin using SEM
NOTE: To understand how septins can be sensitive to micrometer curvatures, an in vitro approach has been used to incubate septin filament complexes with solid-supported lipid bilayers deposited on micrometer-scale wavy undulated patterns.
- Designing wavy NOA (Norland optical adhesive) replica from wavy polydimethylsiloxane (PDMS) patterns
- Use PDMS undulated patterns of 250 nm amplitude and 2 µm lateral periodicity or other dimensions to assay the curvatures suitable for the protein of interest.
NOTE: PDMS undulated patterns are designed and generated as described in Nania et al.28,29. - In a cleanroom environment, deposit 5 µL of liquid NOA on a circular glass coverslip of 1 cm in diameter and place the PDMS template on the drop. Treat with UV light (320 nm) for 5 min to photo-polymerize the liquid NOA into a thin polymer film. Then, gently peel off the PDMS template from the coverslip with the freshly polymerized NOA.
NOTE: NOA is a common optically transparent glue suitable for optical microscopy imaging. In addition, NOA is resistant to the chemical fixation and staining processes carried out prior to SEM imaging. Both NOA 71 and NOA 81 resins can be used with similar outcomes. The initial PDMS pattern could be used several times to produce NOA replicas. The obtained NOA replicas can be kept in a box at room temperature for months.
- Use PDMS undulated patterns of 250 nm amplitude and 2 µm lateral periodicity or other dimensions to assay the curvatures suitable for the protein of interest.
- Generation of a supported lipid bilayer and protein incubation
- Treat the NOA films using an air plasma cleaner for 5 min to make the surface hydrophilic.
- Prepare a solution of small unilamellar vesicles (SUVs) with a molar composition of 57% EggPC, 15% cholesterol, 10% DOPE, 10% DOPS, and 8% brain PI(4,5)P2 at a total lipid concentration of 1 mg·mL-1. Prepare the SUVs by resuspending a dried lipid film in the observation buffer, as described in Step 2.1.3. Gently sonicate the solution using a bath sonicator for 5-10 min until the solution is transparent.
NOTE: The solution of SUVs can be stored frozen for several weeks at -20 °C. - Insert the coverslips supporting the NOA patterns within the wells of cell culture boxes. Deposit 100 µL of 1 mg·mL-1 SUV solution on the freshly glow-discharged NOA patterns (Step 3.2.1.) and incubate for 30 min at room temperature. This step induces the fusion of SUVs with the surface of the NOA pattern to generate a supported lipid bilayer.
- Rinse the slides thoroughly 6x with the septin buffer (50 mM Tris-HCl [pH 8], 50 mM KCl, 2 mM MgCl2) to remove unfused SUV. After each rinse, never let the sample completely dry.
- Dilute the octameric septin stock solution in septin high-salt buffer (50 mM Tris-HCl [pH 8], 300 mM KCl, 2 mM MgCl2) using septin buffer (50 mM Tris-HCl [pH 8], 50 mM KCl, 2 mM MgCl2) to final concentrations ranging from 10 nM to 100 nM, and volumes of 1 mL. Incubate the protein solution on the slides for ~1 h at room temperature.
NOTE: The volume is large enough so that evaporation is not an issue.
- Sample preparation for SEM analysis
NOTE: Various protocols have been developed in electron microscopy to analyze protein structures and their assembly. The current protocol preserves the organization of septin filaments, using a fixation protocol derived from Svitkina et al.30 that is easy to implement. Besides, this protocol optimizes high-resolution SEM observations.- Preparation of reagents and stock solutions
NOTE: This protocol requires several reagents and solutions that can be prepared in advance or just prior to incubation. Follow the provided instructions to avoid any artifacts or lack of chemical reactivity.- Sodium cacodylate 0.2 M stock solution: To prepare this double-strength solution, dissolve 2.14 g of sodium cacodylate powder in ~ 40 mL of distilled water under magnetic stirring. After complete dissolution, adjust the pH to 7.4 by gently adding 0.1 M HCl (~1 mL for 50 mL of solution) and make up the final volume with distilled water. This solution can be stored for 24-48 h at 4°C.
- 2% glutaraldehyde (GA) in 0.1 M sodium cacodylate (fixative solution): Prepare this solution right before use by diluting highly pure EM grade GA with 0.2 M sodium cacodylate solution (see above). Use a 25%-50% commercial GA stock solution because of its optimized storage capacity at 4 °C. For preparing 10 mL of fixative solution, dilute 0.8 mL of 25% commercial GA solution with 4.2 mL of distilled water and 5 mL of 0.2 M sodium cacodylate.
- 1% osmium tetroxide (OsO4) in 0.1 M sodium cacodylate: Prepare this second fixative solution right before use by diluting the commercial 4% OsO4 stock solution with 0.2 M sodium cacodylate (see above). For 4 mL of OsO4 fixative solution, dilute 1 mL of 4% commercial OsO4 solution with 1 mL of distilled water and 2 mL of 0.2 M sodium cacodylate.
NOTE: Use commercial 4% OsO4 stock solution in cleavable glass bulbs because of their storage and handling properties. Note that OsO4 is highly reactive. Make sure that its color is slightly yellow and not dark. - 1% tannic acid (TA) in water: Prepare this solution right before use. Prepare the TA solution to obtain a final concentration of 1% TA in distilled water at room temperature. Dissolve 10 mg in 1 mL of distilled water and vortex for a few minutes. TA solution cannot be stored and must be filtered with a 0.2 µm filter before use.
- 1% uranyl acetate (UA) solution in water Prepare UA solution to obtain a final concentration of 1% UA in distilled water. Dissolve 10 mg in 1 mL of distilled water and vortex for at least 30 min to 1 h by vortexing or shaking at room temperature. This solution can be stored for 1 month at 4 °C but can easily precipitate and must, therefore, be filtered with a 0.2 µm filter before use.
- Graded series of ethanol solutions Prepare 50%, 70%, 95%, and 100% ethanol solutions in water. Prepare the final bath from newly opened bottles of 100% ethanol or from 100% ethanol that has been dehydrated for at least 24 h with molecular sieves (nominal pore diameter = 4 Å) to remove from samples all traces of water that might interfere with drying and coating. Be careful not to shake this solution as it might cause resuspension of silicate particles.
NOTE: Be careful when handling the chemical reagents used in this protocol. Glutaraldehyde, osmium tetroxide, sodium cacodylate, and uranyl acetate are all highly toxic, uranyl acetate being radioactive as well. All manipulation of the reagents and their waste must be done using individual (gloves, lab coats, safety glasses) and collective protection (fume hoods and plexiglass shields), according to the laboratory-specific procedures.
- Sample fixation
- Wash the samples (i.e., NOA slides with fused supported lipid bilayers and incubated protein) with PBS. Replace PBS with the GA fixative solution prewarmed at 37 °C and let the reaction proceed for 15 min. Samples can then be stored at 4 °C.
- Remove the fixative solution and wash the fixed samples 3x with 0.1 M sodium cacodylate (5 min per washing) with gentle shaking.
- Incubate the samples in the OsO4 fixative solution for 10 min with maximum protection from light to allow the fixation of membranous structures and increase the electrical conductivity of the samples. Remove the fixative solution and wash the samples 3x in distilled water (5 min per washing) with gentle shaking.
- Incubate the washed samples in the filtered TA fixative solution for a maximum of 10 min. Remove the TA solution and wash the samples 3x in distilled water (5 min per washing) with gentle shaking.
- Incubate the washed samples in a freshly filtered UA fixative solution for 10 min with maximum protection from light. Remove the UA solution and wash the samples 3x in distilled water (5 min per washing) with gentle shaking.
- Sample dehydration and critical point drying
NOTE: Air-drying is not permitted to avoid damaging samples through unwanted surface tensions when changing from the liquid to the gaseous state. Two methods have been established in EM: the physical method of reaching the supercritical state of CO2 and bypassing its critical point (31 °C, 74 bar), or the chemical method of evaporating hexamethyldisilazane (HMDS), a drying agent with reduced surface tension. CO2 and HMDS are poorly miscible with water. Consequently, all traces of water must be substituted with a transitional solvent (ethanol) to avoid any damage later during the drying processes.- Incubate the samples for 2-3 min in each ethanol solution, starting from 50% to 100% (anhydrous) ethanol bath.
NOTE: As the evaporation of ethanol between baths is fast, the sample handling must also be fast to avoid any air-drying. - Transfer the glass slides inside the critical point dryer prefilled with ethanol and follow the manufacturer's instructions.
NOTE: In this protocol, an automatic apparatus was used, but any system can be used. Protocols may differ for each apparatus but must be optimized to completely remove ethanol (25 baths in this protocol) and reduce speeds for the different solvent exchanges (CO2 inlets and ethanol/CO2 outlets) and the final depressurization (almost 1 h in this protocol). - At the end of the critical point drying, store samples immediately in a desiccator until mounting and coating. As dried samples are highly hygroscopic, coat them (see below) as soon as possible.
NOTE: Alternately, HMDS can offer a cheaper and faster method. Although HDMS has never been tested on our samples, this approach produced good results for the observation of proteins at the inner face of cellular membranes31,32.
- Incubate the samples for 2-3 min in each ethanol solution, starting from 50% to 100% (anhydrous) ethanol bath.
- Sample mounting and coating.
NOTE: Since biological samples present poor electrical conductivity properties, they need to be coated with a conductive metal film before SEM observation. Plasma-magnetron sputtering is thus used.- Attach the coverslip to stubs that will be used for future SEM observations. Use silver paint because of its improved electrical conductivity, as compared with carbon disks. Add a strip of silver paint to the upper face of the coverslip and make sure that the connection with the stub is satisfactory. Avoid any paint agglomerates.
NOTE: The silver paint strip must be thin to avoid any contact between the sample and the objective lens of the SEM when working at high resolutions (i.e., low working distance). - Wait for the solvent to evaporate completely.
NOTE: The duration of this step may vary depending on the amount and the thickness of the silver paint deposit. This step can be shortened by using a bell jar or the coater and a primary vacuum for 10-30 min. - Use an apparatus equipped with a plasma-magnetron sputtering head and a rotary-planetary stage, and follow standard protocols provided by the manufacturer. Here, the apparatus was evacuated to 2.5 x 10-5 mbar, purged 1x with high-quality argon, and then adjusted to 8.0 x 10-3 mbar.
- Perform pre-sputtering (120 mA for 60 s) to remove the oxide layer at the surface. Then, deposit 1.5 nm of tungsten (90 mA, working distance = 50 mm) with the aid of a film thickness monitor.
NOTE: The coater must be perfectly cleaned to ensure the reproducibility of the film evaporation. The coating must be stopped once the targeted thickness has been reached. The final film thickness is calculated and corrected afterward. Films of about 1.5 nm have, on average, a post-correction of 0.7 nm using our apparatus. As those correction values are close to the target, a series of coatings are performed to assess and then subtract this correction from the target value. - Store the samples under vacuum to protect them from the ambient air until and throughout all the SEM analyses.
NOTE: The nature of the metal used for sputtering matters. Pt, despite being a common material, results in a poor-quality coating with a Pt film thickness adapted to septin filaments (1.5 nm). At high resolutions, a 1.5 nm Pt film lacks cohesivity; the size of the Pt clusters and the septin filaments thereby become similar, leading to misinterpretations during the filament segmentation process33 (see Figure 2A, inset). Tungsten is a good alternative to Pt because it displays a smaller grain size, barely visible at high-resolution SEM (see Figure 2B, inset). Nonetheless, pure tungsten is easily oxidized, leading to strong charging effect artifacts during SEM observation if the procedure detailed in Step 3.3.4. is not strictly followed.
- Attach the coverslip to stubs that will be used for future SEM observations. Use silver paint because of its improved electrical conductivity, as compared with carbon disks. Add a strip of silver paint to the upper face of the coverslip and make sure that the connection with the stub is satisfactory. Avoid any paint agglomerates.
- Preparation of reagents and stock solutions
- Acquire the images using a field-emission SEM (FESEM) microscope.
NOTE: SEM technologies have recently been upgraded to improve resolution, and the technologies depend on the manufacturer (e.g., electron optics, beam-deceleration, magnetic lens). This gain in resolution (accessible with several manufacturers), especially at a low accelerating voltage (near the nanometer at 1 kV), is necessary to resolve nanometric structures similar to septin networks.- Achieve high-resolution imaging through the detection of primary secondary electrons (SE1) with the "in-lens" detector using the following settings.
- Set the accelerating voltage to 3 kV. Fix the beam current with the 20 μm aperture (for Zeiss Gemini I columns, or equivalent to 34 pA) or with the 15 μm aperture (for Zeiss Gemini I columns, or equivalent to 18.5 pA), if the suppression of charging effects is needed.
- For observations, use resolutions ranging from 21.25 nm/pixel to 1.224 nm/pixel, and for data analysis, use a resolution of ~5.58 nm/pixel (20,000x magnification according to the Polaroid 545 reference).
- Set the working distance between 1 mm and 2 mm for high-resolution observation, and around 3 mm if an increased depth of field is required. Adjust the scan speed and line integration continuously to ensure a constant signal-to-noise ratio with an acquisition time of around 30-45 s per image.
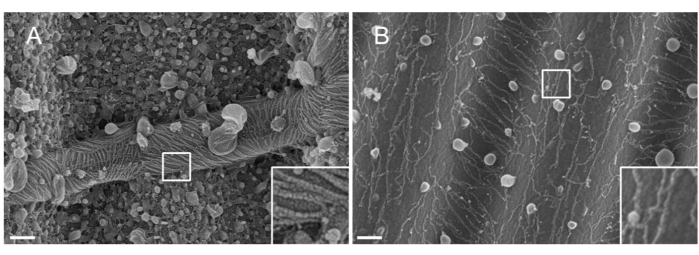
Figure 2: Effect of the material deposited on septin filaments on wavy PDMS patterns. SEM of septin filaments coated by sputtering with either (A) 1.5 nm of platinum, exhibiting the "arid cracked soil" pattern typical of a lack of cohesivity between the clusters of platinum nuclei, or (B) 1.5 nm of tungsten covered with a smooth and cohesive layer. Scale bar = 200 nm. White square boxes represent the magnified views on the lower right. The spherical globules are small lipid vesicles interacting with the septins. Please click here to view a larger version of this figure.
Representative Results
GUVs deformations
Typical confocal fluorescence images of GUVs reshaped after being incubated with septins are displayed in Figure 3, in conditions where septins polymerize. Bare GUVs (Figure 3A) were perfectly spherical. Upon incubation with more than 50 nM budding yeast septin filaments, the vesicles appeared deformed. Up to a concentration of 100 nM budding yeast septin octamers, the vesicles appeared facetted, and the deformations remained static and thus did not fluctuate (Figure 3B). Above 200 nM budding yeast septins, with the membrane being saturated with septins, periodic deformations were observed (Figure 3C). Using budding yeast septins, spikes were visualized with a periodicity varying from 2 µm to 6 µm and with an amplitude of about 1 µm. Hence, septins strongly reshape membranes in a specific fashion. These amplitudes and associated curvatures of the observed deformations strikingly reflect the affinity of septins for micrometer curvatures observed in cells. GUVs, being deformable, are thus appropriate to assay how proteins reshape membranes.
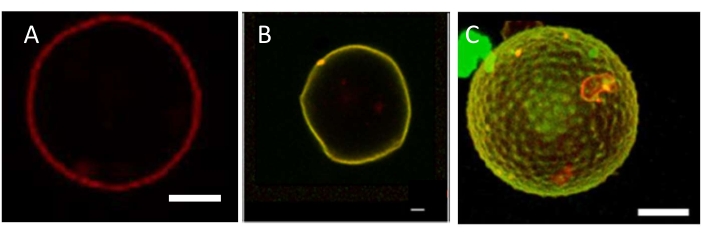
Figure 3: Septin-induced deformation of GUVs. (A) Equatorial plane of control GUVs labeled with 0.5% rhodamine PE and imaged by confocal microscopy. Scale bar = 3 µm. (B) GFP-septins (100 nM) bound to a deformed GUV and imaged by fluorescence confocal microscopy. Scale bar = 1 µm. (C) GFP-septins (600 nM) bound to a deformed GUV and imaged by fluorescence spinning disk microscopy. Scale bar = 10 µm. The GUVs are made of a molar ratio of 56.8% EggPC, 15% cholesterol, 10% DOPE, 10% DOPS, 8% brain PI(4,5)P2, and 0.2% Bodipy-TR-Ceramide. The observation buffer includes 75 mM NaCl and 10 mM Tris (pH = 7.8). Please click here to view a larger version of this figure.
Self-assembly of septin filaments bound to LUVs imaged by cryo-EM
As compared with fluorescence microscopy, cryo-EM provides better spatial resolution. Individual septin filaments and lipid bilayers can thus be discerned unambiguously in images. Figure 4A displays an image of LUVs decorated with budding yeast septin filaments (some of them being highlighted in blue) at 50 nM. Parallel sets of filaments, about 10 nm apart, were observed on the LUVs. This typical image is a 2D projection. Hence, to decipher any 3D deformation and to know whether septin filaments directly interact with the membrane, cryo-electron tomography was carried out (Figure 4B and Figure 4C). Figure 4B displays a slice in a 3D reconstruction, while Figure 4C presents a segmentation of the same area highlighting the membrane in yellow and the septin filaments in blue. As seen on the side view, septin filaments remained essentially straight bound to the liposome and the vesicle was considerably flattened as compared with the naked spherical vesicles.
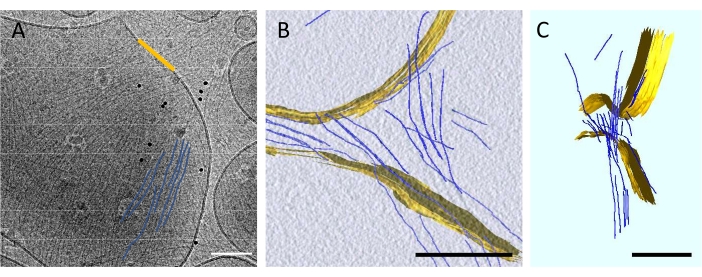
Figure 4: Self-assembly of septin filaments bound to LUVs. (A) Cryo-EM image of septin filaments bound to a vesicle. Some of the filaments are highlighted in blue for visibility. The dark dots are gold fiducials usually used to align data for tomography. Scale bar = 100 nm. (B) and (C) 3D reconstruction obtained from a tilted series. The lipids are highlighted in yellow, while the septins are segmented in blue. Scale bars = 200 nm. Please click here to view a larger version of this figure.
Curvature-dependent arrangement of septin filaments visualized by SEM
Septins localize in cells at sites displaying micrometer curvatures (e.g., at the cell division furrow, the base of cilia, etc.). In addition, in vitro studies show that septins, as seen above, can reshape membranes so they eventually display micrometer curvatures. To assay the curvature sensitivity of septins for both positive (convex) and negative (concave) curvatures, septins were incubated with a supported lipid bilayer fused with the wavy patterns described above. The outcome is displayed in Figure 5. Figure 5A recapitulates the different steps required to obtain the samples. Figure 5B presents a wavy pattern at low magnification, where the periodicity of negative and positive curvatures is clearly visible. Two successive waves are indicated by orange solid lines. Defects approximately orthogonal to the waves are indicated by white arrows. Figure 5C and Figure 5D display the ultrastructural organization of septin filaments at higher magnification. The septins assembled into sets of parallel filaments, whose orientation depended on the curvature (see filaments highlighted in blue). Indeed, the septin filaments were not oriented randomly. Instead, septins can bend when interacting with negative (concave) curvatures to follow the imposed curvature. Conversely, on positive curvatures, the septin filaments remained straight, unbent, and aligned along the convex waves. Hence, septins can interact with both positive and negative curvatures but adopt specific organizations on given curvatures. This methodology thus appears particularly relevant to highlight the curvature-sensitivity of binding and self-assembly of filamentous proteins.
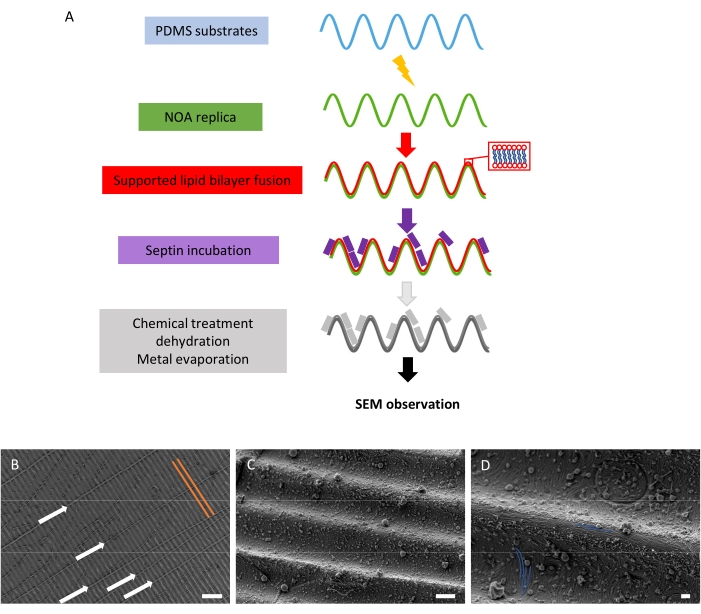
Figure 5: Curvature-dependent arrangement of septin filaments. (A) Schematic representation of the steps required to obtain the "wavy" samples meant to analyze septin curvature sensitivity through SEM. (B) Low magnification SEM image of a wavy pattern with bound septin filaments that are not resolved at this resolution. Two consecutive waves are highlighted in orange. Orthogonal defects are indicated by white arrows. Scale bar = 10 µm. (C) SEM image at 10,000x magnification where septin filaments are visible. Scale bar = 1 µm. (D) SEM image at 20,000x magnification. Some of the filaments are highlighted in blue for visibility. Scale bar = 200 nm. The spherical objects present in the images are small vesicles interacting with the septins and substrate. The septin concentration was set at 200 nM in B–D. Please click here to view a larger version of this figure.
Discussion
As stated above, a lipid mixture has been used that enhances PI(4,5)P2 incorporation within the lipid bilayer and thus facilitates septin-membrane interactions. Indeed, we have shown elsewhere25 that budding yeast septins interact with vesicles in a PI(4,5)P2-specific fashion. This lipid composition was adjusted empirically from screening multiple compositions and is now widely used by the authors. PI(4,5)P2 lipids have to be handled carefully. Stock solutions must be aliquoted in small volumes so that a specific vial is not opened more than twice for pipetting. Besides, lipid aliquots should be stored under argon to prevent lipid oxidation. Finally, once the lipid mixes are solubilized in an aqueous solution, the mixture must be used at once and cannot be stored for further experiments. Notably, after GUV reconstitution or supported lipid bilayer fusion, the interaction between septins and lipids greatly reduced after 2 h.
Some of the results obtained with octameric budding yeast septin complexes (enclosing Cdc10, Cdc11, Cdc12, and Cdc3) have been presented here; however, the experimental procedures followed to analyze the behavior of septin complexes from other species should be identical.
To generate GUVs, we prefer the electro-formation method using platinum wires over electro-formation on ITO plates or PVA swelling. As demonstrated earlier25, this method reproducibly produces unilamellar GUVs of good quality without any defects.
The wavy substrates can be designed with a wide range of curvatures. After screening various designs, patterns with a periodicity of 2 µm and an amplitude of 250 nm, which generate curvatures ranging from -3.5 µm-1 to 3.5 µm-1 (± 0.5 um-1), have been consistently used. Using patterns with a short periodicity and higher amplitude, the lipid bilayers did not nicely conform to the surface. Often, in the case of higher curvature, the bilayer would be suspended in between two convex waves instead of being fully supported on the NOA substrate. With lower amplitudes and larger periodicity, septins organized with random nematic-like orientations, indicating that the patterns were too flat. Besides, using wavy patterns instead of cylindrical tubes20 or spheres20 has the benefit that both positive and negative curvatures can be tested simultaneously. In the future, examining the behavior of filaments on gaussian "saddle-like" curvatures would be relevant to mimic the curvature of more realistic cellular contexts.
In solution, mixing LUVs with septins induced the generation of septin-lipid aggregates even at low concentrations of proteins and lipids. The samples thus were heterogeneous, and it is wise to look for thinner ice domains where smaller and more defined septin-vesicle objects are found. This is particularly crucial when choosing areas of interest where cryo-tomography will be performed. Often, even at low magnifications, vesicles displaying protrusions of strong deformations, as compared with spherical vesicles, are more likely to have a high density of septin filaments bound. This protocol focuses on obtaining a description of the global ultrastructure of filaments on biomimetic membranes. Currently, higher resolutions are reachable. The resolution limitation results from the limited number of views available by imaging. However, this remains beyond the scope of this protocol.
We have shown through this protocol that a combination of complementary methods is essential to understand and dissect how the specific arrangement of cytoskeletal septin filaments can sense and reshape membranes in a curvature-dependent manner.
Disclosures
The authors have nothing to disclose.
Acknowledgements
We thank Patricia Bassereau and Daniel Lévy for their useful advice and discussions. This work benefited from the support of the ANR (Agence Nationale de la Recherche) for funding the project "SEPTIME", ANR-13-JSV8-0002-01, ANR SEPTIMORF ANR-17-CE13-0014, and the project "SEPTSCORT", ANR-20-CE11-0014-01. B. Chauvin is funded by the Ecole Doctorale "ED564: Physique en Ile de France" and Fondation pour lea Recherche Médicale. K. Nakazawa was supported by Sorbonne Université (AAP Emergence). G.H. Koenderink was supported by the Nederlandse Organisatie voor Wetenschappelijk Onderzoek (NWO/OCW) through the ‘BaSyC-Building a Synthetic Cell'. Gravitation grant (024.003.019).We thank the Labex Cell(n)Scale (ANR-11-LABX0038) and Paris Sciences et Lettres (ANR-10-IDEX-0001-02). We thank the Cell and Tissue Imaging (PICT-IBiSA), Institut Curie, member of the French National Research Infrastructure France-BioImaging (ANR10-INBS-04).
Materials
| 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine | Avanti Polar Lipids | 850725 | |
| 1,2-dioleoyl-sn-glycero-3-phospho-L-serine | Avanti Polar Lipids | 840035 | |
| Bath sonicator | Elma | Elmasonic S10H | |
| Bodipy-TR-Ceramide | invitrogen, Thermo Fischer scientific | 11504726 | |
| Chemicals: NaCl, Tris-HCl, sucrose, KCl, MgCl2, B-casein, chloroform, sodium cacodylate, tannic acid, ethanol | Sigma Aldrich | ||
| Confocal microscope | nikon | spinning disk or confocal | |
| Critical point dryer | Leica microsystems | CPD300 | |
| Deionized water generator | MilliQ | F1CA38083B | MilliQ integral 3 |
| Egg L-α-phosphatidylcholine | Avanti Polar Lipids | 840051 | |
| Field Emission Gun SEM (FESEM) | Carl Zeiss | Gemini SEM500 | |
| Glutaraldehyde 25 %, aqueous solution | Thermo Fischer scientific | 50-262-19 | |
| High vacuum grease, Dow corning | VWR | ||
| IMOD software | https://bio3d.colorado.edu/imod/ | software suite for tilted series image alignment and 3D reconstruction | |
| Lacey Formvar/carbon electron microscopy grids | Eloise | 01883-F | |
| Lipids | Avanti Polar Lipids | ||
| L-α-phosphatidylinositol-4,5-bisphosphate | Avanti Polar Lipids | 840046 | |
| Metal evaporator | Leica microsystems | EM ACE600 | |
| NOA (Norland Optical Adhesives), NOA 71 and NOA 81 | Norland Products | NOA71, NOA81 | |
| Osmium tetraoxyde 4% | delta microscopies | 19170 | |
| Osmometer | Löser | 15 M | |
| Plasma cleaner | Alcatel | pascal 2005 SD | |
| Plasma generator | Electron Microscopy Science | ||
| Plunge freezing equipment | leica microsystems | EMGP | |
| Transmission electron microscope | Thermofischer | Tecnai G2 200 kV, LaB6 | |
| Uranyl acetate | Electron Microscopy Science | 22451 | this product is not available for purchase any longer |
| Wax plates, Vitrex | VWR |
References
- Finger, F. P. Reining in cytokinesis with a septin corral. BioEssays: News and Reviews in Molecular, Cellular and Developmental Biology. 27 (1), 5-8 (2005).
- Barral, Y., Kinoshita, M. Structural insights shed light onto septin assemblies and function. Current Opinion in Cell Biology. 20 (1), 12-18 (2008).
- Hu, Q., et al. A septin diffusion barrier at the base of the primary cilium maintains ciliary membrane protein distribution. Science. 329 (5990), 436-439 (2010).
- Lin, Y. -. H., Kuo, Y. -. C., Chiang, H. -. S., Kuo, P. -. L. The role of the septin family in spermiogenesis. Spermatogenesis. 1 (4), 298-302 (2011).
- Addi, C., Bai, J., Echard, A. Actin, microtubule, septin and ESCRT filament remodeling during late steps of cytokinesis. Current Opinion in Cell Biology. 50, 27-34 (2018).
- Spiliotis, E. T., Kesisova, I. A. Spatial regulation of microtubule-dependent transport by septin GTPases. Trends in Cell Biology. 31 (12), 979-993 (2021).
- Spiliotis, E. T., Nakos, K. Cellular functions of actin- and microtubule-associated septins. Current Biology: CB. 31 (10), 651-666 (2021).
- Salameh, J., Cantaloube, I., Benoit, B., Poüs, C., Baillet, A. Cdc42 and its BORG2 and BORG3 effectors control the subcellular localization of septins between actin stress fibers and microtubules. Current Biology: CB. 31 (18), 4088-4103 (2021).
- Ewers, H., Tada, T., Petersen, J. D., Racz, B., Sheng, M., Choquet, D. A septin-dependent diffusion barrier at dendritic spine necks. PloS One. 9 (12), 113916 (2014).
- Myles, D. G., Primakoff, P., Koppel, D. E. A localized surface protein of guinea pig sperm exhibits free diffusion in its domain. The Journal of Cell Biology. 98 (5), 1905-1909 (1984).
- Luedeke, C., Frei, S. B., Sbalzarini, I., Schwarz, H., Spang, A., Barral, Y. Septin-dependent compartmentalization of the endoplasmic reticulum during yeast polarized growth. The Journal of Cell Biology. 169 (6), 897-908 (2005).
- Gilden, J. K., Peck, S., Chen, Y. -. C. M., Krummel, M. F. The septin cytoskeleton facilitates membrane retraction during motility and blebbing. The Journal of Cell Biology. 196 (1), 103-114 (2012).
- Dolat, L., Hu, Q., Spiliotis, E. T. Septin functions in organ system physiology and pathology. Biological Chemistry. 395 (2), 123-141 (2014).
- Angelis, D., Spiliotis, E. T. Septin mutations in human cancers. Frontiers in Cell and Developmental Biology. 4, 122 (2016).
- Takehashi, M., et al. Septin 3 gene polymorphism in Alzheimer’s disease. Gene Expression. 11 (5-6), 263-270 (2004).
- Shuman, B., Momany, M. Septins from protists to people. Frontiers in Cell and Developmental Biology. 9, 824850 (2022).
- Bertin, A., et al. Saccharomyces cerevisiae septins: supramolecular organization of heterooligomers and the mechanism of filament assembly. Proceedings of the National Academy of Sciences of the United States of America. 105 (24), 8274-8279 (2008).
- Iv, F., et al. Insights into animal septins using recombinant human septin octamers with distinct SEPT9 isoforms. Journal of cell science. 134 (15), (2021).
- Beber, A., et al. Membrane reshaping by micrometric curvature sensitive septin filaments. Nature communications. 10 (1), 420 (2019).
- Bridges, A. A., Jentzsch, M. S., Oakes, P. W., Occhipinti, P., Gladfelter, A. S. Micron-scale plasma membrane curvature is recognized by the septin cytoskeleton. The Journal of Cell Biology. 213 (1), 23-32 (2016).
- Patzig, J., et al. Septin/anillin filaments scaffold central nervous system myelin to accelerate nerve conduction. eLife. 5, 17119 (2016).
- Szuba, A., et al. Membrane binding controls ordered self-assembly of animal septins. eLife. 10, 63349 (2021).
- Tanaka-Takiguchi, Y., Kinoshita, M., Takiguchi, K. Septin-mediated uniform bracing of phospholipid membranes. Current Biology: CB. 19 (2), 140-145 (2009).
- Bertin, A., et al. Phosphatidylinositol-4,5-bisphosphate promotes budding yeast septin filament assembly and organization. Journal of Molecular Biology. 404 (4), 711-731 (2010).
- Beber, A., et al. Septin-based readout of PI(4,5)P2 incorporation into membranes of giant unilamellar vesicles. Cytoskeleton. 76 (4,5), 92-103 (2019).
- Mastronarde, D. N., Held, S. R. Automated tilt series alignment and tomographic reconstruction in IMOD. Journal of Structural Biology. 197 (2), 102-113 (2017).
- Kremer, J. R., Mastronarde, D. N., McIntosh, J. R. Computer visualization of three-dimensional image data using IMOD. Journal of Structural Biology. 116 (1), 71-76 (1996).
- Nania, M., Foglia, F., Matar, O. K., Cabral, J. T. Sub-100 nm wrinkling of polydimethylsiloxane by double frontal oxidation. Nanoscale. 9 (5), 2030-2037 (2017).
- Nania, M., Matar, O. K., Cabral, J. T. Frontal vitrification of PDMS using air plasma and consequences for surface wrinkling. Soft Matter. 11 (15), 3067-3075 (2015).
- Svitkina, T. M., Borisy, G. G. Correlative light and electron microscopy of the cytoskeleton of cultured cells. Methods in Enzymology. 298, 570-592 (1998).
- Franck, A., et al. Clathrin plaques and associated actin anchor intermediate filaments in skeletal muscle. Molecular Biology of the Cell. 30 (5), 579-590 (2019).
- Elkhatib, N., et al. Tubular clathrin/AP-2 lattices pinch collagen fibers to support 3D cell migration. Science. 356 (6343), (2017).
- Stokroos, I., Kalicharan, D., Van Der Want, J. J., Jongebloed, W. L. A comparative study of thin coatings of Au/Pd, Pt and Cr produced by magnetron sputtering for FE-SEM. Journal of Microscopy. 189, 79-89 (1998).

