Stepwise Cell Seeding on Tessellated Scaffolds to Study Sprouting Blood Vessels
Summary
Engineered tissues heavily rely on proper vascular networks to provide vital nutrients and gases and remove metabolic waste. In this work, a stepwise seeding protocol of endothelial cells and support cells creates highly organized vascular networks in a high-throughput platform for studying developing vessel behavior in a controlled 3D environment.
Abstract
The cardiovascular system is a key player in human physiology, providing nourishment to most tissues in the body; vessels are present in different sizes, structures, phenotypes, and performance depending on each specific perfused tissue. The field of tissue engineering, which aims to repair or replace damaged or missing body tissues, relies on controlled angiogenesis to create a proper vascularization within the engineered tissues. Without a vascular system, thick engineered constructs cannot be sufficiently nourished, which may result in cell death, poor engraftment, and ultimately failure. Thus, understanding and controlling the behavior of engineered blood vessels is an outstanding challenge in the field. This work presents a high-throughput system that allows for the creation of organized and repeatable vessel networks for studying vessel behavior in a 3D scaffold environment. This two-step seeding protocol shows that vessels within the system react to the scaffold topography, presenting distinctive sprouting behaviors depending on the compartment geometry in which the vessels reside. The obtained results and understanding from this high throughput system can be applied in order to inform better 3D bioprinted scaffold construct designs, wherein fabrication of various 3D geometries cannot be rapidly assessed when using 3D printing as the basis for cellularized biological environments. Furthermore, the understanding from this high throughput system may be utilized for the improvement of rapid drug screening, the rapid development of co-cultures models, and the investigation of mechanical stimuli on blood vessel formation to deepen the knowledge of the vascular system.
Introduction
The field of tissue engineering is rapidly progressing towards the fabrication of engineered constructs to replace missing or damaged organs and tissues1. However, fully functional constructs have yet to be achieved, in part, since generating operational vascular networks for tissue nourishment remains an outstanding challenge. Without proper vascularization, engineered tissues are limited to a passive diffusion transport of oxygen and nutrients, constraining the maximum viable tissue thickness to the diffusion limit, approximately 200 µm2. Such thicknesses are not suitable to repair large tissue defects or for full organ fabrication, which renders the presence of functional vascular network a mandatory characteristic for functional and implantable tissues3.
The vascular system is comprised of a wide variety of blood vessels, with different sizes, phenotypes, and organization, tightly related to the host tissue. Understanding the behavior, response and migration decisions made by the developing and sprouting vessels can instruct their integration in engineered tissues4. Currently, the most common approach for creating in vitro vascular networks is combining endothelial cells (ECs) with support cells (SCs, with the capability to differentiate into mural cells), seeded within a three-dimensional micro-environment. This environment provides chemical and physical cues to allow the cells to attach, proliferate and self-assemble into vessel networks2,5,6,7,8. When co-cultured, SCs secrete extracellular matrix (ECM) proteins while providing mechanical support to the ECs, which form the tubular structures. Furthermore, a cross-interaction between both cell types promote tubulogenesis, vessel sprouting and migration, in addition to the SCs maturation and differentiation into α-smooth muscle actin-expressing (αSMA) mural cells4. Vessel network development is most commonly studied in 3D environments created using hydrogels, porous polymeric scaffolds, or a combination thereof. The latter option equally provides a cell-friendly environment and the required mechanical support for both the cells and the ECM9.
A great amount of work has been carried out to study vascular development, including co-culturing the cells on hydrogels10, hydrogels-scaffold combinations11,12, 2D platforms, and microfluidic devices13. However, hydrogels can be easily deformed by the cell-exerted forces14, while 2D and microfluidics systems fail to recreate a closer-to-nature environment to obtain a more extrapolatable response15,16. Understanding how forming vessels react to their surrounding environment can provide critical insight that might allow for the fabrication of engineered environments with the capability of guiding the vessel development in a predictable manner. Understanding vascular formation phenomena is especially critical to keep pace with the rapid emergence of submicron-to-micron scale fabrication techniques, such as stereolithography, digital projection lithography, continuous liquid interface production, 3D melt-electro jetwriting, solution based 3D electro jet writing, and emerging bioprinting techniques17,18,19,20,21. Aligning the control of these micromanufacturing techniques with a deepened understanding of vascular biology is key to the creation of an appropriate engineered vasculature for a target tissue.
Here, we present a 3D system to study the response of new forming and sprouting vessels to the surrounding scaffold geometry, observing their sprout origin and subsequent migration22. By utilizing 3D scaffolds with tessellated compartment geometries, and a two-step seeding technique, we succeeded to create highly organized vascular networks in a clear and easy to analyze fashion. The tessellated geometries provide a high throughput system with individual units containing vessels that respond to their local environment. Using multicolored ECs, we tracked sprout formation origins and subsequent migration patterns, correlated to the compartment geometry and the SCs location22.
Although the proposed protocol has been prepared to analyze the effects of geometrical cues on vascularization behavior, this approach can be expanded and applied to a variety of new applications. The tessellated scaffold and the easily imageable networks allow for the straightforward analysis of different ECs and SCs interaction, the addition of specific organ cells and their interaction with the vascular networks, drug effect on vascular networks, and more. Our suggested system results very versatile and of simple fabrication and processing.
Protocol
1. Tessellated scaffold fabrication
NOTE: Photolithography is a widespread technique that requires specialized equipment typically housed within a nanofabrication facility/laboratory. The method laid out in this protocol was generalized as much as possible for the audience; however, slight changes to procedures may be necessary depending on equipment available to the reader. We recommend performing these procedures in a clean room at a nanofabrication facility to ensure the highest process quality. Before beginning, obtain access to a mask-aligner (or some UV-exposure set-up), a spin coater, hot plates, a solvent washing station, a photomask, and a plasma cleaner. The solvents and chemicals used in procedure are hazardous, so please take the upmost care to avoid any chemical exposure. When designing the photomask, identify what sized silicon wafers and photomasks are compatible with the spin-coater and mask-aligner. Additionally, the photoresist located toward edges of the silicon wafer is typically deformed from handling; hence, make the designs toward the center area of the wafer.
- Prepare scaffolds using a photolithography technique with the selected geometry of interest.
- Before spin coating, clean the silicon wafer. This can be done with plasma cleaning or a solvent-based technique. If plasma cleaning, follow the standard operating procedure of the instrument for operational details. Spray with compressed nitrogen gas and inspect the wafer to ensure it is free of debris before spin-coating. This wafer will serve as the substrate upon which the scaffolds are created.
NOTE: For best results start with a fresh wafer. Wafers can be re-used but should be free of old photoresist, surface defects and debris. It is helpful to use wafer-tweezers for handling. - Center a silicon wafer onto the spin-coater chuck using a guide. Briefly spin the wafer to ensure it has been properly centered on the chuck, adjust as necessary until properly centered. This must be done every time a wafer is placed on the spin-coater. Dispense 1-4 mL of the lift-off reagent onto the wafer (this will depend on the size of the wafer). Approximately 2 mL of lift-off reagent works well for a 4-inch wafer.
- Set spin-coater spread speed at 500 rpm for 5 seconds, and the spin-speed to 1000 rpm for 30 seconds, then spin the lift-off reagent. Inspect the wafer to ensure it has an even coating across the wafer. Any debris left on the wafer will be very obvious at this point. Any scaffolds in an area with debris will likely be unusable. After spin coating, transfer to a hot plate and bake for 1 minute at 200 °C.
- Repeat the previous step two more times for a total of three coatings.
- Spin-coat the lift-off reagent coated silicon wafer with SU-8 2050 photoresist until obtaining a thickness of approximately 100 µm.
- Dispense 1 mL of resist for every 25 mm of substrate diameter.
NOTE: Be careful to avoid bubbles while pouring the resist. For SU-8 2050, pouring approximately a 2-inch diameter circle, when using a 4-inch silicon wafer works well. - Spin-coat the SU-8 2050. Spread at a speed of 500 rpm for 5 seconds, followed by a spin speed between 1700 to 1800 rpm to achieve approximately 100 µm thickness.
- Optional: After spinning, leave the spin coated wafers to de-gas overnight on a level surface protected from light prior to the pre-bake. This may help to rid the resist of any bubbles and allow defects to level out.
NOTE: The actual thickness of the resist and the resultant scaffolds can vary with user error and equipment parameters. Therefore, the thickness of the scaffolds should be verified later, in step 1.6. Modify spin coating procedures accordingly to achieve the desired thickness.
- Dispense 1 mL of resist for every 25 mm of substrate diameter.
- Prior to exposure, pre-bake the wafers at 65 °C for 10 min and then 95 °C for 40-50 min. Prior to exposure, test the surface of the resist by pressing on the edge with a pair of tweezers to ensure that it is no longer tacky/viscous. It is helpful to pull the wafer off of the hot plate and allow it to cool for a 0.5-1 min before assessing the tackiness.
NOTE: Slow temperature ramp times (heating and cooling) may help to prevent warping of the scaffolds.
- Before spin coating, clean the silicon wafer. This can be done with plasma cleaning or a solvent-based technique. If plasma cleaning, follow the standard operating procedure of the instrument for operational details. Spray with compressed nitrogen gas and inspect the wafer to ensure it is free of debris before spin-coating. This wafer will serve as the substrate upon which the scaffolds are created.
- Expose the photoresist to UV light (350-400 nm) through a photomask with an exposure energy of 215-240 mJ/cm2 for a resist thickness of 85 – 110 µm. Refer to photoresist manufacturer guidelines for recommended exposure energy-thickness correlations.
- Ensure that the exposure is properly calibrated such that the energy is verified. Adjust the exposure time as necessary to achieve desired exposure energy. Furthermore, mask aligners may allow for different exposure modes: vacuum contact, hard contact, soft contact, and proximity contact (specific terms may vary). Generally, these will impact resolution and feature alignment. The authors typically used a "hard contact" mode.
- For small features or multiple layers, where alignment will matter, consider exposure modes and mask aligner procedures more carefully. Be sure to consider the height of the resist when adjusting the thickness value. Refer to the standard operating procedures of your mask-aligner for operation details.
NOTE: The design of the photomask will determine the scaffolds size, compartment geometry, and number of scaffolds obtained per batch. The use of a hard glass or quartz photomask will yield the highest resolution; however, a soft polymer transparent film photomask generally can be used for these large feature sizes (>10 µm). The use of a film-photomask may lead to topographical features along the z-axis of the scaffold pores as a result of poorer feature resolution. This could potentially have an influence on cell behavior. Verify the desired resolution consult with whomever is producing the photomask to ensure the desired resolution is achieved.
- Bake the wafer immediately after UV exposure at 65 °C for 2-5 minutes and then at 95 °C for 8-10 minutes.
- Develop the scaffolds to remove undeveloped resist.
- Immerse the scaffolds using a low volume of SU-8 developer solution for 7-10 minutes to dissolve any undeveloped resist.
- Rinse with isopropyl alcohol (IPA).
- Dry wafer with compressed nitrogen.
NOTE: Be careful during step 1.4 to not cause premature release of the scaffolds. Low volumes of developer and gentle handling are necessary to achieve this.
- Following development, hard bake the wafers at 150 °C for 15 min.
NOTE: After hard bake, turning off the hotplate to allow the wafers to cool very slowly may be advantageous in preventing warping/curling of the final scaffolds. - Optional: Following the hard bake and prior to lift-off, assess scaffold thickness since the scaffolds should still be gently adhered to the surface of the wafer. For this procedure, contact profilometry works well; however, any appropriate method could be employed.
- Lift the scaffolds off the wafer.
- Submerge the scaffolds in SU-8 developer to cause them to immediately lift off the wafer. If scaffolds do not lift off, gently push the scaffolds with a pair of wafer-tweezers.
- Remove excess developer.
- Transfer the scaffolds into new container and submerge in isopropyl alcohol. Rinse in IPA for a minimum of 6 times to ensure all developer has been removed.
- Air dry the scaffolds for a minimum of a week before use.
2. Scaffold fibronectin coating
- For disinfection, submerge the scaffolds in 70% ethanol (v/v) for a minimum of 15 minutes, and wash 2 times in phosphate buffered saline (PBS) before use.
CAUTION: The SU-8 scaffolds are frail and can break easily. Handling is recommended using blunt and curved-tip forceps, and they should be handled with upmost care. Easier handling can be achieved by submerging the scaffolds in liquid, to avoid electrostatic interactions between the scaffolds and the surface. - Prepare a dilution of 50 µg/mL of human fibronectin in PBS and cover the scaffolds by protein adsorption.
- Mix 1.5 µL of fibronectin stock solution (1 mg/mL) with 28.5 µL of PBS per each scaffold to be seeded. Preferably, prepare only one fibronectin dilution to be used for all scaffolds to avoid pipetting errors.
- For 10 scaffolds, mix 15 µL of stock fibronectin solution in 285 µL of PBS, amounting to a total of 300 µL of 50 µg/mL fibronectin dilution.
- Place the scaffolds sparsely on top of a hydrophobic surface (i.e., non-tissue culture (nonTC) 10 cm dish) and cover each scaffold with 30 µL of the fibronectin dilution prepared in step 2.2.1.
CAUTION: Make sure that no bubbles are trapped in the scaffold compartments. If bubbles are present, they can be removed by gently shaking the scaffold within the droplet or light pipetting. - Replace the plate's lid and place the fibronectin-covered scaffolds into an incubator with 37 °C and 100% humidity for a minimum of an hour.
- After incubation, lightly rinse the scaffolds in PBS to remove fibronectin remnants. The scaffolds can be kept in PBS at 4 °C for up to a week before use.
- Mix 1.5 µL of fibronectin stock solution (1 mg/mL) with 28.5 µL of PBS per each scaffold to be seeded. Preferably, prepare only one fibronectin dilution to be used for all scaffolds to avoid pipetting errors.
3. Endothelial cells seeding
- Prepare EC medium by mixing the basal medium with its correspondent medium kit components, including an antibiotic solution (Pen/Strep), fetal bovine serum (FBS), and endothelial cell growth supplements, as indicated by the manufacturer.
- Make a human adipose microvascular endothelial cell (HAMEC) suspension in EC medium with a concentration of 4 x 106 cells/mL.
NOTE: Real time imaging can be performed if the used endothelial cells are previously transfected to express a fluorescent protein, or pre-stained using a non-toxic cytoplasmic membrane dye (furtherly referred as labeled ECs). - Using forceps, place one fibronectin-coated scaffold per well on a nonTC 24-well plate well.
- Cover each scaffold with 25 µL droplets of the HAMEC suspension. Beware not to let the suspension flow away from the scaffold, since it can hinder cell adhesion.
- Put the lid on the plate and place it in an incubator at 37 °C, 5% CO2 and 100% humidity. Let incubate for a minimum of 60 minutes and a maximum of 90 minutes.
NOTE: During incubation, placing the plate on an orbital shaker within the incubator improves cell attachment onto the scaffold's walls. The orbital shaker should be set at no more than 5 rpm. - After incubation, fill each well with 700 µL of EC medium using a pipette.
NOTE: Expect to see ECs at the bottom of the well, since some of the cells do not attach to the scaffold and fall through the scaffold's voids. - Incubate the endothelialized scaffolds until EC confluence can be observed using fluorescent microscopy on the scaffold's walls (when using labeled ECs), or for 3 days (when using non-labeled ECs), which provides enough time to achieve EC confluence. Change the medium every other day.
4. Support cell seeding and co-culture
- Prepare dental pulp stem cell (DPSC) medium (DPSCm) by mixing 500 mL of low-glucose Dulbecco's modified Eagle medium (low-glucose DMEM), 57.5 mL of FBS, 5.75 mL of non-essential amino acids (NEAA), 5.75 mL of GlutaMAX, and 5.75 mL of penicillin-streptomycin-nystatin solution.
- Transfer the endothelialized scaffolds into a new nonTC 24-well plate to discard ECs attached to the wells' bottom using forceps.
- Discard all media from the current plate using a 1 mL pipette, or vacuum suction. Be careful not to apply vacuum straight on the scaffold.
- Place one scaffold per well, preferably at the center of the well to avoid running liquid toward the walls due to surface tension effects. Dry the area surrounding the scaffold with light vacuum but avoid complete drying of the scaffold.
- Prepare a DPSC suspension in fibrin pre-gel solution.
- Dilute thrombin and fibrinogen stock solutions with PBS until obtaining a final concentration of 5 U/mL and 15 mg/mL, respectively.
- Prepare an 8 x 106 DPSC/mL suspension in the 5 U/mL thrombin dilution and distribute in individual eppendorfs 12.5 µL of said suspension per scaffold to be seeded.
- Set a 5-50 µL pipette (or similar) to 12.5 µL and fill it with the 15 mg/mL fibrinogen solution.
- Without removing the tip, set the pipette to 25 µL. The material in the tip should rise and leave an empty volume towards the tip opening.
- Slowly press the plunger button until the liquid reaches the tip opening but does not leak out. Hold the plunger in this position and put the tip into one of the microcentrifuge tubes containing the cells in thrombin suspension, making sure the tip is in contact with the liquid.
CAUTION: The fibrin crosslinking will begin immediately after the thrombin and fibrinogen come into contact. The following step should be done quickly. - Gently release the plunger button and draw the cell suspension into the tip. Thoroughly mix both materials, avoiding bubble formation.
- Quickly dispense the mixed materials on top of an endothelialized scaffold. Repeat the previous steps for each scaffold, making sure to always change tips between uses to avoid unexpected fibrin gel formation within the tip.
- Replace the plate lid and incubate the scaffolds at 37 °C, 5% CO2 and 100% humidity for 30 minutes.
- After incubation, fill each well with 1 mL of 1:1 DPSC and EC medium.
- Culture for 1 week, changing medium every other day.
- During culture, remove the medium from the well and image the constructs using a confocal microscope at different time points to study the vascular development or any other parameter of interest.
NOTE: This step can only be performed if the experiment is performed using labeled ECs. See Step 6 – NOTE for further clarification.
5. Immunofluorescent staining for characteristic vascular markers
NOTE: The following steps can be performed in the same wells where the constructs were cultured. It is critical to always make sure that the solutions completely cover the scaffolds. Moreover, when possible, performing the steps on an orbital shaker is recommended, although not mandatory.
- At day 7, discard the media from the wells and rinse the scaffolds by adding PBS to the wells, and removing it using vacuum.
- Fix the scaffolds by adding 4% paraformaldehyde for 20 minutes. Wash in PBS 3 times, 5 minutes per wash.
- Permeabilize the fixed cells using 0.3% Triton-X in PBS (v/v) for 15 minutes. Wash in PBS 3 times, 5 minutes per wash.
- Prepare a 5% bovine serum albumin (BSA) in PBS (w/v) blocking solution and cover the scaffolds. Leave at room temperature for 1 hour.
NOTE: All the required solution amounts will depend on the number of scaffolds to be stained. Prepare the solutions as needed keeping the stated concentrations. - Prepare and apply the primary antibodies staining solution .
- Dilute rabbit anti-von Willebrand factor (vWF) antibody 1:150 and mouse anti-SMA antibody 1:50 in fresh blocking solution.
NOTE: Other endothelial cells markers, such as CD31 or VE-Cadherin can be used. - Cover the scaffolds with the primary antibodies solution and keep at 4 °C overnight. Wash in PBS three times, 5 minutes per wash.
- Dilute rabbit anti-von Willebrand factor (vWF) antibody 1:150 and mouse anti-SMA antibody 1:50 in fresh blocking solution.
- Prepare and apply the secondary antibodies staining solution.
- Dilute goat anti-mouse Cy3 antibody 1:150 and goat anti-rabbit Alexa-Fluor 488 1:400 in PBS.
- Cover the scaffolds with the secondary antibodies solution and incubate for a minimum of 3 hours at room temperature, or at 4 °C overnight. Wash 3 times in PBS, 5 minutes per wash. Keep stained scaffold in 0.3% PFA in PBS (v/v) for up to a month.
6. Scaffold confocal imaging and vessel development analysis
NOTE: The following steps can be performed on the fixed and stained scaffolds at the chosen final time point or, if fluorescent cells were used, during the cell culture period without the need to terminate the experiment. For the latter, it is recommended to set specific time points; this work shows day 0 (before SCs seeding), and days 1, 3, 5 and 7 after SCs seeding (Figure 3A).
- Image the scaffolds using a confocal microscope with a 5X lens and a zoom of 0.5 and the full z range of the scaffold. This will allow to capture 9 separate compartments for the squared and circular geometries, or 8 full compartments for the hexagonal geometry. Image the fluorescent signal and the transmitted light, to get both the vessel network structure and the compartment geometry (Figure 3A).
- Using ImageJ (or any other image processing software), crop each individual compartment; for this process, only the vascular networks markers are relevant. Remove any vessel not located inside the compartment area. Create a maximum intensity projection and save each individual cropped compartment as a TIFF image.
- In ImageJ, press File > Open and select the desired TIFF image containing several compartments. The image should have at least two channels, the fluorescent vessel network, and the transmitted light image.
- From the toolbar, select the Polygon selection tool. Select the transmitted light channel. Click on one of the corners of the hexagonal compartment, at the interface between the compartment area and the scaffold wall, this will start the mask definition. Continue clicking on the next corners in a sequential order until the initial corner is selected, closing the mask.
NOTE: This explanation applies to a hexagonal compartment. When the compartment is either squared or circular, use the Rectangle or Oval tools, respectively, to create an appropriate mask. Regardless the shape, always fit the mask to follow the inner wall of the scaffold. - Click on Analyze > Tools > ROI manager. In the ROI manager (ROI, region of interest), click the Add button to save the created mask. The new mask will show up in the left side list. This step allows the analysis to be consistent between different images.
NOTE: In the ROI manager, it is possible to save a file with all the created masks by clicking on More > Save. In the Save selection window, choose a name and location, and save the .roi file. To retrieve the mask, in the ROI manager, click on More > Open and choose the desired .roi file. - Select the fluorescent vessel network image and click on Edit > Clear outside. Only the image contained in the mask will remain, while the rest will be eliminated.
- With the cleared network image open, click on Image > Duplicate. In the Duplicate pop-up window, write the name of the image. If the Duplicate hyperstack is selected, unselect it and click OK. A new image containing only the cropped fluorescent vessel networks will be created.
- Save the cropped image by clicking on File > Save as > Tiff. Choose the name and directory and click OK.
- Analyze the vascular development on the cropped images.
- Download and install the analytical free software AngioTool, which provides quantitative tools to quantify a variety of vascular parameters. The software can be obtained from the following address https://ccrod.cancer.gov/confluence/display/ROB2/Home.
- Set the scale value of the images to obtain results in millimeter units and load an image.
- Click on the Setting tab and fill the Distance in pixels and Distance in mm with representative values from the taken images. Using the imaging parameters mentioned above, the values to fill are 400 pixels per 1 mm.
- Press the Open image button and browse for the image of interest.
- Choose the analysis parameters to quantify vessel development in the Analysis tab.
- Under the Vessel diameter and intensity field, deselect any present marker from the top range selector, and activate the marker representing the number 3.
- Under the Vessel diameter and intensity field, on the bottom range selector, move the lower range marker to 60, and leave the high ranger maker at 255.
- Activate the Remove small particles box and set the marker to 100.
- Under the field Saving preferences, select the name and location of the spread sheet file in which the results will be saved and press the Run analysis button. If the Save result image box is selected, Angiotool will generate and save a new jpeg image showing the vessel network quantified parameters.
NOTE: For each performed analysis without changing the spread sheet file name, a new row will be added at the bottom of the file, with the newly acquired results.
- Organize the data and run a statistical analysis.
- For all the processed images, choose the parameter of interest from the spread sheet (e.g. Total vessel length and Vessel area) and input them into a statistical analysis software to properly study the obtained results (Figure 3C).
7. Time lapse imaging for sprouting origin detection and migration tracking
- Alternatively, to culturing the scaffolds in the incubator as described in step 4.6, set a confocal microscope culture chamber to 37 °C, 5% CO2 and 100% humidity.
- Put the plate containing the fully seeded scaffolds in the confocal microscope chamber and set the imaging parameters to the desired values.
- Set the image taking interval to 30 minutes, and the total imaging time of 72 hours. This will enable to run the time lapse completely without the need for a medium change.
CAUTION: In case a longer time lapse imaging is desired, a careful position calibration should be performed to be able to retrieve the original imaging location, since medium changes require removing the plate from the confocal incubator and into the sterile biological hood.
NOTE: ECs detachment from the scaffold walls and tubular structures formation will start within the first hour after SCs seeding (step 4) and may take up to 50 hours to complete. After initial tube formation, new sprouts will start migrating into the compartment from the surrounding vessels. Sprouting and subsequent network remodeling will continue throughout the experiment approximately until day 10, when the vessel networks become stable22. Use this information to choose the time lapse starting point and time step accordingly.
- Save the complete time lapse movie file as a TIFF sequence.
- Open ImageJ and click File > Import > Image sequence and select your files. This will open your TIFF sequence as a movie.
- Open the Manual Tracking plugin by clicking on Plugins > Manual Tracking. This will open the Tracking window.
- Observe the movie and select the vessel to be tracked. Pay attention to its origin and migration.
- Activate the Show parameters box and input the 30 minutes for Time interval. The x/y calibration will depend on the pixel amount selected during imaging.
- Proceed to manually track the sprouting vessels from the scaffold contour.
- Press the Add track button. On the movie's first frame, press the location of the vessel to be tracked. After this, the movie will change to the next frame automatically.
- Click on the image of the second frame where the vessel is located. Again, the movie will change to the next frame automatically.
- Proceed with the same vessel until the full migration tracking has been completed.
- On the Tracking window, click on the Overlay Dots & Lines to imprint a visual path of the vessel migration on the movie. This will open a copy of the movie with a dot representing the current position of the tracker, and a line showing the previous path.
- Click on the End Track button to finalize recording the current vessel movement.
- Click on the Add track button again to start over with a new vessel. When doing this, the line and dot markers will automatically change their color for the new vessel. Repeat until all desired vessels are tracked.
NOTE: The x/y location, vessel velocity and moved distance can be retrieved from the pop-up window Results. Each vessel will be identified by its track number in the leftmost column. This data can be copied and pasted into a spread sheet to perform further analysis.
Representative Results
The presented protocol, using stereolithography techniques, allows for the fabrication of tessellated scaffolds made of SU-8 photoresist. Scaffolds with distinct compartment geometries (squares, hexagons, and circles), and highly accurate and repeatable features were obtained (Figure 1).
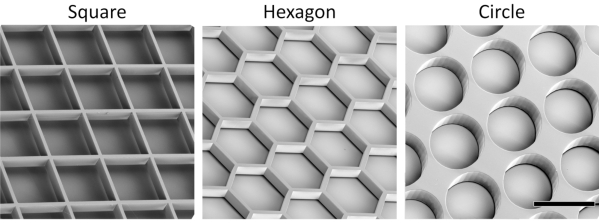

Figure 1: Representative scanning electron microscopy images of the tessellated square, circular, and hexagonal scaffold geometries (scale bar = 500 µm). Please click here to view a larger version of this figure.
With a stepwise cell seeding (steps 2 to 4), the fabricated scaffolds were used to create highly organized vascular networks. When using a traditional simultaneous seeding of both ECs and SCs, the resulting vessels lacked a clear organization. For this, the scaffold fibronectin coating was performed (step 2), the scaffold endothelialization step was skipped (step 3), and the DPSC and HAMEC were simultaneously co-seeded in fibrin gel (step 4). In this fashion, the cells are homogeneously distributed over the scaffold (Figure 2, top row), resulting in unpredictable and disorganized developed vascular networks that do not seem to interact with the surrounding scaffold. Contrarily, firstly seeding the ECs on the scaffold walls provides an accurate initial endothelial cell patterning. The later addition of SCs within a fibrin gel results in a predictable tubulogenesis phenomenon, with forming vessels closely following the shape of the scaffold wall, and sprouting new vessels migrating into the compartment space (Figure 2, bottom row).
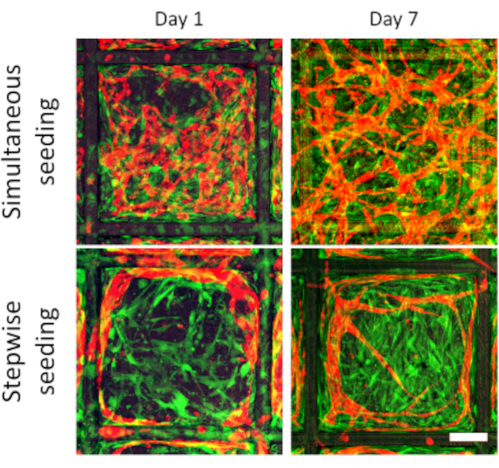
Figure 2: Vascularization comparison between simultaneous vs. stepwise cell seeding. Representative images of vascular development in tessellated scaffolds for a simultaneous (top row) cell seeding of ECs (red) and SCs (green), and step-wise cell seeding (bottom row) at days 1 and 5. The stepwise seeding results in organized vascular networks that follow the scaffold walls and sprout into the compartment space (scale bar: 100 µm). Please click here to view a larger version of this figure.
When using fluorescent ECs, either transfected or dyed, the vessels can be imaged in real time without the need to fix and terminate the experiment for each time point. Red fluorescent protein expressing ECs (RFP-ECs) were cultured on hexagonal scaffolds and imaged after seeding (Figure 3A, day 0). At day 1, the SCs were added and the vascular networks were imaged every other day to quantify the vessel development (Figure 3A, days 1, 3, 5 and 7). For each time point, wide images of the whole scaffold were taken (Figure 3B). For every compartment, the vessels mainly organized and interacted with cells located within their confinement. Hence, each compartment was isolated and the superfluous vessels outside the compartment were removed using ImageJ. The clean, single-compartment images were then analyzed using Angiotool. Angiotool returned a spread sheet file containing several vessel parameters, and a visual representation of the main network characteristics, such as skeleton, intersection points, and vessel surface. The obtained data was analyzed using the statistical analysis software Prism, and a clear vessel growth was observed for total vessel length and area during the experiment time frame (Figure 3C). During a 1-week experiment, vessels are expected to further develop and extended as shown in Figure 3A and Figure 2C. Decreasing vessel length or area, failure to form vessels by day 3 or vessels forming as shown in the top row of Figure 2 can be interpreted as a failed experiments.
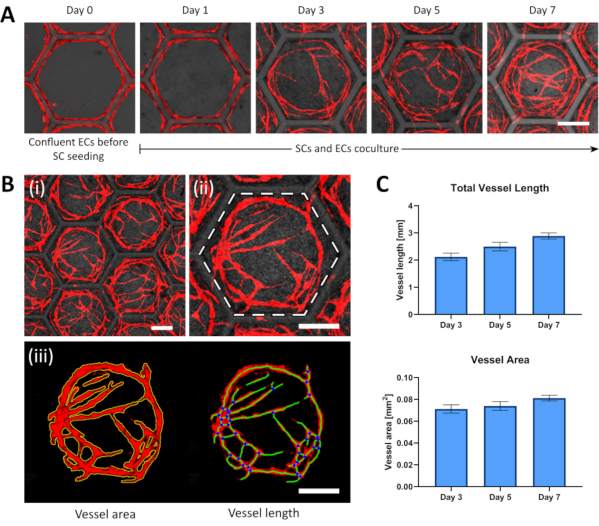
Figure 3: Representative development images and analysis of organized vascular networks. (A) ECs (red) reach confluence on the scaffold wall on which SCs are afterwards seeded; the SCs addition represents day 0 of the experiment. At day 1 after the SCs seeding, the ECs detach to the compartment space and start forming vessels that will continue sprouting and connecting at further days. (B) Confocal image processing steps for vascular network analysis (i) A wide confocal image containing several compartments is taken, (ii) a single compartment is cropped (demarked by the white dashed hexagon), (iii) then the vascular network channel is separated, and all vessels outside the compartment walls are cropped out. The single compartment image is analyzed using Angiotool, returning a list of vascular parameters complemented with visual markers, such as the vessel area (outlined in yellow), the vessels length (displayed with green lines), and the intersection points (marked as blue dots). (C) Comparative results of the total vessel length and the total vessel area within hexagonal compartments at different time points (results are presented as mean ± SD, n > 6; all scale bars: 200 µm). Please click here to view a larger version of this figure.
Using multicolored ECs to facilitate single cell identification, a confocal imaging time lapse was performed to allow single vessel tracking (Supplementary Video 1). The vessels were observed and tracked using the Manual Tracking ImageJ plugin. The tip cell was selected for each frame of the movie (Figure 4A) until the vessel anastomosed with the surrounding vasculature. As a result, the Manual Tracking plugin generated the vessel path in real time, which allowed to observe the vessel migration (Figure 4B).
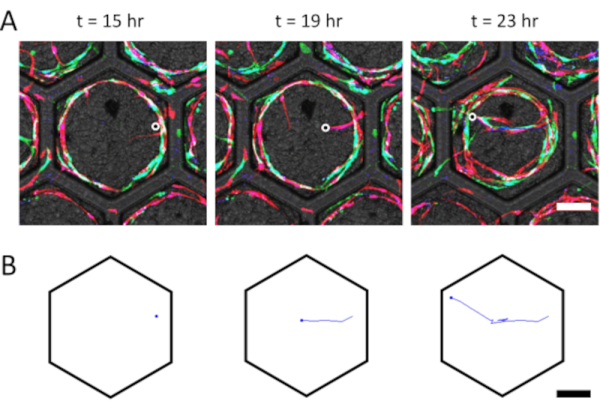
Figure 4: Representative sprouting vessel tracking. (A) A multicolored ECs (green, red, and blue) time lapse is used to facilitate single vessel identification. A sprouting vessel is identified and tracked using the ImageJ plugin Manual Tracking. The end side of the vessel is marked for every time point to track in real time; the black-in-white marker was added to show the selected vessel end point. (B) The resulting 2D tracking of the vessel, as processed by the ImageJ plugin, showing the farthest point with a dot, and the formed path with a line (scale bar = 200 µm). Please click here to view a larger version of this figure.
Vessel maturation, represented by the presence of SMA+ SCs, can be easily observed in the proposed platform. Vasculatures presenting higher numbers of SMA+ SCs represent a more mature network, since SMA expression correlates with vessel stabilization over time23. For the circular, hexagonal and squared compartments, the amount of SMA+ SCs increases over time (Figure 5A). By day 3, all shapes showed scattered SMA+ SCs and uncomplex vessels with few or no sprouts whatsoever. By day 7, all shapes showed a rich and complex vascular network, with a higher presence of SMA+ SCs surrounding the vessels. Furthermore, higher magnification images reveal a denser SMA+ SCs presence co-localized with formed vessels, evidencing the SCs recruitment and differentiation surrounding vascular structures (Figure 5B).
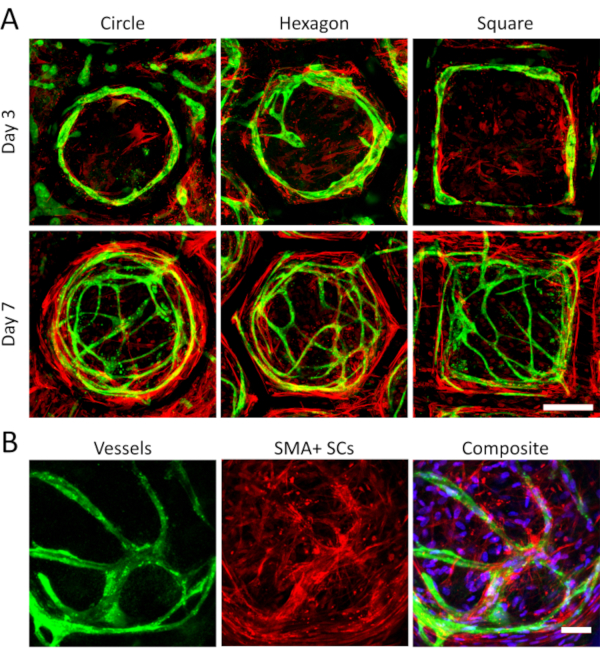
Figure 5: SMA+ SCs and blood vessels increase over time. A) Smooth muscle actin (red) and vWF (vessels, green) are shown for vascular networks in circular, squared and hexagonal compartments at day 3 and day 7. Both vasculature extension and SMA-expressing support cells (SMA+ SCs) increase over time, signifying a higher vessel maturation and complexity (scale bar = 200 µm). B) Representative images of the SMA+ SCs denser accumulation around vessels at day 7. The nuclei (blue) in the composite image reveal the presence of SCs not expressing the SMA protein (scale bar = 50 µm). Please click here to view a larger version of this figure.
Supplementary Video 1: multicolored ECs time lapse for vessel migration tracking. Please click here to download this video.
Discussion
The need for a rich vasculature within embedded in engineered tissues is critical for construct survival and proper function1. Although engineering the vascular system has been the focus of a vast amount of research, much is left to investigate and understand24. In particular, when recreating a specific tissue, the microvasculature should behave and organize accordingly12. The most common approach for microvessels generation is co-seeding endothelial and support cells within a suitable 3D environment compatible for cell attachment, proliferation, and vessel formation25. This methodology often results in greatly unorganized networks, making it difficult to study the microvascular behavior22. Here, the presented protocol provides a new tool that generates highly organized vascular networks in a high-throughput system, with vessels that can be easily tracked and monitored through time, to study their development and behavior. The stepwise seeding of the ECs (step 2-3), followed by the SCs (step 4), are critical steps to achieve such organized networks22. Additionally, this technique presents a real 3D environment for vascular models, in which vessels can migrate in the three dimensions and create relevant structures. In contrast, more popular methods for vascular modelling, such as microfluidics systems, only offer a 2.5D tissue representation26.
The proposed method can be easily modified and applied to study many different factors affecting the vessel network development and behavior. Photolithographic SU-8 resin scaffold fabrication is a common technique with an impressive versatility that enables creating a wide variety of shapes26,27, with the potential to design structures resembling complex native constructs, such as the alveoli or the nephrotic unit. Nonetheless, a drawback of using SU-8 is its low bioabsorbability27, making the platform mainly a research tool, instead of an implantable tissue. However, this could be improved by utilizing biocompatible materials which can crosslink under UV/visible light illumination, and 3D printed using accurate techniques like stereolitography29. The resulting vascularized constructs could then be implanted in an animal model30. Another limitation of the system is the scaffold maximum achievable thickness with high detail accuracy, limited by the stereolithographic technique31. The proposed protocol is suitable for creating thin scaffold which might represent native tissue sizes.
This platform offers the potential to investigate several variables affecting the vascularization phenomena4,32. First, the interactions between different types of ECs and SCs, and their vessel formation, development and maturation capabilities, which are known to differ when using cells from different tissue origins33. Second, the effect of growth factors, small molecules, and inhibitors on the vasculature, allowing for a clear visualization of their impact in real time34,35. Third, the addition of other cell types, spheroids, or organoids and their communication and interaction with the forming vessels36. For this, the proposed system represents a powerful tool for investigating the microvascular system.
Future steps will take advantage of the proposed system to investigate the effect of mechanical stimuli, such as interstitial flow and mechanical stretching37,38, on the developing vasculature. This will hopefully shed light on new aspects that will expand the current knowledge and state-of-the-art research of the vascular mechanobiology.
Disclosures
The authors have nothing to disclose.
Acknowledgements
This research was supported by funding from the University of Michigan – Israel Partnership for Research. The authors would like to thank Uri Merdler, Lior Debbi and Galia Ben David for their great assistance and support, Nadine Wang, Ph.D. and Pilar Herrera-Fierro, Ph.D. of the Lurie Nanofabrication Facility at the University of Michigan, as well as Luis Solorio, Ph.D. for enlightening discussions of photolithography techniques.
Materials
| Angiotool freeware | NIH-CCR | Free download at https://ccrod.cancer.gov/confluence/display/ROB2/Home | |
| Bovine albumin serum Probumin | Millipore | 82-045-1 | |
| Dental pulp stem cells | Lonza | PT-5025 | |
| ECM media + bullet kit | Sciencell | #1001 | |
| Ethanol 96% | Gadot-Group | 64-17-5 | |
| Evicel fibrin sealant | Johnson&Johnson | EVB05IL | Provides both thrombin and fibrinogen (BAC2) solutions |
| GlutaMAX | Gibco | 35050061 | |
| Goat anti-mouse Cy3 antibody | Jackson | 115-166-072 | |
| Goat anti-rabbit Alexa-Fluor 488 | Thermo- Fisher Scientific | A11034 | |
| Human adipose microvascular cells | Sciencell | #7200 | |
| Human fibronectin | Sigma | F0895-5MG | Stock concentration: 1 mg/mL |
| ImageJ | NIH | Free download at https://imagej.nih.gov/ij/download.html | |
| Isopropyl alcohol | Gadot-Group | 67-63-0 | |
| Lift-off reagent | Kayaku Advanced Materials, Inc | G112850 | Commercial name Omnicoat |
| Low-glucose DMEM | Biological Industries | 01-050-1A | |
| Mouse anti-SMA antibody | Dako | M0851 | |
| NEAA | Gibco | 11140068 | |
| Paraformaldehyde solution 4% in PBS | ChemCruz | SC-281692 | |
| Penicillin-Streptomycin-Nystatin Solution | Biological Industries | 03-032-1B | |
| Phospate buffered saline (PBS) | Sigma | P5368-10PAK | |
| Rabbit anti-vWF antibody | Abcam | ab9378 | |
| Silicon wafer | Silicon Valley Microelectronics (SVM) | Wafers 4", Type N-1-10, 500-550 microns thick | |
| SU-8 2050 photoresist | Kayaku Advanced Materials, Inc | Y11058 | |
| SU-8 developer | Kayaku Advanced Materials, Inc | Y020100 | |
| Tryton-X 100 | BioLab LTD | 57836 |
References
- Novosel, E. C., Kleinhans, C., Kluger, P. J. Vascularization is the key challenge in tissue engineering. Advanced Drug Delivery Reviews. 63 (4), 300-311 (2011).
- Landau, S., Guo, S., Levenberg, S. Localization of Engineered Vasculature within 3D Tissue Constructs. Frontiers in Bioengineering and Biotechnology. 6, 2 (2018).
- Griffith, C. K., et al. Diffusion Limits of an in Vitro Thick Prevascularized Tissue. Tissue Engineering. 11 (12), (2005).
- Potente, M., Gerhardt, H., Carmeliet, P. Basic and therapeutic aspects of angiogenesis. Cell. 146 (6), 873-887 (2011).
- Landau, S., et al. Tropoelastin coated PLLA-PLGA scaffolds promote vascular network formation. Biomaterials. 122, 72-82 (2017).
- Lesman, A., et al. Engineering vessel-like networks within multicellular fibrin-based constructs. Biomaterials. 32 (31), 7856-7869 (2011).
- Richards, D., Jia, J., Yost, M., Markwald, R., Mei, Y. 3D Bioprinting for Vascularized Tissue Fabrication. Annals of Biomedical Engineering. 45 (1), 132-147 (2017).
- Levenberg, S., et al. Engineering vascularized skeletal muscle tissue. Nature Biotechnology. 23 (7), 879-884 (2005).
- Rouwkema, J., Khademhosseini, A. Vascularization and Angiogenesis in Tissue Engineering: Beyond Creating Static Networks. Trends in Biotechnology. 34 (9), 733-745 (2016).
- Miller, J. S., et al. Rapid casting of patterned vascular networks for perfusable engineered three-dimensional tissues. Nature Materials. 11, (2012).
- Gariboldi, M. I., Butler, R., Best, S. M., Cameron, R. E. Engineering vasculature Architectural effects on microcapillary-like structure self-assembly. PLOS ONE. 14 (1), 1-13 (2019).
- Blache, U., Guerrero, J., Güven, S., Klar, A. S., Scherberich, A. Microvascular Networks and Models, In vitro Formation. Vascularization for Tissue Engineering and Regenerative Medicine. , 1-40 (2018).
- Wong, K. H. K., Chan, J. M., Kamm, R. D., Tien, J. Microfluidic Models of Vascular Functions. Annual Review of Biomedical Engineering. 14 (1), 205-230 (2012).
- Jansen, K. A., Bacabac, R. G., Piechocka, I. K., Koenderink, G. H. Cells actively stiffen fibrin networks by generating contractile stress. Biophysical Journal. 105 (10), 2240-2251 (2013).
- Pollet, A. M. A. O., den Toonder, J. M. J. Recapitulating the vasculature using Organ-on-Chip technology. 생체공학. 7 (1), (2020).
- Hasan, A., et al. Microfluidic techniques for development of 3D vascularized tissue. Biomaterials. 35 (26), 7308-7325 (2014).
- Jordahl, J. H., et al. 3D Jet Writing: Functional Microtissues Based on Tessellated Scaffold Architectures. Advanced Materials. 30 (14), 1707196 (2018).
- Gauvin, R., et al. Microfabrication of complex porous tissue engineering scaffolds using 3D projection stereolithography. Biomaterials. 33 (15), 3824-3834 (2012).
- Coscoy, S., et al. Microtopographies control the development of basal protrusions in epithelial sheets. Biointerphases. 13 (4), 041003 (2018).
- Kaplan, B., et al. Rapid prototyping fabrication of soft and oriented polyester scaffolds for axonal guidance. Biomaterials. , (2020).
- Steier, A., Muñiz, A., Neale, D., Lahann, J. Emerging Trends in Information-Driven Engineering of Complex Biological Systems. Advanced Materials. 31 (26), 11806898 (2019).
- Szklanny, A. A., et al. High-Throughput Scaffold System for Studying the Effect of Local Geometry and Topology on the Development and Orientation of Sprouting Blood Vessels. Advanced Functional Materials. , 1901335 (2019).
- Welti, J., Loges, S., Dimmeler, S., Carmeliet, P. Recent molecular discoveries in angiogenesis and antiangiogenic therapies in cancer. Journal of Clinical Investigation. 123 (8), 3190-3200 (2013).
- Gui, L., Niklason, L. E. Vascular Tissue Engineering: Building Perfusable Vasculature for Implantation. Current Opinion in Chemical Engineering. 3, 68-74 (2014).
- Blache, U., Ehrbar, M. Inspired by nature: Hydrogels as versatile tools for vascular engineering. Advances in Wound Care. 7 (7), 232-246 (2018).
- Cochrane, A., et al. Advanced in vitro models of vascular biology: Human induced pluripotent stem cells and organ-on-chip technology. Advanced Drug Delivery Reviews. 140, 68-77 (2019).
- Nemani, K. V., Moodie, K. L., Brennick, J. B., Su, A., Gimi, B. In vitro and in vivo evaluation of SU-8 biocompatibility. Materials Science & Engineering. C, Materials for Biological Applications. 33 (7), 4453-4459 (2013).
- Mathew, R., Ravi Sankar, A. A Review on Surface Stress-Based Miniaturized Piezoresistive SU-8 Polymeric Cantilever Sensors. Nano-Micro Letters. 10 (2), 1-41 (2018).
- Knowlton, S., Yenilmez, B., Anand, S., Tasoglu, S. Photocrosslinking-based bioprinting: Examining crosslinking schemes. Bioprinting. 5, 10-18 (2017).
- Redd, M. A., et al. Patterned human microvascular grafts enable rapid vascularization and increase perfusion in infarcted rat hearts. Nature Communications. 10 (1), 1-14 (2019).
- Zhu, Y., et al. SU-8 Photoresist. Encyclopedia of Nanotechnology. , 2530-2543 (2012).
- Zheng, F., et al. Organ-on-a-Chip Systems: Microengineering to Biomimic Living Systems. Small. 12 (17), 2253-2282 (2016).
- Freiman, A., et al. Adipose-derived endothelial and mesenchymal stem cells enhance vascular network formation on three-dimensional constructs in vitro. Stem Cell Research & Therapy. 7 (1), 5 (2016).
- van Duinen, V., et al. Perfused 3D angiogenic sprouting in a high-throughput in vitro platform. Angiogenesis. 22 (1), 157-165 (2019).
- Nguyen, D. -. H. T., et al. Biomimetic model to reconstitute angiogenic sprouting morphogenesis in vitro. Proceedings of the National Academy of Sciences. 110 (17), 6712-6717 (2013).
- Nashimoto, Y., et al. Integrating perfusable vascular networks with a three-dimensional tissue in a microfluidic device. Integrative Biology. 9 (6), 506-518 (2017).
- Rosenfeld, D., et al. Morphogenesis of 3D vascular networks is regulated by tensile forces. Proceedings of the National Academy of Sciences of the United States of America. 113 (12), 3215-3220 (2016).
- Neto, F., et al. and TAZ regulate adherens junction dynamics and endothelial cell distribution during vascular development. bioRxiv. , 174185 (2017).

