Visualizing the Interaction Between the Qdot-labeled Protein and Site-specifically Modified λ DNA at the Single Molecule Level
Summary
Here, we present a protocol to study DNA-protein interactions by total internal reflection fluorescence microscopy (TIRFM) using a site-specifically modified λ DNA substrate and a Quantum-dot labeled protein.
Abstract
The fluorescence microscopy has made great contributions in dissecting the mechanisms of complex biological processes at the single molecule level. In single molecule assays for studying DNA-protein interactions, there are two important factors for consideration: the DNA substrate with enough length for easy observation and labeling a protein with a suitable fluorescent probe. 48.5 kb λ DNA is a good candidate for the DNA substrate. Quantum dots (Qdots), as a class of fluorescent probes, allow long-time observation (minutes to hours) and high-quality image acquisition. In this paper, we present a protocol to study DNA-protein interactions at the single-molecule level, which includes preparing a site-specifically modified λ DNA and labeling a target protein with streptavidin-coated Qdots. For a proof of concept, we choose ORC (origin recognition complex) in budding yeast as a protein of interest and visualize its interaction with an ARS (autonomously replicating sequence) using TIRFM. Compared with other fluorescent probes, Qdots have obvious advantages in single molecule studies due to its high stability against photobleaching, but it should be noted that this property limits its application in quantitative assays.
Introduction
Interactions between protein and DNA are essential to many complex biological processes, such as DNA replication, DNA repair, and transcription. Although conventional approaches have shed light on the properties of these processes, many key mechanisms are still unclear. Recently, with the rapidly developing single molecule techniques, some of the mechanisms have been addressed1,2,3.
The application of single-molecule fluorescence microscopy on visualizing protein-DNA interactions in real-time mainly depends on the development of fluorescence detection and fluorescent probes. For a single molecule study, it is important to label the protein of interest with a suitable fluorescent probe since fluorescence detection systems are mostly available commercially.
Fluorescent proteins are commonly used in molecular biology. However, the low fluorescent brightness and stability against photobleaching restrict its application in many single molecule assays. Quantum dots (Qdots) are tiny light-emitting nanoparticles4. Due to their unique optical properties, Qdots are 10 – 20 times brighter and several thousand times more stable than the widely used organic dyes5. Moreover, Qdots have a large Stokes shift (the difference between the position of excitation and emission peaks)5. Thus, Qdots can be used for long-time observation (minutes to hours) and acquisition of images with high signal-to-noise ratios, while they cannot be utilized in the quantitative assays.
To date, there are two approaches to label a target protein with Qdots site-specifically: labeling with the aid of Qdot-conjugated primary or secondary antibodies6,7,8; or labeling the target protein with Qdots directly, which is based on the strong interaction between biotin and streptavidin9,10,11,12,13. Streptavidin-coated Qdots are commercially available. In our recent study, site-specifically biotinylated proteins in budding yeast with high efficiency were purified by co-overexpression of BirA and Avi-tagged proteins in vivo10. By following and optimizing the single-molecule assays14,15,16,17, we observed the interactions between Qdot-labeled proteins and DNA at the single molecule level using TIRFM10.
Here, we choose the budding yeast origin recognition complex (ORC), which can specifically recognize and bind to the autonomously replicating sequence (ARS), as our protein of interest. The following protocol presents a step-by-step procedure of visualizing the interaction of Qdot-labeled ORC with ARS using TIRFM. The preparation of the site-specifically modified DNA substrate, the DNA biotinylation, the coverslip cleaning and functionalization, the flow-cell assembly, and the single-molecule imaging are described.
Protocol
1. Preparation of λ-ARS317 DNA substrate
- DNA substrate construction and packaging
- Digest native λ DNA using XhoI enzyme; amplify a 543 bp DNA fragment bearing ARS317 from the genomic DNA of budding yeast using primers containing 20 bp homologous sequences of upstream and downstream of XhoI enzyme site on lambda DNA. Add 100 ng of XhoI digested λ DNA and 10 ng of DNA fragment to 10 µL of homologous recombination reaction system, and incubate the reaction at 37 °C for 30 min.
- To package the recombination λ DNA, add 25 µL of lambda packaging extracts into the recombination product, and incubate the reaction at 30 °C for 90 min. Then, add an additional 25 µL of lambda packaging extracts into the reaction tube. Continue to incubate the reaction at 30 °C for 90 min.
- Add 500 µL of sterile dilution buffer (10 mM Tris-HCl (pH 8.3), 100 mM NaCl, 10 mM MgCl2) into the reaction system, and mix gently by turning the tube upside down several times. Add 25 µL of chloroform, mix gently and store at 4 °C.
- Add 100 µL of packaged phage and 100 µL of bacteria LE392MP (cultured at 0.8 – 1.0 using LB medium adding with 10 mM MgSO4) into a new tube. Incubate at 37 °C for 15 minutes.
- Add 200 µL of phage-bacterium mixture into 4 mL oftop agar (LB medium + 0.7% agar + 10 mM MgSO4, cooled to 48 °C). Mix immediately by turning the tube upside down several times and pour it onto a pre-warmed (37 °C) LB plate.
- Incubate the plate at 37 °C overnight and screen the λ-ARS317 plaque using PCR and sequencing.
- DNA substrate purification from liquid lysates11,18
- Pick one plaque into 200 µL of sterile ddH2O with 10 mM MgCl2 and 10 nM CaCl2. Mix with 200 µL of bacteria LE392MP (cultured overnight in LB medium), and incubate at 37 °C for 15 min.
- Add the phage-bacterium mixture into 100 mL of NZCYM (10 g/L NZ-amine, 5 g/L yeast extract, 5 g/L NaCl, 1 g/L Casamino Hydrolysate, and 2 g/L MgSO4·7H2O) medium and culture for 7 h at 37 °C. Add 250 µL of chloroform into the culture and shake for another 10 min.
- Transfer the culture into a 200 mL flask. Add 5.8 g of NaCl (1 M final concentration), stir to dissolve by hand, and incubate on ice for 30 min.
- Centrifuge at 12,000 x g for 10 min at 4 °C to remove the cell debris. Collect the supernatant into a 200 mL flask and add 10% PEG8000 (m/V). Stir to dissolve by magnetic stirring apparatus and incubate on ice for 30 min or longer.
- Centrifuge at 12,000 x g for 10 min at 4 °C to precipitate the bacteriophage. Remove the supernatant.
- Resuspend the precipitate in 2 mL of phage dilution buffer. Transfer it into a 15 mL tube and add 10 µL of RNase (final concentration is 20 µg/mL) and 40 µL of DNase (final concentration is 5 µg/mL). Incubate at 37 °C for 30 min.
- Add 2 mL of 0.3 M Tris-HCl (pH 9.0), 100 mM EDTA + 1.25% SDS, 15 µL proteinase K (final concentration is 10 µg/mL). Incubate at 65 °C for 10 min.
- Add 2 mL of pre-cooled potassium acetate (3 M, pH 4.8), and incubate the tube on ice for 10 min.
- Centrifuge at 8,000 x g for 10 min at 4 °C to remove the insoluble materials.
- Add 0.7x volume of isopropanol into the supernatant. Mix by turning the tube upside down several times, and incubate for 2 min at room temperature (RT).
- Centrifuge at 8,000 x g for 10 min at RT to precipitate DNA. Remove the supernatant.
- Wash the DNA once with 70% ethanol by centrifugation at 8,000 x g for 10 min. Remove the supernatant.
- Elute the DNA using 500 µL TE buffer (10 mM Tris-HCl, 1 mM EDTA, pH 8.0) gently, and transfer the DNA into 1.5 mL tube.
- Extract the DNA twice using phenol: chloroform by centrifugation at 8,000 g for 10 min.
- Precipitate with 0.7x volume isopropanol. Mix by turning the tube upside down gently, and centrifuge at 8,000 x g for 10 min.
- Wash once with 70% ethanol, resuspend in 200 µL of TE. Measure the DNA concentration, and store at -20 °C in 12.5 µg aliquots.
2. λ-ARS317 DNA Biotinylation
- To phosphorylate biotinylated oligonucleotides (5'-AGGTCGCCGCC-TEG-Biotin-3') complementary to the left end of native λ DNA, add 1 µL (100 µM) of oligonucleotides, 7.5 µL of ddH2O, 1 µL of 10x ligase buffer, and 0.5 µL of T4 PNK. Incubate the reaction at 37 °C for 3 h.
- To phosphorylate λ-ARS317, add 12.5 µg of λ-ARS317, 3 µL of 10x ligase buffer, 1 µL of T4 PNK, and ddH2O to total volume of 30 µL. Incubate the reaction at 37 °C for 3 h.
- To anneal oligonucleotides with λ-ARS317, add 0.5 µL of phosphorylated oligonucleotides, 195 µL of ddH2O, 25 µL of 10×ligase buffer into the phosphorylated λ-ARS317 tube. Mix gently by turning the tube upside down and incubate at 65 °C for 5 min. Turn off the block heater, and let the sample cool to below 33 °C in the block.
- To ligate oligonucleotides with λ-ARS317, add 1 µL of T4 DNA ligase and 0.63 µL of ATP (200 mM). Mix gently by turning the tube upside down and incubate at room temperature for 2 h or 4 °C overnight. Store at 4 °C for 1 month.
NOTE: To avoid breaking up the DNA, all the mixing steps should be done by gently turning the tube upside down, instead of mixing using pipettes.
3. Coverslip Cleaning and Functionalization
- Coverslip cleaning
- Place 20 coverslips into 4 staining jars (5 coverslips/jar), sonicate for 30 minutes in ethanol, and rinse the coverslips with ultrapure H2O 3 times.
- Sonicate for 30 minutes with 1 M potassium hydroxide (KOH), and rinse the coverslips with ultrapure H2O 3 times. Repeat the ethanol and KOH sonication once.
- Sonicate with acetone for 30 minutes, and then rinse with ultrapure H2O thoroughly (at least 3 times).
NOTE: Acetone must be removed thoroughly by rinsing with ultrapure H2O, because it may cause an explosion when organic solvent (such as acetone) mixed with the piranha solution mistakenly14. - Place the coverslips into piranha solution (3:1 mixture of H2SO4 and 30% H2O2) and incubate at 95 °C for 1 h.
NOTE: Piranha solution is highly energetic and erosive, so use with caution. When preparing piranha solution, add 50 mL of H2O2 first into a 500 mL glass beaker, and then add 150 mL of H2SO4 slowly into the beaker. - Rinse the coverslips with ultrapure H2O 5 times. Then wash each coverslip with ultrapure H2O thoroughly using 3 beakers filled with ultrapure H2O, and dry the coverslip using paper from the edge of the coverslip. Place the coverslip into the staining jar and dry the coverslip thoroughly in a 110 °C oven for 30 minutes.
- Rinse the coverslip with methanol and place the staining jar into the 110 °C oven again to dry the coverslip thoroughly.
NOTE: Drying the coverslip thoroughly is very important, because APTES will have lots of possible surface structures once it is in the presence of water19.
- Rinse the coverslip with methanol and place the staining jar into the 110 °C oven again to dry the coverslip thoroughly.
- Coverslip functionalization
- Add 70 mL of silane solution (93% methanol, 5% acetic, and 2% APTES) into each jar, screw the cap and leave the jar at room temperature overnight.
- Wash and dry the coverslips as described in step 3.1.5, except dry the coverslips thoroughly using nitrogen gas instead of placing them in the oven.
- Dissolve 150 mg of mPEG (methoxy-polyethylene glycol) and 6 mg of biotin-PEG (biotin-polyethylene glycol) into 1 mL of 0.1 M fresh-made NaHCO3 (pH 8.2) thoroughly. Then centrifuge at 17, 000 x g for 1 min to remove insoluble PEGs.
NOTE: Freshly made NaHCO3 is ready; there is no need to adjust its pH. - Place the silanized coverslips in boxes, and put two small coverslips on the top of two ends of the silanized coverslips. Pipette 100 µL of PEG solution on the center of the silanized coverslip, and place another silanized coverslip on the top.
- Add some ultrapure H2O in the box to keep it humid, and incubate the coverslips with PEG solution for at least 3 h in the dark. An overnight incubation can work as well.
- Separate the coverslip pairs and keep the functionalized surface face up, rinse the coverslips using ultrapure H2O extensively, and dry them with nitrogen gas.
- Mark the functionalized side of the coverslips on one of the corners using a marker pen, keep the functionalized side face up, place them in boxes and store the boxes in the vacuum desiccator for 1 month.
- To store the coverslips for a longer time, place one coverslip into a 50 mL tube drilled with one hole on its cap, then put the tube into a plastic bag, and seal the bag using a vacuum sealer. In this way, the coverslips can be stored at -20 °C about 3 months.
4. Flow Cell Assembly
- Cut a 15 mm × 2 mm channel at the center of a piece of double-sided tape (30 mm × 12 mm) using a puncher.
- Peel off the paper side of the double-sided tape and paste it on the glass slide with two holes. Press to remove air bubbles. Based on our experience, it is easier to remove air bubbles by peeling off the paper side than the plastic side of the double-sided tape.
- Cut a functionalized coverslip (60 mm × 24 mm) into four pieces (30 mm × 12 mm) using a diamond-tipped glass scribe, and remove the debris using nitrogen gas. Remember to keep the functionalized side face up.
- Peel off the plastic side of the double-sided tape, and paste the slide on the functionalized coverslip. Press gently to remove air bubbles between the coverslip and the tape. Removing air bubbles thoroughly can protect the flow cell from leaking while buffers were pumped into it.
- Insert inlet and outlet tubing into small and big holes, respectively. Fix the tubing using epoxy.
- Pump 20 µL of streptavidin (0.2 mg/mL) into flow cell using a syringe manually and incubate at RT for 10 min. Then pump blocking buffer10 into flow cell to replace streptavidin, and keep it at room temperature.
5. Single Molecule Visualization
- Obtain the focal alignment of red and far-red with A5 on the fluorescence microscope test slide #1 by simultaneous excitation using a 532 nm laser and a 640 nm laser. Produce dual wavelength images with splitting optics (See Table of Materials).
- Place the flow cell on the microscope and connect its outlet tubing to a longer tubing connecting with a 10 mL spring installed on an automated infusion/withdrawal programmable pump.
NOTE: Pumping buffers into the flow cell mentioned below means withdrawing buffers from the inlet tubing of the flow cell to the syringe. - Pump blocking buffer at 500 µL/min into the flow cell to remove air quickly and ensure that the buffer in the flow cell is unblocked. Then pump the blocking buffer at 200 µL/min into the flow cell and flip the outlet tubing to remove air bubbles from the inlet tubing and the flow cell thoroughly.
NOTE: All buffers pumped into the flow cell should be degassed in a vacuum desiccator for at least 15 minutes before use. - Add 0.5 µL of biotinylated λ-ARS317 DNA into 80 µL of blocking buffer, and pump it into the flow cell at 25 µL/min for 2 min. Then flush λ-ARS317 DNA using 200 µL of blocking buffer at a rate of 50 µL/min.
- Pump 200 µL of binding buffer10 at a rate of 50 µL/min to remove the blocking buffer in the flow cell.
- Add 0.2 µL of streptavidin-coated Qdot705 (1 µM) and 0.2 µL of biotinylated ORC (1.2 µM) into a tube, and incubate at room temperature for 5 min. Then add 20 µL of binding buffer into the tube, and place it on ice. The final concentration of ORC-Qdot705 is about 10 nM.
- Add 2 µL of ORC-Qdot705 (10 nM), 1 µL of DTT (100 mM), 1 µL of ATP (200 mM) into 96 µL of binding buffer. The final concentration of ORC-Qdot705 is 0.2 nM.
- Pump 20 µL of 0.2 nM ORC-Qdot705 at the rate of 10 µL/min into the flow cell. Flush out of excessive ORC-Qdot705 using 200 µL of binding buffer at the rate of 100 µL/min. Then pump binding buffer with 30 nM SYTOX Orange into the flow cell to stain DNA substrates at a rate of 100 µL/min.
- Excite the ORC-Qdot705 signal and SYTOX Orange stained DNA signal using a 405 nm laser and 532 nm laser, respectively. Observe the signals simultaneously with 100 µL/min flow using a Quad-band bandpass filter (FF01-446/510/581/703-25) and record the images by EM-CCD with 100 ms per frame. Collect 20 image stacks from different fields.
6. Data Analysis
- Crop the images using the Fiji software with a new plugin, which is modified by ourselves based on the Image plugin (OI_cut_RGBmerge) developed by Dr. Ron Vale's lab at University of California, San Francisco.
- Crop a stack of images (512 × 512 pixels) into two stacks of images (256 × 256 pixels). One is a 532 nm laser exciting result, and the other is a 405 nm laser exciting result.
- Process 61 sequential images using Fiji-Image-Stacks-Z project (average intensity).
- Measure the length of DNA (LDNA) and the distance from the site of ORC-Qdot705 (LORC) binding on DNA to the DNA's tethered end manually.
- Calculate the result of LORC/LDNA using excel, analyze the data using the bootstrap method by R, then create the histogram and fit Gaussian distributions.
Representative Results
To visualize the interaction between Qdot-labeled ORC and the ARS, we first constructed the λ-ARS317 DNA substrate. A DNA fragment containing ARS317 was integrated into XhoI site (33.5 kb) of native λ DNA by homologous recombination (Figure 1A). The recombination product was packaged using extracts and the packaged phage particles were cultured on LB plates (Figure 1B). The positive phage plaque was screened by PCR and confirmed by sequencing (Figure 1C). λ-ARS317 DNA was purified from the liquid lysates (Figure 1D).
Single-molecule imaging assays were carried out on the objective-type TIRFM set-up (Figure 2). By labeling the biotinylated ORC with a nearly 1:1 molar ratio streptavidin-coated Qdots rapidly, Qdot-labeled ORC was pumped into the λ-ARS317 tethered flow cell. DNA was stained using SYTOX Orange. The signals of Qdot-labeled ORC and SYTOX-Orange-stained DNA were imaged concurrently using TIRFM (Figure 3A-3C). To determine the distribution of ORC on λ-ARS317, the imaging stack was separated into two stacks: one stack of DNA signal and the other stack of ORC signal as described in step 6.1 (Figure 3B). Based on the formula, Position (kb) = (LORC/LDNA) × 50kb (Figure 3D), 273 binding positions of ORC-Qdot705 molecules on 168 DNA substrates were quantitively analyzed. The data showed that ORC-Qdot705 specially bindson the ARS317 inserted site with an obviously high abundance (Figure 3E). Meanwhile, ORC-Qdot705 also binds on the AT-rich areas located in middle and the free end of λ DNA, which is consistent with the results of the previous study10.
The high-quality image acquisition depends on the molar ratio of protein and Qdots in the labeling assay. Excessive Qdots may be good for the protein labeling efficiency, but they can create background noise. As shown in Figure 4, while labeling the biotinylated ORC with streptavidin-coated Qdots at 1:3 molar ratio, the background noise increased. Thus, it is critical to choose the appropriate molar ratio of biotinylated proteins to streptavidin-coated Qdots.
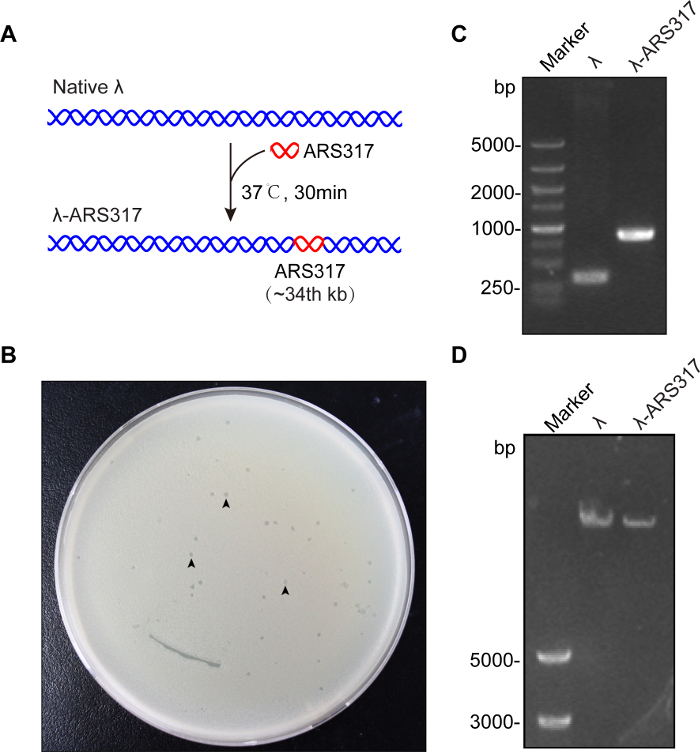
Figure 1: Preparation of λ-ARS317 DNA substrate. (A) Illustration of λ-ARS317 construction. A DNA fragment bearing ARS317 was integrated into λ DNA by homologous recombination. (B) Plaques on LB solid medium. Three of them were marked using black arrows. (C) A λ-ARS317 plaque was identified using PCR. (left) Marker, (middle) native λ DNA, (right) a plaque of λ-ARS317. (D) Purified λ-ARS317 DNA was detected by 0.6% agarose gel electrophoresis. (left) Marker, (middle) native λ DNA (right) purified λ-ARS317. Please click here to view a larger version of this figure.
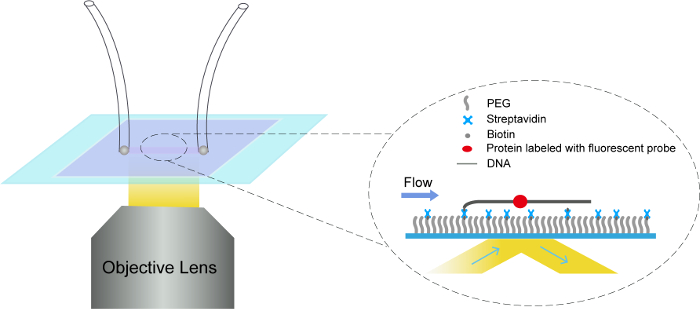
Figure 2: Schematic overview of objective-type TIRF and flow cell. Single molecule imaging assays were carried out based on the TIRFM combining with flow cell system. Our TIRFM was performed on an inverted microscope fitted with a 60X oil objective (numerical aperture = 1.49). DNA (gray line) was tethered on the coverslip in the flow cell through the biotin-streptavidin linkage, and stretched by the flow from left to right. A protein binding on the tethered DNA was indicated by a red-oval dot. Please click here to view a larger version of this figure.
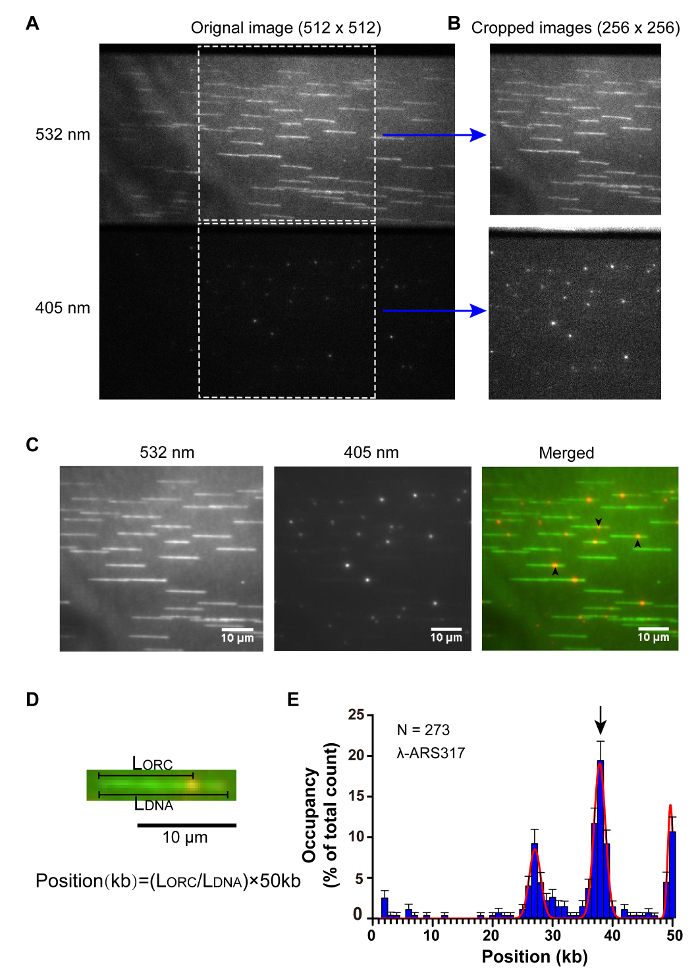
Figure 3: Qdot705-labeled ORC (1:1 molar ratio) binding on λ-ARS317. (A) ORC and λ-ARS317 DNA were observed simultaneously. (top) SYTOX Orange stained DNA was excited using a 532 nm laser and observed using the 550-613 nm transmission band of the quad-band band-pass filter. (bottom) Qdot705-labeled ORC was excited using a 405 nm laser and observed using the 663-743 nm transmission band of the quad-band band-pass filter. (B) Two 256×256 sub-stacks were cropped from the original 512×512 stack. (C) The images acquired using 61 sequential frames in the corresponding stacks. (left) SYTOX Orange stained DNA, (middle) Qdot705-labeled ORC, (right) merged image; three Qdot705-labeled ORC binding at ARS317 site on λ-ARS317 DNA substrates were marked using black arrows. (D) Illustration of the calculation of ORC binding position on DNA. LDNA means the DNA length, LORC means the distance from ORC binding site on DNA to the tethered end of DNA. (E) Histogram of ORC binding distribution on λ-ARS317 DNA. Gaussian fit to the data was indicated using the red solid line. Error bars indicate a 95% confidence interval based on 1000 bootstrap samples. N means the number of ORC molecules. ARS317 inserted site was indicated using a black arrow. Please click here to view a larger version of this figure.
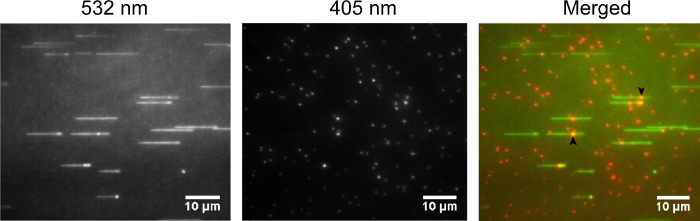

Figure 4: Qdot-labeled ORC binding (3:1 molar ratio) on λ-ARS317. ORC and λ-ARS317 DNA were observed in the same way as in Figure 3A. (left) SYTOX Orange stained DNA was excited using a 532 nm laser. (middle) Qdot-labeled ORC was excited using a 405 nm laser. (right) merged images. Two Qdot-labeled ORC binding at ARS317 site on λ-ARS317 DNA substrates were marked using black arrows. Please click here to view a larger version of this figure.
Discussion
Here, we present a protocol to observe the interaction between the Qdot-labeled protein and the site-specifically modified λ DNA using the TIRFM in a flow-cell. The necessary steps include site-specific modification of DNA substrate, DNA biotinylation, coverslip cleaning and functionalization, flow-cell preparation, and single-molecule imaging. There are two key points that should be noted. First, all the steps involved with λ DNA should be manipulated gently to decrease any possible damage, e.g., avoiding mixing by pipetting up and down repeatedly. Second, it is very important to ensure that the coverslip is clean enough to minimize its background noise. During the coverslip cleaning and functionalization process, water should reach ultrapure level, and the air should be dust-free. It is better to carry out the coverslip functionalization and flow-cell assembly on a clean bench.
In the single molecule assays for studying protein-DNA interaction, λ DNA is an appropriate substrate with several advantages20. λ DNA is long enough to be easily observed and permit recording protein movement on it in a single molecule experiment. Moreover, the ssDNA overhangs allow many designs for tethering λ DNA on a glass coverslip. In this study, we present a method for inserting an exogenous DNA fragment at a specific site of λ DNA. Thus, various user-definable DNA fragments can be integrated into λ DNA. The modified λ DNA can be conveniently purified from liquid lysates in large quantities for single molecule study.
It is difficult to evaluate the labeling efficiency accurately in the streptavidin-coated Qdot labeling assay because there are approximately 5 – 10 streptavidins per Qdot nanocrystal. Accordingly, the appropriate molar ratio of streptavidin-coated Qdots and biotinylated proteins should be optimized in the experiments. A 1:1 molar ratio can be a reference.
It should be noted that Qdot cannot be used in accurate quantitative assays since its high photo-stability is unsuitable for the photo-bleaching assay. Although the high stability of Qdots limits its application in the quantitative assays, it allows easy observation and long-time tracking5. With increasing applications of single-molecule fluorescence microscopy in biology, the protocol described in this paper will greatly contribute to unraveling the mechanisms of DNA metabolism in future.
Declarações
The authors have nothing to disclose.
Acknowledgements
We thank Dr. Hasan Yardimci and Dr.Sevim Yardimci of the Francis Crick Institute for kind help in the single-molecule experiments, Dr. Daniel Duzdevich from Dr. Eric C. Greene's lab of Columbia University, Dr. Yujie Sun of Peking University and Dr. Chunlai Chen of Tsinghua University for useful discussion. This study was supported by the National Natural Science Foundation of China 31371264, 31401059, CAS Interdisciplinary Innovation Team and the Newton Advanced Fellowship (NA140085) from the Royal Society.
Materials
| Lambda DNA | New England Biolabs | N3011 | Store 25 μL aliquots at -20 ºC. |
| XhoI enzyme | Thermo Fisher Scientific | FD0694 | |
| Quick-fusion cloning kit | Biotool | B22611 | |
| MaxPlax Lambda Packaging Extracts |
Epicentre | MP5110 | Bacterial strain LE392MP is included in this package. |
| MgSO4 | Sinopharm Chemical Reagent Co.,Ltd | 10013092 | Any brand is acceptable. |
| Tris | Amresco | 0497-5KG | |
| NaCl | Beijing Chemical works | N/A | Any brand is acceptable. |
| MgCl2 | Sinopharm Chemical Reagent Co.,Ltd | 10012818 | |
| Chloroform | Beijing Chemical works | N/A | Any brand is acceptable. |
| NZ-amine | Amresco | J853-250G | |
| Casamino acids | Sigma-Aldrich | 22090-500G | |
| PEG8000 | Beyotime | ST483 | |
| Magnetic stirring apparatus | IKA | KMO2 basic | |
| 15 mL Eppendorf tube | Eppendorf | 30122151 | 15 mL, sterile, bulk, 500pcs |
| Rnase | SIGMA | R4875-100MG | |
| Dnase | SIGMA | D5319-500UG | |
| Proteinase K | Amresco | 0706-100MG | |
| Biotinylated primers | Thermo Fisher Scientific | N/A | |
| T4 DNA ligase, T4 DNA Ligase, Reaction Buffer (10x) | New England Biolabs | M0202 | |
| Coverslip | Thermo Fisher Scientific | 22266882 | |
| Ethanol | Sinopharm Chemical Reagent Co.,Ltd | 10009259 | |
| Potassium hydroxide (KOH) | Sigma-Aldrich | 306568-100G | |
| Acetone | Thermo Fisher Scientific | A949-4 | |
| H2SO4 | Sinopharm Chemical Reagent Co.,Ltd | 80120891 | sulfuric acid |
| H2O2 | Sinopharm Chemical Reagent Co.,Ltd | 10011218 | 30% Hydrogen peroxide |
| Methanol | Sigma-Aldrich | 322415-2L | |
| Acetic acid | Sigma-Aldrich | V900798 | |
| APTES | Sigma-Aldrich | A3648 | |
| mPEG (methoxy-polyethylene glycol) |
Lysan | mPEG-SVA-5000 | |
| biotin-PEG (biotin-polyethylene glycol) |
Lysan | Biotin-PEG-SVA-5000 | |
| NaHCO3 | Sigma-Aldrich | 31437-500G | |
| Vacuum desiccator | Tianjin Branch Billion Lung Experimental Equipment Co., Ltd. | IPC250-1 | |
| Vacuum sealer | MAGIC SEAL | WP300 | |
| Diamond-tipped glass scribe | ELECTRON MICROSCOPY SCIENCES | 70036 | |
| Glass slide | Sail Brand | 7101 | |
| Inlet tubing | SCI (Scientific Commodties INC.) | BB31695-PE/2 | inner diameter 0.38 mm; outer diameter 1.09 mm. |
| Outlet tubing | SCI (Scientific Commodties INC.) | BB31695-PE/4 | inner diameter 0.76 mm; outer diameter 1.22 mm. |
| Double-sided tape | Sigma-Aldrich | GBL620001-1EA | |
| Epoxy | LEAFTOP | 9005 | five minutes epoxy |
| Streptavidin | Sigma-Aldrich | S4762 | |
| Fluorescence Microscope | Olympus | IX71 | |
| Infusion/withdrawal programmable pump |
Harvard apparatus | 70-4504 | |
| 532 nm laser | Coherent | Sapphire-532-50 | |
| 640 nm laser | Coherent | OBIS-640-100 | |
| EMCCD Camera | Andor | DU-897E-CS0-BV | |
| W-View Gemini Imaging splitting optics |
Hamamatsu photonics K.K. | A12801-01 | |
| TIRF illumination system | Olympus | IX2-RFAEVA2 | |
| 60×TIRF objective | Olympus | APON60XOTIRF | |
| Quad-edge laser dichroic beamsplitter |
Semrock | Di01-R405/488/ 532/635-25×36 |
|
| Quad-band bandpass filter | Semrock | FF01-446/510/ 581/703-25 |
|
| Dichroic beamsplitter | Semrock | FF649-Di01-25×36 | |
| Emission filter | Chroma Technology Corp | ET585/65m | |
| Emission filter | Chroma Technology Corp | ET665lp | |
| FocalCheck fluorescence, microscope test slide #1 | Thermo Fisher Scientific | F36909 | |
| SYTOX Orange | Thermo Fisher Scientific | S11368 | |
| Qdot705 Streptavidin Conjugate | Thermo Fisher Scientific | Q10163MP | Store at 4 ºC, do not freeze. |
| ATP | Amresco | 0220-25G | Prepare 200 mM ATP solution, using ddH2O, adjust pH to 7.0, and store 10 μL aliquots at -20 ºC. |
| DTT | Amresco | M109-5G | Prepare 1 M solution using ddH2O, and store 10 μl aliquots at -20 ºC. |
Referências
- Fu, Y. V., et al. Selective Bypass of a Lagging Strand Roadblock by the Eukaryotic Replicative DNA Helicase. Cell. 146 (6), 930-940 (2011).
- Qi, Z., et al. DNA sequence alignment by microhomology sampling during homologous recombination. Cell. 160 (5), 856-869 (2015).
- Chong, S., Chen, C., Ge, H., Xie, X. S. Mechanism of transcriptional bursting in bacteria. Cell. 158 (2), 314-326 (2014).
- Wegner, K. D., Hildebrandt, N. Quantum dots: bright and versatile in vitro and in vivo fluorescence imaging biosensors. Chemical Society Reviews. 44 (14), 4792-4834 (2015).
- Gao, X., et al. In vivo molecular and cellular imaging with quantum dots. Current opinion in biotechnology. 16 (1), 63-72 (2005).
- Sternberg, S. H., Redding, S., Jinek, M., Greene, E. C., Doudna, J. A. DNA interrogation by the CRISPR RNA-guided endonuclease Cas9. Nature. 507 (7490), 62 (2014).
- Lee, J. Y., Finkelstein, I. J., Arciszewska, L. K., Sherratt, D. J., Greene, E. C. Single-molecule imaging of FtsK translocation reveals mechanistic features of protein-protein collisions on DNA. Molecular cell. 54 (5), 832-843 (2014).
- Wang, H., Tessmer, I., Croteau, D. L., Erie, D. A., Van Houten, B. Functional characterization and atomic force microscopy of a DNA repair protein conjugated to a quantum dot. Nano letters. 8 (6), 1631-1637 (2008).
- Redding, S., et al. Surveillance and processing of foreign DNA by the Escherichia coli CRISPR-Cas system. Cell. 163 (4), 854-865 (2015).
- Xue, H., et al. Utilizing Biotinylated Proteins Expressed in Yeast to Visualize DNA-Protein Interactions at the Single-Molecule Level. Frontiers in microbiology. 8, 2062 (2017).
- Duzdevich, D., et al. The dynamics of eukaryotic replication initiation: origin specificity, licensing, and firing at the single-molecule level. Molecular cell. 58 (3), 483-494 (2015).
- Hughes, C. D., et al. Real-time single-molecule imaging reveals a direct interaction between UvrC and UvrB on DNA tightropes. Nucleic acids research. 41 (9), 4901-4912 (2013).
- Kad, N. M., Wang, H., Kennedy, G. G., Warshaw, D. M., Van Houten, B. Collaborative dynamic DNA scanning by nucleotide excision repair proteins investigated by single-molecule imaging of quantum-dot-labeled proteins. Molecular cell. 37 (5), 702-713 (2010).
- Chandradoss, S. D., et al. Surface Passivation for Single-molecule Protein Studies. Jove-Journal of Visualized Experiments. (86), (2014).
- Chen, X. Q., Zhao, E. S., Fu, Y. V. Using single-molecule approach to visualize the nucleosome assembly in yeast nucleoplasmic extracts. Science Bulletin. 62 (6), 399-404 (2017).
- Yardimci, H., Loveland, A. B., van Oijen, A. M., Walter, J. C. Single-molecule analysis of DNA replication in Xenopus egg extracts. Methods. 57 (2), 179-186 (2012).
- Tanner, N. A., Loparo, J. J., van Oijen, A. M. Visualizing single-molecule DNA replication with fluorescence microscopy. Journal of visualized experiments: JoVE. (32), (2009).
- Lockett, T. J. A bacteriophage λ DNA purification procedure suitable for the analysis of DNA from either large or multiple small lysates. Analytical biochemistry. 185 (2), 230-234 (1990).
- Smith, E. A., Chen, W. How to prevent the loss of surface functionality derived from aminosilanes. Langmuir. 24 (21), 12405-12409 (2008).
- Kim, Y., Torre, A., Leal, A. A., Finkelstein, I. J. Efficient modification of λ-DNA substrates for single-molecule studies. Scientific reports. 7 (1), 2071 (2017).

