Sandwich-like Microenvironments to Harness Cell/Material Interactions
Summary
The following protocol describes the procedure to assemble sandwich-like cultures to be used as an intermediate stage between bi-dimensional (2D) and three-dimensional (3D) cellular environments. The engineered systems can have applications in microscopy, biomechanics, biochemistry and cell biology assays.
Abstract
Cell culture has been traditionally carried out on bi-dimensional (2D) substrates where cells adhere using ventral receptors to the biomaterial surface. However in vivo, most of the cells are completely surrounded by the extracellular matrix (ECM), resulting in a three-dimensional (3D) distribution of receptors. This may trigger differences in the outside-in signaling pathways and thus in cell behavior.
This article shows that stimulating the dorsal receptors of cells already adhered to a 2D substrate by overlaying a film of a new material (a sandwich-like culture) triggers important changes with respect to standard 2D cultures. Furthermore, the simultaneous excitation of ventral and dorsal receptors shifts cell behavior closer to that found in 3D environments. Additionally, due to the nature of the system, a sandwich-like culture is a versatile tool that allows the study of different parameters in cell/material interactions, e.g., topography, stiffness and different protein coatings at both the ventral and dorsal sides. Finally, since sandwich-like cultures are based on 2D substrates, several analysis procedures already developed for standard 2D cultures can be used normally, overcoming more complex procedures needed for 3D systems.
Introduction
Traditionally, cell culture has been carried out on bi-dimensional (2D) substrates, though most of the in vivo cellular microenvironments have a three-dimensional (3D) nature. This unnatural 2D environment triggers changes in cell behavior as a way of self-adaptation to a flat world, which directly impacts cell fate1,2. Hence, results obtained on 2D cell cultures are not always reproducible in vivo. This has encouraged the development of new relevant culture systems seeking to provide more physiological-like conditions to get further insights into any dimension-dependent biological mechanism3,4.
One of the main differences between 2D culture and the 3D in vivo environment is the distribution of cell receptors anchored to the extracellular matrix (ECM): whereas on 2D substrates cells adhere ventrally, the majority of cells in vivo are completely surrounded by the ECM and thus cell adhesion occurs through a 3D distribution of receptors. This triggers different cell adhesion signaling pathways thereby modulating important processes such as cell growth, cell differentiation and gene expression. During the last decades, many different 3D culture systems have been established5-8, though their variability and complexity hinder their standardization in common cell culture procedures. Moreover 3D systems are usually not easy to handle and current experimental procedures on 2D substrates cannot be easily established for 3D cultures. In addition, literature rarely compares 3D cultures with the equivalent 2D condition or other 3D systems, hindering the proper understanding of cell behavior in these models.
Once having the cells adhered on a 2D substrate, the excitation of the dorsal receptors — by overlaying a film of a new material (sandwich-like culture) — can trigger cell responses alike 3D environments. The reason behind this is the simultaneous activation of both dorsal and ventral receptors to adhere and spread within the sandwich environment (Figure 1)9,10. As a consequence, cells undergo important changes with respect to 2D cultures11,12. Thus, cell fate is determined during assembly because of the sandwich culture, since the dorsal stimulation triggers changes in key cellular pathways. Therefore, the cell fate is highly determined by the time when the sandwich-like culture is assembled11.
Due to the nature of the system, a sandwich-like culture is a simple and versatile tool that allows the study of different parameters in cell/material interactions such as chemistry, topography, stiffness and protein coatings at both the ventral and dorsal sides. This offers a higher degree of versatility compared to other 3D systems (Figure 2) due to the independent dorsal and ventral combination of a wide variety of surface conditions. Additionally, different cell lines and different times to assemble the sandwich-like culture can be studied, increasing the wide spectra of possibilities.
A standard protocol of the sandwich-like culture is detailed below using either poly-L-lactic acid (PLLA) electrospun fibers or films as dorsal substrates, glass coverslip as ventral substrate and fibronectin as protein coating. Sandwich-like cultures were assembled just after cell seeding or after 3 hr of 2D culture. However, note that other material systems and proteins could be used; likewise the sandwich-like culture can be assembled at different time points.
Protocol
1. Production of Dorsal Substrates
- Production of a flat film of PLLA by solvent casting.
- Work in a fume hood to prepare a solution of 2% (w/v) PLLA in chloroform (2 g PLLA in 100 ml chloroform) by stirring at room temperature (RT) until completely dissolved (3 hr approximately).
- Place the washers on a glass Petri dish.
Note: Use stainless-steel washers with an inner diameter of 10.5 mm (slightly smaller than the ventral sample diameter) so that the washer is placed on top of the ventral substrate. Other materials such as polytetrafluoroethylene (PTFE) or glass washers could be used as well. - Add 200 µl of the PLLA solution in the washer and let evaporate slowly for 30 min at RT. In order to allow slow evaporation of the solvent, close the Petri dish with aluminum foil with a few holes.
- Heat the samples at 120 °C for 5 min in order to evaporate solvent traces.
- Let samples cool down at RT.
- Once samples have cooled down (30 min approximately), cover with 18.2 MΩ·cm water in the Petri dish and incubate for 8 min at RT.
Note: If samples are not completely cooled down, they may take up 18.2 MΩ·cm water and adopt a rubbery state. Additionally, incubating in 18.2 MΩ·cm water for less than 8 min may result in improper detachment, and for more than 8 min in detachment from the washer. - With a razor blade, pry the washers to peel them off the Petri dish.
- Remove any imperfections from the edge of the washer with tweezers and let the samples air-dry.
- Store the samples in a vacuum desiccator (80 mbar) until the day of the experiment because PLLA is degradable by hydrolysis.
- Production of PLLA fibers by electrospinning.
- Work in a fume hood to prepare a solution of 8% (w/v) PLLA in hexafluoroisopropanol (8 g PLLA in 100 ml hexafluoroisopropanol) by stirring at RT till completely dissolved (3 hr approximately).
- Place the polytetrafluoroethylene (PTFE) washers on an aluminum sheet or on a drum for the electrospinning in order to get random or aligned fibers respectively.
Note: Use polytetrafluoroethylene (PTFE) washers with an inner diameter of 10.5 mm (slightly smaller than the ventral sample diameter) so that the washer is placed on top of the ventral substrate. Other materials such as stainless steel or glass washers could be used as well. The drum should rotate at an angular velocity of 160.7 sec-1 (radius = 7 cm) in order to obtain aligned fibers. A slower speed might not trigger fiber alignment and a faster speed will result in broken fibers. - Load the syringe with the polymer solution and place it on the syringe pump.
- Electrospin the PLLA solution using a 0.9 ml/hr flow rate with a voltage of 30 kV and a collector distance of 12 cm.
- Stop the electrospinning process after 20-30 min, once a good fiber density is achieved (since cells have to interact with several fibers simultaneously).
- Heat the samples at 120 °C for 5 min in order to evaporate solvent traces.
- Store the samples in a vacuum desiccator (80 mbar) until the day of the experiment because PLLA is degradable by hydrolysis.
2. Sandwich Culture
- Assemble the sandwich-like culture once cells are adhered to the ventral 2D substrate.
- Work in a culture hood to ensure sterile conditions and sterilize all materials (glass coverslips, dorsal substrates, tweezers, etc.) by UV exposure for 30 min.
- Coat the ventral and dorsal substrate with fibronectin in order to direct specific cell-protein adhesion. To do this, incubate the samples in 20 µg/ml of fibronectin dissolved in Dulbecco's Phosphate-Buffered Saline (DPBS) containing Ca2+ and Mg2+ (200 µl/sample) for 1 hr.
Note: Due to the fabrication process, dorsal substrates have a designated top and bottom sides because the solvent casted film and electrospun fibers attached to one of the sides of the washer. This side is therefore the one facing the cells (Figure 3).- After the protein coating, wash the samples twice with DPBS in order to remove protein in excess. Then incubate samples in DPBS until their use in order to prevent samples from getting dry since this causes protein denaturalization. Use DPBS containing Ca2+ and Mg2+ since divalent cations regulate protein folding.
- Seed cells on the glass coverslips13.
Note: As the sandwich-like culture is based on 2D substrates, cell seeding will be performed similarly as on a 2D sample. For example, C2C12 myoblast are seeded at 17,500 cells cm-1 for differentiation experiments and NIH3T3 fibroblast are seeded at 7,000 cells cm-1 for isolated cultures to study cell morphology and adhesion. - Place the samples in an incubator at 37 °C with 5% CO2 and allow cells to adhere to the ventral substrate for 3 hr.
- Place the ventral seeded substrate in a new well of a 12 multi-well plate (where washers fit).
Note: Assembling the sandwich-like culture in a new well is one way to get rid of the cells that have adhered to the bottom of the well after the seeding since these will consume nutrients and secrete growth factors and waste that influence cells cultured within the sandwich-like system. This is also a must for cellular extractions in order to lyse only the cells cultured within the sandwich-like environment. - Carefully and with the help of a pair of tweezers, overlay the dorsal substrate on the cells. Refill with medium (1 ml) and incubate at 37 °C with 5% CO2.
Note: The weight of the washer will prevent the dorsal substrate from floating. - Change the medium twice a week as in standard 2D cultures. Do not move the dorsal substrate since this could result in cell damage.
- Assemble the sandwich-like culture just after cell seeding.
- Work in a culture hood to ensure sterile conditions and sterilize all materials (glass coverslips, dorsal substrates, tweezers, etc.) by UV exposure for 30 min.
- Coat the ventral and dorsal substrate with fibronectin in order to direct specific cell-protein adhesion. To do this, incubate the samples in 20 µg/ml of fibronectin dissolved in DPBS containing Ca2+ and Mg2+ (200 µl/sample) for 1 hr.
- After the protein coating, wash the samples twice with DPBS in order to remove protein in excess. Then incubate samples in DPBS until their use in order to prevent samples from getting dry since this causes protein denaturalization. Use DPBS containing Ca2+ and Mg2+ since divalent cations regulate protein folding.
- Seed cells on glass coverslips in a well of a 12 multi-well plate (where washers fit). To do this, use a highly concentrated cellular suspension to avoid cell loss after laying the dorsal substrate.
- Carefully and with the help of a pair of tweezers, overlay the dorsal substrate on the cells. Refill with medium (1 ml) and incubate at 37 °C with 5% CO2.
Note: The weight of the washer will prevent the dorsal substrate from floating. - Change the medium twice a week as in standard 2D cultures.
Note: Do not move the dorsal substrate since this could result in cell damage.
3. Analysis
Note: Sandwich-like cultures are based on 2D substrates, and so can be commonly analyzed by procedures already developed for standard 2D cultures. For example, since PLLA is transparent and cells are constrained to move within the x-y plane, microscopy is done as on 2D substrates. Cell migration can be therefore analyzed as for 2D cultures, without the need of tracking cells in the z-axis as for 3D cultures, which simplifies the experiment and image analysis. To study the wound healing assay by a scratch assay follow this protocol:
- Study cell migration (wound healing assay) within the sandwich-like culture.
- Work in a culture hood to ensure sterile conditions and sterilize all materials (glass coverslips, dorsal substrates, tweezers, etc.) by UV exposure for 30 min.
- Coat the ventral and dorsal substrate with fibronectin in order to direct specific cell-protein adhesion. To do this, incubate the samples in 20 µg/ml of fibronectin dissolved in Dulbecco's Phosphate-Buffered Saline (DPBS) containing Ca2+ and Mg2+ for 1 hr.
- After the protein coating, wash the samples twice with DPBS in order to remove protein in excess. Then incubate samples in DPBS until their use in order to prevent samples from getting dry since this could cause protein denaturalization. Use DPBS containing Ca2+ and Mg2+ since divalent cations regulate protein folding.
- Seed cells on the glass coverslips at a high density13.
- Place the samples in an incubator at 37 °C with 5% CO2 and allow cells to achieve confluence.
- Use a pipette tip to scratch the cell monolayer in order to get 2 cell populations separated by a gap.
- Place the ventral seeded substrate in a new well suitable for the time-lapse acquisition and where dorsal substrates fit (i.e. 12 multi-well plate).
- Carefully and with the help of a pair of tweezers, overlay the dorsal substrate on the cells and then refill with medium (1 ml for a 12 multi-well plate).
Note: The weight of the washer will prevent the dorsal substrate from floating. - Place the sample in the time-lapse microscope (set to 37 °C and 5% CO2) and take images of the gap closure every 20 min.
- Analyze the gap closure using specific software such as the plugin MiToBo from ImageJ.14
Note: Protein and nucleic acid extraction is performed similarly as on 2D substrates. There is only one extra step that consists in disassembling the sandwich-like culture to add the lysis buffer directly on the cells in order to increase the extraction efficiency. For instance, for mRNA extraction:
- Extract mRNA from Sandwich-like Cultures
- Prepare all the buffers from the commercial kit according to the manufacturer's instructions.
- Take the sandwich-like culture out from the incubator.
- Remove the culture media from the samples.
- Wash the samples once with DPBS containing Ca2+ and Mg2+ (1 ml) and remove all the solution.
- With the help of a pair of tweezers, remove the dorsal substrate and add the lysis buffer (350 µl) to the ventral substrate. Overlay again the dorsal substrate on the ventral substrate in order to lyse also the cells adhered to the dorsal substrate.
- Remove the dorsal substrate to isolate the lysis solution.
- Follow manufacturer's instructions to get the mRNA.
Note: Immunodetection of proteins can be also performed as on 2D substrates. Since sandwich-like cultures could hinder the correct diffusion of the antibodies and buffers, incubation times should be increased. Also, the sandwich can be disassembled before starting the staining protocol but in this latter case some cells will remain attached to the dorsal substrate and some to the ventral substrate.
- Immunodetect proteins within a sandwich-like culture
- Wash the sample once with DPBS and fix with 4% formaldehyde in PBS for 20 min at 4 °C.
Note: Dorsal substrate can be removed during the fixation process so that diffusion problems are avoided. - Rinse with DPBS (1 ml) and then permeabilize with 0.5% Triton X-100 in DPBS (1 ml) for 5 min at RT.
- Block non-specific binding sites with 1% albumin in DPBS (1 ml) for 30 min at RT.
- Incubate with primary antibody (2 µg/ml; 1 ml) in blocking solution for 3 hr at RT.
- Wash 3 times for 10 min with DPBS/0.5% Tween 20 (1 ml).
- Incubate 2 hr with secondary antibodies (2 µg/ml) and 1 µg/ml DAPI in blocking solution (1 ml) at RT.
- Wash 3 times for 10 min in DPBS/0.5% Tween 20 (1 ml).
- Wash in DPBS.
Note: Samples can be mounted on a slide with Vectashield and sealed with nail polish when dorsal substrates are removed prior the immunodetection. - Observe under the fluorescence microscope as for standard 2D substrates.
- Wash the sample once with DPBS and fix with 4% formaldehyde in PBS for 20 min at 4 °C.
Representative Results
The stimulation of dorsal receptors within the sandwich-like culture triggers changes in cell morphology, cell adhesion and intracellular signaling pathways (e.g. focal adhesion kinase, FAK)10-12. As an example, fibroblasts cultured within the sandwich-like system overexpressed the α5 integrin subunit compared to the 2D, as observed for other 3D cultures15,16.
Cell fate is highly dependent on the time when the dorsal receptors are stimulated and by the properties of the dorsal interaction, similarly as happens in other 3D systems such as hydrogels. For example, hydrogels where proteins are tightly bound usually show smaller and rounded cells with undeveloped actin cytoskeleton and diffuse focal adhesions. This can be mimicked in a sandwich-like culture using substrates that adsorb proteins tightly, so that cells are not able to mechanically reorganize this layer of proteins and cell spreading is hindered. Similarly, hydrogels where cells can remodel the ECM can be mimicked with the sandwich system by using substrates that adsorb proteins more loosely10.
The stimulation of the dorsal receptors has been shown to modulate C2C12 cell fate. Dorsal electrospun PLLA fibers direct cell alignment when coated with fibronectin but not when coated with bovine serum albumin (a non-adhesive protein). This result points out cells do biologically sense and react to the dorsal inputs (Figure 4)9. Additionally, sandwich-like cultures with plane dorsal PLLA triggered an increase in the level of myogenesis. This depends also on the dorsal biological stimuli since the interaction with different dorsal proteins results in distinct differentiation rates (Figure 5)9.
Cell migration within the sandwich-like culture is also altered when compared to the 2D culture. It has been shown that in a wound-healing assay, cells within the sandwich culture adopt a highly elongated morphology and migrate shorter distances than on 2D substrates (Movies 1 and 2). Cell migration rates are furthermore related to the nature of the ventral and dorsal stimulation12.
Similarly as within 3D fibronectin and collagen gels17,18, sandwich-like culture increases cell-mediated ECM reorganization (i.e. the ventral fibronectin) in respect to the 2D condition (Figure 6)12. This process relies on the dorsal mechanical stimuli and cytoskeleton stability since the use of contractility inhibitors (blebbistatin, Y27632) hindered the process. Interestingly, the ventral fibronectin was also reorganized when using different dorsal protein coatings (i.e. vitronectin and bovine serum albumin) and even if left uncoated12.
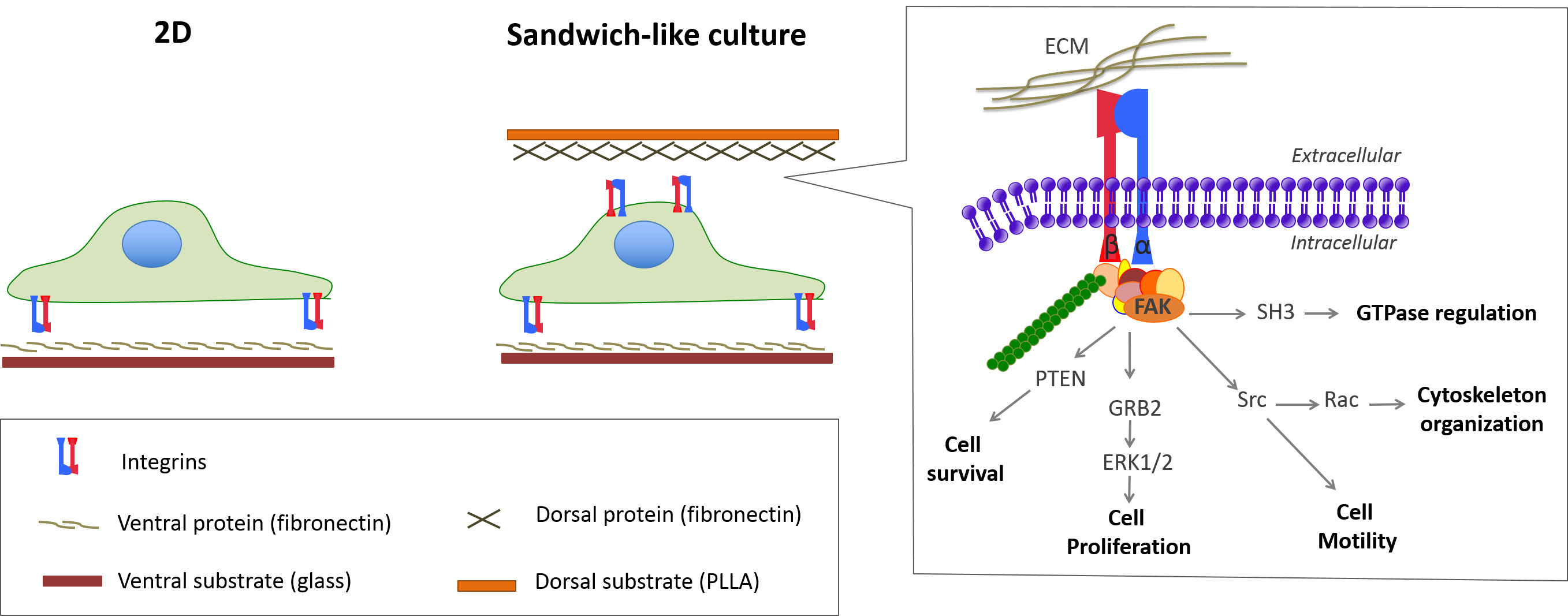
Figure 1: Sketch of standard (2D) and sandwich-like cultures. Stimulation of the dorsal receptors within the sandwich-like culture triggers additional cell adhesion signaling that modulates important cellular processes. Please click here to view a larger version of this figure.
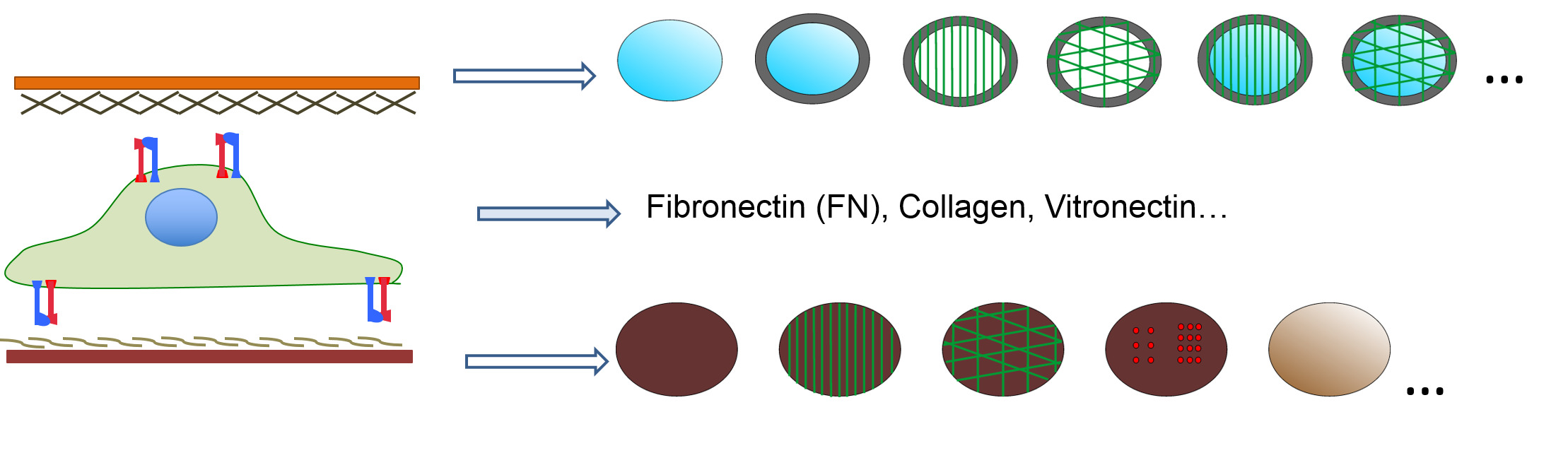
Figure 2: Sandwich-like culture is a versatile system that allows the study of different well-controlled parameters (topographical inputs, stiffness, gradients…) on both the ventral and dorsal substrates.
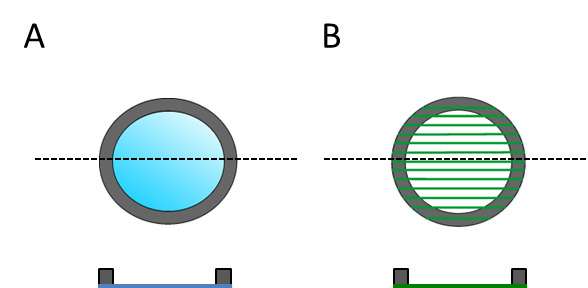
Figure 3: Dorsal substrates are sketched for both the top and cross-section view in order to show that have a designated top and bottom side. (A) Flat PLLA film and (B) electrospun fibers.

Figure 4: C2C12 morphology under different culture conditions including plane (p) and aligned fibers (a) of PLLA that were used as ventral (subscript) or dorsal (superscript) substrates. Dotted lines represent fibers orientation where necessary. Cells cultured on the plane substrate and overlaid with aligned fibers of PLLA (SWpa) sense the dorsal stimuli. Particularly, cells sense the dorsal fibers when coated with fibronectin but not when coated with bovine serum albumin (a non-adhesive protein). Consequently cells adhere to the fibers coated with fibronectin and align in the same direction. Image adapted from reference9. Please click here to view a larger version of this figure.

Figure 5: Cell differentiation in sandwich-like cultures after 4 days in differentiation media. Different culture conditions were analyzed including plane (p) and aligned fibers (a) of PLLA that were used as ventral (subscript) or dorsal (superscript) substrates. Samples were coated with fibronectin in all cases. (A) Fluorescence staining showing sarcomeric myosin positive cells (green) and cell nuclei (red). (B) Differentiated cells orientation as calculated by Fast Fourier Transform. (C) Myogenesis as determined by the percentage of sarcomeric myosin-positive cells. Data is normalized to the gold standard control. Statistically significant differences are indicated with *** P <0.001. Image adapted from reference9. Please click here to view a larger version of this figure.

Figure 6: Reorganization of ventral fibronectin. Sandwich culture triggers ventral FN reorganization by forming new fibronectin fibrils (brush-like labeling; pointed out with white arrows). Ventral fibronectin is reorganized within sandwich-like cultures with different dorsal protein coating (fibronectin, vitronectin and bovine serum albumin) or even when the dorsal substrate is left uncoated. Actin cytoskeleton (green), nuclei (blue) and FN (red) are shown. Please click here to view a larger version of this figure.
Movie 1: Cell migration on 2D. L929 fibroblast migrating on a fibronectin coated glass coverslip in a wound healing assay. Images were acquired for 16 hr (with a frame taken every 20 min). Please click here to view this video.
Movie 2: Cell migration within the sandwich-like culture. L929 fibroblast migrating in a wound healing assay within a sandwich-like culture were the ventral substrate was a fibronectin coated glass coverslip and the dorsal substrate a fibronectin coated PLLA film. Images were acquired for 16 hr (with a frame taken every 20 min). Please click here to view this video.
Discussion
Nowadays, 3D culture is an important topic for the pharmaceutical and biotechnological industry as well as research in cell biology, including cancer and stem cells. As a consequence several 3D culture systems have been developed. Unfortunately, differences between the 3D systems usually result in different cell behavior, hindering the understanding of cell fate. Besides, experimental procedures are usually not as straightforward as for 2D culture systems. Hence developing new culture systems seeking to overcome some of these drawbacks is highly important.
Sandwich-like culture has been shown to strongly influence key cellular processes such as cell differentiation, cell morphology, cell signaling and cell migration. Furthermore cells share similarities with cells cultured in 3D systems, supporting the statement that sandwich-like culture links 2D with 3D culture systems. Physiological tissues have pores in the range of 3 to 14 µm that constraint cells and therefore influence cellular processes such as migration. This is somehow recapitulated using the sandwich-like system as dorsal and ventral stimuli represent per se a constrained environment that limits cell morphology and will necessarily influence cell migration regardless of the protein coatings.
Due to the simultaneous stimulation of dorsal and ventral receptors, sandwich-like culture is a robust technology to investigate the role of dimensionality in cell behavior. Moreover, as it is based on 2D substrates, this culture system is highly versatile to study different material properties and ECM inputs allowing the study of cell behavior under different microenvironments. Besides, the influence of the time at which dorsal receptors are stimulated on cell fate can be investigated by overlaying the dorsal substrate at different time points. Hence, unlike other 3D systems, the sandwich-like system provides a wide spectrum of well-controlled cellular microenvironments. Consequently the sandwich-like system is an interesting cell culture platform to mimic different physiological environments in order to study cell biology and test cell fate under different culture conditions.
As mentioned before, one advantage of this system is that different ventral and dorsal substrates can be studied using different protein coatings. Therefore, important parameters for cell fate such as ligand density can be tuned by controlling the ligand density of each one of the 2D substrates used (e.g. ventral and dorsal substrates). However, note that different substrates may need changes in the protocol. For example, using bigger dorsal substrates may result in the need to set the proper incubation time to peel the samples off the Petri dish. Likewise, bigger ventral substrates could result in hypoxic areas at the center of the sample because of the limited oxygen permeability of the ventral substrate (due to the glass coverslip). Additionally, analysis procedures will depend on the substrate properties. For instance, using opaque dorsal substrates will hinder standard microscopy protocols though it will still allow protein/nucleic acids extraction. Another key factor is the permeability of the dorsal substrate since cells should be allowed to get nutrients from the medium and discard waste.
To sum up, a sandwich-like culture is a simple system that offers the possibility to mimic different 3D-like microenvironments to investigate cell fate.
Disclosures
The authors have nothing to disclose.
Acknowledgements
The support from ERC through HealInSynergy (306990) and the FPU program AP2009-3626 are acknowledged.
Materials
| Ploy(lactic acid) | NatureWorks | 4042D | Reagent |
| Cover glasses (12 mmØ) | Marienfeld | 631-0666 | Equipment |
| Chloroform | Scharlab | CL0200 | Reagent |
| 1,1,1,3,3,3-Hexafluoro-2-propanol (HFIP) | Sigma | 105228 | Reagent |
| Syringe (1mL) | Henke Sass Wolf | 4010-200V0 | Equipment |
| Syringe pump | New Era Pump Systems | NE1000 | Equipment |
| High Voltage DC Power Supply | Glassman High Voltage | Series FC | Equipment |
| Incubator | Hucoa-Herlös | 3111 | Equipment |
| Laminar flow hood | Telstar | AV30/70 | Equipment |
| Human Fibronectin | Sigma | F2006 | Reagent |
| RNeasy Micro Kit | Qiagen | 74004 | Reagent |
| Inverted microscope | Leica Microsystems | DMI 6000 | Equipment |
| Triton X-100 | Sigma-Aldrich | T8787 | Reagent |
| Albumin | Sigma-Aldrich | A7409 | Reagent |
| Tween 20 | Sigma-Aldrich | P2287 | Reagent |
References
- Baker, B. M., Chen, C. S. Deconstructing the third dimension: how 3D culture microenvironments alter cellular cues. J Cell Sci. 125 (Pt 13), 3015-3024 (2012).
- Weiss, P. . Rev. Mod. Phys. 31 (1), 11-20 (1959).
- Ruffner, H., Lichtenberg, J. 3D cell culture systems–towards primary drug discovery platforms: an interview with Heinz Ruffner (Novartis) and Jan Lichtenberg (InSphero). Biotechnol J. 7 (7), 833-834 (2012).
- Horning, J. L., et al. 3-D tumor model for in vitro evaluation of anticancer drugs. Mol Pharm. 5 (5), 849-862 (2008).
- Wolf, K., et al. Compensation mechanism in tumor cell migration: mesenchymal-amoeboid transition after blocking of pericellular proteolysis. J Cell Biol. 160 (2), 267-277 (2003).
- Schor, S. L., et al. A novel ‘sandwich’ assay for quantifying chemo-regulated cell migration within 3-dimensional matrices: wound healing cytokines exhibit distinct motogenic activities compared to the transmembrane assay. Cell Motil Cytoskeleton. 63 (5), 287-300 (2006).
- Lee, E. J., Hwang, C. M., Baek, D. H., Lee, S. H. Fabrication of microfluidic system for the assessment of cell migration on 3D micropatterned substrates. Conf Proc IEEE Eng Med Biol Soc. 2009 (2009), 6034-6037 (2009).
- Zaman, M. H., et al. Migration of tumor cells in 3D matrices is governed by matrix stiffness along with cell-matrix adhesion and proteolysis. Proc Natl Acad Sci U S A. 103 (29), 10889-10894 (2006).
- Ballester-Beltrán, J., Lebourg, M., Salmerón-Sánchez, M. Dorsal and ventral stimuli in sandwich-like microenvironments. Effect on cell differentiation. Biotechnol Bioeng. 110 (11), 3048-3058 (2013).
- Ballester-Beltrán, J., Lebourg, M., Moratal, D., Salmerón-Sánchez, M. Fibronectin-matrix sandwich-like microenvironments to manipulate cell fate. Biomater. Sci. 2 (3), 381-389 (2014).
- Ballester-Beltrán, J., Lebourg, M., Rico, P., Salmerón-Sánchez, M. Dorsal and ventral stimuli in cell-material interactions: effect on cell morphology. Biointerphases. 7 (1-4), 39 (2012).
- Ballester-Beltrán, J., Lebourg, M., Rico, P., Salmerón-Sánchez, M. Cell migration within confined sandwich-like nanoenvironments. Nanomedicine. , (2015).
- Glaß, M., et al. Cell Migration Analysis: Segmenting Scratch Assay Images with Level Sets and Support Vector Machines. Pattern Recogn. 45 (9), 3154-3165 (2012).
- Cukierman, E., Pankov, R., Stevens, D. R., Yamada, K. M. Taking cell-matrix adhesions to the third dimension. Science. 294 (5547), 1708-1712 (2001).
- Huebsch, N., Arany, P. R., Mao, A. S., et al. Harnessing traction-mediated manipulation of the cell/matrix interface to control stem-cell fate. Nat. Mater. 9 (6), 518-526 (2010).
- Petrie, R. J., Gavara, N., Chadwick, R. S., Yamada, K. M. Nonpolarized signaling reveals two distinct modes of 3D cell migration. J Cell Biol. 197 (3), 439-455 (2012).
- Mao, Y., Schwarzbauer, J. E. Stimulatory effects of a three-dimensional microenvironment on cell-mediated fibronectin fibrillogenesis. J Cell Sci. 118 (Pt 19), 4427-4436 (2005).

