Bioorthogonal Chemical Imaging of Cell Metabolism Regulated by Aromatic Amino Acids
Summary
We present a protocol to directly visualize metabolic activities in cells regulated by amino acids using deuterium-oxide (heavy water D2O) probed stimulated Raman scattering (DO-SRS) microscopy, which is integrated with two-photon excitation fluorescence microscopy (2PEF).
Abstract
Essential aromatic amino acids (AAAs) are building blocks for synthesizing new biomasses in cells and sustaining normal biological functions. For example, an abundant supply of AAAs is important for cancer cells to maintain their rapid growth and division. With this, there is a rising demand for a highly specific, noninvasive imaging approach with minimal sample preparation to directly visualize how cells harness AAAs for their metabolism in situ. Here, we develop an optical imaging platform that combines deuterium oxide (D2O) probing with stimulated Raman scattering (DO-SRS) and integrates DO-SRS with two-photon excitation fluorescence (2PEF) into a single microscope to directly visualize the metabolic activities of HeLa cells under AAA regulation. Collectively, the DO-SRS platform provides high spatial resolution and specificity of newly synthesized proteins and lipids in single HeLa cell units. In addition, the 2PEF modality can detect autofluorescence signals of nicotinamide adenine dinucleotide (NADH) and Flavin in a label-free manner. The imaging system described here is compatible with both in vitro and in vivo models, which is flexible for various experiments. The general workflow of this protocol includes cell culture, culture media preparation, cell synchronization, cell fixation, and sample imaging with DO-SRS and 2PEF modalities.
Introduction
Being essential aromatic amino acids (AAAs), phenylalanine (Phe) and tryptophan (Tryp) can be absorbed by the human body to synthesize new molecules for sustaining normal biological functions1. Phe is needed for the synthesis of proteins, melanin, and tyrosine, while Tryp is required for the synthesis of melatonin, serotonin, and niacin2,3. However, excess consumption of these AAAs can upregulate the mammalian target of the rapamycin (mTOR) pathway, inhibit AMP-activated protein kinase, and interfere with the mitochondrial metabolism, collectively altering macromolecule biosynthesis and leading to the production of malignant precursors, such as reactive oxygen species (ROS) in healthy cells4,5,6. Direct visualization of altered metabolic dynamics under excess AAA regulation is essential to understand AAAs' roles in promoting cancer development and the growth of healthy cells7,8,9.
Traditional AAA studies rely on gas chromatography (GC)10. Other methods, such as magnetic resonance imaging (MRI), have limited spatial resolutions, making it hard to perform cellular and sub-cellular analysis of biological samples11. Recently, matrix-assisted laser desorption/ionization (MALDI) has been developed to elucidate the role of AAAs in lipid and protein syntheses in cancer proliferation with noninvasive biomarkers12,13,14. However, this technique still suffers from shallow imaging depths, poor spatial resolution, and extensive sample preparation. At the cellular level, nontoxic stable isotopes, such as nitrogen-15 and carbon-13, can be traced with multi-isotope imaging and nanoscale secondary ion mass spectrometry to understand their incorporation into macromolecules. However, these methods are destructive to living biological samples15,16. Atomic force microscopy (AFM) is another powerful technique that can visualize metabolic dynamics17. The slow rate of scanning during AFM imaging, on the other hand, may cause image distortion of the result from thermal drift.
We developed a noninvasive biorthogonal imaging modality by coupling deuterium-oxide (D2O) probed stimulated Raman scattering (DO-SRS) microscopy and label-free two-photon excitation fluorescence microscopy (2PEF). This modality achieves a high spatial resolution and chemical specificity when imaging biological samples18,19,20,21,22,23,24. This protocol introduces the applications of DO-SRS and 2PEF to examine the metabolic dynamics of lipids, protein, and redox ratio changes during cancer progressions. With D2O being a stable isotope form of water, cellular biomolecules can be labeled with deuterium (D) due to its quick compensation with total body water in cells, forming carbon-deuterium (C-D) bonds through enzymatic exchange21. The C-D bonds in newly synthesized macromolecules, including lipids, proteins, DNA/RNA, and carbohydrates, can be detected in the cell silent region of the Raman spectrum20,21,22,25,26,27. With two synchronized laser pulses, C-D bonds of newly synthesized lipids and proteins can be displayed on single cells via hyperspectral imaging (HSI) without extracting or labeling them with cytotoxic agents. In addition, SRS microscopy has the capability to construct three-dimensional (3D) models of selected regions of interest in biological samples by capturing and combining a set of cross-sectional images22,26. With hyperspectral and 3D volumetric imaging, DO-SRS can obtain spatial distributions of newly synthesized macromolecules in single cells, along with the type of organelles that facilitate the process of promoting cancer growth under AAA regulation22. Furthermore, using 2PEF, we can obtain autofluorescence signals of Flavin and nicotinamide adenine dinucleotide (NADH) with high resolution, deep penetration depth, and low-level damage in biological samples21,23,24. Flavin and NADH autofluorescence signals have been used to characterize redox homeostasis and lipid peroxidation in cancer cells22,26. As such, not only does the coupling of DO-SRS and 2PEF provide subcellular analysis of AAA-regulated metabolic dynamics in cancer cells with high spatial distribution, chemical specificity information, and minimal sample preparation, but the method also reduces the need to extract or label endogenous molecules with toxic reagents. In this protocol, we first present the procedures of D2O and amino acid preparation, as well as cancer cell culture. Then, we show the protocols of DO-SRS imaging and 2PEF imaging. Finally, we present the representative results of SRS and 2PEF imaging, which demonstrate AAA-regulated metabolic changes of lipids and protein, and redox ratio changes in cancer cells. A detailed illustration of the process is highlighted in Figure 1.
Protocol
1. Media preparation
- Prepare 10 mL of control and excess AAAs in Dulbecco's modified Eagle medium (DMEM) containing 50% D2O.
- For the control media, measure and mix 10 mg of DMEM powder with 4.7 mL of double distilled water (ddH2O) in a 15 mL conical tube. The DMEM powder contains all the amino acids at standard concentrations. Thoroughly vortex and invert the tube to ensure the solution is well mixed. Add 4.7 mL of D2O, 0.5 mL of fetal bovine serum (5% FBS), and 0.1 mL of penicillin/streptomycin (1%). Thoroughly vortex and invert the tube to ensure the solution is well mixed.
- For the 15x Phe treatment medium, measure and mix 10 mg of DMEM powder with 4.7 mL of ddH2O, 0.5 mL of FBS (5%), and 0.1 mL of penicillin/streptomycin (1%) in a 15 mL conical tube. Thoroughly vortex and invert the tube to ensure the solution is well mixed. Add excess Phe powder at a concentration of 924 mg/L to the medium. Thoroughly vortex and invert the tube to ensure the solution is well mixed.
- For the 15x Tryp treatment medium, measure and mix 10 mg of DMEM powder with 4.7 mL of ddH2O, 0.5 mL of FBS (5%), and 0.1 mL of penicillin/streptomycin (1%) in a 15 mL conical tube. Thoroughly vortex and invert the tube to ensure the solution is well mixed. Add excess Tryp powder at a concentration of 224 mg/L to the medium. Thoroughly vortex and invert the tube to ensure the solution is well mixed.
- Add methionine and threonine at 30.0 mg/mL and 95.0 mg/mL, respectively, to all conical tubes of the previous steps. Thoroughly vortex and invert the tube. Sodium pyruvate is not required to be added to the culture media.
- Filter the control and treatment media with a 25 mm syringe filter with 0.22 µm polyethersulfone membrane filters.
- Seal all the media-contained conical tubes with parafilm and store at 4 °C for up to 3 weeks.Prior to treating the cells with media, warm all the media to 37°C.
NOTE: The powder and liquid reagents should be measured and combined based on the total volume of treatment and control media.
2. Cell sample preparation
- Maintain cervical cancer (HeLa) cells in a culture flask (i.e., T10, T25, etc.) using standard DMEM supplemented with 10% FBS and 1% penicillin/streptomycin at 37 °C and 5% carbon dioxide (CO2).
- Sub-culture the HeLa cells at a split ratio of 1:10 until reaching 80% or higher cell viability.
- Once the cells achieve 80% or higher confluency, proceed to seed 2 x 105 cells per well onto a 24-well plate using DMEM supplemented with 0.5% FBS and 1% penicillin/streptomycin.
- Prior to cell seeding, prepare a 24-well plate with sterilized, poly-d-lysin, laminin-coated, round coverslips, 12 mm in diameter. Submerge the coverslips in 70% ethanol and dry gently using lint-free wipes. Place the clean coverslips in the appropriate wells of the plate.
- Once the cells reach 80% confluency in step 2.2, wash the cells once with 1x phosphate buffered saline (PBS) without magnesium and calcium ions.
- Add 0.25% trypsin to dissociate the adherent cells from the flask and incubate at 37 °C and 5% CO2 for 3 min.
- Add DMEM supplemented with 10% FBS and 1% penicillin/streptomycin, gently pipette up and down, and collect the cells by centrifugation at 300 x g at room temperature (RT) for 3 min.
- Resuspend the cells in DMEM supplemented with 0.5% FBS and 1% penicillin/streptomycin.
- Count the cells using a hemocytometer, light microscope, and trypan blue, and seed a density of 2 x 105 cells per well of a 24-well plate. Alternatively, an automated cell counter can be used to count the cells.
- Return the cells to the incubator at 37 °C and 5% CO2, and gently shake the plate to distribute the cells evenly.
- After 8 h, replace the old culture media with 50% D2O/excess Phe or 50% D2O/excess Tryp media. Return the cells to the incubator at 37 °C and 5% CO2 for 36 h.
- After 36 h, fix the cells on microscope slides.
- Prior to fixing, prepare microscope slides with imaging spacers 9 mm in diameter and place 15 µL of 1x PBS supplemented with calcium and magnesium ions.
- Rinse the cells with 1x PBS supplemented with calcium and magnesium ions.
- Add 0.5 mL of 4% methanol-free paraformaldehyde (PFA) solution and leave the plate under the biosafety hood for approximately 15 min.
- Aspirate the PFA solution and rinse with 1x PBS supplemented with calcium and magnesium ions twice.
- Add 1.5 mL of 1x PBS supplemented with calcium and magnesium ions in each well.
- Use sterile tweezers to gently grab the coverslips out of each well. Invert and place the cell-containing side of the coverslips onto the imaging spacers to contact the PBS prepared in step 2.5.1. Ensure that the non cell-containing side is exposed to air.
- Using transparent nail polish, seal the outer layer of the coverslip with the imaging spacer.
- Store the microscope slides at 4 °C when not imaging.
3. Spontaneous Raman spectroscopy measurement (Figure 2)
- Use a confocal Raman microscope to obtain spontaneous Raman spectra (Figure 2A).
- Turn on the laser by turning the key to the ON position.
- Double-click on the LabSpec6 software icon to open the acquisition software.
- Load the biological sample onto the sample holder directly underneath the objective lens.
- On the software, click the Camera icon to turn on the video camera.
- Adjust the focus plane to identify a region of interest with the 50x objective lens. Move the joystick to control the planer translation of the XY stage and rotate the joystick to change the depth of focus.
- After choosing the position for measurement, switch to the 100x objective lens with 40 mW of excitation power. Click the red Stop icon to turn off the video camera.
- Select a grating of 1800 lines/mm, and set the acquisition range from 400 cm-1 to 3,150 cm-1.
- Set the Acquisition Time to 90 s, the Accumulation to 3, and the Binning to 4 for the least noise and most accurate spectrum.
NOTE: These imaging parameters can be optimized to fit the experiment. - Click the Arrow icon to turn on the real-time display.
- Tune the Raman shift (cm-1) to a value of choice to fine-tune the focus plane.
NOTE: In this experiment, the Raman laser was focused onto the lipid droplets, which contained a high concentration of CH2 bonds. Therefore, a Raman shift of 2,850 cm-1 that matched the stretching modes of the CH2 bond was selected for the purpose. - Adjust the focus plane with the joystick to optimize the best signal-to-noise ratio.
- After choosing the focus plane, click the red Stop icon for a real-time display.
- Select the Circle icon to start the spectrum acquisition.
- Once the cellular region is taken, use the same focus plane to measure the background with PBS. Subtract the background spectrum from each subcellular target spectrum using a separate computer software application, such as Origin (Figure 1B).
4. Imaging experiments with 2PEF and SRS
NOTE: Detailed descriptions of SRS laser alignment can be found in a prior report28. This protocol focuses on the operation of a multimodal SRS and 2PEF imaging system (Figure 2C, D).
- Click on Start to warm up the laser.
- Follow the order to set the main switches of the control unit and the monitor on: 1) press the main switch of the control box IX3-CBH to On, 2) press the main switch of the touch panel controller to On, 3) press the main switch of the AC adapter of the power supply for LD OBIS 6 Laser Remote connected to the main laser combiner FV31-SCOMB to On, and 4) press the main switch of the AC adapter of the power supply for LD OBIS 6 Laser Remote connected to the sub laser combiner FV31-SCOMB to On.
- Press the main switches of the Si photodiode detector and lock-in amplifier On.
- Set the Stokes Beam at 1,031 nm, Pulse Width at 6 ps, and Repetition Rate at 80 MHz.
- Coupled into the microscope, set the picoEmerald system supplied with the synchronized pump beam with a Tunable Wavelength from 720-990 nm, a Pulse Width of 5-6 ps, and a Repetition Rate of 80 MHz. The average excitation power of both the pump and Stokes is 450 mW. Optimize the excitation power to minimize photodamage for the sample.
- Use a high-numerical aperture (NA) oil condenser (i.e., 1.4 NA) to collect the Stokes and pump beams where the sample is mounted by emitting a few drops on the condenser.
- Mount the sample on the oil and place a large water droplet on top of the microscope slide where the sample is fixed. This is for the water-immersion objective lens.
- Set the lock-in amplifier to 20 MHz. Process the image now using the super-resolution software module.
- Select 512 pixels x 512 pixels and 80 µs/pixel for the dwell time.
- Once the desired cellular region is focused properly, acquire the image. Using the acquisition software of the microscope, save the images as an Olympus .oir graphic file.
- Acquire a background image at 1,900 cm-1 and subtract it from all cellular images.
- To perform 2PEF imaging, use the same tunable picosecond laser, and set the label-free autofluorescence of Flavin and NADH to 800 nm and 780 nm, respectively.
- Use a 460 nm/515 nm filter cube to collect the Flavin and NADH autofluorescence backscattered emission.
- Adjust the dwell time to 8 µs/pixel and the pixel size to 512 pixels x 512 pixels. Set the laser shutter power parameter to 150 mW.
- For 3D image reconstruction, tune the stimulated Raman loss to 2,850 cm-1 (797 nm).
- Once the desired single cells have been identified, register the laser to scan from the top focus plane to detect the top and bottom layers of those cells.
NOTE: 3D models are generated and saved as .oir files.
5. Spectra and image analysis
- Spectra analysis
- Use the Origin software or other relevant applications to subtract the background spectrum from all subcellular target spectra.
- Use MATLAB's mathematical modeling operations to process the Raman spectra with the software's built-in functions. Python can also be used for spectra processing.
- Spectral pre-processing begins with converting the files into an array. The spectra are interpolated at every reciprocal centimeter.
- For validation, graph the raw data. Perform the baseline correction on the raw spectra using the built-in function msbackadj in MATLAB. In brief, the built-in function estimates baseline points at separation-unit values identified by users and returns the distance between adjacent windows. From the baseline points, estimate and draw a regression line. Apply Smoothing to smooth the regression line to subtract the baseline from each individual raw spectrum.
- Perform min-max normalization by dividing the Raman intensity of the protein at 2,930 cm-1 by the intensity of each Raman shift. The 2,930 cm-1 signal is typically the highest intensity on the Raman spectrum. With this normalization, the maximum value gets transformed into a 1 while the minimum value gets transformed into a 0. Every other value gets transformed into a decimal between 0 and 1.
- Average the individual spectra of the same condition into a single spectrum to reduce noise.
- Once processed, use applications, such as Origin, to display all the spectra simultaneously. The peaks displayed in the spectra represent a specific chemical bond or functional group.
- 3D image analysis of lipid droplets
- Use FIJI-ImageJ software to process all 3D images
- Perform the functions Bandpass Filters and Smoothing for the 3D image stacks.
- For 3D image analysis, assign each lipid droplet to a spherical score calculated by measuring the distance between its center of mass to the surface. Compare each spherical score with a spherical score of a perfect sphere on the same Euclidean plan. Discard lipid droplets with low spherical scores and evaluate the remaining lipid droplets for their volume and counts.
- Analyze the volume and counts of lipid droplets between different treatments for statistical significance on an appropriate statistical software application. Here, GraphPad Prism was used.
- 2D hyperspectral image analysis
- Use FIJI-ImageJ software to process all the 2D hyperspectral images.
- Select the 2,930 cm-1 channel to make a mask image that contains 0 and 1 values. Assign the background to 0 and the cellular region to 1.
- Subtract the deuterium-labeled protein channel (2,175 cm-1) to the background channel (1,900 cm-1) to remove the fluorescent effects.
- Divide the resulting deuterium-labeled protein channel by the protein channel (2,930 cm-1) to obtain protein turnover rate ratiometric images.
- Then, multiply the mask image made in step 5.3.2 by the ratiometric images to remove any remaining background noises. As a result, a dark background is generated in each ratiometric image.
- Use the Freehand Selection tool in FIJI-ImageJ to manually segment and calculate pixel intensities displayed on single cells. The pixel intensities correspond to the concentrations of the chemical bond that is being visualized.
- Analyze the pixel intensities for statistical significance on an appropriate statistical software application. Here, GraphPad Prism was used. Repeat this analysis with the remaining ratiometric analysis.
Representative Results
The addition of excess AAAs at 15x concentrations to the 50% D2O-containing cell culture media produced distinct C-D Raman bands of newly synthesized lipids and proteins in HeLa cells (Figure 2B). Previous experiments were performed with different concentration levels, such as 2x and 5x, and although the data is not presented, the 15x concentration produced the most distinct C-D Raman bands of newly synthesized lipids and proteins. Specifically, by investigating lipid droplets (LDs), we noticed that both 15x Phe and 15x Tryp induced newly synthesized lipids and protein signals at 2,143 cm-1 and 2,172 cm-1, respectively. DO-SRS was subsequently used to visualize the spatial distribution of C-D signals on single cells (Figure 3). Using ImageJ, pixel intensities of individual cells were manually segmented and calculated, as indicated by dotted-white borders in Figure 3A. The control cells display moderate C-D lipid and protein bands; however, the 15x Phe and 15x Tryp display stronger C-D lipid and protein bands. Quantitative analysis indicates that excess AAAs may upregulate lipid synthesis by 10%-17% but downregulate protein synthesis by 10% (Figure 3C,D). The results infer the possibility of a lack of autophagy that accumulates newly synthesized lipids, promotes mitochondrial dysfunction, and induces oxidative imbalance under excess AAA regulation7.
Label-free multimodal SRS and 2PEF imaging of unsaturated lipid (~3,011 cm-1), saturated lipid (~2,880 cm-1), and NADH, Flavin were acquired to understand the effects of AAAs on cancer metabolism. Similarly, pixel intensities of individual cells were manually segmented and calculated using ImageJ, as indicated by dotted-white borders in Figure 3B. Ratiometric analysis of AAA-treated cells displays a 10% increase in unsaturated lipid/saturated lipid and a 50% increase in Flavin/Flavin + NADH (Figure 3E,F). In the electron transport chain of mitochondria, ROS can be generated from the transfer of electrons from NADH and FADH2 to molecular oxygen species. The accumulation of ROS in many cancer cells results in an oxidative imbalance that oxidizes unsaturated fatty acids, promotes saturated fatty acid synthesis, and depletes NADH autofluorescence signals. Therefore, the observed increase in Flavin/Flavin + NADH is an indicator of accumulated ROS that reduces NADH autofluorescence signals. In response to oxidative imbalance, HeLa cells may upregulate their unsaturated lipid synthesis to replace the oxidized ones. This response is not observed in other cancer cell lines29, which signifies the metabolic heterogeneity of cancer cells under an excess AAA diet30.
In addition to multimodal imaging, SRS can reconstruct label-free 3D images of LDs in control and AAA-treated HeLa cells. In brief, microscopy generates a set of cross-sectional images throughout a selected region of interest. In this study, the stimulated Raman loss (SRL) was tuned to 2,845 cm-1 and scanned from the top layer to the bottom layer with a step size of 1 µm (Figure 4A). Quantitative analyses of 3D lipid droplets reveal that LDs reduced in size but increased in counts in AAA-treated cells compared to the control (Figure 4B,C). The increased presence of bulky hydrophobic amino acids, such as Tryp and Phe may impair the function of LD-coating proteins. This ultimately reduces lipolysis, which accumulates numerous small LDs. The label-free 3D SRS volumetric imaging results of this study corroborate with previous studies by visualizing that excess AAA-treated cells exhibit numerous smaller LDs31,32.
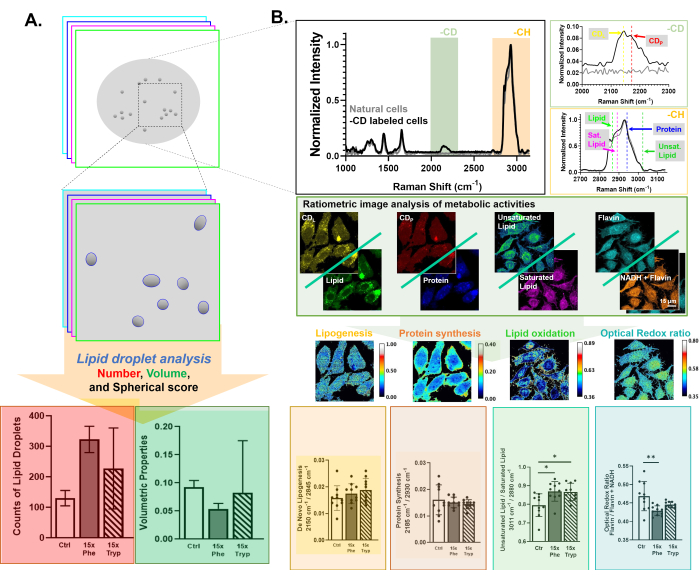
Figure 1: An illustration of the image acquisition and analysis with DO-SRS and 2PEF. (A) A 3D image reconstruction and analysis to acquire lipid droplet number, volume, and spherical score. (B) Hyperspectral imaging and analysis of deuterium-labeled lipid (CDL; 2,145 cm-1), lipid (2,845 cm-1), deuterium-labeled protein (CDP; 2,175 cm-1), protein (2,940 cm-1), unsaturated lipid (3,011cm-1), saturated lipid (2,880 cm-1), NADH, and Flavin. Please click here to view a larger version of this figure.
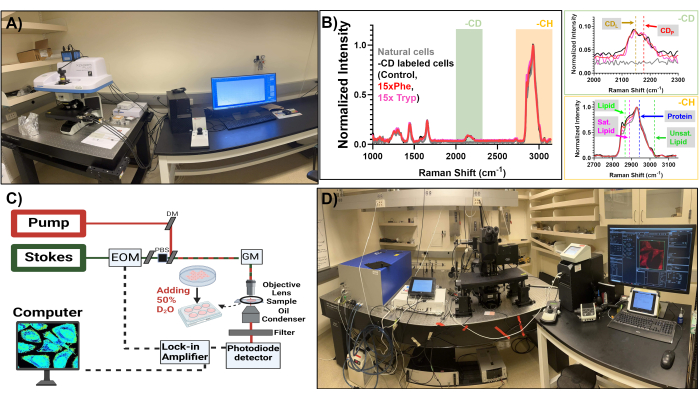
Figure 2: Physical setup of spontaneous Raman spectroscopy and stimulated Raman scattering microscopy. (A) Spontaneous Raman spectroscopy setup used for this study. (B) Sample spontaneous Raman spectra for CD label (red), without CD label (black), control group (blue, solid line), 15x Phe-treated group (red, dotted line), and 15x Tryp-treated group (pink, dotted line). (C) Schematic diagram of the stimulated Raman scattering microscopy setup used for this investigation. (D) Stimulated Raman scattering microscopy setup used as the DO-SRS and 2PEF platform. Please click here to view a larger version of this figure.
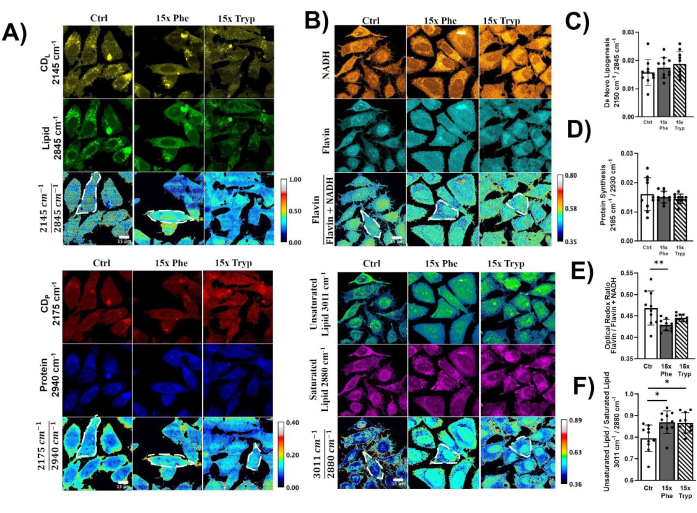
Figure 3: Visualizing metabolic dynamics in HeLa cells using DO-SRS and 2PEF microscopy. (A) Deuterium-labeled lipid (2,145 cm-1), lipid (2,845 cm-1), deuterium-labeled protein (2,175 cm-1), protein (2,940 cm-1) visualized in HeLa cells under control (Ctrl), 15x phenylalanine (15x Phe), and 15x tryptophan (15x Tryp) with the DO-SRS platform. Lipid turnover rate and protein turnover rate were calculated as 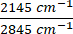 and
and 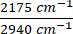 . Raw images were first subtracted by the PBS signal to remove background intensity and masked to remove intensity outside of the cells using ImageJ. Pixel-wise division was performed for the ratiometric analysis. (B) NADH and Flavin channels visualized with 2PEF microscopy, and unsaturated lipid (3,011 cm-1) and saturated lipid (2,880 cm-1) visualized with label-free SRS microscopy. Optical redox ratio and saturation ratio were calculated with
. Raw images were first subtracted by the PBS signal to remove background intensity and masked to remove intensity outside of the cells using ImageJ. Pixel-wise division was performed for the ratiometric analysis. (B) NADH and Flavin channels visualized with 2PEF microscopy, and unsaturated lipid (3,011 cm-1) and saturated lipid (2,880 cm-1) visualized with label-free SRS microscopy. Optical redox ratio and saturation ratio were calculated with 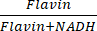 and
and 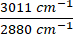 . (C–F) Quantification of ratiometric intensities for each HeLa cell under control, 15x Phe, and 15x Tryp conditions. The statistical difference was used to compare excess AAA conditions with the control conditions. ****p < 0.0001, ***p < 0.001, **p < 0.01, *p < 0.05 were calculated from a two-way ANOVA test. Please click here to view a larger version of this figure.
. (C–F) Quantification of ratiometric intensities for each HeLa cell under control, 15x Phe, and 15x Tryp conditions. The statistical difference was used to compare excess AAA conditions with the control conditions. ****p < 0.0001, ***p < 0.001, **p < 0.01, *p < 0.05 were calculated from a two-way ANOVA test. Please click here to view a larger version of this figure.
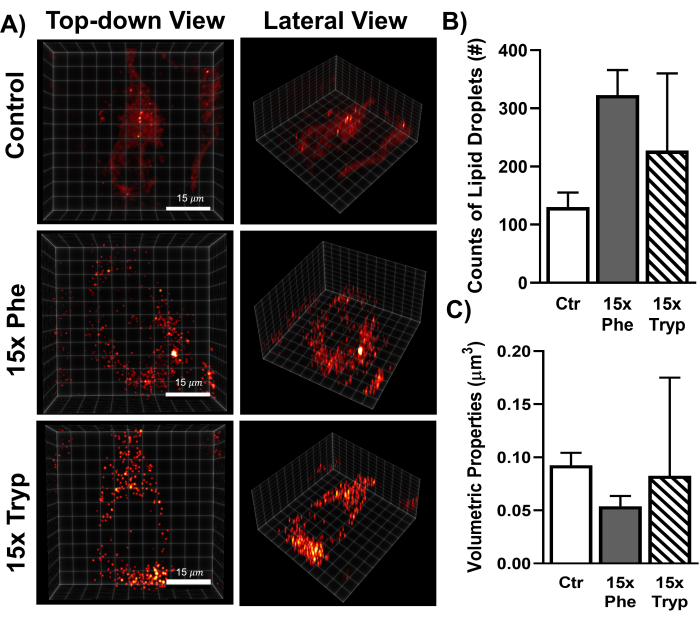
Figure 4: Visualization of 3D lipid droplet distribution of a single HeLa cell using a label-free SRS microscope. (A) SRS 3D lipid droplet volume projection in HeLa cells under control and excess AAA conditions. The threshold was defined prior to the analysis, revealing lipid droplet signal distribution. (B,C) Quantification of lipid droplet volume and counts within individual HeLa cells under control group and excess AAA conditions using the two-way ANOVA test. Please click here to view a larger version of this figure.
Discussion
DO-SRS and 2PEF imaging have been applied to investigate metabolic dynamics in various ex vivo models, including Drosophila and human tissues21,22,23,24,26,27,33. The imaging modality used in this study integrates DO-SRS and 2PEF microscopy, which can outpace other molecule-specific imaging methods by eliminating the need for molecule extraction or labeling with cytotoxic reagents and requiring minimal sample preparation. Specifically, DO-SRS microscopy enables us to probe de novo lipogenesis and protein synthesis in animals and cells and visualize their metabolic dynamics in situ23,27,34. 2PEF captures the autofluorescence signals of naturally occurring biomolecules, such as Flavin and NADH, in biological samples35,36. The imaging penetration depth of SRS can achieve up to 200-500 µm in less scattering samples37. With tissue-clearing methods such as urea, the penetration depth can increase more than tenfold37. The coupling of SRS with 2PEF enables us to image endogenous fluorophores such as Flavin and NADH in biological samples, which further eliminates the need for exogenous biomarkers for detection. 2PEF can achieve a penetration depth of 500 µm38. Compared to other multiplex imaging modalities, such as multi-color single-photon confocal fluorescence microscopy, both DO-SRS and 2PEF can achieve a significantly larger penetration depth without cytotoxic exogenous biomarkers.
To increase the spatial resolution of DO-SRS and 2PEF, we developed an adaptive moment estimation (Adam)-based pointillism deconvolution (A-PoD) image post-processing tool34. A-PoD can convert diffraction-limited DO-SRS and 2PEF images to super-resolved ones without the need for any hardware improvements. Additionally, the Raman spectrum contains many specific chemical vibration modes that can be harnessed to increase the number of detectable molecules. Taking this advantage, our lab has further generated a penalized reference matching (PRF) method to distinguish multiple lipid species in cell and tissue models36. These two technical developments open new avenues for studying biological processes in more detail at cellular and sub-cellular levels, which have been used to characterize metabolic changes in brain, cancer, and aging processes26,27,35.
One main limitation of this protocol is the concentration and time incubation of cellular samples with D2O to produce distinct, quantifiable C-D Raman bands. More robust, metabolically active cell lines, such as HeLa cells, can incorporate deuterium atoms into newly synthesized macromolecules at a faster rate than non-diseased mammalian cell lines, such as human embryonic kidney (HEK) cells, leading to various C-D intensities observed for the same concentration and time incubation with D2O in two different cell lines21. Furthermore, the administration of D2O at a concentration of 80% or above for 48 h may introduce toxicity to cellular samples. An optimization experiment to identify the optimal D2O concentration and time incubation is necessary to achieve desirable results.
A critical aspect of the protocol is the integration of DO-SRS and 2PEF imaging. Typically, DO-SRS and 2PEF modalities share the same picosecond excitation laser, as its narrow bandwidth can be optimized with SRS and 2PEF signals. Although our technology is homebuilt, this platform is commercially available39. Additional equipment, such as two synchronized laser pulses with a modulator, photodiode detector, a lock-in amplifier, and a scanning microscope's joint efforts are required to detect the Raman signal. D2O can be readily purchased from biotechnology vendors. Therefore, researchers can access this technology very easily.
Disclosures
The authors have nothing to disclose.
Acknowledgements
We thank Dr. Yajuan Li and Anthony Fung for their technical support, and the Fraley lab for the cell line. We acknowledge the start-up funds from UCSD, NIH U54CA132378, NIH 5R01NS111039, NIH R21NS125395, NIHU54DK134301, NIHU54 HL165443, and Hellman Fellow Award.
Materials
| 10 mL Serological Pipettes | Avantor (by VWR) | 75816-100 | https://us.vwr.com/store/product?keyword=75816-100 |
| 15 mL Conical Centrifuge Tube | VWR | 89039-664 | https://mms.mckesson.com/product/1001859/VWR-International-89039-664 |
| 16% Formaldehyde, Methanol-free | ThermoFisher Scientific | 28906 | https://www.thermofisher.com/order/catalog/product/28906 |
| 24-well plate | Fisherbrand | FB0112929 | https://www.fishersci.com/shop/products/24-well-tc-multidish-100-cs/FB012929#?keyword=FB012929 |
| 25 mm Syringe Filter, 2 μm PES | Foxx Life Sciences | 381-2216-OEM | https://www.foxxlifesciences.com/collections/pes-syringe-filters/products/381-2216-oem?variant=16274336003 |
| 460 nm Filter Cube | Olympus | OCT-ET460/50M32 | |
| AC Adapters of the Power Supply for LD OBIS 6 Laser Remote | Olympus | Supply power to the laser | |
| Band-pass Filter | KR Electronics | KR2724 | 8 MHz |
| BNC 50 Ohm Terminator | Mini Circuits | STRM-50 | |
| BNC Cable | Thorlabs | 2249-C | Coaxial Cable, BNC Male/Male |
| Broadband Dielectric Mirror | Thorlabs | BB1-E03 | 750 – 1100 nm |
| Centrifuge | |||
| Condenser | Olympus | ||
| Cover Glass | Corning | 2850-25 | https://ecatalog.corning.com/life-sciences/b2b/NL/en/Glassware/Cover-Glass/Corning%C2%AE-Square-%231%C2%BD-Cover-Glass/p/2850-25 |
| DC power supply | TopWard | 6302D | |
| Dichroic Mount | Thorlabs | KM100CL | |
| Dimethyl Sulfoxide Cell Culture Reagent | mpbio | 196055 | https://www.mpbio.com/0219605525-dimethyl-sulfoxide-cf |
| Dulbecco's Modified Eagle’s Medium without Methionine, Threonine, and Sodium Pyruvate | MilliporeSigma | 38210000 | https://www.usbio.net/media/D9800-22/dulbeccorsquos-mem-dmem-wsodium-bicarbonate-wo-methionine-threonine-sodium-pyruvate-powder With Sodium Bicarbonate and without Methionine, Threonine, and Sodium Pyruvate |
| Dulbecco’s Modified Eagle’s Medium | Corning | MT10027CV | https://www.fishersci.com/shop/products/dmem-dulbecco-s-modified-eagle-s-medium-4/MT10027CV#:~:text=Dulbecco's%20Modified%20Eagle's%20Medium%20 |
| FIJI ImageJ | ImageJ | Version 1.53t 24 August 2022 | https://imagej.net/software/fiji/downloads |
| Heavy Water (Deuterium Oxide) | Cambridge Isotope Laboratories, Inc. | 7732-18-5 | https://shop.isotope.com/productdetails.aspx?itemno=DLM-4-1L |
| Hela Cells | ATCC | CCL-2 | https://www.atcc.org/products/ccl-2 |
| Hemocymeter | MilliporeSigma | Z359629-1EA | https://www.sigmaaldrich.com/US/en/product/sigma/z359629?gclid=Cj0KCQiA37KbBhDgARIsAI zce15A5FIy0WS7I6ec2KVk QPXVMEqlAnYis_bKB6P6lr SIZ-wAXOyAELIaAhhEEAL w_wcB&gclsrc=aw.ds |
| High O.D. Bandpass Filter | Chroma Technology | ET890/220m | Filter the Stokes beam and transmit the pump beam |
| HyClone Fetal Bovine Serum (FBS) | Cytiva | SH300880340 | https://www.fishersci.com/shop/products/hyclone-fetal-bovine-serum-u-s-standard-4/SH300880340 |
| HyClone Trypsin 0.25% (1x) Solution | Cytiva | SH30042.02 | https://www.cytivalifesciences.com/en/us/shop/cell-culture-and-fermentation/reagents-and-supplements/cell-disassociation-reagents/hyclone-trypsin-protease-p-00445 |
| Integrated SRS Laser System | Applied Physics & Electronics, Inc. | picoEMERALD | picoEMERALD provides an output pulse at 1031 nm with 6-ps pulse width and 80-MHz repetition rate, which serves as the Stokes beam. The frequency doubled beam at 532 nm is used to synchronously seed a picosecond optical parametric oscillator (OPO) to produce a mode-locked pulse train with five~6 ps pulse width (the idler beam of the OPO is blocked with an,interferometric filter). The output wavelength of the OPO is tunable from 720–950 nm, which serves as the pump beam. The intensity of the 1031 nm Stokes beam is modulated sinusoidally by a built-in EOM at 8 MHz with a modulation depth of more than 90%. The pump beam is spatially overlapped with the Stokes beam by using a dichroic mirror inside picoEMERALD. The temporal overlap between pump and Stokes pulses are achieved with a built-in delay stage and optimized by the SRS signal of pure D2O at the microscope. |
| Inverted Laser-scanning Microscope | Olympus | FV1200MPE | |
| IX3-CBH Control box | Olympus | Control the laser-scanning microscope | |
| Kinematic Mirror Mount | Thorlabs | POLARIS-K1-2AH | 2 Low-Profile Hex Adjusters |
| L-Phenalynine | Sigma | P5482-25G | https://www.sigmaaldrich.com/US/en/product/sigma/p5482 |
| L-Tryptophan | Sigma | T8941-25G | https://www.sigmaaldrich.com/US/en/product/sigma/t8941 |
| LabSpec 6 | Horiba XploRA | N/A | https://www.horiba.com/gbr/scientific/products/detail/action/show/Product/labspec-6-spectroscopy-suite-software-1843/ |
| Lock-In Amplifier | Zurich Instruments | N/A | https://www.zhinst.com/americas/en/products/shfli-lock-in-amplifier |
| Long-pass Dichroic Beam Splitter | Semrock | Di02-R980-25×36 | 980 nm laser BrightLine single-edge laser-flat dichroic beamsplitter |
| MATLAB | MathWorks | Version: R2022b | https://www.mathworks.com/products/new_products/latest_features.html |
| Microscope Slides | Fisherbrand | 12-550-003 | https://www.fishersci.com/shop/products/fisherbrand-selectfrost-microscope-slides-9/12550003#?keyword=12-550-003 |
| Microscopy Imaging Software | Olympus | FluoView | |
| MPLN 100x, Olympus | Olympus | MPLAPON | https://www.olympus-ims.com/en/microscope/mplapon/#!cms[focus]=cmsContent11364 |
| MPLN 50x, Olympus | Olympus | MPLAPON | https://www.olympus-ims.com/en/microscope/mplapon/#!cms[focus]=cmsContent11363 |
| NA Oil Condenser | Olympus | 6-U130 | https://www.hitechinstruments.com/Product-Details/olympus-achromatic-aplanatic-high-na-condneser |
| Nail Polish | Wet n Wild | B01EO2G5O4 | https://www.amazon.com/dp/B01EO2G5O4/ref=cm_sw_r_api_i_E609VVDWW HHQP38FXXDC_0 |
| Origin | OriginLab | Origin 2022b (9.95) | https://www.originlab.com/index.aspx?go=PRODUCTS/Origin |
| Parafilm | Fisher Scientific | S37440 | https://www.fishersci.com/shop/products/parafilm-m-wrapping-film-3/p-2379782 |
| PBS 1x (Dulbecco's Phosphate Buffered Saline) | Thermofischer – Gibco | 14040117 | https://www.thermofisher.com/order/catalog/product/14040117?SID=srch-hj-14040117 |
| Penicillin/Streptomycin | Thermofischer – Gibco | 15140122 | https://www.thermofisher.com/order/catalog/product/15140122 |
| Periscope Assembly | Thorlabs | RS99 | Includes the top and bottom units, Ø1" post, and clamping fork. |
| picoEmerald System | A.P.E | N/A | https://www.ape-berlin.de/en/cars-srs/ |
| Shielded Box with BNC Connectors | Pomona Electronics | 2902 | Aluminum Box with Cover, BNC Female/Female |
| Si Photodiode Detector | Home Built | N/A | DYI series |
| Silicon Wafer | |||
| Spacers | Grace Bio-Labs | 654008 | https://gracebio.com/product/secureseal-imaging-spacers-654008/ |
| Spontaneous Raman spectroscopy | Horiba XploRA | N/A | https://www.horiba.com/int/products/detail/action/show/Product/xploratm-plus-1528/ |
| Stimulated Raman Scattering Microscopy | Home Built | N/A | |
| Touch Panel Controller | Olympus | Control the X-Y direction of the laser-scanning microscope | |
| Trypan Blue 0.4% (0.85% NaCl) | Lonza | 17-942E | https://bioscience.lonza.com/lonza_bs/US/en/Culture-Media-and-Reagents/p/000000000000181876/Trypan-Blue%2C-0-4%25-Solution" |
| Tweezers | Kaverme – Amazon | B07RNVXXV1 | https://www.amazon.com/Precision-Anti-Static-Electronics-Laboratory-Jewelry-Making/dp/B07RNVXXV1" |
| Two Photon Excitation Fluorescence Microscopy | Home Built | N/A | |
| Weighing Paper | VWR | 12578-165 | https://us.vwr.com/store/product/4597617/vwr-weighing-paper |
| Zurich LabOneQ Software | Zurich Instruments | Control the Zurich lock-in amplifier |
References
- Wu, G. Functional amino acids in nutrition and health. Amino Acids. 45 (3), 407-411 (2013).
- Wei, Z., Liu, X., Cheng, C., Yu, W., Yi, P. Metabolism of amino acids in cancer. Frontiers in Cell and Developmental Biology. 8, 603837 (2020).
- Parthasarathy, A., et al. A three-ring circus: Metabolism of the three proteogenic aromatic amino acids and their role in the health of plants and animals. Frontiers in Molecular Biosciences. 5, 29 (2018).
- Wang, H., et al. l-tryptophan activates mammalian target of rapamycin and enhances expression of tight junction proteins in intestinal porcine epithelial cells. The Journal of Nutrition. 145 (6), 1156-1162 (2015).
- Saxton, R. A., Sabatini, D. M. mTOR signaling in growth, metabolism, and disease. Cell. 168 (6), 960-976 (2017).
- Mossmann, D., Park, S., Hall, M. N. mTOR signalling and cellular metabolism are mutual determinants in cancer. Nature Reviews. Cancer. 18 (12), 744-757 (2018).
- Kimura, T., Watanabe, Y. Tryptophan protects hepatocytes against reactive oxygen species-dependent cell death via multiple pathways including Nrf2-dependent gene induction. Amino Acids. 48 (5), 1263-1274 (2016).
- Ma, Q., et al. Dietary supplementation with aromatic amino acids decreased triglycerides and alleviated hepatic steatosis by stimulating bile acid synthesis in mice. Food and Function. 12 (1), 267-277 (2021).
- Cheng, C., et al. Treatment implications of natural compounds targeting lipid metabolism in nonalcoholic fatty liver disease, obesity and cancer. International Journal of Biological Sciences. 15 (8), 1654-1663 (2019).
- Lubes, G., Goodarzi, M. GC-MS based metabolomics used for the identification of cancer volatile organic compounds as biomarkers. Journal of Pharmaceutical and Biomedical Analysis. 147, 313-322 (2018).
- Di Gialleonardo, V., Wilson, D. M., Keshari, K. R. The potential of metabolic imaging. Seminars in Nuclear Medicine. 46 (1), 28-39 (2016).
- Bowman, A. P., et al. Evaluation of lipid coverage and high spatial resolution MALDI-imaging capabilities of oversampling combined with laser post-ionisation. Analytical and Bioanalytical Chemistry. 412 (10), 2277-2289 (2020).
- Murphy, R. C., Hankin, J. A., Barkley, R. M. Imaging of lipid species by MALDI mass spectrometry. Journal of Lipid Research. 50, 317-322 (2009).
- Pirman, D. A., et al. Changes in cancer cell metabolism revealed by direct sample analysis with MALDI mass spectrometry. PLoS One. 8 (4), e61379 (2013).
- Li, Z., et al. Single-cell lipidomics with high structural specificity by mass spectrometry. Nature Communications. 12 (1), 2869 (2021).
- Miyagi, M., Kasumov, T. Monitoring the synthesis of biomolecules using mass spectrometry. Philosophical Transactions. Series A, Mathematical, Physical and Engineering Sciences. 374 (2079), 20150378 (2016).
- Wang, T., Shogomori, H., Hara, M., Yamada, T., Kobayashi, T. Nanomechanical recognition of sphingomyelin-rich membrane domains by atomic force microscopy. 생화학. 51 (1), 74-82 (2012).
- Fung, A. A., Shi, L. Mammalian cell and tissue imaging using Raman and coherent Raman microscopy. Wiley Interdisciplinary Reviews. Systems Biology and Medicine. 12 (6), e1501 (2020).
- Shi, L., Fung, A. A., Zhou, A. Advances in stimulated Raman scattering imaging for tissues and animals. Quantitative Imaging in Medicine and Surgery. 11 (3), 1078-1101 (2021).
- Yamakoshi, H., et al. Imaging of EdU, an alkyne-tagged cell proliferation probe, by Raman microscopy. Journal of the American Chemical Society. 133 (16), 6102-6105 (2011).
- Shi, L., et al. Optical imaging of metabolic dynamics in animals. Nature Communications. 9 (1), 2995 (2018).
- Bagheri, P., Hoang, K., Fung, A. A., Hussain, S., Shi, L. Visualizing cancer cell metabolic dynamics regulated with aromatic amino acids using DO-SRS and 2PEF microscopy. Frontiers in Molecular Biosciences. 8, 779702 (2021).
- Li, Y., et al. Direct imaging of lipid metabolic changes in drosophila ovary during aging using DO-SRS microscopy. Frontiers in Aging. 2, 819903 (2022).
- Li, Y., Zhang, W., Fung, A. A., Shi, L. DO-SRS imaging of metabolic dynamics in aging Drosophila. Analyst. 146 (24), 7510-7519 (2021).
- Zhang, L., et al. Spectral tracing of deuterium for imaging glucose metabolism. Nature Biomedical Engineering. 3 (5), 402-413 (2019).
- Fung, A. A., et al. Imaging sub-cellular methionine and insulin interplay in triple negative breast cancer lipid droplet metabolism. Frontiers in Oncology. 12, 858017 (2022).
- Li, Y., Zhang, W., Fung, A. A., Shi, L. DO-SRS imaging of diet regulated metabolic activities in Drosophila during aging processes. Aging Cell. 21 (4), e13586 (2022).
- Shi, L., Wei, M., Min, W. Highly-multiplexed tissue imaging with raman dyes. Journal of Visualized Experiments. (182), e63547 (2022).
- Rysman, E., et al. De novo lipogenesis protects cancer cells from free radicals and chemotherapeutics by promoting membrane lipid saturation. 암 연구학. 70 (20), 8117-8126 (2010).
- Lisec, J., Jaeger, C., Rashid, R., Munir, R., Zaidi, N. Cancer cell lipid class homeostasis is altered under nutrient-deprivation but stable under hypoxia. BMC Cancer. 19 (1), 501 (2019).
- Thiam, A. R., Dugail, I. Lipid droplet-membrane contact sites – from protein binding to function. Journal of Cell Science. 132 (12), (2019).
- Schott, M. B., et al. Lipid droplet size directs lipolysis and lipophagy catabolism in hepatocytes. The Journal of Cell Biology. 218 (10), 3320-3335 (2019).
- Hoang, K., et al. Subcellular resolution DO-SRS and 2PEF imaging of metabolic dynamics regulated by L-methionine in amyotrophic lateral sclerosis. Optical Biopsy XXI: Toward Real-Time Spectroscopic Imaging and Diagnosis. SPIE. 1237303, 6-13 (2023).
- Jang, H., et al. Super-resolution stimulated Raman scattering microscopy with A-PoD. bioRxiv. , (2022).
- Li, Y., et al. Optical metabolic imaging uncovers sex- and diet-dependent lipid changes in aging drosophila brain. bioRxiv. , (2022).
- Zhang, W., et al. Multi-molecular hyperspectral PRM-SRS imaging. bioRxiv. , (2022).
- Wei, M., et al. Volumetric chemical imaging by clearing-enhanced stimulated Raman scattering microscopy. Proceedings of the National Academy of Sciences. 116 (14), 6608-6617 (2019).
- Chang, T., et al. Non-invasive monitoring of cell metabolism and lipid production in 3D engineered human adipose tissues using label-free multiphoton microscopy. Biomaterials. 34 (34), 8607-8616 (2013).
- Leica TCS SP8 CARS CARS Microscope – Label Free Imaging. Leica Microsystems Available from: https://www.leica-microsystems.com/products/confocal-microscopes/p/leica-tcs-sp8-cars/downloads/ (2023)

