Assessing Energy Substrate Oxidation In Vitro with 14CO2 Trapping
Summary
This protocol describes an easy-to-use method to examine substrate oxidation by tracking 14CO2 production in vitro.
Abstract
Mitochondria host the machinery for the tricarboxylic acid (TCA) cycle and electron transport chain (ETC), which generate adenosine triphosphate (ATP) to maintain energy homeostasis. Glucose, fatty acids, and amino acids are the major energy substrates fueling mitochondrial respiration in most somatic cells. Evidence shows that different cell types may have a distinct preference for certain substrates. However, substrate utilization by various cells in the skeleton has not been studied in detail. Moreover, as cellular metabolism is attuned to physiological and pathophysiological changes, direct assessments of substrate dependence in skeletal cells may provide important insights into the pathogenesis of bone diseases.
The following protocol is based on the principle of carbon dioxide release from substrate molecules following oxidative phosphorylation. By using substrates containing radioactively labeled carbon atoms (14C), the method provides a sensitive and easy-to-use assay for the rate of substrate oxidation in cell culture. A case study with primary calvarial preosteoblasts versus bone marrow-derived macrophages (BMMs) demonstrates different utilization of the main substrates between the two cell types.
Introduction
Oxidative phosphorylation (OXPHOS) in eukaryotes is the process by which nutrients are broken down inside mitochondria to release chemical energy in the form of ATP through consumption of oxygen. Catabolism of various substrates inside mitochondria through the tricarboxylic acid (TCA) cycle generates few ATP molecules directly, but rather stores energy through reduction of the electron carriers nicotinamide adenine dinucleotide (NAD+) and flavin adenine dinucleotide (FAD+). The reduced carriers are then oxidized by ETC located on the inner membrane of mitochondria to generate a proton concentration gradient across the membrane. The protons eventually flow down their gradient back into the mitochondrial matrix through ATP synthase to produce ATP. OXPHOS is the most efficient means of ATP production from energy substrates and generally preferred in aerobic environments. Previously, aerobic glycolysis-production of lactate from glucose while oxygen is present-was thought to be pathophysiological, often a hallmark of cancer cells. More and more, it is being discovered that some normal cell types use aerobic glycolysis for reasons that have yet to be fully deciphered.
Metabolic flexibility is the capacity for cells or organisms to adapt to changing energy demands and available fuel sources. For example, the energetic demand of skeletal muscle is mainly met by OXPHOS at steady state but by anaerobic glycolysis during high-intensity exercise1. As the exercise duration increases, glucose and fatty acid oxidation contribute more to overall energy production2. However, substrate use is not dependent only on availability, as substrates antagonistically compete during oxidation. Most notably, fatty acid oxidation has been shown to inhibit glucose utilization by skeletal muscle in a phenomenon known as the Randle effect3. A reciprocal effect was demonstrated by subsequent studies4,5. In addition, many diseases are associated with a change in substrate preference and the development of metabolic inflexibility in cells. For instance, fatty acid oxidation is reduced in the skeletal muscle of type II diabetic patients compared to normal control subjects6. The metabolic changes in disease settings are a subject of intense investigation as they may contribute to pathogenesis.
Energy metabolism in skeletal cell types is relatively understudied but has gained attention in recent years7. Previous work has shown that aerobic glycolysis is the dominant energy pathway in calvarial osteoblasts, while glucose oxidation through the TCA cycle plays a role in osteoclast formation8,9. Others have provided evidence for fatty acids as an energy source for osteoblasts10. Glutamine catabolism has also been shown to support osteoblast differentiation from the progenitors11,12. However, a comprehensive understanding of substrate utilization by various skeletal cell types is still lacking. In addition, changes in cellular metabolism during cell differentiation or in response to pathological signals are expected to alter fuel substrate utilization. Described below is an easy-to-use protocol for assaying substrate oxidation in vitro.
Protocol
The use of radioactive materials (RAM) requires prior approval by a designated safety committee at each institution. RAMs used in this protocol have been approved by Environmental Health & Radiation Safety (EHRS) at the University of Pennsylvania. The use of animals requires prior approval by the Institutional Animal Care and Use Committee (IACUC) at the home institution. The following study was approved by IACUC at The Children's Hospital of Philadelphia.
1. Preparation of stock solutions for 14C-labeled substrates
- Make 4 mM bovine serum albumin (BSA) stock solution by mixing 11 g of BSA and 33.1 mL of Dulbecco's phosphate-buffered saline (DPBS) and shaking it gently at room temperature for ~3 h until BSA is fully dissolved. Warm the BSA stock solution in a 70 °C water bath before use.
- Air-dry 50 µCi 14C-oleate (50 µCi/µmol in ethanol) in the vial with the lid off for 8 h in a designated RAM working hood. Add 312.5 µL of H2O and then 3.5 mg (11.5 µmol) of sodium oleate. Mix thoroughly.
- Warm the resulting solution to 70 °C in a water bath. Add 0.9375 mL of the prewarmed BSA stock solution to yield 10 mM hot oleate stock solution (containing a 1:11.5 ratio of 14C-oleate to unlabeled oleate). Aliquot the stock solution and store it at -20 °C for long-term use.
- Use 14C-glucose and 14C-glutamine directly after thawing.
2. Preparation of medium containing 14C-labeled substrates
NOTE: To ensure reliable concentrations of energy substrates in the media, custom-made media must be used with fresh substrates added shortly before use. Here, a custom-made Minimum Essential Medium (MEMα) without glucose, pyruvate, glutamine, phenol red, or sodium bicarbonate is used. However, an optimal medium should be determined for each cell type.
- Reconstitute 8.67 g of the medium powder in 1 L of purified water. Add 2.2 g of sodium bicarbonate to achieve a pH of 7.4 and filter the solution using 0.22 µm vacuum filtration.
- Add fresh ingredients to obtain the complete medium (cMEMα) with final concentrations of 5.5 mM glucose, 2 mM glutamine, 1 mM pyruvate, and 10% fetal bovine serum (FBS). Further supplement cMEMα with 100 mM L-carnitine and 10 µM HEPES to facilitate fatty acid oxidation.
- Immediately before use, add 14C-labeled substrates to freshly prepared cMEMα to make hot media. To make the oleate hot medium, add the 10 mM hot oleate stock solution to cMEMα for a final concentration of 100 µM oleate (1:100 dilution, final radioactivity 0.4 µCi/mL).
NOTE: The 10 mM hot oleate stock containing a 1:11.5 ratio of hot:cold oleate (see step 1.3) is used to achieve a physiological level of 100 µM total oleate in the media while minimizing the amount of radioactive material. - Make 10 mM unlabeled oleate stock by adding 24.36 mg of sodium oleate to 2 mL of H2O in a water bath at 70 °C for ~20 min until completely dissolved before mixing quickly with 6 mL of the 4 mM BSA stock solution prewarmed to 70 °C. Transfer to a 37 °C water bath for 1 h and filter through a 0.22 µm filter. Aliquot and store at -20 °C for long-term use.
- To make glucose hot medium, add 14C-glucose to a final concentration of 1.333 µM (1:4,125 dilution, final radioactivity 0.4 µCi/mL). To make glutamine hot medium, add 14C-glutamine to the final concentration of 2 µM (1:1,000 dilution, final radioactivity 0.4 µCi/mL).
- Supplement the glucose and glutamine hot medium with a 10 mM stock solution of unlabeled oleate from step 2.4 to achieve the final concentration of 100 µM oleate, the same as that in the oleate hot medium.
3. Preparation of cells
NOTE: Calvarial preosteoblasts and bone marrow macrophages are used as examples here. Users should prepare their cell type of choice according to appropriate protocols. Optimize the concentration of collagenase II to be used for digestion in pilot experiments as the enzymatic activity may vary among different lots.
- Isolation and culture of calvarial preosteoblasts
- Sacrifice postnatal day 3-5 (P3-5) pups by decapitation and transfer them to ice-cold DPBS containing Penicillin-Streptomycin (P/S).
- Expose the calvaria by removing the skin and soft tissue. Collect the middle region by cutting away the surrounding tissues from back to front in DPBS (Figure 1A). Scratch the inner and outer surfaces of the calvaria gently using tweezers to help release the cells upon subsequent digestion.
- Prepare a 2 mg/mL and a 4 mg/mL solution of Collagenase type II in DPBS and filter each solution with a fresh 0.22 µm filter. Digest the cleaned calvaria first with the 2 mg/mL collagenase solution for 15 min and discard the digestion solution. Digest the cleaned samples in 4 mg/mL collagenase solution 3 times for 15 min each time, pooling and saving the digestion solution.
- Filter the digestion solution through 70 µm cell strainers and centrifuge the filtrate at 300 × g for 5 min. Resuspend the cell pellet in cMEMα (see step 2.2) containing 10% FBS and P/S.
- Count and seed the cells at 4 × 104 cells/cm2 in 10 cm plates. Culture the cells in a 37 °C incubator with 5% CO2 for 3 days.
- Dissociate the cells at 37 °C with 0.25% trypsin-EDTA. Count and seed the cells in 24-well cell culture plates with cMEMα containing 10% FBS and P/S at 7.5 × 105/cm2. Seed at least 4 wells per cell type per substrate and at least three extra wells for each cell type for cell counting before the assay.
- Culture the cells at 37 °C overnight before the substrate oxidation assays.
- Isolation and culture of BMMs
NOTE: Culture of BMMs requires macrophage colony-stimulating factor (M-CSF), which can be purchased commercially as a recombinant protein. Conditioned medium from the cell line CMG14-12 engineered to express M-CSF is an economical alternative used here13.- Collect femurs and tibias from 8-week-old mice after removing the muscle and connective tissues. Cut and discard both ends of the bones with sharp scissors.
- Flush out the bone marrow from one femur and one tibia into a 10 cm Petri dish with a syringe fitted with a 23 G needle containing 15 mL of cMEMα with 10% FBS, P/S, and 10% CMG14-12 conditioned medium (BMM medium). Culture the cells in the Petri dish in a 37 °C incubator with 5% CO2.
- After 3 days, discard the culture media and rinse with DPBS. Dissociate the attached cells at 37 °C with 0.25% trypsin-EDTA for 5 min. Centrifuge and resuspend the cell pellet in the BMM medium.
- Count and seed the cells in 24-well cell culture plates at 7.5 × 105/cm2 in the same way as described in 3.1.6. Culture at 37 °C overnight in the BMM medium before starting the assays.
4. Substrate oxidation assay with CO2 trap
NOTE: An appropriate seeding density should be determined for each cell type to achieve 80-90% confluence before the start of the assay. Note that cell density can affect the metabolic state of the cells.
- Wash the cells in the extra wells twice with DPBS. Dissociate the cells with 0.25% trypsin-EDTA and mixed 20 µL cells resuspended in DPBS with 20 µL of acridine orange/propidium iodide (AO/PI) ready-to-use commercial dye solution. Determine the number of live cells with an automated cell counter and record the number for final calculations.
- Wash the cells in the assay wells twice with DPBS. Add 500 µL of hot medium to each assay well in the RAM-designated tissue culture hood. Seal the plates with parafilm and incubate the cells in a RAM-designated incubator at 37 °C for 4 h.
- During incubation, cut filter paper into circular pieces slightly bigger than the area inside the cap of 1.5 ml microcentrifuge tubes and insert the paper snugly into the cap (Figure 1B). Add 200 µL of 1 M perchloric acid to each tube and 20 µL of sodium hydroxide to the filter paper fitted inside the cap.
- After incubation of the cells, transfer 400 µL of culture medium from each well to the prepared tubes and close the caps immediately. Leave the tubes in a tube rack at room temperature for 1 h.
- During incubation, set up a scintillation vial for each tube and fill it with 4 mL of scintillation fluid. Transfer each piece of filter paper to a scintillation vial and incubate at room temperature for 30 min.
- Perform wipe tests on the tissue culture hood, the water bath (used for 14C-oleate preparation), refrigerator, incubator, sink, ground, and any other working area for potential RAM contamination. Put the paper wipes into scintillation vials containing scintillation fluid.
- Measure 14C radioactivity in the scintillation vials with a scintillation counter. Record the reading results. Decontaminate the working environment according to radiation safety guidelines if necessary.
5. Data analysis
NOTE: Assuming that each substrate is fully oxidized to release CO2, the substrate oxidation rate can be calculated from the trapped CO2 radioactivity.
- Calculate the substrate oxidation rate using Eq (1) and the X, Y, and Z values given in Table 1 for different substrates.
Substrate oxidation rate (µmol/cell/h) =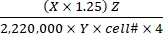 (1)
(1)
- Calculate the total disintegrations per min (DPM) for each reaction well. For X, use the DPM value (scintillation counter reading) from 0.4 mL of the reaction medium used for CO2 trapping out of 0.5 mL of the total reaction medium. Therefore, the total DPM from each reaction well is X × 1.25.
- Divide the total DPM by a factor of 2,220,000 to convert to µCi.
- Convert the radioactivity in µCi to the number of 14C-labeled molecules. Divide the µCi value by the specific activity (Y) of each labeled substrate. Use the following Y values (Table 1): 14C-glucose: 300 mCi/mmol; 14C-glutamine: 200 mCi/mmol; 14C-oleate: 50 mCi/mmol.
- Calculate the total number of oxidized molecules from the oxidized 14C-labeled molecules by multiplying the latter with the hot-to-total dilution factor (Z). Use the following Z values (Table 1): Glucose: 4,125; glutamine: 1,000; oleate: 12.5.
- Divide the resulting product by cell number (cell#) and 4 (for the reaction time of 4 h) to determine the substrate oxidation rate per cell. Use the cell# determined in the parallel wells at the beginning of the assay.
Representative Results
In this example, the CO2 trapping method is used to compare substrate oxidation by primary calvarial preosteoblasts versus BMMs, which are frequently used for in vitro osteoblast or osteoclast differentiation, respectively. After the primary cells are passaged and cultured in cMEMα overnight, they typically reach 80-90% of confluence and exhibit their characteristic morphology. The calvarial preosteoblasts are notably larger than BMMs (Figure 2). Each cell type is subjected to oxidation assays for glucose, glutamine, and oleate by following steps 4.1 through 4.7. The results are analyzed by using Eq (1).
The results show that the oxidation rate for each substrate is significantly higher in calvarial preosteoblasts than BMMs, likely indicating greater energy production by OXPHOS in the preosteoblasts (Figure 3). Previous studies with Seahorse technology have shown that OXPHOS accounts for ~60% of ATP production in preosteoblasts but markedly decreases in mature osteoblasts, where aerobic glycolysis is responsible for ~80% of the energy9. The current results demonstrate that all three major substrates contribute to OXPHOS in preosteoblasts. However, further investigation is necessary to determine whether utilization of one or all the substrates is reduced in the mature osteoblasts.
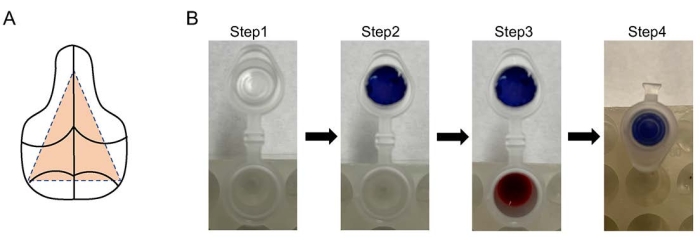

Figure 1: Diagrams for CO2 trapping assay and calvaria dissection. (A) The middle region of the calvaria, indicated by the orange dashed triangle, is harvested for cell isolaton. (B) 1.5 mL microcentrifuge tubes are used for CO2 trapping (step 1). Filter papers (blue paper for illustration purpose) are cut to fit into tube caps (step 2). Add 200 µL of 1 M perchloric acid to 400 µL of hot media from the cell culture in each tube and 20 µL of NaOH to the filter paper (step 3) before closing the lid (step 4). Please click here to view a larger version of this figure.
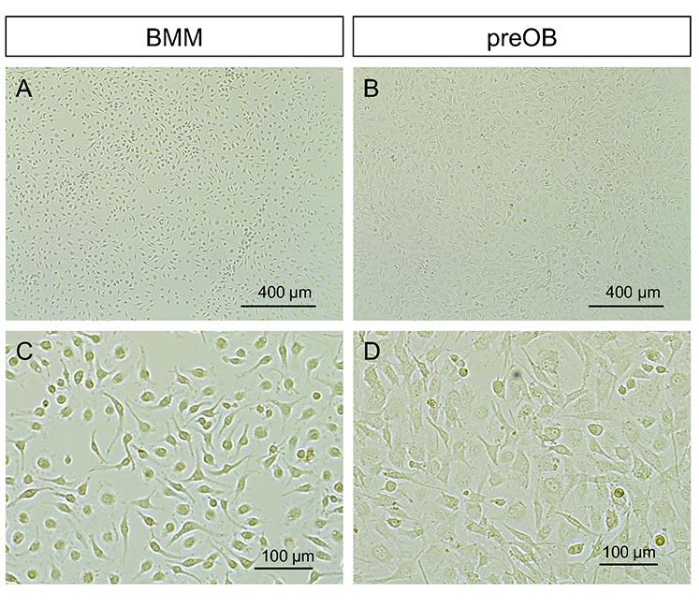
Figure 2: Typical morphology of BMMs and calvarial preosteoblasts. (A, B) Representative images at lower magnification. (C, D) Representative images at higher magnification. Scale bars = 400 µm (A, B), 100 µm (C, D). Abbreviations: BMMs = bone marrow-derived macrophages; preOBs = preosteoblasts. Please click here to view a larger version of this figure.
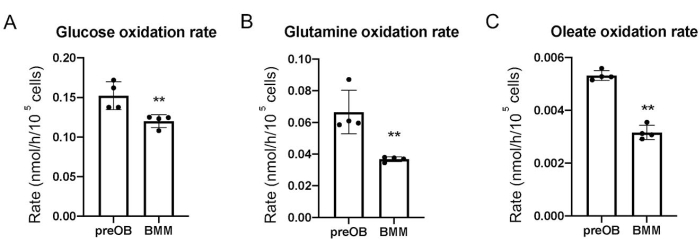
Figure 3: Substrate oxidation rates in calvarial preosteoblasts and BMMs. (A) Glucose. (B) Glutamine. (C) Oleate. The calculations assume that each substrate molecule is fully oxidized. Abbreviations: BMM = bone marrow-derived macrophage; preOB = preosteoblast. Please click here to view a larger version of this figure.
| Substrate | Radioactivity (X) | Specific Activity (Y), mCi/mmol | Dilution Factor (Z) |
| Glucose | DPM1 | 300 | 4,125 |
| Glutamine | DPM2 | 200 | 1,000 |
| Oleate | DPM3 | 50 | 12.5 |
Table 1: Parameters used in data analysis.
Discussion
The protocol provides an easy-to-use method to determine the oxidation rate of major energy substrates. It is a simpler alternative to other protocols that use flasks containing a central well and capped with rubber stoppers14,15,16. Although the example study here is performed with cell culture, the method can be easily adapted for tissue explants or tissue homogenates containing intact mitochondria, as previously described17.
Some key steps to consider in this protocol involve the preparation of the reagents and the set up of the tubes for 14CO2 trapping. Ensuring appropriate conjugation of oleate to BSA is crucial to prevent free fatty acids (FFAs) from damaging the cell membranes and to allow efficient uptake by the cell for metabolism. Incubate the BSA solution just long enough for complete dissolution, as long incubations with BSA at 70 °C lead to irreversible solidification.
Fresh cMEMα must be made for each experimental setup due to the short half-lives of substrates such as L-glutamine. Finally, the filter paper should be large enough to fit securely in the cap to avoid erroneous readings. An accidental fall of the filter paper into the reaction liquid will lead to abnormally high DPM values, which should be omitted from further analysis. Four to five replicated wells can be set in case of potential contamination. If the final DPM values are too low, for example, less than 100, increasing the amount of radioactive substrate or the incubation time can be helpful.
There are caveats for using oleate as the surrogate substrate for fatty acid oxidation. Oleate is the most abundant monounsaturated, long-chain fatty acid constituting 32% of all fatty acids in circulation and tissues. However, other long-chain fatty acids, such as palmitate (28%) and linoleate (14%), are also major substrates in vivo18,19. Therefore, supplementation of the cell culture media with oleate alone may not accurately reflect the oxidation rate of fatty acids when other species are also present. When performing the assay, this concern can be ameliorated by including those other fatty acids in the culture media.
It should be noted that cell metabolism is highly context-dependent and can exhibit great plasticity depending on substrate availability and oxygen tension, among other factors. It is therefore important to evaluate in vitro results in the context of physiological conditions. Nonetheless, it is increasingly clear that cells tend to maintain at least some aspects of their metabolic characteristics in vitro, likely due to the cell-specific metabolic programming and memory. Thus, straightforward in vitro methods, such as this one, offer an important tool for gaining insights into not only the metabolic diversity among cell types but also potential dysregulation in disease conditions.
Declarações
The authors have nothing to disclose.
Acknowledgements
The work was supported in part by NIH grant R01 AR060456 (FL). We thank Dr. Michael Robinson and Elizabeth Krizman (The Children's Hospital of Philadelphia) for their generous help with the scintillation counter.
Materials
| 0.22 µm filters | Sigma-Aldrich | SLGVM33RS | Used to filter BSA solution |
| 0.25% Trypsin-EDTA | Gibco | 25200056 | Dissociate cells from cell culture plates |
| 1.5 mL Eppendorf tubes | PR1MA | PR MCT17 RB | Used for reaction incubation |
| 10 cm plates | TPP | 93100 | Used for cell culture |
| 10 mL syringe | BD | 302995 | Used to flush marrow from long bones |
| 10% FBS | Atlanta biologicals | S11550 | For Cell culture medium preparation |
| 14C-Glucose | PerkinElmer | NEC042X050UC | Used to make hot media |
| 14C-glutamine | PerkinElmer | NEC451050UC | Used to make hot media |
| 14C-oleate | PerkinElmer | NEC317050UC | Used to make hot media |
| 23 G needle | BD | 305120 | Used to flush marrow from long bones |
| 24-well plates | TPP | 92024 | Used for cell culture |
| 70 μm cell strainers | MIDSCI | 70CELL | Used to filter supernatant during cavarial digestion |
| Acridine Orange/Propidium Iodide (AO/PI) dye | Nexcelom Biosciences | CS2-0106 | Stains live cells to determine seed density |
| Bovine Serum Ablumin | Proliant Biologicals | 68700 | Used for fatty acid conjugation |
| Cellometer Auto 2000 | Nexcelom Biosciences | Determine the number of viable cells | |
| Centrifuge | Thermo Fisher | Legend Micro 21R | Used to pellet cells |
| Collagenase type II | Worthington | LS004176 | Dissociate cells from tissue |
| Custom MEM alpha | GIBCO | SKU: ME 18459P1 | Used to create custom hot media |
| Dulbecco's Phosphate-Buffered Saline | Gibco | 10010023 | Used to dissolve and dilute reagents, and wash culture dishes |
| Filter Paper | Millipore-Sigma | WHA1001090 | Traps CO2 with sodium hydroxide |
| Glucose | Sigma-Aldrich | g7528 | Used to make custom media |
| HEPES | Gibco | 15630080 | Traps CO2 during cell culture |
| L-carnitine | Sigma-Aldrich | C0283 | Supplemented for fatty acid oxidation |
| L-Glutamine | Sigma-Aldrich | g3126 | Used to make custom media |
| MEM alpha | Thermo | A10490 | Cell culture medium |
| Parafilm | Pecheney Plastic Packaging | PM998 | Used to seal cell culture dishes |
| Penicillin-Streptomycin | Thermo Fisher | 15140122 | Prevents contamination in cell culture |
| Perchloric Acid | Sigma-Aldrich | 244252 | Releases CO2 during metabolic assay |
| Pyruvate | Sigma-Aldrich | p5280 | Used to make custom media |
| Scintillation Counter | Beckman Coulter | LS6500 | Determines radioactivity from the filter paper |
| Scintillation Fluid | MP Biomedicals | 882453 | Absorb the energy emitted by RAMs and re-emit it as flashes of light |
| Scintillation Vial | Fisher Scientific | 03-337-1 | Reaction containers for scintillation fluid |
| Sodium carbonate | Sigma-Aldrich | S5761 | Balance buffer for medium |
| Sodium Hydroxide | Sigma-Aldrich | 58045 | Traps CO2 during metaboilc assay |
| Sodium oleate | SANTA CRUZ | SC-215879 | BSA conjugated fatty acid preparation |
| Vaccum filtration 1000 | TPP | 99950 | Filter cMEMα |
Referências
- Hargreaves, M., Spriet, L. L. Skeletal muscle energy metabolism during exercise. Nature Metabolism. 2 (9), 817-828 (2020).
- Jeukendrup, A. E. Regulation of fat metabolism in skeletal muscle. Annals of the New York Academy of Sciences. 967, 217-235 (2002).
- Randle, P. J., Garland, P. B., Hales, C. N., Newsholme, E. A. The glucose fatty-acid cycle. Its role in insulin sensitivity and the metabolic disturbances of diabetes mellitus. Lancet. 1 (7285), 785-789 (1963).
- Randle, P. J., Newsholme, E. A., Garland, P. B. Regulation of glucose uptake by muscle. 8. Effects of fatty acids, ketone bodies and pyruvate, and of alloxan-diabetes and starvation, on the uptake and metabolic fate of glucose in rat heart and diaphragm muscles. Biochemical Journal. 93 (3), 652-665 (1964).
- Taegtmeyer, H., Hems, R., Krebs, H. A. Utilization of energy-providing substrates in the isolated working rat heart. Biochemical Journal. 186 (3), 701-711 (1980).
- Cha, B. S., et al. Impaired fatty acid metabolism in type 2 diabetic skeletal muscle cells is reversed by PPARgamma agonists. American Journal of Physiology. Endocrinology and Metabolism. 289 (1), 151-159 (2005).
- Lee, W. C., Guntur, A. R., Long, F., Rosen, C. J. Energy metabolism of the osteoblast: implications for osteoporosis. Endocrine Reviews. 38 (3), 255-266 (2017).
- Li, B., et al. Both aerobic glycolysis and mitochondrial respiration are required for osteoclast differentiation. FASEB Journal. 34 (8), 11058-11067 (2020).
- Lee, W. C., Ji, X., Nissim, I., Long, F. Malic enzyme couples mitochondria with aerobic glycolysis in osteoblasts. Cell Reports. 32 (10), 108108 (2020).
- Kim, S. P., et al. Fatty acid oxidation by the osteoblast is required for normal bone acquisition in a sex- and diet-dependent manner. JCI Insight. 2 (16), 92704 (2017).
- Yu, Y., et al. Glutamine metabolism regulates proliferation and lineage allocation in skeletal stem cells. Cell Metabolism. 29 (4), 966-978 (2019).
- Karner, C. M., Esen, E., Okunade, A. L., Patterson, B. W., Long, F. Increased glutamine catabolism mediates bone anabolism in response to WNT signaling. The Journal of Clinical Investigation. 125 (2), 551-562 (2015).
- Takeshita, S., Kaji, K., Kudo, A. Identification and characterization of the new osteoclast progenitor with macrophage phenotypes being able to differentiate into mature osteoclasts. Journal of Bone and Mineral Research. 15 (8), 1477-1488 (2000).
- Itoh, Y., et al. Dichloroacetate effects on glucose and lactate oxidation by neurons and astroglia in vitro and on glucose utilization by brain in vivo. Proceedings of the National Academy of Sciences of the United States of America. 100 (8), 4879-4884 (2003).
- Oba, M., Baldwin, R. L. 4. t. h., Bequette, B. J. Oxidation of glucose, glutamate, and glutamine by isolated ovine enterocytes in vitro is decreased by the presence of other metabolic fuels. Journal of Animal Science. 82 (2), 479-486 (2004).
- Esen, E., Lee, S. Y., Wice, B. M., Long, F. PTH promotes bone anabolism by stimulating aerobic glycolysis via IGF signaling. Journal of Bone and Mineral Research. 30 (11), 1959-1968 (2015).
- Huynh, F. K., Green, M. F., Koves, T. R., Hirschey, M. D. Measurement of fatty acid oxidation rates in animal tissues and cell lines. Methods in Enzymology. 542, 391-405 (2014).
- Hodson, L., Skeaff, C. M., Fielding, B. A. Fatty acid composition of adipose tissue and blood in humans and its use as a biomarker of dietary intake. Progress in Lipid Research. 47 (5), 348-380 (2008).
- Abdelmagid, S. A., et al. Comprehensive profiling of plasma fatty acid concentrations in young healthy Canadian adults. PLoS One. 10 (2), 0116195 (2015).

