Rapid Assembly of Multi-Gene Constructs using Modular Golden Gate Cloning
Summary
The goal of this protocol is to provide a detailed, step-by-step guide for assembling multi-gene constructs using the modular cloning system based on Golden Gate cloning. It also gives recommendations on critical steps to ensure optimal assembly based on our experiences.
Abstract
The Golden Gate cloning method enables the rapid assembly of multiple genes in any user-defined arrangement. It utilizes type IIS restriction enzymes that cut outside of their recognition sites and create a short overhang. This modular cloning (MoClo) system uses a hierarchical workflow in which different DNA parts, such as promoters, coding sequences (CDS), and terminators, are first cloned into an entry vector. Multiple entry vectors then assemble into transcription units. Several transcription units then connect into a multi-gene plasmid. The Golden Gate cloning strategy is of tremendous advantage because it allows scar-less, directional, and modular assembly in a one-pot reaction. The hierarchical workflow typically enables the facile cloning of a large variety of multi-gene constructs with no need for sequencing beyond entry vectors. The use of fluorescent protein dropouts enables easy visual screening. This work provides a detailed, step-by-step protocol for assembling multi-gene plasmids using the yeast modular cloning (MoClo) kit. We show optimal and suboptimal results of multi-gene plasmid assembly and provide a guide for screening for colonies. This cloning strategy is highly applicable for yeast metabolic engineering and other situations in which multi-gene plasmid cloning is required.
Introduction
Synthetic biology aims to engineer biological systems with new functionalities useful for pharmaceutical, agricultural, and chemical industries. Assembling large numbers of DNA fragments in a high-throughput manner is a foundational technology in synthetic biology. Such a complicated process can break down into multiple levels with decreasing complexity, a concept borrowed from basic engineering sciences1,2. In synthetic biology, DNA fragments usually assemble hierarchically based on functionality: (i) Part level: "parts" refers to DNA fragments with a specific function, such as a promoter, a coding sequence, a terminator, an origin of replication; (ii) Transcription units (TU) level: a TU consists of a promoter, a coding sequence, and a terminator capable of transcribing a single gene; (iii) Multi-gene level: a multi-gene plasmid contains multiple TUs frequently comprised of an entire metabolic pathway. This hierarchical assembly pioneered by the BioBrick community is the foundational concept for the assembly of large sets of DNAs in synthetic biology3.
In the past decade4,5,6,7, the Golden Gate cloning technique has significantly facilitated hierarchical DNA assembly2. Many other multi-part cloning methods, such as Gibson cloning8, ligation-independent cloning (SLIC)9, uracil excision-based cloning (USER)10, the ligase cycling reaction (LCR)11, and in vivo recombination (DNA Assembler)12,13, have also been developed so far. But Golden Gate cloning is an ideal DNA assembly method because it is independent of gene-specific sequences, allowing scar-less, directional, and modular assembly in a one-pot reaction. Golden Gate cloning takes advantage of type IIS restriction enzymes that recognize a non-palindromic sequence to create staggered overhangs outside of the recognition site2. A ligase then joins the annealed DNA fragments to obtain a multi-part assembly. Applying this cloning strategy to the modular cloning (MoClo) system has enabled the assemble of up to 10 DNA fragments with over 90% transformants screened containing the correctly assembled construct4.
The MoClo system offers tremendous advantages that have accelerated the design-build-test cycle of synthetic biology. Firstly, the interchangeable parts enable combinatorial cloning to test a large space of parameters rapidly. For example, optimizing a metabolic pathway usually requires cycling through many promoters for each gene to balance the pathway flux. The MoClo system can easily handle such demanding cloning tasks. Secondly, one needs to sequence the part plasmid but typically not the TU or the multi-gene plasmids. In most cases, screening by colony PCR or restriction digestion is sufficient for verification at the TU and multi-gene plasmid level. This is because cloning the part plasmid is the only step requiring PCR, which frequently introduces mutations. Thirdly, the MoClo system is ideal for building multi-gene complex metabolic pathways. Lastly, because of the universal overhangs, the part plasmids can be reused and shared with the entire bioengineering community. Currently, MoClo kits are available for plants14,15,5,16,17, fungi6,18,19,20,21,22, bacteria7,23,24,25,26,27, and animals28,29. A multi-kingdom MoClo platform has also been introduced recently30.
For Saccharomyces cerevisiae, Lee et al.6 have developed a versatile MoClo toolkit, an excellent resource for the yeast synthetic biology community. This kit comes in a convenient 96-well format and defines eight types of interchangeable DNA parts with a diverse collection of well-characterized promoters, fluorescent proteins, terminators, peptide tags, selection markers, origin of replication, and genome editing tools. This toolkit allows the assembly of up to five transcription units into a multi-gene plasmid. These features are valuable for yeast metabolic engineering, in which partial or entire pathways are over-expressed to produce targeted chemicals. Using this kit, researchers have optimized the production of geraniol, linalool31, penicillin32, muconic acid33, indigo34, and betalain35 in yeast.
Here we provide a detailed, step-by-step protocol to guide the use of the MoClo toolkit to generate multi-gene pathways for either episomal or genomic expression. Through extensive use of this kit, we have found that the accurate measurement of DNA concentrations is key to ensuring the equimolar distribution of each part in the Golden Gate reaction. We also recommend the T4 DNA ligase over the T7 DNA ligase because the former works better with larger numbers of overhangs36. Lastly, any internal recognition sites of BsmBI and BsaI must be removed or domesticated prior to assembly. Alternatively, one may consider synthesizing parts to remove multiple internal sites and to achieve simultaneous codon optimization. We demonstrate how to use this toolkit by expressing a five-gene pathway for β-carotene and lycopene production in S. cerevisiae. We further show how to knock out the ADE2 locus using the genome-editing tools from this kit. These color-based experiments were selected for easy visualization. We also demonstrate how to generate fusion proteins and to create amino acid mutations using Golden Gate cloning.
Protocol
NOTE: The hierarchical cloning protocol offered in this toolkit can be divided into three major steps: 1. Cloning part plasmids; 2. Cloning transcription units (TUs); 3. Cloning multi-gene plasmids (Figure 1). This protocol starts from the primer design and ends with applications of the cloned multi-gene plasmid.
1. Primer design for cloning the part plasmid (pYTK001):
- Design the forward and reverse primers containing flanking nucleotides TTT at the 5' end,a BsmBI recognition site with an additional nucleotide (CGTCTCN), a 4-nucleotides (nt) overhang (TCGG) complementary to that of the entry vector, a BsaI recognition site with an additional nucleotide (GGTCTCN), and a 4-nt part-specific overhang, in addition to the template-specific sequence (Figure 2A). GoldenBraid 4.037 and GoldenMutagenesis38 are some of the online software that can be used for Golden Gate specific primer design.
- If BsaI or BsmBI recognition sites are present in any part, use a domestication step to mutate these sites prior to Golden Gate assembly39. For integrative plasmids (see step 4), domesticate any NotI recognition site. To domesticate a part with one undesirable recognition site, divide the part into two subparts near the undesirable site (Figure 2A):
- Design the forward primer of the first subpart in the same manner as in step 1.1. but design the reverse primer with the BsmBI site and a 4-nt gene-specific overhang only.
- Design the second sub-part forward primer with the BsmBI site and the 4-nt gene-specific overhang only that overlaps with the reverse primer of the first sub-part. Design the reverse primer of the second sub-part in the same manner as in step 1.1.
- Introduce the desired mutation(s) in either the reverse (for step 1.2.1) or forward primer (step 1.2.2) at the gene-specific region of the primer.
NOTE: Alternatively, a BsmBI- and BsaI-free and codon-optimized part can be synthesized commercially. For coding sequences (CDS), a synonymous mutation can be readily incorporated at the third nucleotide of an amino acid codon. For promoters and terminators, however, checking the mutated promoter or terminator activity using a reporter assay is recommended39. If there is an undesirable restriction site toward the end of the sequence, it can be mutated using a longer reverse primer. If multiple undesirable sites are present, site-directed mutagenesis allowing mutating multiple sites may be performed40.
- Occasionally, fusing two proteins as a single part with a linker in between is desirable (Figure 2A). The linker helps to ensure the structural integrity of the two individual proteins41.
- The forward primer for the first gene is the same as in step 1.1. In the reverse primer, include a BsmBI site, a 4-nt gene-specific overhang, and a linker sequence. The 4-nt overhang can be either the linker or second gene's first few nucleotides.
- For the second gene, design the forward primer such that it has a BsmBI recognition site and a 4-nt overhang complementary to that of the reverse primer of the first gene. Design the reverse primer of the second gene as in step 1.1.
- To amplify the parts by polymerase chain reaction (PCR)42, use a high-fidelity DNA polymerase to amplify the parts from either a genomic DNA, a cDNA, or a plasmid. Check the PCR product on a 1% agarose gel followed by gel-purification. Using purified DNA is strongly recommended, if gel purification is laborious, use at least a spin column to purify the PCR product.
2. Cloning parts into the entry vector (pYTK001) to create part plasmids (Figure 2B)
- To set up the Golden Gate reaction mix, add 20 fmol of each PCR product and the entry vector (pYTK001), 1 μL of 10X T4 ligase buffer, 0.5 μL of Esp3I (a highly efficient isoschizomer of BsmBI), and 0.5 μL of T4 ligase. Add ddH2O to bring the total volume to 10 μL.
- To set up the cloning reaction, run the following program in a thermocycler: 25-35 cycles of 37 °C for 5 min (digestion) and 16 °C for 5 min (ligation), followed by a final digestion at 50 °C for 10 min and enzyme inactivation at 80 °C for 10 min.
NOTE: 35 cycles of digestion/ligation are recommended when cloning multiple DNA pieces into the entry vector simultaneously, for example, during the cloning of fusion genes or domesticating a gene. - Transform the entire reaction mix into the DH5α strain or equivalent Escherichia coli chemically competent cells by heat shock. Transforming the entire 10 μL cloned product into 35 μL chemically competent E. coli cells (2 X 105 cfu/mL, cfu is calculated from transforming 5 ng pYTK001 into 100 μL of the competent cells) is recommended. Spread on a lysogeny Broth (LB) plate with 35 μg/mL chloramphenicol (Cm). Incubate at 37 °C overnight.
- After 16-18 h, take the plate out from the incubator and keep the plate at 4 °C for about 5 h to let the super folder green fluorescent protein (sfGFP) develop for a more intense green color.
- For easier screening, place the plate on an ultraviolet (UV) or a blue light transilluminator. The sfGFP containing colonies will fluoresce under the UV light.
- The green colonies are negative because they contain the uncut pYTK001. The white colonies are likely positive. The cloning is usually successful if there are ~30-100% white colonies. Perform further screening of a few white colonies by either colony PCR or restriction digestions (suggested enzyme: BsaI-HFv2).
- Purify plasmids from a few of the potentially correct colonies and confirm the sequences by Sanger sequencing.
3. Assembling part plasmids into "cassette" plasmids
NOTE: A cassette plasmid contains a user-defined transcription unit (TU) consisting of a promoter, a CDS, and a terminator. A cassette plasmid allows the expression of a single gene. If the cassette plasmids will be assembled into a multi-gene plasmid, then the first step is to determine the number and the order of TUs in the multi-gene plasmid. These will determine which connectors to use in the cassette plasmids since connectors link TUs in the multi-gene plasmid. The first TU's left connector should be ConLS, and the right connector of the last TU should be ConRE. They will overlap with ConLS' and ConRE' of multi-gene plasmids. The rest of the connectors should be in the increasing numerical order. For example, if the multi-gene plasmid contains four TUs, the connector combinations would be ConLS-TU1-ConR1, ConL1-TU2-ConR2, ConL2-TU3-ConR3, and ConL3-TU4-ConRE (Figure 1).
- Before assembling transcription units, assembling an intermediate vector with the following six parts is recommended: the left connector, the sfGFP dropout (pYTK047), the right connector, a yeast selection marker, a yeast origin of replication and the part plasmid with an mRFP1, an E. coli origin and the ampicillin-resistant gene (pYTK083) (Figure 3).
- Purify the above six plasmids. Record their concentrations using a UV-Vis spectrophotometer or a fluorescence-based assay and dilute each plasmid with ddH2O so that 1 μL has 20 fmol of DNA. Calculate the DNA molar concentration by using an online calculator.
NOTE: It is very important to measure the DNA concentrations accurately and to pipet precisely for the assembly to work, especially for assemblies with five to seven part plasmids. Small errors in the DNA concentration of each plasmid can cause a significant decrease in cloning efficiency. - Add 1 μL of each plasmid, 1 μL of 10X T4 ligase buffer, 0.5 μL of BsaI-HFv2 (a highly efficient version of BsaI), and 0.5 μL of T4 ligase. Make up the volume to 10 μL by adding ddH2O.
- To set up the cloning reaction, run the following program in the thermocycler: 25-35 cycles of 37 °C for 5 min (digestion) and 16 °C for 5 min (ligation). Omit the final digestion and heat inactivation steps as the BsaI sites need to be retained in the intermediate vector (Figure 3).
- Transform the entire reaction mix into the DH5α strain or other E. coli competent cells. Spread on an LB plate with 50 μg/mL carbenicillin (Cb) or ampicillin. Incubate at 37 °C overnight.
NOTE: Carbenicillin is a stable analog of ampicillin. - After 16-18 h, take the plate out of the incubator. The plate will contain both pale red and pale green colonies (Figures 6C and 6D). Keep the plate at 4 °C for about 5 h to let the mRFP1 and sfGFP mature. Use a UV or a blue light transilluminator to identify the green colonies, which contain the potentially correct intermediate vector.
- Streak out the green colonies on an LB + Cb plate and incubate at 37 °C overnight. The next day, streak out again on an LB + Cm plate and incubate at 37 °C overnight. The colonies growing on LB + Cm plates contain misassembled plasmids because Cm resistant part vectors are retained.
- Pick the colonies that do not grow on the LB + Cm plate and perform restriction digestions (suggested enzymes: BsaI-HFv2, Esp3I) to confirm the correctly assembled plasmid. Alternatively, use colony PCR for screening.
- Purify the above six plasmids. Record their concentrations using a UV-Vis spectrophotometer or a fluorescence-based assay and dilute each plasmid with ddH2O so that 1 μL has 20 fmol of DNA. Calculate the DNA molar concentration by using an online calculator.
- Once the intermediate vector has been successfully assembled, the next step is to assemble transcription units. This a 4-piece assembly with the following parts: the intermediate vector, a promoter, a CDS, and a terminator.
- Purify the four part plasmids. Record their concentrations and dilute each of plasmid so that 1 μL has 20 fmol of DNA.
- To set up the reaction mix, follow Step 3.1.2.
- To set up the cloning reaction, follow Step 2.2.
- Transform the entire cloning reaction mix into the DH5α or equivalent E. coli competent cells and plate on LB + Cb. Incubate at 37 °C overnight.
- After 16-18 h, take the plate out from the incubator. White and pale green colonies will appear (Figures 6E and 6F). Keep the plate at 4 °C for about 5 h to let the sfGFP mature. Use a UV or a blue light transilluminator to identify the non-fluorescent white colonies. These contain the potentially correct transcription units.
- Streak out and grow 8-10 white colonies and perform a colony PCR. Purify plasmids from the colonies that test positive from colony PCR. Carry out restriction digestion (suggested enzyme: Esp3I) to further confirm the assembly.
NOTE: Sequencing transcription units is typically not necessary because the cloning involves only restriction digestion and ligation. All sequences of interest have been confirmed at the part plasmid level.
4. Assembling cassette plasmids into "multi-gene" plasmids:
NOTE: Multi-gene plasmids allow the expression of more than one gene. Depending on the downstream application, multi-gene plasmids could be replicative or integrative. Replicative plasmids have the yeast origin of replication; therefore, it can be stably maintained when yeast cell divides. Integrative plasmids do not have the yeast origin of replication. Instead, they have 5' and 3' homology arms allowing the integration of multiple genes into specific loci of the genome through homologous recombination.
- For multi-gene plasmids, assemble an intermediate vector first.
- To assemble replicative intermediate vectors (Figure 4A), assemble the following six parts: the left connector (ConLS'-pYTK008), the sfGFP dropout (pYTK047), the right connector (ConRE'-pYTK072), a yeast selection marker, a yeast origin of replication, and the part plasmid with mRFP1, an E. coli origin of replication, and the kanamycin-resistant gene (pYTK084).
- For assembly, follow the steps from 3.1.1 to 3.1.3.
- Transform the entire cloning reaction mix into DH5α or equivalent E. coli competent cells, and plate on LB plus 50 μg/mL kanamycin (Km). Incubate at 37 °C overnight.
- For red/green color-based screening, follow step 3.1.5.
- For screening of misassemblies, streak and grow the green colonies on an LB + Km plate. Then follow step 3.1.6.
- For integrative multi-gene vectors (Figure 4B), determine the genomic locus of interest first, then design approximately 500 base pairs of 5' and 3' homology arms for integrating to that locus.
- Clone the 5' and 3' homology arms from yeast genomic DNA into the entry vector-pYTK001. Follow steps 1 and 2.
- Assemble the following seven plasmids: the left connector (ConLS'-pYTK008), the sfGFP dropout (pYTK047), the right connector (ConRE'-pYTK072), a yeast selection marker, the 3' homology arm, the part plasmid with mRFP1, E. coli origin of replication, and the kanamycin-resistant gene (pYTK090), and the 5' homology arm.
- For assembly and screening, follow step 4.1.1.
- To assemble replicative intermediate vectors (Figure 4A), assemble the following six parts: the left connector (ConLS'-pYTK008), the sfGFP dropout (pYTK047), the right connector (ConRE'-pYTK072), a yeast selection marker, a yeast origin of replication, and the part plasmid with mRFP1, an E. coli origin of replication, and the kanamycin-resistant gene (pYTK084).
- Assembly of the multi-gene plasmid
- Purify plasmids of the intermediate vector obtained in step 4.1 and the cassette plasmids from step 3. Record their concentrations using a UV-Vis spectrophotometer or a fluorescence-based assay. Dilute each in ddH2O so that 1 μL has 20 fmol DNA.
- Add 1 μL of intermediate vector, 1 μL of each transcription unit, 1 μL of 10x T4 ligase buffer, 0.5 μL of Esp3I, and 0.5 μL of T4 ligase. Bring the volume to 10 μL of using ddH2O.
- To set up the cloning reaction, follow step 2.2.
- Transform the entire cloning reaction mix into DH5α or equivalent E. coli competent cells, and plate on LB + Km. Incubate at 37 °C overnight.
- Perform the green/white screening as in step 3.2.5.
- Purify plasmids from a few white colonies and perform restriction digestions. Using the NotI-HF enzyme is recommended because there are two NotI sites at the E. coli origin and Km selection marker part, respectively (Figure 4B). If the assembled plasmid is very large, then another restriction site can be chosen for further confirmation. Alternatively, screen with colony PCR before proceeding to restriction digestion.
5. Applying multi-gene plasmids for chromosomal or plasmid-based gene expression
- Integrating multi-gene plasmid into the yeast genome for chromosomal gene expression (Figure 5)
- Design a guide RNA (gRNA) for the desired locus. Using multiple online resources, such as Benchling, CRISPRdirect43, and CHOPCHOP44, are recommended to determine the maximum on-target specificity.
- Clone the synthesized 20-nt gRNA into the pCAS plasmid45 by Gibson cloning. Linearize the pCAS plasmid by PCR using a reversed primer pair binding to the 3' of the HDV ribozyme and the 5' of the tracrRNA respectively (Figure 5).Alternatively, clone the gRNA into the sgRNA dropout plasmid (pYTK050) and assemble the dropout plasmid into a cassette plasmid with linkers. Then assemble the Cas9 TU with the Cas9 part plasmid (pYTK036). Lastly, assemble the Cas9 TU and the sgRNA TU into a replicative multi-gene plasmid.
- Linearize 5-15 μg of integrative multi-gene plasmid with 1 μL of NotI-HF enzyme overnight. Transform 1 μg of pCAS-gRNA and the linearized integrative multi-gene plasmid into S. cerevisiae. Prepare competent cells using either a commercially available yeast transformation kit or following the protocol by Geitz and Schiestl, 200746.
NOTE: It is unnecessary to purify the linearized multi-gene plasmid after the NotI digestion. - Pellet the cells after recovery, discard the supernatant, wash with an equal volume of water. Plate yeast cells on the complete synthetic medium (CSM) dropout plate or the yeast extract peptone dextrose medium (YPD) plate with antibiotics, depending on the yeast selective marker. Incubate at 37 °C for two days for colonies to form. In case no colony is observed, incubate for an additional one to two days at 30°C.
- Screen yeast colonies for integration by colony PCR47.
- To cure the pCAS, streak out the colony with the correct integration onto a non-selective YPD plate. Grow at 30 °C overnight. Streak one colony from the YPD plate onto a fresh YPD plate. Again, grow at 30 °C overnight. Streak a colony from the second YPD plate to a YPD plus 100 μg/mL nourseothricin, the selection marker of pCAS. Successful curing occurs when yeast cells fail to grow on the selective plate.
NOTE: If the pCAS plasmid is not cured in two rounds of non-selective YPD, streak again onto a fresh YPD for another 1-2 rounds.
- Transforming replicative multi-gene plasmid for plasmid-based gene expression
- Transform 100 ng-1 μg pure multi-gene plasmid into S. cerevisiae competent cells.
- Plate yeast cells immediately after transformation onto the CSM dropout plate or YPD plus antibiotic plate, depending on the yeast selection marker used. Incubate at 30°C for 2-3 days for colonies to form.
Representative Results
Here the results of four replicative multi-gene plasmids for β-carotene (yellow) and lycopene (red) production. One integrative multi-gene plasmid for disrupting the ADE2 locus was constructed, the colonies of which are red.
Cloning CDSs into the entry vector (pYTK001)
ERG20 was amplified from the yeast genome and the three carotenoid genes crtE, crtYB, crtI from the plasmid pLM49448 into the entry vector pYTK001 as described. Yeast promoters pENO2, pTIP1, pPYK1 and pPDC1 and terminators tTDH2, tHSP26, tADH2, tACS2 were cloned as part plasmids. A point mutation was introduced into CrtYB (G247A) for producing lycopene and a BTS1-ERG20 fusion construct was created to better channel the prenyl intermediate to carotenoids. Figures 6A and 6B show a representative plate of the successful cloning of a part plasmid and provide an example of the green/white screening with 90.35 ± 4.22% total colonies being potentially correct (white).
Assembly of transcription units (cassette plasmids)
Before assembling the cassette plasmid, the design of the multi-gene plasmid was finalized. For the four carotenoid TUs, four intermediate vectors with different connectors were cloned first following step 3.1. Figures 6C and 6D show a representative plate from a successful assembly of the intermediate vector and provide an example of the red/green screening, with 17.56 ± 3.32% total colonies being positive (green). Although this ratio is relatively low, the screening is greatly facilitated by the green fluorescence.
Next, the TUs for BTS1-ERG20, ERG20, crtE, crtYB, crtYB(G247A), and crtI were assembled following step 3.2 (Table 1). Figures 6E and 6F show a representative plate of a successful assembly of the TUs and provide an example of a green/white screening method, with 65.02 ± 4.99% total colonies being positive (white).
Assembly of multi-gene plasmids
Four replicative and one integrative multi-gene plasmids were assembled. For replicative multi-gene plasmids, ERG20, BTS1-ERG20, crtE, crtYB, crtYB (G247A) and crtI were assembled in four different combinations, creating four plasmids: two for β-carotene and two for lycopene production (Table 2). The ratio of potentially correct colonies (green) for the intermediate vector cloning was 1.83 ± 0.15% (Figures 7A and 7B). Although this number seems low, the screening was made easy by detecting the green fluorescence (Figure 7B). Once the intermediate plasmid was cloned, the success rate of assembling multi-gene plasmids (white) from the intermediate was 93.77± 1.65% (Figures 7C and 7D). Figures 7E and 7F show a suboptimal assembly of multi-gene plasmids, as the numbers of positive colonies (white) were negligible. After transforming into yeast, colonies producing β-carotene (yellow) and lycopene (red) grew on day three. Four colonies from each plate were streaked out onto fresh plates and grown for two more days. Figures 8A and 8B show that fusing the BTS1-ERG20 directs more geranylgeranyl-pyrophosphate toward the carotenoid production, as seen by darker colors than using the ERG20 alone. Upon extraction49 and quantification of the carotenoids by UV-Vis spectrophotometry with authentic standards, it is seen that fusion of BTS1-ERG20 leads to the production of 0.729 µg/mg β-carotene, which is ~35 fold higher than 0.021 μg/mg β-carotene produced by the strain with ERG20 alone. Likewise, the production of lycopene is ~16.5 fold higher in the strain with BTS1-ERG20 (1.126 μg/mg) compared to ERG20 (0.068 μg/mg) alone (Figure 8C).
Replicative multi-gene plasmids transformed into yeast with (A) BTS1-ERG20 fusion TU and (B) ERG20 TU. On each plate, yeast on the left side has plasmids containing crtE TU, crtYB TU and crtI TU for the production of β–carotene and yeast on the right side has plasmids containing crtE TU, crtYB (G247A)TU, and crtI TU for the production of lycopene.
For the integrative plasmid, the ConLS', sfGFP dropout, ConRE', HIS3 (yeast selection marker), ADE2 5' and 3' homology arms were assembled following step 4.1.2. The 5' and 3' homology arms were 500 bp apart, deleting ~180 amino acids after integration. Additionally, the 5' homology arm had six stop codons towards its 3' end. The mutated ADE2 resulted in red colonies50. The ADE2 gRNA 5'-ATTGGGAC GTATGATTGTTGAGG-3'51 were used and followed step 5.1 for genomic integration. After 3-4 days, red colonies were observed on the YPD plate + 100 μg/mL nourseothricin, indicating that ADE2 had been successfully disrupted (Figures 8D and 8E).
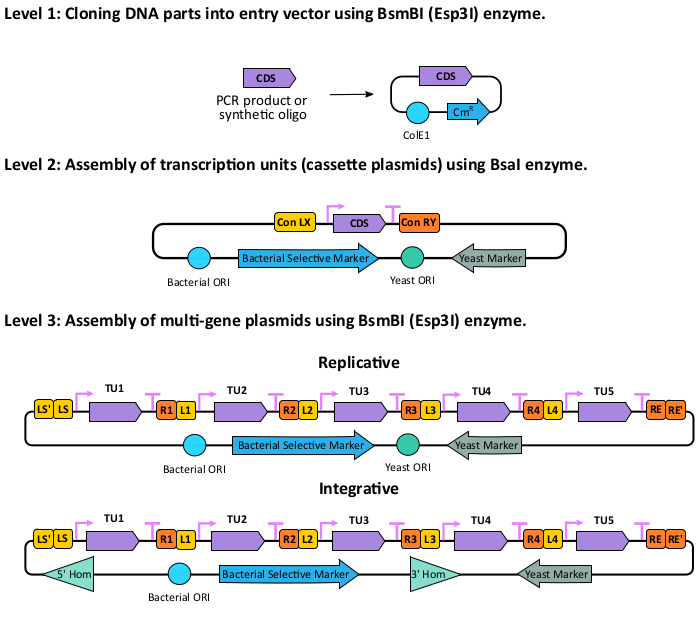
Figure 1: Overview of multi-gene assembly. Assembly takes place in three levels. In0 level 1, the CDS, or any other part, is amplified from the genome or synthesized, and then cloned into the pYTK001 entry vector using the BsmBI (or Esp3I) enzyme. ColE1: E. coli origin of replication; CmR: Chloramphenicol resistant gene. In level 2, the transcription unit (TU) containing a promoter, a CDS, and a terminator is assembled using the BsaI enzyme. In level 3, up to five transcription units are assembled into a multi-gene plasmid using the BsmBI (or Esp3I) enzyme. The multi-gene plasmid can be either replicative or integrative. Please click here to view a larger version of this figure.
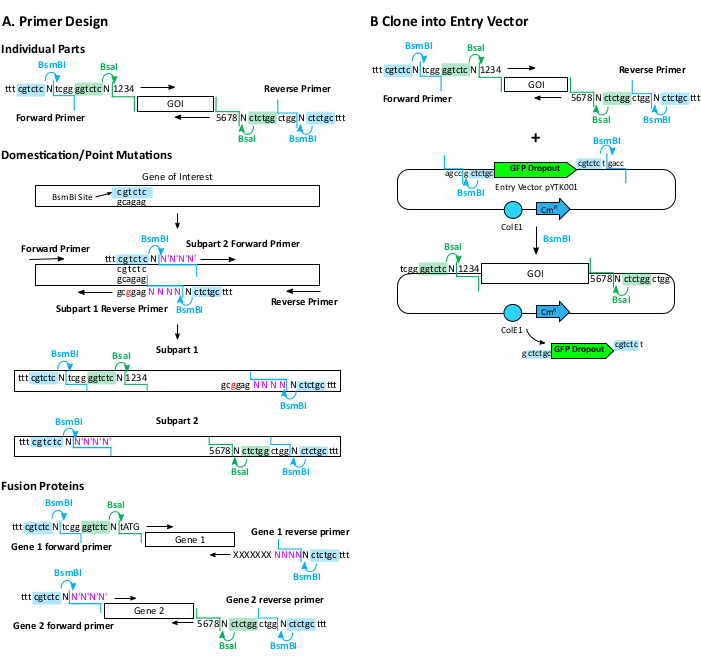
Figure 2: Primer design and cloning of part plasmids. (A) Primer design for amplifying individual parts, domesticating genes or creating point mutations, and assembling fusion proteins. Primers include BsmBI and BsaI recognition and cut sites and the MoClo overhang for proper assembly. MoClo overhangs are represented as 1, 2, 3, 4, and 5, 6, 7, 8. Internal primers for domestication or creating fusion protein contains the BsmBI but not the BsaI sites. The overhangs for these are customized internal sequences (NNNN and N'N'N'N' in purple). Terminal "ttt" are included for optimal enzyme digestion. GOI: gene of interest. (B) Cloning amplified parts into the pYTK001 entry vector using BsmBI (or Esp3I). Complementary overhangs lead to the integration of the part and the removal of the BsmBI recognition site. ColE1: E. coli origin of replication; CmR: Chloramphenicol resistant gene. Please click here to view a larger version of this figure.
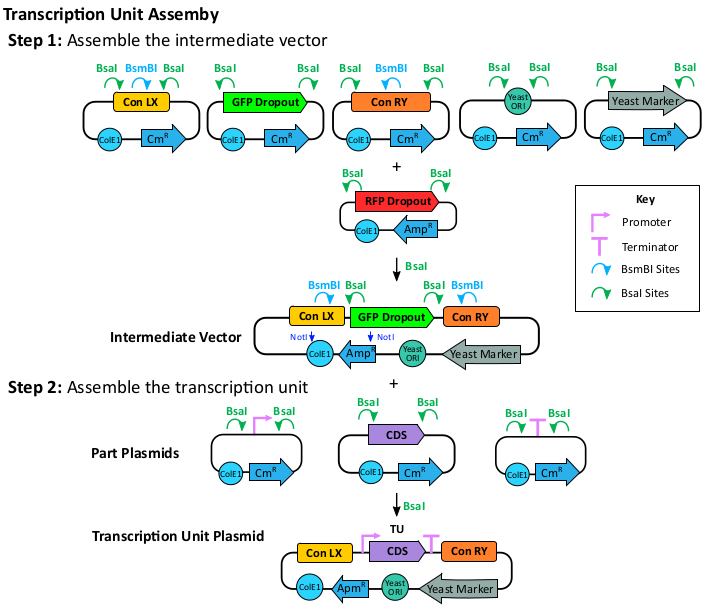
Figure 3: Transcription unit assembly. To assemble the TU plasmids, the assembly of an intermediate vector is recommended to facilitate combinatorial TU assembly. To assemble the intermediate vector, clone the Con LX (X: one of the five left connectors), the sfGFP dropout, the Con RY (Y: one of the five right connectors), a yeast ORI (origin of replication), and a yeast marker part into the mRFP1 dropout vector using the BsaI enzyme. The intermediate plasmid is resistant to ampicillin. The BsaI recognition sites are retained for TU plasmid cloning. To clone the TU plasmid, a promoter, a CDS, and a terminator are assembled into the intermediate vector using BsaI. The cloned TU will have BsmBI sites at the ConLX and ConRY regions for the next step multi-gene assembly. The cloned TU is also resistant to ampicillin. Please click here to view a larger version of this figure.
| Name | ConLX (left connector) | Promoter | CDS | Terminator | ConRY (right connector) |
| BTS/ERG20 TU | LS | pENO2 | BTS1/ERG20 | tTDH2 | R1 |
| ERG20 TU | LS | pENO2 | ERG20 | tTDH2 | R1 |
| crtE TU | L1 | pTIP1 | crtE | tHSP26 | R2 |
| crtYB TU | L2 | pPDC1 | crtYB | tADH2 | R3 |
| crtYBG247A TU | L2 | pPDC1 | crtYBG247A | tADH2 | R3 |
| crtI TU | L3 | pPYK1 | crtI | tACS2 | RE |
Table 1: Transcription units used in this study. Promoters and terminators were amplified from S. cerevisiae. BTS1 (geranylgeranyl diphosphate synthase) and ERG20 (farnesyl pyrophosphate synthetase) were amplified from S. cerevisiae. The genes crtE (geranylgeranyl diphosphate synthase), crtYB (bifunctional lycopene cyclase/phytoene synthase), and crtI (phytoene desaturase) were from Xanthophyllomyces dendrorhous.
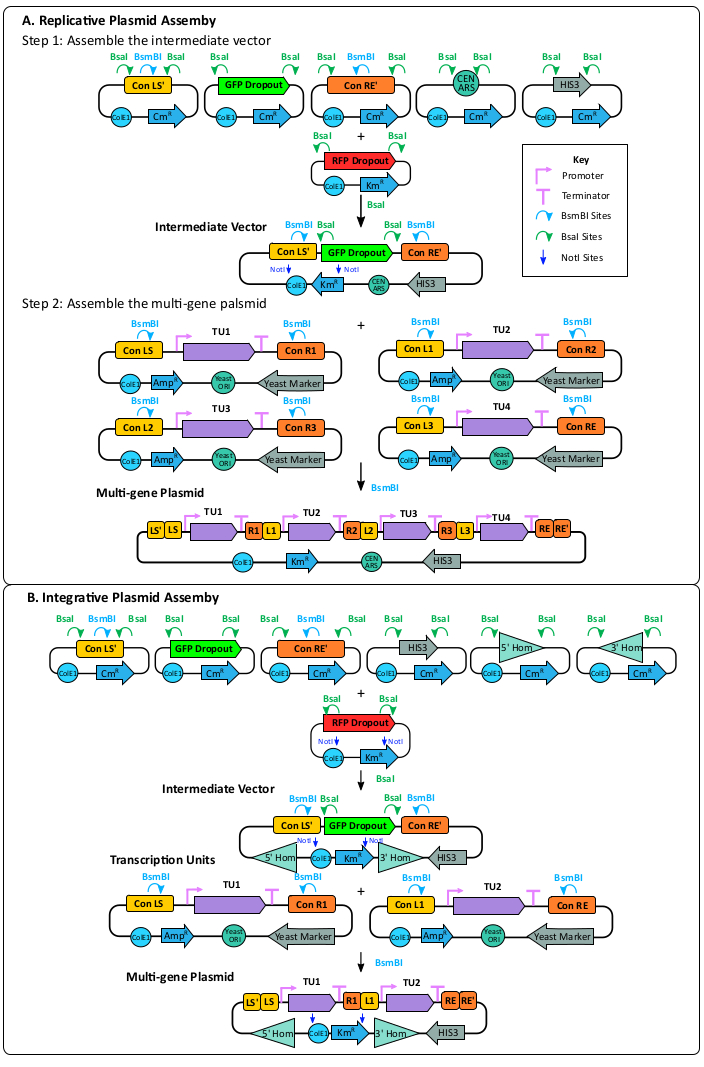
Figure 4: Multi-gene plasmid assembly. (A) Replicative plasmid assembly. Assembly of the replicative intermediate vector includes cloning the Con LS', the sfGFP dropout, the Con RE', a yeast ORI, a yeast marker, and an E. coli origin and marker on the mRFP1 dropout vector using the BsaI enzyme. The ConLS' and ConRE' sites introduce BsmBI recognition sites to the vector. Potentially correct assemblies can be screened by looking for green colonies on a selective plate with kanamycin. The previously assembled TUs can then be cloned into the intermediate vector using the BsmBI enzyme. This plasmid contains a yeast ORI allowing it to replicate in a yeast host. (B) Integrative plasmid assembly. Assembly of the integrative intermediate vector includes cloning the Con LS', the sfGFP dropout, the Con RE', a 5' homology arm, a 3' homology arm, a yeast marker, and the E. coli origin and marker into the RFP dropout vector using the BsaI enzyme. Correct assemblies should appear green on a selective plate with kanamycin. Transcription units previously made can be cloned into the replicative intermediate vector using the BsmBI enzyme. This vector does not have a yeast ORI and will be integrated into the target locus through CRISPR and homologous recombination. Please click here to view a larger version of this figure.
| Name | TU1 | TU2 | TU3 | TU4 | Product |
| B/E-β-Carotene | BTS1/ERG20 | crtE | crtYB | crtI | β-carotene |
| B/E-Lycopene | BTS1/ERG20 | crtE | crtYBG247A | crtI | Lycopene |
| E-β-Carotene | ERG20 | crtE | crtYB | crtI | β-carotene |
| E-Lycopene | ERG20 | crtE | crtYBG247A | crtI | Lycopene |
Table 2: Multi-gene Plasmids used in this study.
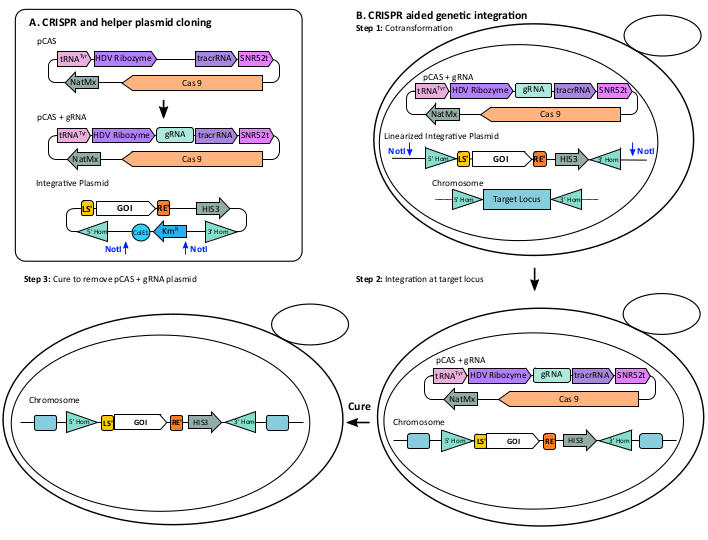
Figure 5: CRISPR integration. (A) CRISPR and helper plasmid cloning. The pCAS plasmid contains the Cas9 endonuclease and components (tRNA promoter, SNR52 terminator, HDV ribozyme, and tracrRNA) for optimal expression of a gRNA. Clone the pCAS+gRNA plasmid by assembling the synthetic gRNA with the linearized pCAS using Gibson cloning. (B) CRISPR aided genetic integration. Step 1: Cotransformation: pCAS +gRNA was co-transformed into yeast with the integrative plasmid containing the gene(s) of interest (GOI), a yeast selective marker, and 5' and 3' homology region targeting the genomic locus. For optimal integration, linearize the integrative plasmid with NotI. Step 2: Integration at target locus: Growing the transformed yeast on a plate selective for the yeast marker, either antibiotic or auxotrophic. Perform genotyping to confirm the integration. Step 3: Cure the pCAS + gRNA plasmid by streaking yeast on a non-selective plate. Please click here to view a larger version of this figure.
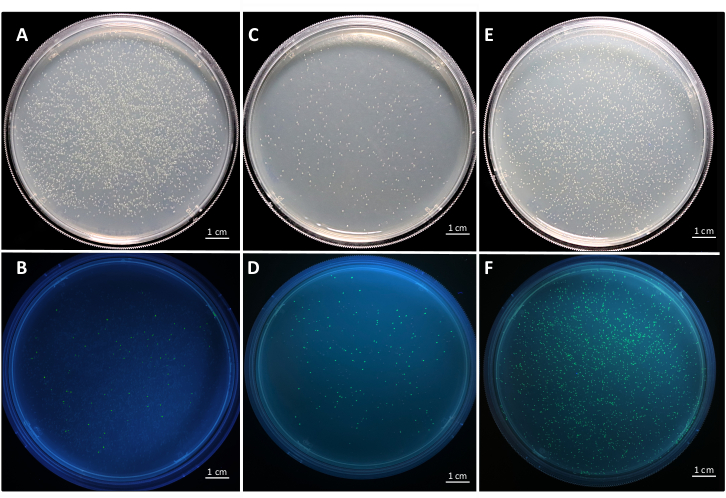
Figure 6: Representative plates of entry vector and transcription unit level cloning in E. coli. Representative plates of the successful cloning of a gene into the entry vector pYTK001 under (A) visible light and (B) UV light. Positive colonies are white and negative colonies are green. The ratio of positive colonies overall colonies is 90.35 ± 4.22 %. Successful assembly of the intermediate vector for transcription unit level assembly and green/red selection under (C) visible light and (D) UV light. Positive colonies are green. The ratio of positive colonies overall colonies is 17.56 ± 3.32 %. Successful assembly of transcription unit from the intermediate vector and green/white screening under (E) visible light and (F) UV light. Positive colonies are white and negative colonies are green.The ratio of positive colonies overall colonies is: 65.02 ± 3.32 %. Data are from three biological replicates. Please click here to view a larger version of this figure.
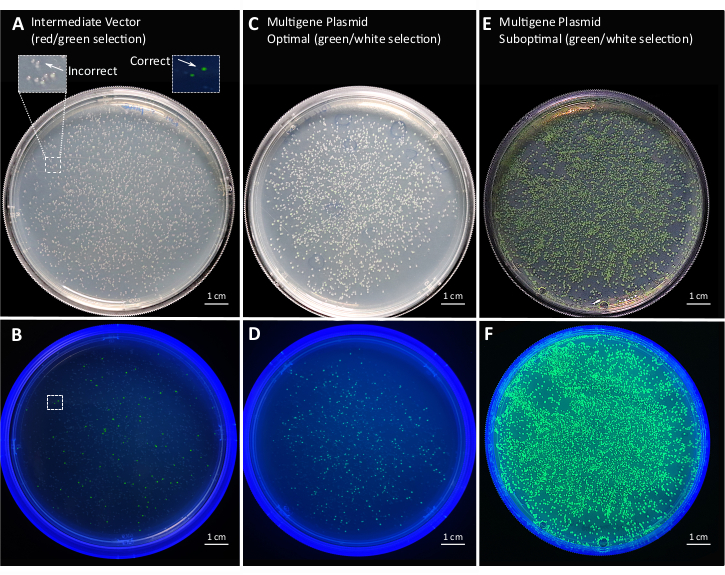
Figure 7: Representative plates of multi-gene plasmid cloning in E. coli. Assembly of the intermediate vector for multi-gene level assembly and red/green selection under (A) visible light and (B) UV light. Positive colonies are green and negative colonies are red. The ratio of positive colonies overall colonies is 1.83 ± 0.15%. Successful assembly of multiple TUs from the intermediate vector and green/white selection (C) under visible light and (D) UV light. Positive colonies are white. The ratio of positive colonies overall colonies is 93.77 ± 1.65%. Suboptimal assembly of multi-gene plasmid under (E) visible light and (F) UV light. The number of positive colonies is negligible. Data are from three biological replicates. Please click here to view a larger version of this figure.
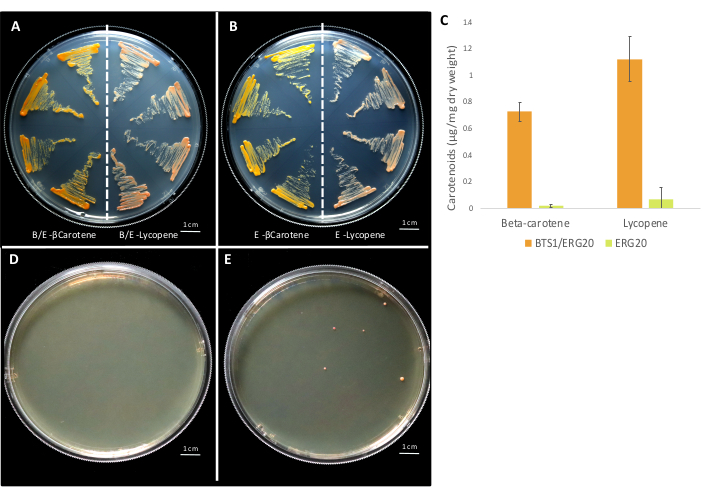
Figure 8: Representative plates of integrative and replicative plasmids in yeast. Replicative multi-gene plasmids transformed into yeast with (A) BTS1-ERG20 fusion TU and (B) ERG20 TU. On each plate, yeast on the left side has plasmids containing crtE TU, crtYB TU and crtI TU for the production of β–carotene and yeast on the right side has plasmids containing crtE TU, crtYB (G247A)TU, and crtI TU for the production of lycopene. (C) quantifying β-carotene and lycopene from yeast extract with four multi-gene plasmids using a UV-vis spectrometer. The maximal absorbance was recorded at 450 nm and 470 nm for β-carotene and lycopene respectively. The absolute quantification was performed using authentic standards (Supplementary Figure S3). Fusion of BTS1-ERG20 leads to the production of ~35-fold higher β-carotene and ~16.5-fold higher lycopene compared to ERG20 alone. Representative plates for the disruption of the ADE2 locus by the integration of a multi-gene integrative plasmid with a gRNA and no helper DNA (D) and with gRNA and a multi-gene integrative plasmid as helper DNA (E). Please click here to view a larger version of this figure.
Supplementary Materials. Please click here to download this file.
Discussion
The MoClo based cloning kit developed by Lee et al. provides an excellent resource for quick assembly of one to five transcription units into a multi-gene plasmid either for replication or integration into the yeast genome. The use of this kit eliminates the time-consuming cloning bottleneck that frequently exists for expressing multiple genes in yeast.
We tested five different conditions for the digestion/ligation cycles of Golden Gate cloning with T4 DNA ligase. We found that 30 cycles of digestion at 37 °C for 5 min and ligation at 16 °C for 5 min followed by a final digestion step at 50 °C for 10 min and a protein inactivation step at 80 °C for 10 min resulted in 99.5% colonies that were potentially correct in a 4-piece assembly (Supplementary Figure S1 and Table S3). Ligation at 16 °C ensures optimal ligase activity and overhang annealing. Final digestion at 50 °C prevents digested products from re-ligation. This cycle was followed for all of the assembly reactions unless otherwise specified.
We have also found some critical steps that need special attention for optimal results. We strongly recommend assembling intermediate vectors with the sfGFP dropout before assembling transcription units. In theory, all seven parts can be cloned into the E. coli vector with the mRFP1, followed by red/white screening. However, in practice, mRFP1 develops color very slowly and it is challenging to distinguish between pale red and pale-yellow E. coli colonies under visible light. As sfGFP absorbs at the UV range52, using a UV or a blue light transilluminator facilitates the screening for bright green colonies. Also, cloning an intermediate vector allows more facile assemble of the various promoter, CDS, and terminator combinations, enabling an easier combinatorial library creation, since an assembly with four parts usually results in more positive colonies than an assembly with more parts. Supplementary Figure S2 shows a progressive decrease in the ratio of potentially correct colonies from a 6-piece to an 8-piece assembly.
The concentrations of all of the parts must be measured meticulously to ensure the equimolar concentration of each part. Quantifying parts using a UV-Vis spectrometer is usually sufficient but more sensitive methods, such as fluorescence-based assays, may give better results. Failure to accurately quantify all of the parts often results in a low assembly efficiency. One should select the highly efficient Esp3I and BsaI-HFv2, respectively. We have observed that T4 works well for both overhangs in the kit and customized overhangs are necessary to fuse two parts, which is consistent with literature36, although the original paper recommended T7 ligase6. Increasing the number of cycles can increase the number of potentially correct colonies to some extent. For example, 25 cycles of digestion and ligation are enough to assemble three to four parts but 30-35 cycles give better results for more parts.
The MolClo strategy is advantageous over alternative multi-part assembly methods8,9,10,11,12 because it allows modular and highly versatile cloning. However, the primary limitation is the domestication step since parts sometimes have multiple BsmBI, BsaI, or NotI sites. Mutating all of the parts can be time-consuming. In this case, one may consider synthesizing the CDS without these restriction sites. However, for promoters and terminators, mutating even a single nucleotide may change their functionality. Therefore, validating the activity of these parts using a reporter assay is recommended39. Another limitation is that this kit only allows up to five transcription units in a multi-gene plasmid. If more transcription units are desired, additional connectors may be constructed by selecting a set of highly compatible overhangs36.
The kit provides 27 promoters, six terminators, seven yeast selection markers, and two yeast origin of replications. Customized parts, such as promoters and terminators, either native or synthetic, can be cloned into the entry vector following this protocol. The yeast MoClo kit has been used primarily for overexpressing multi-gene metabolic pathways to produce high-value chemicals in yeast. This protocol can also be used when different switches for biological circuits are desired in yeast. There is also tremendous potential to apply this kit for the investigation of basic biological questions about protein-protein interactions, protein localization, and enzyme activities. Overall, this protocol is extremely flexible and reliable and can support any demanding cloning in yeast biology.
Disclosures
The authors have nothing to disclose.
Acknowledgements
This work was funded by the Research Foundation for the State University of New York (Award #: 71272) and the IMPACT Award of University at Buffalo (Award #: 000077).
Materials
| 0.5 mm Glass beads | RPI research products | 9831 | For lysing yeast cells |
| Bacto Agar | BD & Company | 214010 | Component of the yeast complete synthetic medium (CSM) |
| Bacto Peptone | BD& Company | 211677 | Component of the yeast extract peptone dextrose medium (YPD) |
| BsaI-HFv2 | New England Biolabs | R3733S | a highly efficient version of BsaI restriction enzyme |
| Carbenicillin | Fisher Bioreagents | 4800-94-6 | Antibiotic for screening at the transcription unit level |
| Chloramphenicol | Fisher Bioreagents | 56-75-7 | Antibiotic for screening at the entry vector level |
| CSM-His | Sunrise Sciences | 1006-010 | Amino acid supplement of the yeast complete synthetic medium (CSM) |
| Dextrose | Fisher Chemical | D16-500 | Carbon source of the yeast complete synthetic medium (CSM) |
| Difco Yeast Nitrogen Base w/o Amino Acids | BD & Company | 291940 | Nitrogen source of the yeast complete synthetic medium (CSM) |
| dNTP mix | Promega | U1515 | dNTPs for PCR |
| Esp3I | New England Biolabs | R0734S | a highly efficient isoschizomer of BsmBI |
| Frozen-EZ Yeast Trasformation II Kit | Zymo Research | T2001 | For yeast transformation |
| Hexanes | Fisher Chemical | H302-1 | For carotenoid extraction from yeast cells |
| Kanamycin | Fisher Bioreagents | 25389-94-0 | Antibiotic for screening at the multigene plasmid level |
| LB Agar, Miller | Fisher Bioreagents | BP1425-2 | Lysogenic agar medium for E. coli culturing |
| LB Broth, Miller | Fisher Bioreagents | BP1426-2 | Lysogenic liquid medium for E. coli culturing |
| Lycopene | Cayman chemicals | NC1142173 | For lycopene quantification |
| MoClo YTK | Addgene | 1000000061 | Depositing Lab: John Deuber |
| Monarch Plasmid Miniprep Kit | New England Biolabs | T1010L | For plasmid purification from E.coli |
| Nanodrop Spectrophotometer | Thermo Scientific | ND2000c | For measuring accurate DNA concentrations |
| NotI-HF | New England Biolabs | R3189S | Restriction enzyme for integrative multigene plasmid linearization |
| Nourseothricin Sulphate | Goldbio | N-500-100 | Antibiotic Selection marker for the pCAS plasmid used in this study |
| Phusion HF reaction Buffer (5X) | New England Biolabs | B0518S | Buffer for PCR using Phusion polymerase |
| Phusion High Fidelity DNA Polymerase | New England Biolabs | M0530S | High fidelity polymerase for all the PCR reactions |
| pLM494 | Addgene | 100539 | Plasmid used to amplify crtI, crtYB and crtE used in this study |
| Quartz Cuvette | Thermo Electron | 10050801 | For quantifing carotenoids |
| T4 ligase | New England Biolabs | M0202S | Ligase for Golden Gate cloning |
| Thermocycler | BIO-RAD | 1851148 | For performing all the PCR and cloning reactions |
| Tissue Homogenizer | Bullet Blender | Model: BBX24 | For homogenization of yeast cells |
| UV-Vis Spectrophotometer | Thermo Scientific | Genesys 150 | For quantifing carotenoids |
| Yeast Extract | Fisher Bioreagents | BP1422-500 | Component of the yeast extract peptone dextrose medium (YPD) |
| β-carotene | Alfa Aesar | AAH6010603 | For β-carotene quantification |
References
- Endy, D. Foundations for engineering biology. Nature. 438 (7067), 449-453 (2005).
- Engler, C., Gruetzner, R., Kandzia, R., Marillonnet, S. Golden gate shuffling: a one-pot DNA shuffling method based on type IIs restriction enzymes. PLoS One. 4 (5), 5553 (2009).
- Shetty, R. P., Endy, D., Knight, T. F. Engineering BioBrick vectors from BioBrick parts. Journal of Biological Engineering. 2, 5 (2008).
- Peccoud, J., et al. A Modular cloning system for standardized assembly of multigene constructs. PLoS ONE. 6 (2), (2011).
- Sarrion-Perdigones, A., et al. GoldenBraid: an iterative cloning system for standardized assembly of reusable genetic modules. PLoS One. 6 (7), 21622 (2011).
- Lee, M. E., DeLoache, W. C., Cervantes, B., Dueber, J. E. A Highly characterized yeast toolkit for modular, multipart assembly. ACS Synthetic Biology. 4 (9), 975-986 (2015).
- Andreou, A. I., Nakayama, N. Mobius assembly: A versatile Golden-Gate framework towards universal DNA assembly. PLoS One. 13 (1), 0189892 (2018).
- Gibson, D. G., et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nature Methods. 6 (5), 343-345 (2009).
- Li, M. Z., Elledge, S. J. Harnessing homologous recombination in vitro to generate recombinant DNA via SLIC. Nature Methods. 4 (3), 251-256 (2007).
- Geu-Flores, F., Nour-Eldin, H. H., Nielsen, M. T., Halkier, B. A. USER fusion: a rapid and efficient method for simultaneous fusion and cloning of multiple PCR products. Nucleic Acids Research. 35 (7), 55 (2007).
- de Kok, S., et al. Rapid and reliable DNA assembly via ligase cycling reaction. ACS Synthetic Biology. 3 (2), 97-106 (2014).
- Shao, Z., Zhao, H., Zhao, H. DNA assembler, an in vivo genetic method for rapid construction of biochemical pathways. Nucleic Acids Research. 37 (2), 16 (2009).
- Gibson, D. G. Synthesis of DNA fragments in yeast by one-step assembly of overlapping oligonucleotides. Nucleic Acids Research. 37 (20), 6984-6990 (2009).
- Engler, C., et al. A golden gate modular cloning toolbox for plants. ACS Synthetic Biology. 3 (11), 839-843 (2014).
- Gantner, J., et al. Peripheral infrastructure vectors and an extended set of plant parts for the Modular Cloning system. PLoS One. 13 (5), 0197185 (2018).
- Ordon, J., et al. Generation of chromosomal deletions in dicotyledonous plants employing a user-friendly genome editing toolkit. Plant Journal. 89 (1), 155-168 (2017).
- Sarrion-Perdigones, A., et al. GoldenBraid 2.0: a comprehensive DNA assembly framework for plant synthetic biology. Plant Physiology. 162 (3), 1618-1631 (2013).
- Agmon, N., et al. Yeast Golden Gate (yGG) for the efficient assembly of S. cerevisiae transcription units. ACS Synthetic Biology. 4 (7), 853-859 (2015).
- Hernanz-Koers, M., et al. FungalBraid: A GoldenBraid-based modular cloning platform for the assembly and exchange of DNA elements tailored to fungal synthetic biology. Fungal Genetics and Biology. 116, 51-61 (2018).
- Prielhofer, R., et al. GoldenPiCS: a Golden Gate-derived modular cloning system for applied synthetic biology in the yeast Pichia pastoris. BMC Systems Biology. 11 (1), 123 (2017).
- Celinska, E., et al. Gate Assembly system dedicated to complex pathway manipulation in Yarrowia lipolytica. Microbial Biotechnology. 10 (2), 450-455 (2017).
- Pullmann, P. K., et al. A modular two yeast species secretion system for the production and preparative application of fungal peroxygenases. bioRxiv. , (2020).
- Wu, D., Schandry, N., Lahaye, T. A modular toolbox for Golden-Gate-based plasmid assembly streamlines the generation of Ralstonia solanacearum species complex knockout strains and multi-cassette complementation constructs. Molecular Plant Pathology. 19 (6), 1511-1522 (2018).
- Moore, S. J., et al. EcoFlex: A multifunctional MoClo kit for E. coli synthetic biology. ACS Synthetic Biology. 5 (10), 1059-1069 (2016).
- Iverson, S. V., Haddock, T. L., Beal, J., Densmore, D. M. CIDAR MoClo: Improved MoClo assembly standard and new E. coli part library enable rapid combinatorial design for synthetic and traditional biology. ACS Synthetic Biology. 5 (1), 99-103 (2016).
- Vasudevan, R., et al. CyanoGate: A modular cloning suite for engineering cyanobacteria based on the plant MoClo syntax. Plant Physiology. 180 (1), 39-55 (2019).
- Leonard, S. P., et al. Genetic Engineering of bee gut microbiome acteria with a Ttoolkit for modular assembly of broad-host-range plasmids. ACS Synthetic Biology. 7 (5), 1279-1290 (2018).
- Schwartz, M. L., Jorgensen, E. M. SapTrap, a toolkit for high-throughput CRISPR/Cas9 gene modification in Caenorhabditis elegans. 유전학. 202 (4), 1277-1288 (2016).
- Martella, A., Matjusaitis, M., Auxillos, J., Pollard, S. M., Cai, Y. EMMA: An extensible mammalian modular assembly toolkit for the rapid design and production of diverse expression vectors. ACS Synthetic Biology. 6 (7), 1380-1392 (2017).
- Chiasson, D., et al. A unified multi-kingdom Golden Gate cloning platform. Scientific Reports. 9 (1), 10131 (2019).
- Denby, C. M., et al. Industrial brewing yeast engineered for the production of primary flavor determinants in hopped beer. Nature Communications. 9 (1), 965 (2018).
- Awan, A. R., et al. Biosynthesis of the antibiotic nonribosomal peptide penicillin in baker’s yeast. Nature Communications. 8, 15202 (2017).
- Leavitt, J. M., et al. Biosensor-Enabled Directed evolution to improve muconic acid production in Saccharomyces cerevisiae. Biotechnology Journal. 12 (10), (2017).
- Hsu, T. M., et al. Employing a biochemical protecting group for a sustainable indigo dyeing strategy. Nature Chemical Biology. 14 (3), 256-261 (2018).
- Grewal, P. S., Modavi, C., Russ, Z. N., Harris, N. C., Dueber, J. E. Bioproduction of a betalain color palette in Saccharomyces cerevisiae. Metabolic Engineering. 45, 180-188 (2018).
- Potapov, V., et al. Comprehensive profiling of four base overhang ligation fidelity by T4 DNA ligase and application to DNA assembly. ACS Synthetic Biology. 7 (11), 2665-2674 (2018).
- Vazquez-Vilar, M., et al. Software-assisted stacking of gene modules using GoldenBraid 2.0 DNA-assembly framework. Methods in Molecular Biology. 1284, 399-420 (2015).
- Pullmann, P., et al. Mutagenesis: An efficient multi-site-saturation mutagenesis approach by Golden Gate cloning with automated primer design. Scientific Reports. 9 (1), 10932 (2019).
- Engler, C., Sylvestre, M., Valla, S., Lale, R. . DNA Cloning and Assembly Methods. 1116, 119-131 (2014).
- Liu, H., Naismith, J. H. An efficient one-step site-directed deletion, insertion, single and multiple-site plasmid mutagenesis protocol. BMC Biotechnology. 8, 91 (2008).
- Chen, X., Zaro, J. L., Shen, W. C. Fusion protein linkers: property, design and functionality. Advanced Drug Delivery Reviews. 65 (10), 1357-1369 (2013).
- Mullis, K., et al. Specific enzymatic amplification of DNA in vitro: the polymerase chain reaction. Cold Spring Harbor Symposia on Quantitative Biology. 51, 263-273 (1986).
- Naito, Y., Hino, K., Bono, H., Ui-Tei, K. CRISPRdirect: software for designing CRISPR/Cas guide RNA with reduced off-target sites. Bioinformatics. 31 (7), 1120-1123 (2015).
- Labun, K., et al. CHOPCHOP v3: expanding the CRISPR web toolbox beyond genome editing. Nucleic Acids Research. 47 (1), 171-174 (2019).
- Ryan, O. W., et al. Selection of chromosomal DNA libraries using a multiplex CRISPR system. Elife. 3, (2014).
- Gietz, R. D., Schiestl, R. H. High-efficiency yeast transformation using the LiAc/SS carrier DNA/PEG method. Nature Protocols. 2 (1), 31-34 (2007).
- Heintz, N., Gong, S. Small-scale preparations of yeast DNA. Cold Spring Harbor Protocols. 2020 (10), (2020).
- Hochrein, L., Mitchell, L. A., Schulz, K., Messerschmidt, K., Mueller-Roeber, B. L-SCRaMbLE as a tool for light-controlled Cre-mediated recombination in yeast. Nature Communications. 9 (1), 1931 (2018).
- Verwaal, R., et al. High-level production of beta-carotene in Saccharomyces cerevisiae by successive transformation with carotenogenic genes from Xanthophyllomyces dendrorhous. Applied and Environmental Microbiology. 73 (13), 4342-4350 (2007).
- Jones, E., Strathern, J. N., Jones, E. W., Broach, J. R., et al. . The Molecular Biology of the Yeast Saccharomyces: Metabolism and Gene Expression. 11, 181 (1982).
- Sehgal, N., et al. CRISPR gene editing in yeast: an experimental protocol for an upper-division undergraduate laboratory course. Biochemistry and Molecular Biology Education. 46 (6), 592-601 (2018).
- Stepanenko, O. V., Stepanenko, O. V., Kuznetsova, I. M., Verkhusha, V. V., Turoverov, K. K. Sensitivity of superfolder GFP to ionic agents. PLoS One. 9 (10), 110750 (2014).

