Generation of Brown Fat-Specific Knockout Mice Using a Combined Cre-LoxP, CRISPR-Cas9, and Adeno-Associated Virus Single-Guide RNA System
Summary
In this protocol, we describe the technical procedures to generate brown adipose tissue (BAT)-specific knockout mice leveraging a combined Cre-LoxP, CRISPR-Cas9, and adeno-associated virus (AAV) single-guide RNA (sgRNA) system. The described steps include the design of the sgRNAs, the preparation of the AAV-sgRNA particles, and the microinjection of AAV into the BAT lobes.
Abstract
Brown adipose tissue (BAT) is an adipose depot specialized in energy dissipation that can also serve as an endocrine organ via the secretion of bioactive molecules. The creation of BAT-specific knockout mice is one of the most popular approaches for understanding the contribution of a gene of interest to BAT-mediated energy regulation. The conventional gene targeting strategy utilizing the Cre-LoxP system has been the principal approach to generate tissue-specific knockout mice. However, this approach is time-consuming and tedious. Here, we describe a protocol for the rapid and efficient knockout of a gene of interest in BAT using a combined Cre-LoxP, CRISPR-Cas9, and adeno-associated virus (AAV) single-guide RNA (sgRNA) system. The interscapular BAT is located in the deep layer between the muscles. Thus, the BAT must be exposed in order to inject the AAV precisely and directly into the BAT within the visual field. Appropriate surgical handling is crucial to prevent damage to the sympathetic nerves and vessels, such as the Sultzer’s vein that connects to the BAT. To minimize tissue damage, there is a critical need to understand the three-dimensional anatomical location of the BAT and the surgical skills required in the technical steps. This protocol highlights the key technical procedures, including the design of sgRNAs targeting the gene of interest, the preparation of AAV-sgRNA particles, and the surgery for the direct microinjection of AAV into both BAT lobes for generating BAT-specific knockout mice, which can be broadly applied to study the biological functions of genes in BAT.
Introduction
Obesity is increasing at a significant rate worldwide, leading to a broad spectrum of metabolic diseases1,2,3. The adipose tissue is key to these pathologies. The two functionally distinct types of adipose tissue that exist are white adipose tissue (WAT), which stores excess calories, and brown adipose tissue (BAT) and its related beige/brite fat, which dissipate energy for thermogenesis. While BAT has been recognized for its energy-dissipating function, it also has endocrine functions via the production of bioactive molecules that regulate metabolism in distal organs4,5. Numerous studies in rodents have demonstrated that increasing the amount or activity of brown or beige fat leads to increased energy expenditure and improved insulin sensitivity. In humans, people with detectable BAT have a significantly lower prevalence of cardiometabolic diseases6. Thus, BAT holds excellent therapeutic potential for obesity-related metabolic sequelae7,8,9.
To investigate the physiology and pathophysiology of BAT development and function and to elucidate the molecular mechanisms involved in these processes, the BAT-specific transgenic mouse model is a method of choice10. The Cre-LoxP recombination system is the most commonly used means to produce conditional knockout mice by editing the mouse genome. This system has enabled the modification (overexpression or knockout) of genes of interest in a tissue/cell-specific manner11. It can also be utilized to label a specific cell type by expressing a selective fluorescent reporter gene.
Recently, the Cre-LoxP system approach has been further developed by combining the CRISPR-Cas9 technology and the adeno-associated virus (AAV) single-guide RNA (sgRNA) system12. The CRISPR-Cas9 system is a specific and efficient gene-editing tool to modify, regulate, or target precise regions of the genome13. CRISPR-Cas9-based genome editing allows for rapid genetic manipulation of genomic loci because it does not require homologous recombination with a gene-targeting vector. Combined Cre-LoxP, CRISPR-Cas9, and AAV-sgRNA techniques enable researchers to understand gene functions more precisely by allowing the investigation of the role of genes of interest at desired times in tissues/cells. Additionally, these combined techniques reduce the time and effort required to generate transgenic mice and allow the temporal control of CRISPR-Cas9 activity for inducible genome editing in mice if an inducible Cre line is used14.
AAV vectors are safe and effective in vivo gene delivery systems. However, AAVs targeting adipose tissue have lagged behind applications in other tissues, such as the brain, heart, liver, and muscle15. Due to the relatively low transduction efficiency and tropism with naturally occurring serotype vectors, AAV-guided gene delivery to adipose tissue is still challenging15. Over the last 5 years, we and others have successfully established effective and minimally invasive ways to deliver AAV-guided genes into adipose tissue and created mouse models that allow us to gain an understanding of the genes involved in the regulation of BAT function16,17,18,19. For example, by using AAV8 to deliver sgRNA targeting Alox12, which encodes 12-lipoxygenase (12-LOX), into the BAT of the Ucp1-Cre/Cas9 mice, we have discovered that activated BAT produces 12-LOX metabolites, namely 12-hydroxy-eicosapentaenoic acid (12-HEPE) and 13R, 14S-dihydroxy docosahexaenoic acid (maresin 2), to regulate glucose metabolism and resolve obesity-associated inflammation, respectively16,17. Here, we provide a step-by-step protocol on the technical procedures, particularly the surgery for the direct microinjection of AAV-sgRNA into the BAT lobes, to generate BAT-specific knockout mice using the combined Cre-LoxP, CRISPR-Cas9, and AAV-sgRNA system.
Protocol
All the animal experiments and care procedures were approved by the Institutional Animal Care and Use Committee at Joslin Diabetes Center.
1. Screening effective sgRNAs in cultured cells
NOTE: To make the assay cost-effective, before packaging the sgRNAs into AAV particles, we recommend testing different sgRNAs in cultured cells via a lentivirus-based system for Cas9/sgRNA expression (Figure 1) and selecting the sgRNAs that give the highest knockdown efficiency for in vivo experiments.
- CRISPR-Cas9 sgRNA design
- Design sgRNAs using online tools, such as the Broad sgRNA design tool (https://portals.broadinstitute.org/gppx/crispick/public)20,21, the CRISPOR online tool (http://crispor.tefor.net/)22, and other available tools.
- Enter gene names or DNA target sequences in the gene name box, and select NGG as the protospacer adjacent motif (PAM) for SpCas9 to generate potential sgRNA sequences.
- Select the sgRNAs with high predicted on-target efficiency and low off-target activity. For instance, sgRNAs with a specificity score of at least 50 on the CRISPOR output are recommended for use. Among the specific sgRNAs that pass the filter, pick the ones with high efficiency scores23.
NOTE: In general, three or four sgRNAs are picked to ensure the identification of effective sgRNAs.
- Construction of the CRISPR-Cas9 sgRNA plasmid
- Synthesize sgRNA oligo pairs encoding 20 nucleotide (nt) targeted sequences with overhangs (both 5' and 3') from the BsmBI restriction site (Table 1) through a DNA synthesis platform.
- Ligate annealed oligo pairs with BsmBI-linearized lentiCRISPR v2 vector.
- Transform the ligation product into the Stbl3 E. coli strain, and confirm the sgRNA insertions by Sanger DNA sequencing using U6 forward primer.
NOTE: See the detailed cloning procedure in Addgene's protocol entitled LentiCRISPRv2 and lentiGuide-Puro: lentiviral CRISPR/Cas9 and single guide RNA24.
- Producing lentiviral particles
- Approximately 24 h before transfection, plate 7 x 105 HEK-293 cells in 4 mL of complete growth medium in a 6 cm tissue culture plate.
- Add 15 µL of LT1 transfection reagent to 250 µL of serum-free medium, and incubate at room temperature for 5 min.
- Prepare a transfection cocktail for each sgRNA as per Table 2 in 250 µL of serum-free medium. Mix the transfection reagent prepared in step 1.3.2 with the plasmid cocktail, and incubate for 20-30 min at room temperature.
- Replace the complete growth medium with 3.5 mL of fresh growth medium for HEK-293 cells, and then add the DNA transfection reagent mixture dropwise to the HEK-293 cells cultured in the 3.5 mL of fresh growth medium.
- After 12-15 h of transfection, change the medium to remove the transfection reagent, and replace it with 4 mL of fresh growth medium.
- After 24 h of incubation, collect the cell culture medium that contains lentiviral particles, and filter the medium through a 0.45 µm filter to remove any HEK-293 cells. The viruses may be stored at 4 °C for a few days. For long-term storage, the viruses should be frozen at −80 °C.
NOTE: Follow biosafety guidelines when preparing lentiviral particles, and work in an environment (e.g., BL2+) suitable for handling lentivirus. See the detailed lentivirus preparation procedure in Addgene's protocol entitled pLKO.1 – TRC Cloning Vector (https://www.addgene.org/protocols/plko/#G).
- Infection of brown preadipocytes and determination of sgRNA-mediated knockdown
- Plate 1 x 106 mouse immortalized brown preadipocytes in 1 mL of fresh medium containing 8 µg/mL polybrene per well in a 6-well plate.
NOTE: In this study, the immortalized brown preadipocytes were generated using SV40 T antigen25. Brown preadipocytes are cultured in high-glucose DMEM with 10% FBS. - Add 1 mL of lentiviral particle solution from step 1.3 to infect the cells. Maintain one uninfected well of cells in parallel to serve as the antibiotic selection control.
- Change to fresh medium 24 h after infection. Add the corresponding antibiotics (e.g., puromycin with a final concentration of 1 µg/mL) to the medium to kill the uninfected cells and select the infected ones. Change to fresh medium containing the selected antibiotic every other day until all the uninfected control cells are dead.
- Collect protein from the virus-infected cells, and determine the knockdown efficiency of the sgRNAs by western blot analysis. Follow the detailed western blotting procedure in Eslami and Lujan26. Due to the varying turnover rate among proteins, test multiple time points (e.g., from day 6 to day 12 after virus infection) to observe the loss of protein signal.
NOTE: If an antibody that recognizes the protein encoded by the gene of interest is not available, Sanger sequencing can alternatively be utilized to determine the genome editing efficiency of each sgRNA. Briefly, amplify the genomic DNA using primers flanking the sgRNA-targeted region to generate PCR amplicons of ~700 bp in length. The projected break site should preferably be ~200 bp downstream from the sequencing start site. Then, subject the PCR product to Sanger sequencing. Analyze the sequencing results using the online tools, such as Tracking of Indels by DEcomposition (TIDE), to determine the frequency of small indels generated by each sgRNA in a pool of cells27.
- Plate 1 x 106 mouse immortalized brown preadipocytes in 1 mL of fresh medium containing 8 µg/mL polybrene per well in a 6-well plate.
2. Construction of the AAV-sgRNA plasmid
NOTE: In this step, the effective sgRNAs identified from the above screen are cloned into the pAAV-U6-BbsI-gRNA-CB-EmGFP vector28 (Figure 2). In the meantime, a non-targeting sgRNA (e.g., TCTGATAGCGTAGGAGTGAT29) that does not recognize any sequence in the mouse genome is also cloned into the same vector to serve as a non-edited control.
- Linearize pAAV-U6-BbsI-gRNA-CB-EmGFP backbone using BbsI, which generates identical overhangs with synthesized sgRNA oligos.
- Ligate the annealed oligo pairs with linearized pAAV vector, and confirm the sgRNA insertions as described in step 1.2.
NOTE: The cloning procedure for lentiCRISPRv2-sgRNA is also applied for the AAV-sgRNA plasmid construction.
3. AAV packaging
- Package the AAV vectors generated in section 2 into AAV serotype 8. Prepare high-titer AAV according to a previous protocol30, or request an AAV packaging service from viral cores or commercial services. Dilute the virus titer to 1 x 1013 genome copies/mL.
NOTE: The modified AAV2 genome expressing an sgRNA is packaged into AAV serotype 8. This serotype was chosen due to its high efficiency for adipose tissue18,31. To prevent immune response induced by contaminants, such as cell debris and small amounts of medium components, ultra-purified AAV is required for in vivo injection. Ultra-purification can be achieved by iodixanol gradients and ultracentrifugation30.
4. Preparation of Ucp1 Cre/Cas9 mice
- Purchase the Ucp1-Cre mouse strain (see Table of Materials) that expresses Cre recombinase under the control of the Ucp1 promoter. Purchase homozygous Rosa26-floxed STOP-Cas9 knock-in mice32 (see Table of Materials), in which the expression of Cas9 is regulated in a Cre recombinase-dependent manner.
- Cross these strains to generate Ucp1-Cre/Cas9 mice, as previously described16,17. Keep all the mice in a temperature- and humidity-controlled room (23 °C, 30% humidity) on a 12 h light-dark cycle (lights on at 6:30 am; lights off at 6:30 pm) with free access to chow diet and water.
- Treat the resulting Ucp1-Cre/Cas9 mice with AAVs carrying either control sgRNA or sgRNA targeting the gene of interest through the method described below. In this study, 12-15 week old male Ucp1-Cre/Cas9 mice weighing approximately 30 g were used.
5. Surgery for AAV injection into the BAT in mice
- Anesthetize the mice with continuous inhalation of 2.5% isoflurane for induction and maintenance.
- Lubricate both eyes to prevent drying. Then, give analgesics (banamine, 2.5 mg/kg body weight [BW], subcutaneous injection, see Table of Materials) to the mice to minimize and prevent postoperative pain and distress.
- Shave the mouse fur on the interscapular area using a shaver and sterilize the surgical area with at least three alternating rounds of an iodine-based or chlorhexidine-based scrub followed by 70% ethanol. Then, apply sterile surgical drapes around the incision site.
- Incise the skin between the scapulae using a scalpel (Figure 3A-B), and then peel the shaved skin off to expose the fat tissues on the neck using surgical scissors (Figure 3C-D). The size of the incision is dependent on the body size, but generally, the incision should be approximately 2 cm to expose the fat tissues on the interscapular region.
- Cut the fat tissues on the border between the distal region of the fat tissues and muscles using a single incision (Figure 3E-F). The size of the incision should be approximately 1 cm to just open the entrance for the insertion of the surgical scissors for blunt peeling of the fat tissues.
- Using the dominant hand, peel the fat tissues off bluntly and vertically with surgical scissors to expose the BAT while pinching the fat tissues, including the BAT, with the non-dominant hand (Figure 3G). Use the Sultzer's vein as a landmark to indicate the precise BAT location.
NOTE: The BAT in Figure 3H was completely exposed for the purpose of a picture demonstration. Excess dissection may cause damage to the nerves of the surrounding BAT. Incorrect direction of the scissors may hurt the surrounding muscles and the Sultzer's vein draining the BAT and could lead to excess bleeding. - For each mouse, inject 40 µL of AAV slowly into both BAT lobes (20 µL per one lobe of the BAT) using a sharp needle (32 G, see Table of Materials) connected to a Hamilton syringe (100 µL, see Table of Materials; Figure 3I). Check for successful injection as indicated by no leakages from the injected site during AAV administration.
NOTE: The BAT in Figure 3I was completely exposed for the purpose of a picture demonstration. The volume of AAV solution to be administered into a BAT lobe is determined by the size of the BAT of the recipient mouse. We have determined that 20 µL of AAV solution is appropriate for one BAT lobe weighing approximately 0.05 g in a regular C57BL6/J mouse weighing 30 g. If necessary, the viral solution may need to be concentrated to achieve the desired volume. - Confirm no bleeding from the injected BAT or surrounding tissues. Press gauze soaked with hydrogen peroxide solution (see Table of Materials) onto a bleeding site for hemostasis.
- Place the exposed fat tissues back to their normal position. Stitch up the edge of the fat tissues and muscle using a 5-0 absorbable monofilament thread (Figure 3J). Suture the open skin with 5-0 coated vicryl undyed braided threads (Figure 3K-L).
NOTE: Figure 4 describes the steps for stitching the peeled fat tissues and muscles. - Maintain the animals on a heating pad following the surgery until complete recovery. Provide the animals with access to food, water, and ample bedding for nesting. Monitor them regularly for any signs of distress or illness for 7 days of recovery prior to the beginning of the experiment.
- Confirm the efficiency of the targeted gene deletion by western blot analysis in the BAT dissected from mice, as in Sugimoto et al.16.
Representative Results
Throughout the above procedures, the precise delivery of AAV-sgRNA to the BAT is crucial to the success of the protocol. To maximize the effect of AAV-sgRNA and minimize the tissue damage during surgery, it is essential to understand the three-dimensional (3D) anatomical location of the BAT. As shown in Figure 3E, it is hard to identify the precise location of the BAT without exposing the tissue. However, excess BAT exposure may damage the tissue (Figure 3H, I); thus, the procedure should be performed to inject the AAV solution within a limited anatomical space (Figure 3G). Figure 5A is critical to ensure the success of the AAV injection and the rapid recovery of the mice after the surgery. Figure 5B–D indicates potential flaws that may lead to unsuccessful injections. These include injecting the AAV solution into the incorrect region (Figure 5B), needle penetration through the tissue (Figure 5C), and tissue ballooning due to an excessive volume of the AAV solution (Figure 5D). Lastly, the knockdown efficiency of the gene of interest needs to be verified by the protein levels in the BAT dissected from the injected mice16. For example, the protein levels of 12-LOX in the BAT measured by western blot demonstrate efficient knockdown due to AAV8 delivering sgRNA targeting Alox12 (the gene that encodes 12-LOX) into the BAT of Ucp1-Cre/Cas9 mice (see Figure 7b in the previously published study16).
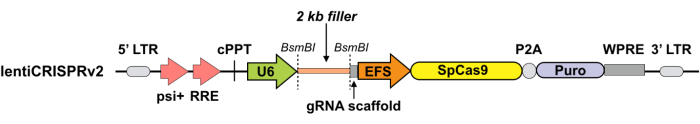
Figure 1: Schematic for the lentiviral plasmid expressing Cas9 and sgRNA. Lentiviral expression vector for Cas9 and sgRNA (lentiCRISPRv2). Abbreviations: Puro = puromycin selection marker; psi+ = psi packaging signal; RRE = rev response element; cPPT = central polypurine tract; EFS = elongation factor-1α short promoter; P2A = 2A self-cleaving peptide; WPRE = posttranscriptional regulatory element; LTR = long terminal repeat. LentiCRIPSRv2 can be digested using BsmBI, and a pair of annealed sgRNA oligos can be cloned into the single-guide RNA scaffold. Please click here to view a larger version of this figure.
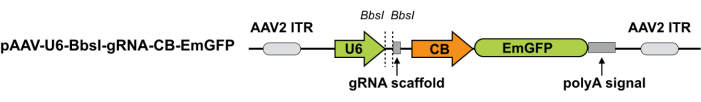
Figure 2: Schematic for the AAV plasmid expressing sgRNA. The AAV vector for sgRNA. Abbreviations: ITR = I terminal repeat; CB = hybrid CMV enhancer/β-actin promoter. pAAV-U6-BbsI-gRNA-CB-EmGFP can be digested using BbsI, and a pair of annealed sgRNA oligos can be cloned into the single-guide RNA scaffold. Please click here to view a larger version of this figure.
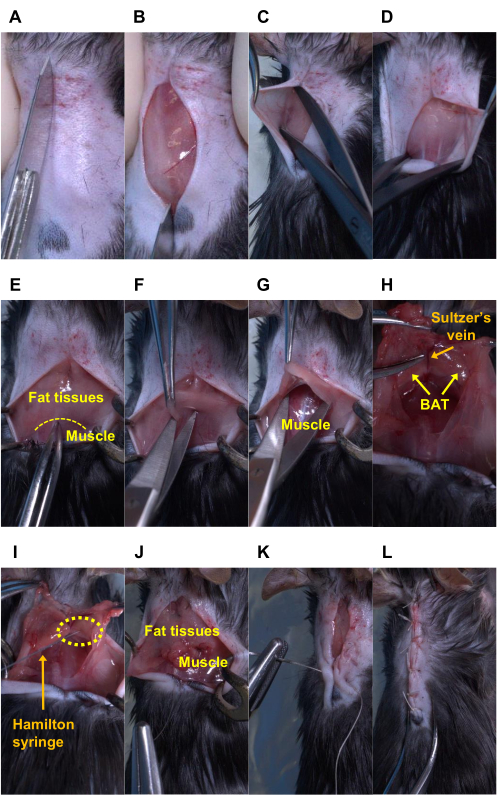
Figure 3: Sequential surgery for AAV injection into the bilateral BAT lobes in mice. An anesthetized mouse has undergone the following procedures: (A-B) cutting the shaved skin, (C-D) bluntly peeling the shaved skin off, (E-F) cutting the fat tissues, (G) bluntly peeling the fat tissues off, (H-I) injecting the AAV solution into the BAT lobes using a Hamilton syringe, (J) suturing between the fat tissues and muscle and (K-L) between both skin flaps. Note: In panels (H) and (I), the Sultzer’s vein is a landmark to confirm the precise location of the BAT. The BAT in those two panels was exposed entirely for picture demonstration. The yellow dotted lines indicate the border between fat tissues and muscle in panel E, and the tip of the needle (i.e., the injection site) in panel I. Please click here to view a larger version of this figure.
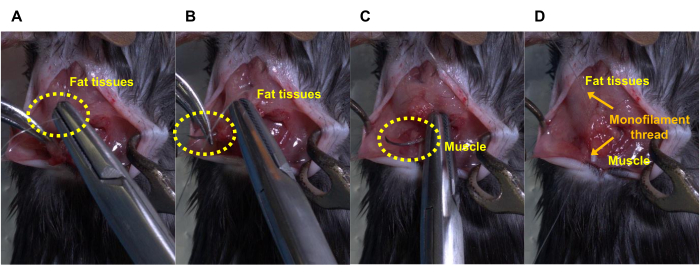
Figure 4: Steps for stitching the peeled fat tissues and muscles. (A) Insert a needle thread into the peeled fat tissues for injection, (B) penetrate the fat tissues, (C) hook a muscle, (D) pull a thread up to stitch the peeled fat tissues and muscle, and then ligate the threads. Please click here to view a larger version of this figure.
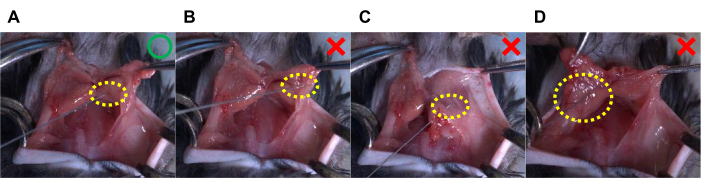
Figure 5. Representative pictures of proper and inappropriate means of AAV injection into the BAT lobes in mice. (A) This figure represents the suggested AAV injection, in which a unilateral BAT lobe receives an injection of 20 µL of AAV solution. (B–D) The following methods of AAV injection can lead to failure. (B) Incorrect injection site (i.e., missing the BAT lobes); (C) the needle penetrates through the tissues; and (D) tissue ballooning due to a large amount of AAV solution, in which a unilateral BAT lobe receives an 80 µL injection of AAV solution. The yellow dotted circles indicate the tip of the needle (i.e., the injection site). Please click here to view a larger version of this figure.
| Forward oligo: 5' CACC-20bp gRNA sequence-3' |
| Reverse oligo: 5' AAAC-20bp gRNA reverse-complement sequence-3' |
| For example, if the target sequence is TCTGATAGCGTAGGAGTGAT, the oligos would be: |
| Forward oligo: 5' CACC TCTGATAGCGTAGGAGTGAT 3' |
| Reverse oligo: 5' AAAC ATCACTCCTACGCTATCAGA 3' |
Table 1: Example of sgRNA design. Example of an sgRNA sequence with overhangs corresponding to the BsmBI restriction site.
| Name of Material | Company | Catalog Number | Amount |
| lentiCRISPER v2 plasmid | Addgene | #52961 | 5 μg |
| psPAX2 packaging plasmid | Addgene | #12260 | 3.75 μg |
| pMD2.G envelope plasmid | Addgene | #12259 | 1.5 μg |
| OPTI-MEM serum-free medium | Invitrogen | #31985 | 250 μl |
Table 2: LentiCRISPR. Plasmid cocktail for the transfection of the sgRNA containing the LentiCRISPRv2 plasmid into HEK-293 cells to produce CRISPR-sgRNA lentivirus.
Discussion
The existing published methods for AAV-mediated gene delivery to the interscapular BAT do not contain detailed pictures and videos describing the surgical techniques and approach for direct AAV injection into the BAT. In most of the published methods33,34, the AAV is injected into the fat tissues surrounding the interscapular BAT instead of the BAT itself. Hence, there are several key steps in this protocol that determine the success of the study. These include the design of the sgRNA, the AAV packaging and concentration, the generation of the BAT-specific Cas9 mice, and the surgical procedures. Careful and outstanding surgical skills are required to complete the in vivo procedures. In particular, minimizing the tissue damage while exposing the BAT and administering the AAV is critical. Due to multiple genetic and environmental factors, such as housing temperature, exercise training, and diets, the volume of white fat surrounding the BAT and the composition of BAT itself can vary among animals. For example, obese mice display vast white fat tissues surrounding the BAT, and mice housed at thermoneutrality (30 °C) exhibit whitened BAT. Hence, it is challenging to precisely determine the 3D location of the BAT without opening the skin. Minimally revealing the BAT is a challenging but crucial step for AAV injection. Excessive exposure of the BAT during this procedure can cause severe damage to the sympathetic nerves and vessels, particularly the Sultzer's vein connected to the BAT.
Of note, the volume of AAV injected into the tissue is also a limiting factor for success. An excessive volume of the solution can be harmful to the cells and interfere with an efficient viral infection to deliver the sgRNA into the adipocytes. As BAT size varies with the age, sex, and genetic background of the recipient mice, optimization experiments may be required16,17. In addition to knocking down genes of interest, this technique can be additionally applied to overexpress genes of interest.
One potential pitfall of this technique is the low efficiency of gene knockout using one sgRNA in vivo, even with sgRNAs that provide efficient knockout in vitro. As a potential solution, combining two or more unique sgRNAs into a single injection can improve the efficiency of gene knockdown in vivo.
In conclusion, the combined Ucp1-Cre-LoxP, CRISPR-Cas9, and AAV-sgRNA technologies offer a robust and efficient system to generate mice with BAT-specific gene manipulation. This method can be adapted for other tissues using selected Cre lines and modified AAV injection into different tissues.
Declarações
The authors have nothing to disclose.
Acknowledgements
This work was supported in part by U.S. National Institutes of Health (NIH) grants R01DK122808, R01DK102898, and R01DK132469 (to Y.-H.T.), as well as P30DK036836 (to Joslin Diabetes Center's Diabetes Research Center). T. T. was supported by the SUNSTAR Research Fellowship (Hiroo Kaneda Scholarship, Sunstar Foundation, Japan) and the American Heart Association grant 903968. Y. Z. was supported by the Charles A. King Trust Fellowship. We thank Sean D. Kodani for kindly proofreading the manuscript.
Materials
| Chemicals, Peptides and Recombinant Proteins | |||
| OPTI-MEM serum-free medium | Invitrogen | 31985 | |
| Polybrene Infection / Transfection Reagent | Sigma | TR-1003-G | |
| TransIT-LT1 Transfection Reagent | Mirus | MIR 2300 | |
| Recombinant DNA | |||
| lentiCRISPR v2 vector | Addgene | 52961 | |
| pAAV-U6-BbsI-gRNA-CB-EmGFP | Addgene | 89060 | |
| pMD2.G envelope plasmid | Addgene | 12259 | |
| psPAX2 packaging plasmid | Addgene | 12260 | |
| Experimental Models: Cell Lines | |||
| HEK-293 cells | ATCC | CRL-1573 | |
| WT-1 cells | Developed in Tseng lab | PMID: 14966273 | |
| Experimental Models: Organisms/Strains | |||
| Cas9 knockin mice | Jackson Laboratories | 24857 | |
| Ucp1-CRE mice | Jackson Laboratories | 24670 | |
| Others | |||
| Banamine | Patterson | 07-859-1323 | |
| Hamilton syringe | Hamilton | Model 710 RN Syringe | |
| Hydrogen peroxide solution | Fisher Scientific | H325-500 | |
| Needle | Hamilton | 32 gauge, small hub RN needle, needle point style 2 | |
| Suture 5-0 Unify PCL25 P-3 Undyed 18 inch Monofilament 12/Bx | Henry Schein | 1273504 | |
| Suture 5-0 Unify Silk P-3 Black 18 inch 12/Bx | Henry Schein | 1294522 |
Referências
- Kopelman, P. G. Obesity as a medical problem. Nature. 404 (6778), 635-643 (2000).
- Finkelstein, E. A., et al. Obesity and severe obesity forecasts through 2030. American Journal of Preventive Medicine. 42 (6), 563-570 (2012).
- Craft, S. Insulin resistance and Alzheimer’s disease pathogenesis: Potential mechanisms and implications for treatment. Current Alzheimer Research. 4 (2), 147-152 (2007).
- Villarroya, F., Cereijo, R., Villarroya, J., Giralt, M. Brown adipose tissue as a secretory organ. Nature Reviews Endocrinology. 13 (1), 26-35 (2017).
- Scheele, C., Wolfrum, C. Brown adipose crosstalk in tissue plasticity and human metabolism. Endocrine Reviews. 41 (1), 53-65 (2020).
- Becher, T., et al. Brown adipose tissue is associated with cardiometabolic health. Nature Medicine. 27 (1), 58-65 (2021).
- Kajimura, S., Spiegelman, B. M., Seale, P. Brown and beige fat: Physiological roles beyond heat generation. Cell Metabolism. 22 (4), 546-559 (2015).
- Scheja, L., Heeren, J. The endocrine function of adipose tissues in health and cardiometabolic disease. Nature Reviews Endocrinology. 15 (9), 507-524 (2019).
- Darcy, J., Tseng, Y. H. ComBATing aging-does increased brown adipose tissue activity confer longevity. Geroscience. 41 (3), 285-296 (2019).
- da Silva Xavier, G., Hodson, D. J. Mouse models of peripheral metabolic disease. Best Practice & Research Clinical Endocrinology & Metabolism. 32 (3), 299-315 (2018).
- Kim, H., Kim, M., Im, S. K., Fang, S. Mouse Cre-LoxP system: General principles to determine tissue-specific roles of target genes. Laboratory Animal Research. 34 (4), 147-159 (2018).
- Wang, D., Zhang, F., Gao, G. CRISPR-based therapeutic genome editing: Strategies and in vivo delivery by AAV vectors. Cell. 181 (1), 136-150 (2020).
- Sander, J. D., Joung, J. K. CRISPR-Cas systems for editing, regulating and targeting genomes. Nature Biotechnology. 32 (4), 347-355 (2014).
- Dow, L. E., et al. Inducible in vivo genome editing with CRISPR-Cas9. Nature Biotechnology. 33 (4), 390-394 (2015).
- Bates, R., Huang, W., Cao, L. Adipose tissue: An emerging target for adeno-associated viral vectors. Molecular Therapy. Methods & Clinical Development. 19, 236-249 (2020).
- Sugimoto, S., et al. Brown adipose tissue-derived MaR2 contributes to cold-induced resolution of inflammation. Nature Metabolism. 4 (6), 775-790 (2022).
- Leiria, L. O., et al. 12-lipoxygenase regulates cold adaptation and glucose metabolism by producing the omega-3 lipid 12-HEPE from brown fat. Cell Metabolism. 30 (4), 768-783 (2019).
- Shamsi, F., et al. FGF6 and FGF9 regulate UCP1 expression independent of brown adipogenesis. Nature Communications. 11, 1421 (2020).
- Romanelli, S. M., et al. BAd-CRISPR: Inducible gene knockout in interscapular brown adipose tissue of adult mice. Journal of Biological Chemistry. 297 (6), 101402 (2021).
- Doench, J. G., et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR-Cas9. Nature Biotechnology. 34 (2), 184-191 (2016).
- Sanson, K. R., et al. Optimized libraries for CRISPR-Cas9 genetic screens with multiple modalities. Nature Communications. 9, 5416 (2018).
- Concordet, J. P., Haeussler, M. CRISPOR: Intuitive guide selection for CRISPR/Cas9 genome editing experiments and screens. Nucleic Acids Research. 46, 242-245 (2018).
- Haeussler, M., et al. Evaluation of off-target and on-target scoring algorithms and integration into the guide RNA selection tool CRISPOR. Genome Biology. 17 (1), 148 (2016).
- Sanjana, N. E., Shalem, O., Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nature Methods. 11 (8), 783-784 (2014).
- Tseng, Y. H., Kriauciunas, K. M., Kokkotou, E., Kahn, C. R. Differential roles of insulin receptor substrates in brown adipocyte differentiation. Molecular and Cellular Biology. 24 (5), 1918-1929 (2004).
- Eslami, A., Lujan, J. Western blotting: Sample preparation to detection. Journal of Visualized Experiments. (44), e2359 (2010).
- Brinkman, E. K., Chen, T., Amendola, M., van Steensel, B. Easy quantitative assessment of genome editing by sequence trace decomposition. Nucleic Acids Research. 42 (22), 168 (2014).
- Jarrett, K. E., et al. Somatic genome editing with CRISPR/Cas9 generates and corrects a metabolic disease. Scientific Reports. 7, 44624 (2017).
- Zhang, Y., et al. Optimized RNA-targeting CRISPR/Cas13d technology outperforms shRNA in identifying functional circRNAs. Genome Biology. 22 (1), 41 (2021).
- Fripont, S., Marneffe, C., Marino, M., Rincon, M. Y., Holt, M. G. Production, purification, and quality control for adeno-associated virus-based vectors. Journal of Visualized Experiments. (143), e58960 (2019).
- Jimenez, V., et al. In vivo adeno-associated viral vector-mediated genetic engineering of white and brown adipose tissue in adult mice. Diabetes. 62 (12), 4012-4022 (2013).
- Platt, R. J., et al. CRISPR-Cas9 knockin mice for genome editing and cancer modeling. Cell. 159 (2), 440-455 (2014).
- Huang, W., Queen, N. J., Cao, L. rAAV-mediated gene delivery to adipose tissue. Methods in Molecular Biology. 1950, 389-405 (2019).
- Xue, K., Wu, D., Qiu, Y. Specific and efficient gene knockout and overexpression in mouse interscapular brown adipocytes in vivo. STAR Protocols. 3 (4), 101895 (2022).

