Subcellular Fractionation for ERK Activation Upon Mitochondrial-derived Peptide Treatment
Summary
This protocol describes how to stimulate cells with mitochondrial-derived peptides and assess the signaling cascade and localization of phospho-proteins.
Abstract
Mitochondrial-derived peptides (MDPs) are a new class of peptides that are encoded by small open reading frames within other known genes of the mitochondrial genome. MDPs have a wide variety of biological effects such as protecting neurons from apoptosis, improving metabolic markers, and protecting cells from chemotherapy. Humanin was the first MDP to be discovered and is the most studied peptide among the MDP family. The membrane receptors and downstream signaling pathways of humanin have been carefully characterized. Additional MDPs such as MOTS-c and SHLP1-6 have been more recently discovered and the signaling mechanisms have yet to be elucidated. Here we describe a cell culture based method to determine the function of these peptides. In particular, cell fractionation techniques in combination with western blotting allow for the quantitative determination of activation and translocation of important signaling molecule. While there are other methods of cell fractionation, the one described here is an easy and straightforward method. These methods can be used to further elucidate the mechanism of action of these peptides and other therapeutic agents.
Introduction
Emerging studies show that mitochondrial-derived peptides (MDPs) play important roles in cytoprotection and metabolism1,2,3. Understanding the signal transduction pathway in the presence of MDPs gives us insight into the mechanism by which MDPs modulate various functions. The first identified MDP, humanin, has been shown to increase extracellular signal-regulated kinase 1/2 (ERK1/2) phosphorylation through its receptor binding4,5. However, the downstream effect of ERK1/2 activation is still underexplored.
The ERK1/2 cascade serves as an essential mediator in a variety of cellular processes including proliferation, cell migration, cellular metabolism, survival, and apoptosis6,7,8. To orchestrate all these distinct cellular processes, the activity and subcellular localization of ERK1/2 are tightly regulated by phosphatases and scaffold proteins9,10. In addition to post-translational modification, the dynamic shuttling of ERK1/2 also regulates its signaling function, activity, and specificity11,12. ERK1/2 is primarily localized in the cytosol13. A set of anchoring and scaffold proteins help retain ERK1/2 in cytoskeletal elements, on the surface of organelles, or diffused in the cytoplasm13. Upon stimulation, ERK1/2 is phosphorylated and becomes dissociated from its anchoring proteins, allowing the translocation of ERK1/2 to other subcellular compartments, including the nucleus, mitochondria, Golgi, and lysosomes14,15,16.
Although humanin is known to activate the ERK1/2 signaling pathway, the activation of ERK1/2 is only observed in the total cell lysates. As described previously, since ERK1/2 subcellular localization plays a crucial role in its downstream effect, the analysis of both the subcellular localization and the total levels of phosphorylated ERK1/2 is necessary to provide a comprehensive understanding of humanin-induced ERK1/2 activation and the activation of downstream targets.
To understand the target organelles of activated ERK1/2, subcellular fractionation followed by western blotting for phosphorylated ERK1/2 was performed. This method can be easily implemented as it utilizes standard laboratory equipment and reagents. The isolated subcellular compartments are of high purity, allowing the results to be straightforwardly interpreted. Immunostaining of ERK1/2 may produce similar results. However, certain subcellular compartments are relatively hard to visualize and require special fixation and permeabilization methods. ERK1/2 levels vary in subcellular compartments, and this variation could cause false positive and false negative signals when looking at whole cell lysates. Therefore, an immunoblot using isolated subcellular compartments gives us a better understanding of ERK1/2 localization.
The versatility of the method enables modifications of the protocol to investigate effects of other stimulants including other MDPs or translocation of other signaling molecules such as STAT3. Recently discovered small humanin-like peptides (SHLPs) are encoded from 16S rRNA region where humanin is encoded, and they have similar but distinct properties compared to HN17. For example, SHLP2 and SHLP3 activate ERK1/2 after 8 h although humanin activates ERK1/2 within 5 min. Examining subcellular localization of ERK1/2 in response to different peptides will give us a better understanding of the biology of these peptides. Emerging evidence showed that the subcellular localization of signaling molecules plays a crucial role in their downstream effects. For instance, STAT3 traditionally is known to be primarily localized in the cytosol in resting cells, and then it translocates into the nucleus to activate gene expression in response to cytokines18. STAT3 also translocates to mitochondria and regulates the TCA cycle and ATP production19. Regarding autophagy regulation, different subcellular localization of STAT3 modulates autophagy in various ways20. For example, nuclear STAT3 transcriptionally regulates autophagy-related genes and acts as an autophagy modulator. Cytoplasmic STAT3 constitutively inhibits autophagy by interacting with autophagy signaling molecules. Mitochondrial STAT3 inhibits and prevents mitophagy by suppressing oxidative stress induced autophagy. Therefore, this subcellular compartment isolation method is crucial for understanding the role of other signaling molecules as well as ERK1/2.
Protocol
1. Peptide Treatment to Cells
- Plate two million SH-SY5Y or HEK293 cells (2 x 106) into a 10 cm dish and grow them for 2 days
- (Optional) The next day, wash the cells with serum-free Dulbecco's Modified Eagle Medium (DMEM) media once and incubate with serum free DMEM overnight if the treatment needs to be done in serum free media.
- On day 3, dissolve S14G-humanin peptides in 0.2 µm filtered, distilled water, and reconstitute them as 1mM stock solution.
NOTE: The peptides should be dissolved in the appropriate solvent, which can be determined by the sequence characteristics of the peptide (e.g., overall charge) and may be provided by the manufacturer if the peptides are commercially available. For example, S14G-humanin, SHLP2, and SHLP6, and MOTS-c dissolve in water. Aliquot the peptides in an appropriate volume (e.g., 50 μL) to avoid freeze and thaw cycles. Once thawed, do not use them again. - Aliquot the pre-warmed serum-containing or serum-free media in a 50-mL conical tube and add room temperature 1 mM stock solution to make it into a working concentration (e.g., 1 µM, 10 µM).
- Replace the media with 6 mL peptide solution from step 1.4. and incubate the cells for the appropriate amount of time.
NOTE: Choose the incubation time depending on your condition which activates ERK.
2. Subcellular Fractionation for Cytosol and Nucleus
- Wash the cells twice with 10 mL of ice-cold PBS, scrape the cells off the plate in 5 mL of ice-cold PBS, and transfer the cell suspension into a 15 mL conical tube.
- Centrifuge the cells at 500 x g for 5 min at 4 °C.
- Aspirate the PBS and re-suspend the pellet in 200 µL of ice-cold, fractionation buffer (10 mM HEPES pH = 7.6, 3 mM MgCl2, 10 mM KCl, 5% (v/v) glycerol, 1% non-ionic surfactant (C13H22O(C2H4O)n), protease/phosphatase inhibitors).
NOTE: To keep the proteins in their phosphorylation status, the phosphatase inhibitor as well as protease inhibitor should be freshly added and samples should be kept on ice at all times. Repeating freeze and thaw cycles of samples should be avoided. Aliquot samples into small amount (e.g. 50 μg) before freezing them at -80 °C. - Incubate the re-suspended pellet on ice for 15 min and then centrifuge at 250 x g for 5 min at 4 °C.
- Collect the supernatant as the cytoplasmic fraction and keep the pellet for the nuclear fraction.
- Centrifuge the supernatant to remove cellular debris and other contaminants at 18,000 x g for 10 min at 4 °C and then transfer the supernatant into a new microcentrifuge tube. This is the cytosol fraction.
- Resuspend the pellet in 200 μL ice-cold wash buffer (10 mM HEPES pH = 7.6, 1.5 mM MgCl2, 420 mM NaCl, 25% (v/v) glycerol, 0.2 mM EDTA, protease/phosphatase inhibitors) and centrifuge at 250 x g for 5 min at 4 °C
- Remove the supernatant and resuspend the pellet in 100 μL ice-cold, nuclear extraction buffer (20 mM HEPES pH = 7.6, 1.5 mM MgCl2, 420 mM NaCl, 25% (v/v) glycerol, 0.2 mM EDTA, protease/phosphatase inhibitors) and sonicate 10 times (5 s on, 10 s off, 30% amp.) on ice.
- Centrifuge at 18,000 x g for 10 min at 4 °C.
- Transfer the supernatant to a new microcentrifuge tube. This is the nuclear fraction.
- Quantify the protein amount using a BCA assay
3. Crude Mitochondrial Fraction
- Wash the cells in each 10 cm dish with 10 mL of ice-cold PBS, add 5 mL ice-cold PBS, and detach the cells using a cell scraper.
NOTE: Use three 10 cm dishes of cells to get a good yield of mitochondria for Western Blotting. - Transfer the cell suspension to a 15 mL conical tube, and combine all from three 10 cm dishes into one 15 ml conical tube.
- Centrifuge the cells at 600 x g for 10 min at 4 °C.
- Aspirate the PBS and re-suspend the pellet in 1 mL of ice-cold mitochondria isolation buffer (10 mM Tris-MOPS, 1 mM EGTA/Tris, 200 mM Sucrose, adjust to pH = 7.4).
- Homogenize the cells with 25 strokes of a 2 mL homogenizer with polytetrafluoroethylene coated pestle on ice.
NOTE: This step is critical to maintain mitochondrial integrity and maximize the yield of the mitochondrial fraction. The number of strokes should be optimized for each cell type. Precool the homogenizer before starting the procedure. - Transfer the homogenate to a microcentrifuge tube and centrifuge it at 600 x g for 10 min at 4 °C to remove nuclei and unbroken cells.
- Collect the supernatant, transfer it to a new microcentrifuge tube, and centrifuge it at 7,000 x g for 10 min at 4 °C.
NOTE: The pellet is loose, collect the supernatant with care and try not to disturb the pellet. - Remove the supernatant, re-suspend the pellet with 200 μL of ice-cold mitochondria isolation buffer, and transfer the solution to a new microcentrifuge tube. Centrifuge the tube at 7,000 x g for 10 min at 4 °C.
- Repeat step 3.8 to wash the pellet. It is not necessary to transfer the supernatant to a new microcentrifuge tube in this step.
- Remove the supernatant from the washed pellet and re-suspend the pellet containing mitochondria with 50 μL of RIPA buffer.
- Incubate the suspension on ice for 10 min.
- Centrifuge the suspension at 16,000 x g for 15 min at 4 °C.
- Transfer the supernatant to a new microcentrifuge tube. This is the mitochondrial fraction.
- Quantify the protein amount using a BCA assay.
4. Western Blotting for Phospho-Specific Proteins
- Perform an SDS-polyacrylamide gel electrophoresis (8-16% premade gel) and transfer the protein to a PVDF membrane4.
- Incubate the membrane with 5% BSA in TBST (0.1% Polysorbate 20) for 30 min at room temperature to block background non-specific binding sites.
NOTE: For phosphorylated protein detection, block the membrane with BSA not Milk. Milk contains casein, an abundant phosphoprotein, which results in high nonspecific signal. - Incubate the membrane with the primary antibody (anti-phospho-ERK1/2) at 4 °C overnight.
- The next day, wash the membrane with TBST (0.1% polysorbate 20) three times for 5 min at room temperature.
- Incubate the membrane with secondary antibody (anti-rabbit HRP) for 1 h at room temperature.
- Wash the membrane with TBST (0.1% polysorbate 20) three times for 5 min at room temperature.
- Incubate the membrane with ECL solution and image the membrane in the image analyzer.
- Strip the membrane with stripping buffer for 15 min at room temperature and wash the membrane with TBST three times for 5 min.
- Repeat steps 4.2 to 4.7 using the anti-total ERK1/2 antibody.
- To check the purity of each fraction, run SDS-PAGE and perform immunoblots using antibodies recognizing markers for each compartment (e.g., Lamin B1 for nucleus, GAPDH for cytosol, and TOM20 for mitochondria).
Representative Results
Using the procedure presented here, we treated HEK293 and SH-SY5Y cells with 1 μM and 100 μM S14G-humanin, a potent humanin analog21, respectively, in complete media for the indicated time periods (Figure 1A and Figure 1B). We then examined the total and phosphorylated form of ERK1/2 at Thr202/Tyr204 from total protein extracts. S14G-humanin treatment showed increase in phosphorylation of ERK1/2 at its regulatory Thr202/Tyr204 Site. The subcellular localization of ERK1/2 plays an important role in its signaling function, particularly in getting its specificity. To understand the target of S14G-humanin-mediated ERK1/2 activation, we analyzed the subcellular localization of total and phosphorylated ERK1/2 following S14G-humanin treatment. In HEK293 cells, the phosphorylated ERK increased in the both cytoplasm and nucleus, but not in the mitochondria (Figure 2A). Interestingly, phosphorylated ERK decreased in the cytoplasm, nucleus, and mitochondria in SH-SY5Y cells (Figure 2B). These results suggest that S14G-humanin-mediated ERK phosphorylation may play a different role in different cell types and different conditions. Therefore, further studies are required to analyze the localization of S14G-humanin-mediated phosphorylation of ERK1/2 in other subcellular compartments, as phosphorylated ERK1/2 translocates into the nucleus, mitochondria, endosomes/lysosomes, and various membranes.
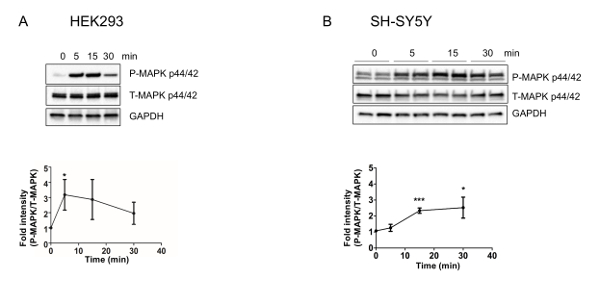
Figure 1: S14G-humanin increases the phosphorylation of ERK1/2 in HEK293 and SH-SY5Y cells. Quantification and representative western blots of ERK activation in (A) HEK293 and (B) SH-SY5Y cells. Total cell lysates following (A) 1 µM or (B) 100 µM S14G-humanin treatment in DMEM supplemented with 10% FBS for the indicated time periods were immunoblotted using anti-phospho- and total-ERK 1/2 (Thr202/Tyr204). This figure has been modified from Kim et al.4 Please click here to view a larger version of this figure.
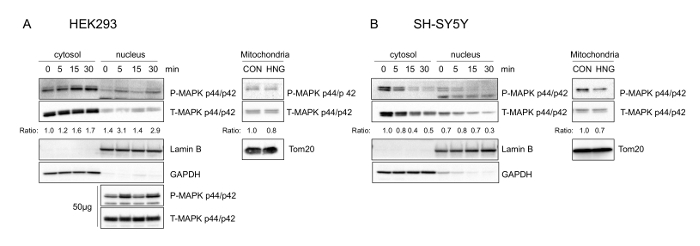
Figure 2: Phosphorylated ERK1/2 is localized to cytosol, nucleus, and other subcellular compartments upon S14G-humanin treatment in HEK293 and SH-SY5Y cells. Representative western blots showing ERK1/2 subcellular localization (n = 3). (A) HEK293 cells were treated with 1 µM S14G-humanin. Cytosolic (15 µg), nuclear (15 µg and 50 µg), and mitochondrial (15 μg) lysates were immunoblotted using phospho- and total-ERK antibody. (B) SH- SY5Y cells were treated with 100 µM S14G-humanin. Mitochondrial fractions were obtained 30 min after S14G-humanin treatment. Cytosolic, nuclear, and mitochondrial (15 µg) lysates were immunoblotted using phospho- and total-ERK antibody. Anti-GAPDH, Anti-Lamin B, anti-Tom20 antibody were used as the cytosolic, nuclear, mitochondrial fraction markers, respectively. This figure has been modified from Kim et al.4. Please click here to view a larger version of this figure.
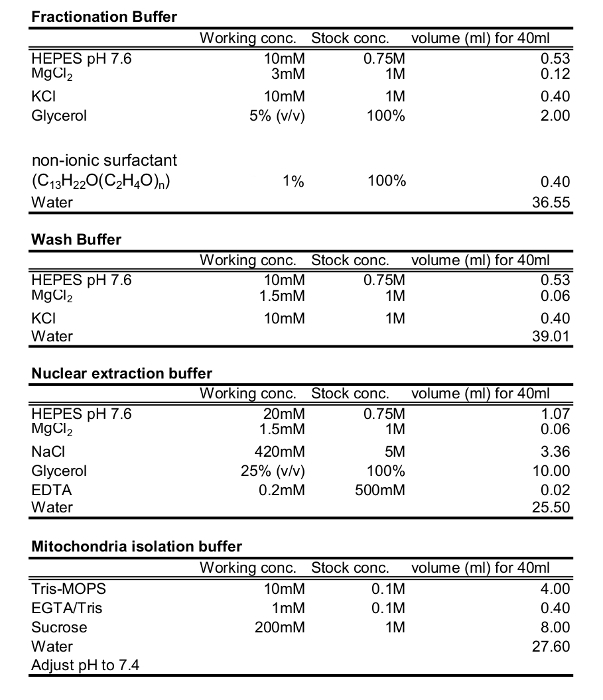
Table 1: Buffers for cytosol, nuclear, and Mitochondrial fraction.
Discussion
Here, we demonstrated that humanin peptide-mediated ERK1/2 activation occurs in two different cell types, and the subcellular localization of activated ERK1/2 can be different depending on the conditions (e.g., dose of peptide, time point, and cell type). It has been shown that humanin signals through two different receptors22,23, which may explain the differences in signaling between the two cell lines as well as the requirement for different doses of humanin. The physiological implications of this have not been fully studied but it may suggest one possible mechanism for tissue dependent responses to HN.
Phosphorylation of ERK1/2 plays a crucial role in ERK1/2 signaling and the phosphorylation status of ERK1/2 shows dynamic changes. It is important to optimize the conditions to find when ERK1/2 is phosphorylated under a specific condition/stimulus. To achieve this, try different stimulation conditions (e.g., dose and medium) and conduct time courses to find when the highest level of phosphorylation is achieved. A majority of studies have only focused on ERK1/2 phosphorylation in total cell lysates. However, the results may provide limited functional information as ERK1/2 can interact with different targets and modulate different downstream cascades in specific subcellular compartments. Therefore, examining the subcellular localization of phosphorylated ERK1/2 gives a better understanding of the role of ERK1/2 activation.
Subcellular fractionation after ERK1/2 activation can be used as a source of further study (e.g., co-immunoprecipitation) to identify novel downstream targets because the subcellular fractionation can enrich for downstream targets of ERK1/2. This method can be applied to other signaling molecules that are translocated into different subcellular compartments upon stimuli after optimization of the conditions.
Although various methods to isolate mitochondria have been proposed in the past decades, there is partial contamination from other subcellular compartments. The crude mitochondrial isolation method proposed here also contains limited amount of cytoplasm contaminants. As the amount of contamination is small, we consider that it is still a valid method for detecting the localization of ERK1/2 into mitochondria upon activation. As peptides are not necessarily stable in solution, it is sometimes difficult to maintain consistency between experiments. Be sure to make small aliquots of peptide solution to avoid freeze-thaw effects and to improve the consistency of peptide activity.
Declarações
The authors have nothing to disclose.
Acknowledgements
This work was supported by an Ellison/AFAR Postdoctoral Fellowship in Aging Research Program to SJK, and a Glenn Foundation Award and NIH grants to PC (1P01AG034906, 1R01GM 090311, 1R01ES 020812). All authors appear in the film.
Materials
| p44/42 MAPK (ERK1/2) | Cell signaling | 9102 | Dilution 1:1,000 |
| phospho-p44/42 MAPK (ERK1/2)(Thr202/Tyr204) | Cell signaling | 4370 | Dilution 1:1,000 |
| Lamin B1 | Cell signaling | 12586 | Nuclear Marker, Dilution 1:1,000 |
| GAPDH | Cell signaling | 5174 | Cytoplasmic marker, Dilution 1:2,000 |
| Tom20 | Santa cruz | SC-17764 | Mitochondria marker, Dilution 1:2,000 |
| anti-Rabbit-HRP conjugated | Cell signaling | 7074 | Dilution 1:30,000 |
| RIPA Lysis and Extraction Buffer | ThermoFisher SCIENTIFIC | 89900 | |
| 100 mm Culture Dish | ThermoFisher SCIENTIFIC | 12556002 | |
| HNG peptide | Genescript | ||
| 25mm sylinge filter | ThermoFisher SCIENTIFIC | 09-719A | |
| HEPES | Sigma | H3375 | |
| MgCl2 | Sigma | M8266 | |
| KCl | Sigma | P9333 | |
| Glycerol | Sigma | G9012 | |
| Triton X-100 | ThermoFisher SCIENTIFIC | BP151-100 | |
| EDTA | Sigma | 3609 | |
| MOPS | Sigma | M1254 | |
| EGTA | Sigma | E3889 | |
| Sucrose | Sigma | S7903 | |
| Tris-base | ThermoFisher SCIENTIFIC | BP152-1 | |
| HCL | Sigma | H1758 | |
| PBS | Lonza | 17-512F | |
| Cell Scraper | FALCON | 353085 | |
| Halt™ Protease and Phosphatase Inhibitor Cocktail (100X) | ThermoFisher SCIENTIFIC | 78440 | |
| Thomas Pestle Tissue Grinder Assemblies with Smooth Pestles | Thomas Scientific | 3432S90 | |
| Tween-20 | ThermoFisher SCIENTIFIC | BP337-500 | |
| BSA | ThermoFisher SCIENTIFIC | BP1600-100 | |
| 8-16% Mini-PROTEAN TGX Precast Protein Gels | BIO RAD | 4561104 | |
| Mini Trans-Blot Module | BIO RAD | 1658030 | |
| Trans-Blot Turbo Transfer System | BIO RAD | 1704150 | |
| Trans-Blot Turbo RTA Mini PVDF Transfer Kit | BIO RAD | 1704272 | |
| Clarity Western ECL Blotting Substrates | BIO RAD | 1705060 | |
| Restore Western blot stripping buffer | ThermoFisher SCIENTIFIC | 21059 | |
| Dulbecco's Modified Eagle Medium | ThermoFisher SCIENTIFIC | 11965-092 | |
| Sonicator, Medel: FB120 | ThermoFisher SCIENTIFIC | 695320-07-12 |
Referências
- Yen, K., Lee, C., Mehta, H., Cohen, P. The emerging role of the mitochondrial-derived peptide humanin in stress resistance. J. Mol. Endocrinol. 50 (1), 11-19 (2013).
- Muzumdar, R. H., Huffman, D. M., et al. Humanin: a novel central regulator of peripheral insulin action. PloS one. 4 (7), 6334 (2009).
- Lee, C., Yen, K., Cohen, P. Humanin: a harbinger of mitochondrial-derived peptides. Trends Endocrinol. Metab. 24 (5), 222-228 (2013).
- Kim, S. J., Guerrero, N., et al. The mitochondrial-derived peptide humanin activates the ERK1/2, AKT, and STAT3 signaling pathways and has age-dependent signaling differences in the hippocampus. Oncotarget. 7 (30), 46899-46912 (2016).
- Ying, G., Iribarren, P., et al. Humanin, a newly identified neuroprotective factor, uses the G protein-coupled formylpeptide receptor-like-1 as a functional receptor. J. Immunol. 172 (11), 7078-7085 (2004).
- Zhang, W., Liu, H. T. MAPK signal pathways in the regulation of cell proliferation in mammalian cells. Cell Res. 12 (1), 9-18 (2002).
- Huang, C., Jacobson, K., Schaller, M. D. MAP kinases and cell migration. J. Cell Sci. 117, 4619-4628 (2004).
- Cagnol, S., Chambard, J. -. C. ERK and cell death: mechanisms of ERK-induced cell death–apoptosis, autophagy and senescence. FEBS J. 277 (1), 2-21 (2010).
- Raman, M., Chen, W., Cobb, M. H. Differential regulation and properties of MAPKs. Oncogene. 26 (22), 3100-3112 (2007).
- Morrison, D. K., Davis, R. J. Regulation of MAP kinase signaling modules by scaffold proteins in mammals. Annu. Rev. Cell Dev. 19 (1), 91-118 (2003).
- Wainstein, E., Seger, R. The dynamic subcellular localization of ERK: mechanisms of translocation and role in various organelles. Curr. Opin. Cell Biol. 39, 15-20 (2016).
- Zehorai, E., Yao, Z., Plotnikov, A., Seger, R. The subcellular localization of MEK and ERK–a novel nuclear translocation signal (NTS) paves a way to the nucleus. Mol. Cell. Endocrinol. 314 (2), 213-220 (2010).
- Kolch, W. Coordinating ERK/MAPK signalling through scaffolds and inhibitors. Nat. Rev. Mol. Cell Biol. 6 (11), 827-837 (2005).
- Horbinski, C., Chu, C. T. Kinase signaling cascades in the mitochondrion: a matter of life or death. Free Radic. Biol. Med. 38 (1), 2-11 (2005).
- Aebersold, D. M., Shaul, Y. D., et al. Extracellular signal-regulated kinase 1c (ERK1c), a novel 42-kilodalton ERK, demonstrates unique modes of regulation, localization. Mol Cell Biol. 24 (22), 10000-10015 (2004).
- Zhu, J. -. H., Guo, F., Shelburne, J., Watkins, S., Chu, C. T. Localization of phosphorylated ERK/MAP kinases to mitochondria and autophagosomes in Lewy body diseases. Brain Pathol. 13 (4), 473-481 (2003).
- Cobb, L. J., Lee, C., et al. Naturally occurring mitochondrial-derived peptides are age-dependent regulators of apoptosis, insulin sensitivity, and inflammatory markers. Aging. 8 (4), 796-809 (2016).
- Ihle, J. N. The Stat family in cytokine signaling. Curr. Opin. Cell Biol. 13 (2), 211-217 (2001).
- Xu, Y. S., Liang, J. J., et al. STAT3 Undergoes Acetylation-dependent Mitochondrial Translocation to Regulate Pyruvate Metabolism. Sci. Rep. 6 (1), 39517 (2016).
- You, L., Wang, Z., et al. The role of STAT3 in autophagy. Autophagy. 11 (5), 729-739 (2015).
- Hashimoto, Y., Niikura, T., et al. Detailed characterization of neuroprotection by a rescue factor humanin against various Alzheimer’s disease-relevant insults. The J Neurosci. 21 (23), 9235-9245 (2001).
- Ying, G., Iribarren, P., et al. Humanin, a newly identified neuroprotective factor, uses the G protein-coupled formylpeptide receptor-like-1 as a functional receptor. J. Immunol. 172 (11), 7078-7085 (2004).
- Hashimoto, Y., Kurita, M., et al. Humanin inhibits apoptosis in pituitary tumor cells through several signaling pathways including NF-κB activation. Mol. Biol. Cell. 20 (12), 2864-2873 (2009).

