Improved 3D Hydrogel Cultures of Primary Glial Cells for In Vitro Modelling of Neuroinflammation
Summary
Herein, we present a protocol for the 3D culture of rat brain-derived glia cells, including astrocytes, microglia, and oligodendrocytes. We demonstrate primary cell culture, methacrylated hyaluronic acid (HAMA) hydrogel synthesis, HAMAphoto-polymerization and cell encapsulation, and sample processing for confocal and scanning electron microscopic imaging.
Abstract
In the central nervous system, numerous acute injuries and neurodegenerative disorders, as well as implanted devices or biomaterials engineered to enhance function result in the same outcome: excess inflammation leads to gliosis, cytotoxicity, and/or formation of a glial scar that collectively exacerbate injury or prevent healthy recovery. With the intent of creating a system to model glial scar formation and study inflammatory processes, we have generated a 3D cell scaffold capable of housing primary cultured glial cells: microglia that regulate the foreign body response and initiate the inflammatory event, astrocytes that respond to form a fibrous scar, and oligodendrocytes that are typically vulnerable to inflammatory injury. The present work provides a detailed step-by-step method for the fabrication, culture, and microscopic characterization of a hyaluronic acid-based 3D hydrogel scaffold with encapsulated rat brain-derived glial cells. Further, protocols for characterization of cell encapsulation and the hydrogel scaffold by confocal immunofluorescence and scanning electron microscopy are demonstrated, as well as the capacity to modify the scaffold with bioactive substrates, with incorporation of a commercial basal lamina mixture to improved cell integration.
Introduction
Inflammation of the central nervous system (CNS) has long been considered a hallmark of acute (e.g., ischemic stroke, traumatic brain and spinal cord injury) and chronic (e.g. Alzheimer's, Parkinson's, and Huntington's diseases) CNS injury, but is increasingly recognized as a causal contributor to neurodegenerative and neuropsychiatric disorders. Sustained or inappropriate inflammation can cause neural injury and demyelination (e.g. multiple sclerosis), and negatively influence brain development (e.g., schizophrenia, autism) and mood states (e.g., depression, anxiety, bipolar disorder). Further, novel therapeutic strategies using implantable devices (e.g.,brain-computer-interfaces1,2,3, deep brain stimulation4,5, intraspinal microstimulation6,7,8,9,10) generate a predictable inflammatory response at the interface between the device and the CNS resulting in a protective tissue response that can cause loss of efficacy or device failure over the lifetime of the implant11. Inflammation in the CNS is typically initiated by microglia, which function as the resident immune cells of the CNS responsible for tissue surveillance and mounting the foreign body response (reviewed12). Depending on the severity of an insult, the microglia signal and recruit additional cell types to an injury site. Specifically, the microglia activate the astrocytes, which in turn act as secondary inflammatory cells and form a dense protective barrier to contain an injury site13,14. Microglia can also initiate an activity cascade in the cells of the peripheral immune system, which can result in the breakdown of the BBB to permit immune infiltration (reviewed in reference15).
In the case of devices implanted into the CNS, tissue damage resulting from device insertion as well as the continued presence of the foreign device may initiate a process termed glial scarring. In this process, the microglia migrate to and proliferate at the site of injury. They also initiate the release of inflammatory factors to neutralize potential threats and recruit additional glial cells. Subsequently, activated astrocytes become hypertrophic and begin encapsulating the implanted device to form a continuous fibrous barrier16. Inflammatory signalling also serves to promote withdrawal of neuronal processes from the vicinity of the implant and eventually recruits fibroblasts to reinforce the developing glial scar17. The oligodendrocytes, responsible for sheathing neurons in myelin to enhance conductance, do not survive this process and distant cells are partitioned from the implant by the scar18. Glial scarring greatly reduces the function and lifetime of implanted devices, particularly for recording electrodes, and ultimately serves to limit the functionality of neural interfaces19.
Several approaches have been exploited to increase the biocompatibility and interface activity of implanted devices in the CNS20,21,22,23. An extensive review is available on the biocompatible design of these neural interfaces24.The most prominent strategies include surrounding the electrode with compatible coatings such as polyelthyleneglycol (PEG), polylactic-co-glycolic acids (PLGA)25, or enhancing the electrode with conductive polymers such as poly(ethylene dioxythiophene) (PEDOT), and polypyrrole (PPy)26,27,28,29,30,31. Bioactive coatings have also been employed to provide cues for neural tissue growth using ligands derived from extracellular matrices including collagens, fibronectins, and hyaluronic acids32,33,34,35,36,37. The bioactivity of these coatings has been further explored with growth factor release systems to emulate natural cell secretions30,38,39,40,41,42,43,44,45,46,47,48,49,50. Simultaneously, some research groups have opted to remodel the electrode geometry, flexibility, and composition to decrease the mechanical mismatch between device and tissue51,52,53,54,55,56,57. Altogether, these strategies have lead to many promising improvements in next-generation neural interfacial devices, however the long-term compatibility is an on-going issue and progress may be hampered by complex and time-consuming in vivo models.
Animal model-based approaches can limit the experimental throughput and increase costs of testing electrode biocompatibility. In vitro approaches using conventional cell culture techniques offer a more cost-effective alternative but fail to recapitulate much of the complexity of the interaction between device and tissue58. In particular, testing of surface coatings using 2D cell culture limits the modeling of electrode geometry and the influence of mechanical mismatch and micromotion thought to contribute to generating a host response contributing to device failure59,60.
To overcome problems associated with 2D cell culture, hydrogel cultures of neural cells have been developed for a wide variety of applications, pharmacological studies61, to direct neural cell differentiation62, to understand disease pathways63,64, or layered in co-culture with other cell types to model cell migration, neuroprotection, or to model tissue microenvironments61. Hydrogels are readily formed at different sizes and geometries can incorporate numerous types of primary or immortalized cell cultures, and are highly amenable to analysis by commonly used techniques such as confocal fluorescence microscopy. To create a model of the glial scarring process, we have recently developed and characterised a hyaluronic-acid based 3D hydrogel system for high-throughput testing of the glial response to implanted electrodes (Figure 1)65. This system has several distinct advantages: 1) primary glial cells (microglia, astrocytes, and oligodendrocytes) are encapsulated in a 3D matrix composed of polymers of hyaluronic acid, which is an endogenous extracellular matrix component; 2) the matrix stiffness can be 'tuned' to recreate the mechanical properties of brain or spinal cord tissue; and 3) cells can be encapsulated in the matrix in a rapid bench-top approach using photopolymerization with green light, limiting toxicity during encapsulation. This system enables key features of in vivo biocompatibility: devices are inserted into the hydrogel in a comparable manner to tissue, and the cellular response to implanted devices are monitored for a wide range of parameters65. These include mechanical mismatch between devices and the hydrogel coatings of various structures and electrical stimulation pulses. This system also includes oligodendrocyte and related precursors, which are often present and recruited in glial scars. Their damage, death, and phagocytosis by microglia are highly indicative of inflammatory injury and as a model reduced scarring or recovery, they have the capacity to demonstrate re-myelination of neurons66.
Herein we describe a method for synthesis and formation of hybrid hyaluronic acid hydrogels combined with commercially available basement membrane formulations to improve cell incorporation. Further, we will demonstrate the incorporation of primary cultured glial cells (microglia, astrocytes, and oligodendrocytes) and analysis of culture growth using immunocytochemistry and confocal microscopy.
Protocol
The protocol for the brain tissue extraction from day 1 Sprague Dawley rat pups, euthanized by decapitation, was approved by the Animal Care and Use Committee at the University of Alberta.
1. Microglia and Astrocyte Isolations67,68
NOTE: All media for isolation and cell culture is preheated to 37 °C in a water bath. Hank's balanced salt solution (HBSS) has 1% penicillin-streptomycin (PS). All Dulbecco's modified Eagle's media with Ham's F12 nutrient mixture (DMEM/F12) is supplemented with 10% fetal bovine serum (FBS) and 1% penicillin-streptomycin (PS). Only mixtures with trypsin lack FBS. All materials in this protocol are sterile (filter, alcohol, purchased, and autoclaved) and every step subsequent to step number 1.8 is performed in a biosafety cabinet with aseptic technique.
- Per brain, pre-heat 15 mL of HBSS and 12.5 mL DMEM/F12 to 37 °C in 50 mL conical centrifuge tubes, and approximately 2 mL of 0.25% trypsin with 1 mM ethylenediaminetetraacetic acid (EDTA) in a 15 mL conical centrifuge tube. Heat an additional 25 mL of DMEM/F12. Warm trypsin about 10 min prior to use to prevent loss of enzymatic activity.
- Pour enough warm HBSS into sterile, 6 and 10 cm culture dishes to cover the surface. Prepare one 10 cm dish for every two brains plus an additional 6 cm dish.
- Place up to 2 brains in each 10 cm2 dish (Figure 2A).
- Under a dissection microscope, use curved and straight forceps to separate the cortices along the midline, and the cerebellum with brain stem. This yields three sections of tissue per brain (Figure 2B).
- Peel the thin, semi-transparent, layer (meninges) of tissue containing blood vessels from each section of tissue and discard, transferring the remaining tissue into a separate culture dish. Grasp the thin layers exposed by the tissue cuts, with the forceps, and carefully pull until major portions are 'hanging off' the brain tissue. Finally remove this portion by gently tugging. See Figure 2C-2E for representative images of before, during, and after this step.
- In a biosafety cabinet, remove the remaining HBSS from the culture dish, carefully retaining the tissue. Macerate the tissue with a scalpel and transfer tissue to the pre-warmed 15 mL tube with trypsin.
- Incubate this tube for 25 min in a 37 °C bath to digest the tissue enzymatically.
- Centrifuge the tube down at 500 x g for 2 min at room temperature, and pour out the supernatant.
- Using a 10 mL serological pipette, add 10 mL of warm DMEM/F12 to inactivate the trypsin and triturate the solution 5 times to break up the tissue.
- Centrifuge the tube down at 500 x g for 2 min at room temperature, and pour out the supernatant.
- Using a 10 mL serological pipette, re-suspend pellet into 10 mL of warm DMEM/F12 and transfer the solution to a 50 mL Conical centrifuge tube.
- Using a 10 mL syringe with an 18 gauge needle, triturate the solution 3 times.
- Distribute this solution into increments of 12.5 mL of warm DMEM/F12 per brain.
- Pipette 1 mL increments of the triturated solution into a 12 well plate (1 per brain).
- Incubate at 37 °C and 5% CO2in a humidified cell culture incubator. Replace media after three days, and refresh media every subsequent three days. After 2 weeks, the cells can be collected for experiments.
2. Macromer Synthesis69
- Weigh 375 mg of hyaluronic acid (HA) salt in a 50 mL conical centrifuge tube and add 37.5 mL of distilled water.
- Flush the solution with a vortexer for 1 min and sonicate the mixture until it appears homogenous and loses its gel-like viscosity. This process may take 6 h.
- Transfer solution to a 100 mL beaker with a magnetic stir bar (38 mm), and stir vigorously at room temperature.
- Carefully titrate 5.0 M NaOH into the mixing HA and measure with pH paper until the pH stabilizes between 8 and 12.
- Collect 3 mL of fresh methacrylic anhydride (MA, 94%), blanketed by nitrogen during storage.
NOTE: MA is exposed to the atmosphere for long durations (days), it will lose its reactivity.
Caution: All use of MA should be done in a fume hood. - Add 200 µL of MA into the mixing HA solution. The reaction lowers the pH of the solution below 8.
- Repeat steps 2.4 to 2.6 until the 3 mL of MA have been added to the solution. This process may take up to 1-2 h.
- Allow the reaction to continue overnight at 4 °C, without mixing.
- Transfer solution to 400 mL of cold ethanol (EtOH) in a 500 mL beaker and allow precipitation for 24 – 72 h at 4 °C.56
- Carefully decant the major portion of the solution into a separate beaker for disposal, and collect the precipitate in a 50 mL conical centrifuge tube. This may be distributed in 2-4 tubes, if necessary.
- Centrifuge the tubes at 200 x g in a centrifuge for 2 min at room temperature and dispose of the supernatant.
- Top up the tube with cold EtOH, mix the solution with a vortexer for 1 min, and repeat step 2.10-2.11.
- Add 25 mL of sterile deionized water, mix the solution with a vortexer for 1 min, sonicate (>20 kHz) for 1 h, and incubate at 4 °C overnight.
- Place the solution in a -80 °C freezer for 15 min or until frozen or flash freeze in liquid nitrogen.
CAUTION: When using liquid nitrogen, use protective gloves, a face shield, and an apron. - Freeze dry the tube (below 0.1 mBar and -30 °C) for 24 – 48 h until a snow-like powder is produced.
- At this point, test the sample for purity via 1H nuclear magnetic resonance, where the amount of methacrylate protons (peaks at 6.1 and 5.6 ppm), relative to methyl protons (1.9 ppm), on the HA backbone can be confirmed. A minimum ratio of 95% methacrylate to methyl protons is acceptable.69
3. Gel Formation and 3D Encapsulation65
- Coverslip and Mold Preparation
- Make a 50 mL distilled water solution of 2% 3-(Trimethoxysilyl)propyl methacrylate in a 50 mL conical centrifuge tube.
- Place 18 mm glass coverslips in solution and rock for 1 h at room temperature. Up to 50 coverslips can be added at once.
- Rinse coverslips by serially dipping each in 3 beakers of 100 mL deionized water.
- Dry the coverslips in a vacuum at 40 °C overnight with a desiccator and oven.
- Quickly immerse individual coverslips in 70% EtOH with forceps.
- Without drying, drop each coverslip into a well of a 12 well plate.
- Wash and aspirate each well with sterile deionized water (1 mL per well).
- Add 1 mL of sterile 2 µg /mL poly-L-lysine (PLL) to each well and incubate for >2 h.
- Aspirate PLL solution and allow the coverslips to air dry.
- Immerse polydimethyl siloxane (PDMS) molds (see Figure 1) in 70% EtOH, and place one upon the center of each coverslip, and allow to air dry and create a seal between the mold and glass.
- Prepare PDMS molds by casting PDMS premix reagents in a 10:1 ratio in a flat-bottomed polystyrene dish. The quantity of PDMS prepared is selected to yield a sheet approximately 1 mm thick. To form wells in the PDMS, cut the sheets with a circle punch with an inner diameter of 10.
- Cell Preparation
NOTE: All steps in the protocol are performed with sterile technique in a biosafety cabinet. The phosphate buffered saline (PBS) solution is adjusted to pH 7.4 for all steps.- 24 h prior to cell encapsulation in HAMA, refresh the medium for the 2 week primary cultures (from Step 1.15) in the 12 well plates. Add 1 mL of DMEM/F12 (10% FBS and 1% PS) medium per well.
- For each 12-well plate of cells prepare 12.5 mL of dilute trypsin (0.25% trypsin-EDTA diluted 30% with DMEM/F12 media), and 25 mL DMEM/F12 (10% FBS and 1% PS) and warm in at 37 °C in a water bath.
- Collect conditioned medium from plates using a 10 mL serological pipette (> 0.5 mL per well), thoroughly aspirate remaining liquid, and replace with 1 mL per well of dilute trypsin (0.075% trypsin-EDTA) for 20-30 min in an incubator (37 °C, 5% CO2) until the confluent cell layer detaches from the plate.
- Recover suspended cell material with a 1 mL pipette (appears as a single floating piece) and collect in a 15 mL conical centrifuge tube. Dilute with an equivalent volume of DMEM/F12 (10% FBS and 1% PS), and centrifuge at 200 x g for 2 min at room temperature, discarding the supernatant.
- Resuspend the cell pellet in 10 mL warm DMEM/F12 (10% FBS and 1% PS), triturate the cells 5 times with a 10 mL pipette, and centrifuge at 200 x g for 2 min.
- Decant the supernatant, re-suspend the cells in 5 mL warm DMEM/F12, and transfer the cells to a 50 mL conical centrifuge tube.
- Using a 10 mL syringe with a 18 gauge needle, triturate the cell solution 3 times, and filter the suspension through a 40 µm cell sieve into a new conical centrifuge tube.
- Collect 10 µL of cell suspension, dilute 1:100 in warm DMEM/F12, and count cells with an automated cell counter
- Incubate in a 37 °C water bath until encapsulation step.
- Cell Encapsulation
NOTE: All steps in the protocol are performed with sterile technique in a biosafety cabinet.- For each 12-well plate of 3D hydrogels warm 12.5 mL of DMEM/F12 and 12.5 mL of the conditioned DMEM/F12 media collected during cell preparation.
- Weigh the quantity of methacrylated hyaluronic acid (HAMA) required for a final concentration of 0.5% wt/vol. Dissolve this HAMA in sterile filtered PBS at a concentration of (2% wt/vol); a concentration 4 times higher than the final concentration is utilized to accommodate the addition of cells and other reagents. One 12 well plate of hydrogels requires ~350 µL PBS with 7 mg HAMA.
- Sonicate (>20 kHz) solutions until HAMA is fully dissolved for 60 min.
- Prepare individual 10% solutions of both triethanolamine (TEA) and 1-vinyl-2-pyrrolidinone (NVP), and a 1 mM solution of Eosin Y (EY) in 1 mL aliquots of sterile PBS pH 7.4.
- For one 12 well plate, make a final 1.4 mL mixture of 1 x 107 cells, 0.5% wt/vol HAMA (350 µL of 2% wt/vol), 0.1% TEA (14 µL of 10%), 0.1% NVP (14 µL of 10%), and 0.01 mM EY (14 µL of 1 mM), and 20% basal lamina mixture (280 µL of stock). The remaining volume is PBS.
- Gently mix the solution and pipette 100 µL into each PDMS mold with a 1 mL tip.
- Expose samples to a high intensity green LED light (~520 nm, 60 mW, in an enclosed 20 cm x 20 cm x 20 cm box) for 5 min at room temperature.
- Add 1 mL per well of warm conditioned DMEM/F12 to a 12 well plate.
- Grasp each coverslip between thumb and forefinger, slowly peel the PDMS mold off with curved tweezers being careful not to displace the gel, and place it into a well with conditioned medium.
- Add an additional 1 mL warm DMEM/F12 to each well.
- Incubate plates at 37 °C with 5% CO2 for 2 weeks, refreshing cell media every week by pipetting 1 mL of media from each wells and replacing with 1 mL DMEM/F12.
4. Microscopy
- Immunocytochemistry
NOTE: An inverted confocal microscope was used to image these samples and ImageJ was used to process and display the images. Microglia, astrocytes, oligodendrocytes, and nuclei, will be labeled after 3 weeks in the HAMA culture.- Preheat 10% PBS-buffered formalin (pH 7.0) in a 37 °C bath.
Caution: Although formalin can be used outside a fume hood, this material is reactive with tissue, and should be handle with care and gloves always. It should be dispose separately. - Carefully aspirate media from the plates and pipette 1 mL of 10% formalin to each well.
- Incubate the plates at 37 °C in a 5% CO2 for 20 min.
- Aspirate liquid from each plate, pipette 2 mL of PBS (pH 7.4) into each well, and incubate for 15 min.
- Repeat step 4.1.4 twice.
- Aspirate liquid from the plates, add 1 mL per well of PBS with 10% normal horse serum (NHS) and 0.5% triton X-100, and gently rock at minimum speed) for 1 h.
- Repeat step 4.1.4 three times.
- Aspirate liquid and add 1 mL per well of the following mixture in PBS: rabbit ionized calcium-binding adapter molecule (Iba1) 1:1,000, chicken glial fibrillary protein (GFAP) 1:5,000, mouse 2',3'-cyclic-nucelotide 3'-phophodiesterase (CNPase) 1:1,000, 1% NHS, and 0.1% cell perforating surfactant. Incubate at 4 °C overnight, rocking the plate as gently as possible.
- Repeat step 4.1.4 three times with 1 h incubations in between.
- Aspirate liquid and add 1 mL per well of the following mixture in PBS: 647 nm excited donkey anti-rabbit antibody 1:200, 546 nm excited goat anti-chicken antibody 1:200, 488 nm excited donkey anti-mouse antibody 1:200, 1% NHS, 1:1,000 blue nuclear stain. Incubate at room temperature for 1 h, gently rocking the plate.
- Repeat step 4 three times with 1 h incubations in between.
- To preserve the hydrogels, aspirate buffer, and immerse them in 0.5 mL of cell mounting medium for 1 min. Pipette out the mounting medium and gently place a coverslip on top of the gel, creating a layer between the glass. Add a small amount of mounting medium to the uncovered side of the coverslips to prevent drying of the hydrogel.
NOTE: For troubleshooting purposes, gels can be imaged at step 4.1.11. to see proper labeling before further processing.
- Preheat 10% PBS-buffered formalin (pH 7.0) in a 37 °C bath.
- Scanning Electron Microscopy (SEM)
NOTE: To visualize the hydrogel, a scanning electron microscope was used to image these samples.- Pre-heat a fixative mixture of 2.5% glutaraldehyde and 2% paraformaldehyde in PBS (0.1 M) in a 37 °C water bath.
Caution: These reagents are highly reactive with tissues, and should be handled with care and gloves. They should be handled in a fume hood as much as possible and disposed separately. - Aspirate medium from the cell culture plates and add 1 mL of the pre-heated fixative mixture.
- Incubate the plates at 37 °C in a 5% CO2 for 20 min.
- Place the plates in 4 °C (fridge/cooler) overnight.
- Aspirate the fixative solution from the wells and add 1 mL of PBS.
- Aspirate liquid from each plate, add 2 mL of PBS into each well, and incubate for 15 min.
- Repeat step 4.2.6 two more times.
- Aspirate the PBS and add 1 mL of 1% osmium tetraoxide buffered in PBS to each well.
NOTE: Until samples are dried, this step and all subsequent steps are performed in a fume hood. - Allow fixation to occur for 30 min. Osmium vapor from the liquid can fix other samples in the same plate; therefore, partition samples accordingly before this step.
CAUTION: Osmium tetraoxide is highly reactive with tissue and volatile. It is dangerous and should be handled in the fume hood with gloves. It and any immediate washed should be disposed separately. - Repeat step 4.2.6 three times.
- Aspirate the liquid and add the dehydrating mixtures in the list below, in order, and incubate (room temperature) for the indicated time. Start by gradually adding ethanol (EtOH) followed then hexamethyldisilazane (HMDS). See Table 2 for each increment.
NOTE: When adding HMDS, the coverslips can strongly adhere to the plastic and become very difficult to remove. This can be prevented by placing a small plastic platform under the glass slip after the first application of HMDS. After the second EtOH incubation, the sample can be taken out and quenched in liquid nitrogen for few seconds to freeze fracture the sample. - Carefully remove the dried samples and mount the samples on scanning electron microscope stubs with double sided conductive tape.
- Sputter coat the samples with a minimum amount of gold. Place the samples in a holder and operate the machine for 2 min at 15 mA.
- Pre-heat a fixative mixture of 2.5% glutaraldehyde and 2% paraformaldehyde in PBS (0.1 M) in a 37 °C water bath.
Representative Results
To model the neural tissue host response and the glial scar in a high throughput, in vitro system requires a 3D cell scaffold with a matrix material that is biocompatible, does not incur a cytotoxic event during in situ formation, and is modifiable with bioactive components to guide benevolent response. To this end, we have created a 3D cell scaffold system based on hyaluronic acid, and encapsulated a primary mixed glial cell population to study cell-cell interactions and glial bioreactivity. A brief visual study of cells and scaffold was performed. Morphology of cell subpopulations, including oligodendrocytes (CNPase in green), microglia (Iba1 in red), and astrocytes (GFAP in yellow), are shown by confocal immunofluorescent microscopy (Figure 3Ai-ii -Ci-ii). Combined scaffold and cell morphology are also shown by SEM (Figure 3Aiii-iv – Ciii-iv). In these images, both low (i and iii), and high (ii and iv) resolution representative images are shown. A 2D PLL coverslip coating (Figure 3A) was compared to a 3D 0.5% w/v HAMA scaffold (Figure 3B), and a mixture of 20% v/v basal lamina mixture in 0.5% w/v HAMA (Figure 3C). This was done to demonstrate the morphological differences in 2D and 3D cell culture, as well as the effect of introducing a biological lamina to the scaffold. A 3D reconstruction is also available in the Supplemental Information.
Cell morphology of each cell type changed with the introduction of a 3D platform. In 2D adherent culture (PLL-coated glass coverslips), oligodendrocytes appeared typically round and formed small networks usually connecting with lightly labeled GFAP processes. Microglia were smaller than other cells and had small branching processes that did not extend far from their cell bodies (<20 µm). Astrocytes were more diversely shaped, some having a radial morphology with smaller cell bodies and extensive thin, branching processes, while others were fibrous with a flattened appearance and few broad processes that coat the surface. In the 3D HAMA culture, oligodendrocytes appeared in clusters having branched off into smaller spheres. Microglia were round with few processes, and astrocytes were either rounded or branched off radially into networks from a single cluster. Generally, these cells appeared to have outgrown from single points or remain in areas of the HAMA without making contact with other cells, which is unlike the 2D culture and is suggestive of limited mobility through the matrix. When 20% v/v of a basal lamina mixture was added to the photo-polymerization mixture, all glial subtypes integrated with the 3D platform, and formed more extensive, interactive branched structures. Oligodendrocytes extended bulbous processes radially similar to the non-basal lamina mixture morphologies, but additionally appeared to extend processes along the thin interconnected networks of astrocyte processes branching throughout the scaffold. Microglia were dispersed among these cell networks, having spread out with thin processes more consistent with their morphology in tissue. Overall, the morphologies suggest all three types of glia integrated into the HAMA-basal lamina mixture freely, potentially through either increased adherence to the mixed substrate or through increased ability to remodel the scaffold.
When observed by SEM, cell morphology was seen to corroborate the confocal micrographs. The HAMA matrix appeared as a smooth surface with 10-50 µm pores seen faintly beneath the surface. Cells were adherent to the surface of the HAMA and were also faintly visible in layers beneath. These glia did not form comparable morphologies to those noted in 2D culture, however with the addition of a basal lamina mixture, extensive networking was observed. Thick astrocyte fibers, with smaller clusters of microglia were observed both on the HAMA-basal lamina mixture surface and extending through the matrix in a seamless manner. This can be seen faintly as the matrix folded around the cells, suggesting that the cells may be altering the scaffold to formed into their desired conformations.
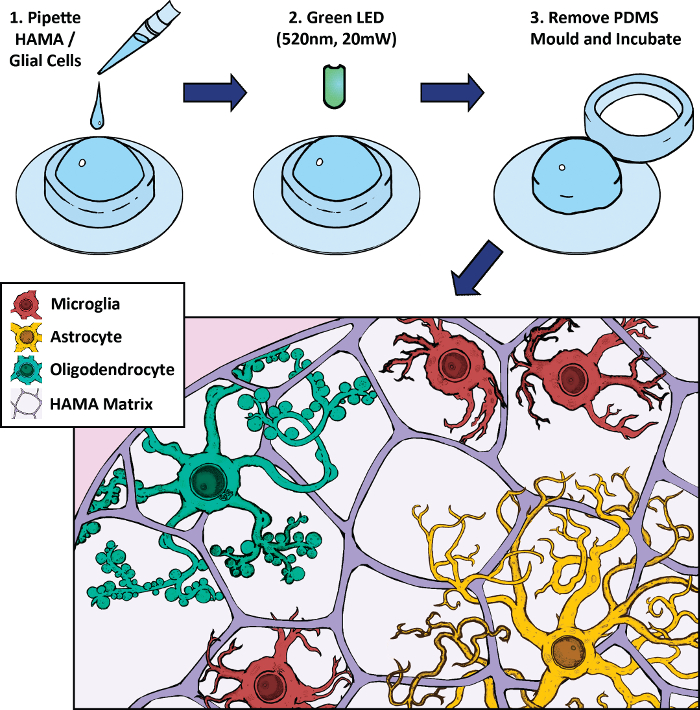
Figure 1: Schematic of methacrylated hyaluronic acid (HAMA)-based 3D cell culture and photopolymerization protocol. Glial cells (2 week primary rat brain culture) are mixed with HAMA and pipetted into a polydimethylsiloxane (PDMS) mold on a glass coverslip. This mixture is exposed to a high intensity green LED light to polymerize the hydrogel for 5 min. The mold is removed and the gel is incubated in media. Representative cells include microglia (red), astrocytes (yellow), and oligodendrocytes (green). Please click here to view a larger version of this figure.
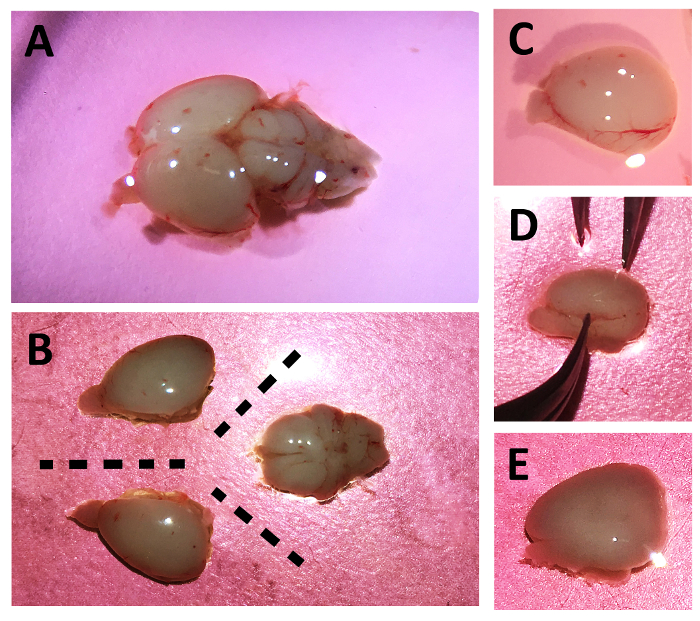
Figure 2: Dissection of day 1 Sprague Dawley rat pup brain. Images of whole brain (A), dissection of cortices and cerebellum (B), a single cortex with meninges (C), peeling of the meninges with forceps (D), and the meninge-free cortex (E). Please click here to view a larger version of this figure.
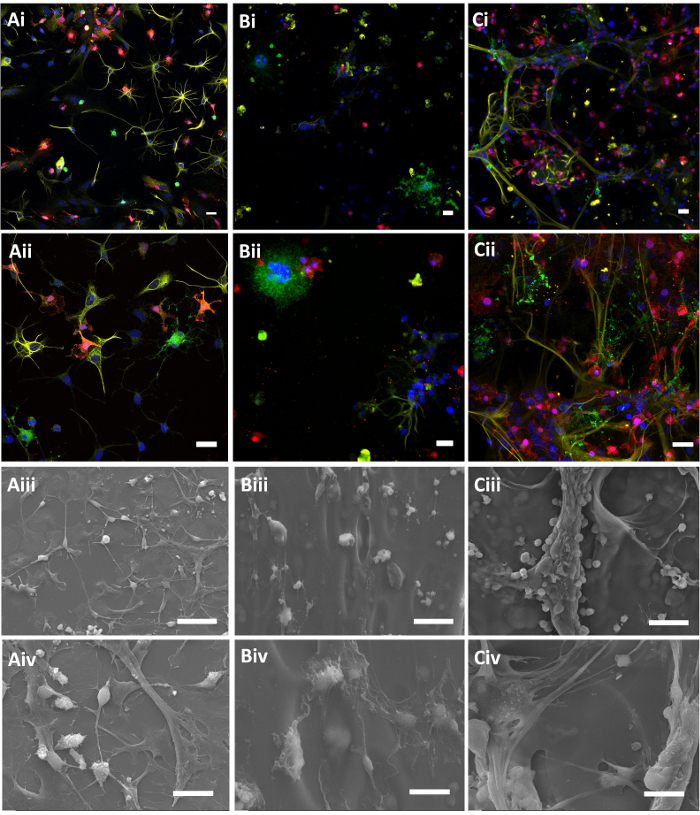
Figure 3: Immunofluorescent confocal (i-ii) and scanning electron (iii-iv) micrographs of mixed glia cell populations on 2D PLL coatings (A), HAMA (B), and HAMA-basal lamina mixture (C). Lower (i, iii) and higher (ii, iv) magnification images are shown. Labels include Iba1 in red for microglia, GFAP in yellow for astrocytes, CNPase in green for oligodendrocytes, and a Hoechst nuclear stain. Scale bars = 25 µm for confocal images and 50 (iii) and 20 µm (iv) for scanning electron micrographs. Hydrogels were 0.5% w/v, and basal lamina mixture was 20% by volume. Please click here to view a larger version of this figure.
Supplemental Figure 1: 3D immunofluorescent micrographic reconstruction of HAMA-basal lamina mixture with mixed glia cell populations HAMA-basal lamina mixture. Labels include Iba1 in red for microglia, GFAP in yellow for astrocytes, CNPase in green for oligodendrocytes, and a Hoechst nuclear stain. Hydrogels were 0.5% w/v, and Getrex was 20% by volume. Please click here to download this file.
| HAMA | Basal Lamina | NVP | TEA | EY | |
| Working concentration | 0.5% w/v | 20% v/v | 0.10% | 0.10% | .01 mM |
| 12-well Plate | 2% w/v | 100% w/v | 10% v/v | 10% v/v | 1 mM |
| 1.4 mL Total | 350 µL | 280 µL | 14 µL | 14 µL | 14 µL |
Table 1: Example of Cell Encapsulation.
| Component | Content | Incubation Time |
| EtOH | 30% | 30 min |
| EtOH | 50% | 30 min |
| EtOH | 70% | 30 min |
| EtOH | 90% | 20 min |
| EtOH | 100% | 20 min |
| EtOH | 100% | 20 min |
| EtOH:HMDS | 75:25 | 20 min |
| EtOH:HMDS | 50:50:00 | 20 min |
| EtOH:HMDS | 25:75 | 20 min |
| HMDS | 100% | 20 min |
| HMDS | 100% | Overnight |
Table 2: Gradual Increments of EtOH and HMDS in Dehydration.
Discussion
Towards the goal of generating a 3D culture system to model glial bioreactivity and the glial scarring process, we have developed a system that can support primary cultured microglia, astrocytes and oligodendrocytes and enables robust characterization of cell morphology and cell-cell interactions. From the micrographs shown, the morphology of each cell type was distinctly different with 2D, 3D-HAMA, and 3D HAMA-Basal lamina mixture platforms. In the 2D system, morphology was distinctly biased along the plane of the surface, but when compared to the 3D HAMA, microglia and astrocytes were generally smaller inside the matrix, with the exception of oligodendrocyte clusters. In the scanning electron micrographs, cells were generally more round shaped on the HAMA. This may be largely due to the cells being encapsulated, with limited capacity to modify the matrix for locomotion and free movement and growth of processes. Cell outgrowth in the HAMA was typically radial for both astrocytes and oligodendrocytes. While these cells extended processes, their organization suggest that their movement throughout the matrix and ability to interact with other cells was limited. In addition, previous work has shown freeze-fractured HAMA to be porous without interconnected networking, which may explain this lack of outgrowth65,70.It may be possible that some of these cells did not survive the photopolymerization, remaining preserved in the matrix, however viability was systematically assessed in our previous work, and remained 80% (0.5% w/v) over the course of a week65.
To improve the cell interaction with the matrix in 3D culture, we introduced a basal lamina mixture as a biocompatible and modifiable matrix component. Cell integration with the matrix was found to be markedly improved, with all cell types showing extensive processes compared to both the 2D culture and 3D HAMA. The basal lamina mixture is comprised of a variety basal lamina components and is primarily formulated to support glial differentiation of neural stem cell precursors71. It was not unexpected that the glial cells responded to the basal lamina mixture, and subsequently the surrounding 3D matrix, but the extent of integration suggests a healthy tissue-like environment. Comparing HAMA to HAMA-basal lamina mixture, the hydrogel morphology does not appear distinctly different, therefore cells may be more actively remodelling it. Considering the simplicity of adding this bioactive component, it is not unimaginable to use different laminas and/or protein cues to favour culture conditions for neurons. In a potential future direction, this system could be used to differentiate neurons from neural progenitors in conjunction with mixed glia for a variety of impactful myelination or neuro-inflammatory injury models.
In the event of serious and damaging CNS injury or biomaterial rejection, the resident tissue initiates inflammatory reactions capable of exacerbating neurodegeneration and demyelination. This typically results in the formation of a glial scar to partition the site of injury, which prevents regeneration. In this study, the methods were detailed for a methacrylated photo-polymerized hyaluronic acid-based 3D scaffold with the intention of modelling the initial cellular response behind glial scar formation; microglial reactivity, astrocyte recruitment and hypertrophy, and oligodendrocyte injury and withdrawal. A 3D cell scaffold was designed to incorporate all three glial cell types, and was further modified with a multi-component basal lamina mixture. It was determined, by confocal and scanning electron microscopy imaging, that that cells were encapsulated in the HAMA and were better able to integrate when the basal lamina mixture was introduced. These results could lead to several novel applications. HAMA alone may serve to create co-culture systems where the intention is to limit cell to cell interaction or study distil drug delivery using the gel as a drug loaded diffusion limited matrix. Hydrogels incorporating the basal lamina mixture into the HAMA matrix allow for a more consistent tissue-like environment capable of modeling acute inflammation, gliosis, or glial scarring, and enables further high-throughput characterization of glial bioreactivity to implants, drug and device development, and other strategies to reduce and recover inflammatory injury.
Declarações
The authors have nothing to disclose.
Acknowledgements
The authors are grateful for financial support from NSERC, CFI, AIHS, Alberta Health Services, and the Davey Endowment for Brain Research.
Materials
| 1. Materials for HAMA synthesis and photopolymerization | |||
| Hyaluronic acid (HA) | Sigma-Aldrich | 53747-10G | Streptococcus equi, MW: 1.5 – 1.8 X 10^6 |
| Methacrylic anhydride (MA) | Sigma-Aldrich | 275585-100ML | |
| Sodium hydroxide (NaOH) | Sigma-Aldrich | 221465-25G | |
| Ethanol (EtOH) | Commerical Alcohols Inc. | Anhydrous | |
| Phosphate buffered saline (pH 7.4) tablets | Fisher Scientific | 18912014 | |
| Triethanolamine (TEA) | Sigma-Aldrich | 90279-100ML | |
| 1-Vinyl-2pyrrolidinone (NVP) | Sigma-Aldrich | V3409-5G | |
| EosinY (EY) | Sigma-Aldrich | E6003-25G | |
| Polydimethylsiloxane (PDMS) Sylgard 184 Silicone Elastomer Kit | Dow Corning | ||
| 3-(Trimethoxysilyl)propyl methacrylate | Sigma-Aldrich | 440159-100ML | |
| Beaker (100 mL) | Corning | 1000-100 | |
| Beaker (500 mL) | Corning | 1000-600 | |
| pH paper (Labstick) | Sigma-Aldrich | 9580 | |
| Name | Company | Catalog Number | Comments |
| 2. Materials for glial cell isolation and cell culture | |||
| P1-2 Sprague Dawley rat pups | Charles River | CD Sprague Dawley rat strain code 001 | |
| Dissector scissors – slim blades (small) | Fine Science Tools | 14081-09 | |
| Surgical scissors – Toughcut (large) | Fine Science Tools | 14130-17 | |
| Fine forceps (Dumont #5) | Fine Science Tools | 11521-10 | |
| Curved fine forceps (Dumont #7) | Fine Science Tools | 11271-30 | |
| Hank's balanced salt solution (HBSS) | Gibco | 14170-112 | |
| Dulbecco's modified Eagle's medium and Ham's nutrient mixture F-12 (DMEM/F12) | Gibco | 11320-033 | |
| Penicillin-streptomycin (PS) | Gibco | 15140-122 | |
| Fetal bovine serum (FBS) | Gibco | 12483-020 | |
| 0.25% Trypsin-ethylenediaminetetraacetic acid (EDTA) | Gibco | 25200-072 | |
| Poly-L-lysine (PLL) | Sigma-Aldrich | P-6282 | |
| 50 mL conical centrifuge tube | Fisher Scientific | 05-539-13 | |
| 15 mL conical centrifuge tube | Fisher Scientific | 05-539-5 | |
| 12 well Tissue culture treated plates (Cellstar) | Greiner Bio-One | 665 108 | |
| 10 mL serological pipette | Fisher Scientific | 13-676-10F | |
| 25 mL serological pipette | Fisher Scientific | 12-676-10K | |
| Petri dish (60 mm X 15 mm) | Fisher Scientific | FB0875713A | |
| Petri dish (100 mm X 15 mm) | Fisher Scientific | FB0875712 | |
| Microscope Coverslip (18 mm) | Fisher Scientific | 12-545-100 18CIR | |
| Name | Company | Catalog Number | Comments |
| 3. Materials for microscopy (confocal and scanning electron microscopy) | |||
| Mouse monoclonal anti-CNPase | abcam | ab6319 | |
| Rabbit anti-Iba1 | Wako Laboratory Chemicals | 019-17741 | |
| Chicken anti-GFAP | abcam | ab4674 | |
| Hoechst 33342 | Fisher Scientific | 62249 | |
| Fluoromount-G | Fisher Scientific | 00-4958-02 | |
| Formalin | Sigma Aldrich | HT501128-4L | Buffered (10%) |
| Triton X-100 | Fisher Scientific | BP151-500 | |
| Horse Serum | Gibco | 16050-122 | |
| Paraformaldehyde | Electon Microscopy Sciences | 157-8 | Buffered (8%) |
| Guteraldehyde | Electon Microscopy Sciences | 16019 | Buffered (8%) |
| Osmium tetraoxide | Electon Microscopy Sciences | 19152 | Buffered (2%) |
| Hexamethyldilazane (HMDS) | Electon Microscopy Sciences | 16700 | |
| Ethanol (EtOH) | Electon Microscopy Sciences | 15055 | Anhydrous |
| Microscope Slide (25 X 75 X 1 mm) | VWR International | 48311-703 |
Referências
- Shih, J. J., Krusienski, D. J., Wolpaw, J. R. Brain-Computer Interfaces in Medicine. Mayo Clin. Proc. 87, 268-279 (2012).
- Kennedy, P. R., Bakay, R. A. Restoration of neural output from a paralyzed patient by a direct brain connection. Neuroreport. 9, 1707-1711 (1998).
- Kennedy, P. R., Bakay, R. A., Moore, M. M., Adams, K., Goldwaithe, J. Direct control of a computer from the human central nervous system. IEEE Trans. Rehabil. Eng. Publ. IEEE Eng. Med. Biol. Soc. 8, 198-202 (2000).
- Mayberg, H. S., et al. Deep Brain Stimulation for Treatment-Resistant Depression. Neuron. 45, 651-660 (2005).
- Dougherty, D. D., et al. A Randomized Sham-Controlled Trial of Deep Brain Stimulation of the Ventral Capsule/Ventral Striatum for Chronic Treatment-Resistant Depression. Biol. Psychiatry. 78, 240-248 (2015).
- Bamford, J. A., Marc Lebel, R., Parseyan, K., Mushahwar, V. K. The Fabrication, Implantation, and Stability of Intraspinal Microwire Arrays in the Spinal Cord of Cat and Rat. IEEE Trans. Neural Syst. Rehabil. Eng. Publ. IEEE Eng. Med. Biol. Soc. 25, 287-296 (2017).
- Holinski, B. J., et al. Intraspinal microstimulation produces over-ground walking in anesthetized cats. J. Neural Eng. 13, 056016 (2016).
- Toossi, A., Everaert, D. G., Azar, A., Dennison, C. R., Mushahwar, V. K. Mechanically Stable Intraspinal Microstimulation Implants for Human Translation. Ann. Biomed. Eng. 45, 681-694 (2017).
- Saigal, R., Renzi, C., Mushahwar, V. K. Intraspinal microstimulation generates functional movements after spinal-cord injury. IEEE Trans. Neural Syst. Rehabil. Eng. Publ. IEEE Eng. Med. Biol. Soc. 12, 430-440 (2004).
- Lau, B., Guevremont, L., Mushahwar, V. K. Strategies for generating prolonged functional standing using intramuscular stimulation or intraspinal microstimulation. IEEE Trans. Neural Syst. Rehabil. Eng. Publ. IEEE Eng. Med. Biol. Soc. 15, 273-285 (2007).
- Polikov, V. S., Tresco, P. A., Reichert, W. M. Response of brain tissue to chronically implanted neural electrodes. J. Neurosci. Methods. 148, 1-18 (2005).
- Sierra, A., et al. Surveillance, phagocytosis, and inflammation: how never-resting microglia influence adult hippocampal neurogenesis. Neural Plast. 2014, 610343 (2014).
- Holm, T. H., Draeby, D., Owens, T. Microglia are required for astroglial Toll-like receptor 4 response and for optimal TLR2 and TLR3 response. Glia. 60, 630-638 (2012).
- Gao, Z., et al. Reciprocal modulation between microglia and astrocyte in reactive gliosis following the CNS injury. Mol. Neurobiol. 48, 690-701 (2013).
- Jin, X., Yamashita, T. Microglia in central nervous system repair after injury. J. Biochem. (Tokyo). 159, 491-496 (2016).
- Griffith, R. W., Humphrey, D. R. Long-term gliosis around chronically implanted platinum electrodes in the Rhesus macaque motor cortex. Neurosci. Lett. 406, 81-86 (2006).
- Biran, R., Martin, D. C., Tresco, P. A. Neuronal cell loss accompanies the brain tissue response to chronically implanted silicon microelectrode arrays. Exp. Neurol. 195, 115-126 (2005).
- Fawcett, J. W., Asher, R. A. The glial scar and central nervous system repair. Brain Res. Bull. 49, 377-391 (1999).
- Szarowski, D. H., et al. Brain responses to micro-machined silicon devices. Brain Res. 983, 23-35 (2003).
- Park, D. -. W., et al. Graphene-based carbon-layered electrode array technology for neural imaging and optogenetic applications. Nat. Commun. 5, 5258 (2014).
- McAllister, J. P., et al. Biocompatibility of Penetrating Recording Electrode Arrays Implanted Chronically in the Feline Visual Cortex. Invest. Ophthalmol. Vis. Sci. 46, 1528 (2005).
- Schmid, C. D., et al. Differential gene expression in LPS/IFNgamma activated microglia and macrophages: in vitro versus in vivo. J. Neurochem. 109, 117-125 (2009).
- Chung, H., et al. In vivo Biocompatibility and Stability of Polyimide Microelectrode Array for Retinal Stimulation. Invest. Ophthalmol. Vis. Sci. 44, 5072 (2003).
- Aregueta-Robles, U. A., Woolley, A. J., Poole-Warren, L. A., Lovell, N. H., Green, R. A. Organic electrode coatings for next-generation neural interfaces. Front. Neuroengineering. 7, (2014).
- Drury, J. L., Mooney, D. J. Hydrogels for tissue engineering: scaffold design variables and applications. Biomaterials. 24, 4337-4351 (2003).
- Yang, J., et al. Ordered surfactant-templated poly(3,4-ethylenedioxythiophene) (PEDOT) conducting polymer on microfabricated neural probes. Acta Biomater. 1, 125-136 (2005).
- Ludwig, K. A., et al. Poly(3,4-ethylenedioxythiophene) (PEDOT) polymer coatings facilitate smaller neural recording electrodes. J. Neural Eng. 8, 014001 (2011).
- Kim, D. -. H., Abidian, M., Martin, D. C. Conducting polymers grown in hydrogel scaffolds coated on neural prosthetic devices. J. Biomed. Mater. Res. A. 71, 577-585 (2004).
- George, P. M., et al. Fabrication and biocompatibility of polypyrrole implants suitable for neural prosthetics. Biomaterials. 26, 3511-3519 (2005).
- Stauffer, W. R., Cui, X. T. Polypyrrole doped with 2 peptide sequences from laminin. Biomaterials. 27, 2405-2413 (2006).
- Green, R. A., Lovell, N. H., Wallace, G. G., Poole-Warren, L. A. Conducting polymers for neural interfaces: challenges in developing an effective long-term implant. Biomaterials. 29, 3393-3399 (2008).
- Gumbiner, B. M. Cell adhesion: the molecular basis of tissue architecture and morphogenesis. Cell. 84, 345-357 (1996).
- Chan, G., Mooney, D. J. New materials for tissue engineering: towards greater control over the biological response. Trends Biotechnol. 26, 382-392 (2008).
- Green, R. A., Lovell, N. H., Poole-Warren, L. A. Impact of co-incorporating laminin peptide dopants and neurotrophic growth factors on conducting polymer properties. Acta Biomater. 6, 63-71 (2010).
- Green, R. A., Lovell, N. H., Poole-Warren, L. A. Cell attachment functionality of bioactive conducting polymers for neural interfaces. Biomaterials. 30, 3637-3644 (2009).
- Koss, K. M., Unsworth, L. D. Neural tissue engineering: Bioresponsive nanoscaffolds using engineered self-assembling peptides. Acta Biomater. 44, 2-15 (2016).
- Koss, K. M., et al. Brain biocompatibility and microglia response towards engineered self-assembling (RADA)4 nanoscaffolds. Acta Biomater. 35, 127-137 (2016).
- Evans, A. J., et al. Promoting neurite outgrowth from spiral ganglion neuron explants using polypyrrole/BDNF-coated electrodes. J. Biomed. Mater. Res. A. 91, 241-250 (2009).
- Yu, X., Bellamkonda, R. V. Tissue-engineered scaffolds are effective alternatives to autografts for bridging peripheral nerve gaps. Tissue Eng. 9, 421-430 (2003).
- Houweling, D. A., Lankhorst, A. J., Gispen, W. H., Bär, P. R., Joosten, E. A. Collagen containing neurotrophin-3 (NT-3) attracts regrowing injured corticospinal axons in the adult rat spinal cord and promotes partial functional recovery. Exp. Neurol. 153, 49-59 (1998).
- Wells, M. R., et al. Gel matrix vehicles for growth factor application in nerve gap injuries repaired with tubes: a comparison of biomatrix, collagen, and methylcellulose. Exp. Neurol. 146, 395-402 (1997).
- Barras, F. M., Pasche, P., Bouche, N., Aebischer, P., Zurn, A. D. Glial cell line-derived neurotrophic factor released by synthetic guidance channels promotes facial nerve regeneration in the rat. J. Neurosci. Res. 70, 746-755 (2002).
- Fine, E. G., Decosterd, I., Papaloïzos, M., Zurn, A. D., Aebischer, P. GDNF and NGF released by synthetic guidance channels support sciatic nerve regeneration across a long gap. Eur. J. Neurosci. 15, 589-601 (2002).
- Burdick, J. A., Ward, M., Liang, E., Young, M. J., Langer, R. Stimulation of neurite outgrowth by neurotrophins delivered from degradable hydrogels. Biomaterials. 27, 452-459 (2006).
- Piantino, J., Burdick, J. A., Goldberg, D., Langer, R., Benowitz, L. I. An injectable, biodegradable hydrogel for trophic factor delivery enhances axonal rewiring and improves performance after spinal cord injury. Exp. Neurol. 201, 359-367 (2006).
- Jain, A., Kim, Y. -. T., McKeon, R. J., Bellamkonda, R. V. In situ gelling hydrogels for conformal repair of spinal cord defects, and local delivery of BDNF after spinal cord injury. Biomaterials. 27, 497-504 (2006).
- Taylor, S. J., Sakiyama-Elbert, S. E. Effect of controlled delivery of neurotrophin-3 from fibrin on spinal cord injury in a long term model. J. Control. Release Off. J. Control. Release Soc. 116, 204-210 (2006).
- Jhaveri, S. J., et al. Release of nerve growth factor from HEMA hydrogel-coated substrates and its effect on the differentiation of neural cells. Biomacromolecules. 10, 174-183 (2009).
- Lee, A. C., et al. Controlled release of nerve growth factor enhances sciatic nerve regeneration. Exp. Neurol. 184, 295-303 (2003).
- Matsumoto, K., et al. Neurite outgrowths of neurons with neurotrophin-coated carbon nanotubes. J. Biosci. Bioeng. 103, 216-220 (2007).
- Khaled, I., et al. A Flexible Base Electrode Array for Intraspinal Microstimulation. IEEE Trans. Biomed. Eng. 60, 2904 (2013).
- David-Pur, M., Bareket-Keren, L., Beit-Yaakov, G., Raz-Prag, D., Hanein, Y. All-carbon-nanotube flexible multi-electrode array for neuronal recording and stimulation. Biomed. Microdevices. 16, 43-53 (2014).
- Hsu, H. -. L., et al. Flexible UV-ozone-modified carbon nanotube electrodes for neuronal recording. Adv. Mater. Deerfield Beach Fla. 22, 2177-2181 (2010).
- Lacour, S. P., et al. Flexible and stretchable micro-electrodes for in vitro and in vivo neural interfaces. Med. Biol. Eng. Comput. 48, 945-954 (2010).
- Lin, C. -. M., Lee, Y. -. T., Yeh, S. -. R., Fang, W. Flexible carbon nanotubes electrode for neural recording. Biosens. Bioelectron. 24, 2791-2797 (2009).
- Richter, A., et al. A simple implantation method for flexible, multisite microelectrodes into rat brains. Front. Neuroengineering. 6, 6 (2013).
- Rousche, P. J., et al. Flexible polyimide-based intracortical electrode arrays with bioactive capability. IEEE Trans. Biomed. Eng. 48, 361-371 (2001).
- Polikov, V. S., Block, M. L., Fellous, J. -. M., Hong, J. -. S., Reichert, W. M. In vitro model of glial scarring around neuroelectrodes chronically implanted in the CNS. Biomaterials. 27, 5368-5376 (2006).
- Sohal, H. S., Clowry, G. J., Jackson, A., O’Neill, A., Baker, S. N. Mechanical Flexibility Reduces the Foreign Body Response to Long-Term Implanted Microelectrodes in Rabbit Cortex. PloS One. 11, e0165606 (2016).
- Moshayedi, P., et al. The relationship between glial cell mechanosensitivity and foreign body reactions in the central nervous system. Biomaterials. 35, 3919-3925 (2014).
- Hopkins, A. M., DeSimone, E., Chwalek, K., Kaplan, D. L. 3D in vitro modeling of the central nervous system. Prog. Neurobiol. 125, 1-25 (2015).
- Pöttler, M., Zierler, S., Kerschbaum, H. H. An artificial three-dimensional matrix promotes ramification in the microglial cell-line, BV-2. Neurosci. Lett. 410, 137-140 (2006).
- Weigelt, B., Ghajar, C. M., Bissell, M. J. The need for complex 3D culture models to unravel novel pathways and identify accurate biomarkers in breast cancer. Adv. Drug Deliv. Rev. , 42-51 (2014).
- Edmondson, R., Broglie, J. J., Adcock, A. F., Yang, L. Three-Dimensional Cell Culture Systems and Their Applications in Drug Discovery and Cell-Based Biosensors. Assay Drug Dev Technol. 12, 207-218 (2014).
- Jeffery, A. F., Churchward, M. A., Mushahwar, V. K., Todd, K. G., Elias, A. L. Hyaluronic acid-based 3D culture model for in vitro testing of electrode biocompatibility. Biomacromolecules. 15, 2157-2165 (2014).
- Fitch, M. T., Silver, J. CNS Injury, Glial Scars, and Inflammation. Exp. Neurol. 209, 294-301 (2008).
- Churchward, M. A., Todd, K. G. Statin treatment affects cytokine release and phagocytic activity in primary cultured microglia through two separable mechanisms. Mol. Brain. 7, 85 (2014).
- Lai, A. Y., Todd, K. G. Differential regulation of trophic and proinflammatory microglial effectors is dependent on severity of neuronal injury. Glia. 56, 259-270 (2008).
- Hachet, E., Van Den Berghe, H., Bayma, E., Block, M. R., Auzély-Velty, R. Design of biomimetic cell-interactive substrates using hyaluronic acid hydrogels with tunable mechanical properties. Biomacromolecules. 13, 1818-1827 (2012).
- Eslami, M., Javadi, G., Agdami, N., Shokrgozar, M. A. Expression of COLLAGEN 1 and ELASTIN Genes in Mitral Valvular Interstitial Cells within Microfiber Reinforced Hydrogel. Cell J (Yakhteh). , (2015).
- Shaltouki, A., Peng, J., Liu, Q., Rao, M. S., Zeng, X. Efficient Generation of Astrocytes from Human Pluripotent Stem Cells in Defined Conditions. STEM CELLS. 31, 941-952 (2013).

