Noninvasive Monitoring of Lesion Size in a Heterologous Mouse Model of Endometriosis
Summary
Here, we present a protocol for live-imaging of fluorescently labeled human endometrial fragments grafted in mice. The method allows studying the effects of drugs of choice on endometriotic lesion size through monitoring and quantification of fluorescence emitted by the fluorescent reporter on real time
Abstract
Here, we describe a protocol for the implementation of a heterologous mouse model in which progression of endometriosis can be assessed in real time through noninvasive monitoring of fluorescence emitted by implanted ectopic human endometrial tissue. For this purpose, biopsies of human endometrium are obtained from donor women ongoing oocyte donation. Human endometrial fragments are cultured in the presence of adenoviruses engineered to express cDNA for the reporter fluorescent protein mCherry. Upon visualization, labeled tissues with an optimal rate of fluorescence after infection are subsequently chosen for the implantation in recipient mice. One week prior to the implantation surgery, recipient mice are oophorectomized, and estradiol pellets are placed subcutaneously to sustain the survival and growth of lesions. On the day of surgery mice are anesthetized, and peritoneal cavity accessed through a small (1.5 cm) incision by the linea-alba. Fluorescently labeled implants are tweezed, briefly soaked in glue and attached to the peritoneal layer. Incisions are sutured, and animals left to recover for a couple of days. Fluorescence emitted by endometriotic implants is usually non-invasively monitored every 3 days for 4 weeks with an in vivo imaging system. Variations in the size of endometriotic implants can be estimated in real time by quantification of the mCherry signal and normalization against the initial time-point showing maximal fluorescence intensity.
Traditional preclinical rodents of models of endometriosis do not allow non-invasive monitoring of lesion in real time but rather allow evaluation of the effects of drugs assayed at the end point. This protocol allows one to track lesions in real time and is more useful to explore the therapeutic potential of drugs in preclinical models of endometriosis. The main limitation of the model thus generated is that non-invasive monitoring is not possible over long periods of time due to the episomal expression of Ad-virus.
Introduction
Endometriosis is a chronic gynecologic disorder initiated by the implantation of the functional endometrium outside the uterine cavity. Ectopic lesions grow and induce inflammatory processes leading to chronic pelvic pain and infertility1. It is estimated that up to 10–15% of women of reproductive age are affected by endometriosis2, and it is present in approximately 40–50% of infertile women3. Current pharmacological treatments for endometriosis are unable to completely eradicate lesions and not free of side effects4,5. The research for more efficient therapies requires of the refinement of the existing animal models of endometriosis in such a way that human lesions can be appropriately mimicked, and the effects of compounds on lesion size among others can be closely assessed.
Primate models have been used to mimic endometriosis by implanting ectopic lesions histologically identical and at similar sites as in humans6,7,8; however, ethical concerns and the high economic costs related to experimentation with primates limit their use9. Consequently, the use of small animals, especially rodents, for the implementation of in-vivo models of endometriosis continues to be favored as it allows studies with larger numbers of individuals10,11. Endometriosis can be induced in these animals by transplanting either pieces of rodent uterine horns ("homologous models")12,13 or human endometrial/endometriotic tissue to ectopic sites (heterologous models)14. In contrast to humans, rodents do not shed their endometrial tissue and thereby endometriosis can not be developed spontaneously in these species. Therefore, homologous mouse models of endometriosis have been criticized due to the fact that implanted ectopic mouse uterine tissue does not reflect the characteristics of human endometriotic lesions15.
Appropriate physiology of endometriosis can be mimicked in the heterologous models of endometriosis where fresh human endometrial fragments are implanted into immunodeficient animals. In conventional heterologous models, the therapeutic effects of compounds of interest are commonly assessed at the end point by the assessment of lesion size with the use of calipers16. An obvious limitation is that, as such, endpoint animal models do not allow studying implantation dynamics or endometriotic lesion development over time. An additional limitation is that the use of calipers does not allow accurate measurements of lesion size. Indeed, the standard error provided by calipers is in the same range (i.e., millimeters) as the size of the lesions implanted in mice, thus restricting the capacity of these tools to detect actual variations in size.
In order to overcome such limitations, herein, we describe the generation of a heterologous mouse model of endometriosis in which implanted human tissue is engineered to express a reporter m-Cherry fluorescent protein. Detection of the fluorescent signal with an appropriate image system enables non-invasive monitoring of lesion status with simultaneous quantification of its size in real time. Thus, our model provides clear advantages when compared to conventional endpoint models as it brings the opportunity of real-time non-invasive monitoring and the possibility to perform more objective and accurate estimation of variations in lesion size.
Protocol
The use of human tissue specimens was approved by the Institutional Review Board and Ethics Committee of the Hospital Universitario La Fe. All patients provided written informed consent. The study involving animals was approved by the Institutional Animal Care Committee at the Centro de Investigacion Principe Felipe de Valencia, and all procedures were performed following the guidelines for the care and use of mammals from the National Institutes of Health.
1. Endometrial Tissue Collection and Pre-processing
- Obtain a good quality biopsy of human endometrial aspirate by using a cannula attached to a suction device. Pour the biopsy into a flask containing 10 mL of sterile saline and wash with gentle manual agitation.
NOTE: Procedures on how to obtain good quality biopsies have been previously described17. - Wash the biopsy of any remaining blood or mucus. Repeat the process by pouring the tissue to flasks containing fresh saline as many tines as required until tissue is observed clean.
- Transfer biopsy fragments with a healthy appearance into a solution containing 10 mL of complete Dulbecco's Modified Eagle's medium (DMEM) medium with 10% Fetal Bovine Serum (FBS) and 1% antibiotic-antimycotic solution.
- Pour the content in a 10 cm Petri dish and proceed to chop the tissue into 5–10 mm3 pieces with a pair of scalpels.
2. Adenoviral Transfection of Endometrial Fragments
NOTE: All the materials that are going to be employed in the process should be introduced in the hood in advance. Take out everything that is not going to be used in the process and place a flask with bleach. All material that comes into contact with the adenoviral vector must be disinfected with the bleach before discarding it in the biohazard container.
- Once the biopsy has been chopped, take a new Petri dish. Pipette multiple 30–50 µL DMEM drops of spread throughout the whole dish. Leave enough space between drops so they do not come into contact.
- Aided with a needle attached to syringe, place a single piece of fragment inside each of the medium drops.
NOTE: Each drop should contain a single piece of endometrium. - Prepare the Ad-mCherry working solution by diluting 1:20 of the mCherry adenoviral stock solution [1 x 1010 pfu/mL]) into DMEM medium WITHOUT antibiotics (DMEM + 10% filtered FBS).
- Dispense 100–200 μL of the Ad-mCherry solution per well on a 96-well plate, filling as many wells as fragments are available in the Petri dish drops (step 2.2). In addition, fill at least 3 wells with adenovirus free DMEM medium as a negative control.
- Aided with a needle attached to syringe, transfer each of the fragments in the Petri dish (step 2.2) to each of the wells containing ad-mCherry solution in the 96-well plate.
- Place the 96 well-plate containing the endometrial fragments with Ad-mCherry solution into an incubator at 37 °C with 5% CO2 for 16 h.
- Wash out the remaining adenoviruses in the medium by transferring tissues into a new 96-well plate filled with complete DMEM medium (WITH antibiotics-antimycotics, free of adenovirus) and incubate for 37 °C with 5% CO2 for 24–48 h.
NOTE: Remember to cover with bleach all well plates and tips that came in contact with Ad-virus before discarding into the biohazard container. - Take out the plate from the incubator and place it under a fluorescence microscope at 568 nm (red channel) to test for optimal labeling.
- Choose the most brilliant fragments and place them into a new 96-well plate with fresh complete DMEM medium (WITH antibiotics-antimycotics, free of adenovirus).
- Seal the plate well with a plastic paraffin film and transport it to into the specific-pathogen-free animal area for implantation of endometrial fragments into recipient animals.
3. Generation of the Endometriosis Mouse Model
NOTE: Use 6–8 week-old athymic nude (or similar immunocompromised strains) female mice housed in specific pathogen–free conditions, as recipient animals. To avoid hormonal cycle-dependent variations and simultaneously fuel lesion growth with estradiol, animals are ovariectomized and placed with 60-day release capsules containing 18 mg of 17 βeta-Estradiol (17β-E2). Oophorectomy and pellet placement have to be performed at least one week in advance of grafting endometrial fragments into the recipient animals.
- Oophorectomy
NOTE: Prepare surgical sterilized material in a hood. Prepare anesthesia equipment and a post-surgical recovery zone ready in the same room.- Perform a subcutaneous injection of morphine derivative at a dose of 5 mg/kg per mouse. Let the mice rest for 30 min after injection so as analgesics effects of drug can be manifested.
- Connect the inhalation anesthesia equipment and let oxygen and isoflurane (2% mg/kg) flow for a few min into a sealed anesthesia chamber.
- Introduce the animal into the isoflurane anesthesia chamber. Wait for 3–5 min and check that animals are fully anesthetized by pressing one of its paws. Transfer the animal to the surgery area and maintain anesthesia by placing a mask with continuous flow of isoflurane gas covering the respiratory airways.
- After disinfecting the area with chlorohexidine, perform a transverse 0.5 cm costal incision approximately at the height of the hip with sharp scissors.
- Separate the skin from the muscle to get access to the abdominal cavity, identify the white fat pad that surrounds the ovary and retract the ovary with dissection forceps.
- Tie a knot around the oviduct with absorbable suture and tighten it to ensure appropriate hemostasis before excising the ovary.
- Close the muscular layer with 6-0 absorbable suture, and then close the skin with 6-0 non-absorbable suture. Clean the area again with antiseptic solution.
- Repeat the procedure to remove the contra lateral ovary.
- Estradiol pellet implant
NOTE: Take care that the animal is anesthetized during the oophorectomy surgery to place pellets at that point.- Immediately after the completion of the oophorectomy procedure, clean the skin with antiseptic solution surrounding the neck and make a transverse subcutaneous small (0.5 cm) incision with sharp scissors in the nape.
- Use the scissors to dissect the skin from the muscle, making a pocket large enough to allow placing the pellets.
- Insert the pellet containing 18 mg of 17β-E2 and suture the skin with a 6-0 non-absorbable suture. Clean the area again with antiseptic solution.
- Place the animal in the recovery zone and administer an optimal dose of long-lasting analgesia to ease the recovery period.
- Endometrial implant surgery
NOTE: Allow at least a period of seven days quarantine to allow full recovery of animals after oophorectomy before starting endometrial fragment implantation surgery. For optimal synchronization with labeling of tissue, collect the biopsy 2–3 days before implantation surgery so as to avoid long-term culture of explants.- Get the surgical room in the specific pathogen free zone ready in advance. Prepare the hood with all required surgical material, the anesthesia equipment and the post-surgical recovery zone also.
- Bring the animals to the room, perform a subcutaneous injection of a morphine derivative at a dose of 5 (mg/kg) in each mouse. Let the mice rest for 30 min after injection so analgesics effects of drug can be manifested.
- Connect the inhalation anesthesia equipment and let oxygen and isoflurane (2% mg/kg) flow for a few min into a sealed anesthesia chamber.
- Before starting surgery, move the plate containing fluorescently labeled fragments (from step 2.10) into the hood, unseal it and pour the fragments into a Petri dish for easier handling.
- Introduce the animal into the isoflurane anesthesia chamber. Wait for 3–5 min and check that animals are fully anesthetized by pressing one of its paws. Transfer the animal to the surgery area and maintain anesthesia by placing a mask with continuous flow of isoflurane gas covering the respiratory airways.
- Place the anesthetized animal face up. Disinfect the ventral area. Perform a longitudinal 1.5 cm incision in abdomen with sharp scissors and separate the skin from the muscle. Then, perform a longitudinal 1.5 cm incision in the muscle to access the peritoneal cavity.
- Hold the left edge of the abdomen muscular wall with mini-forceps and fold it trying to expose the inner face of the peritoneum on the outside.
- Take an endometrial implant with mini tweezers, soak it briefly in an n-butyl-ester cyanoacrylate adhesive and place it to the peritoneum where it will get attached. Let it dry for a few seconds. Repeat steps 3.3.6 and 3.3.7 to place an implant on the contralateral side of the peritoneum.
- Close the muscular layer with an absorbable 6-0 suture, and then close the skin with a non-absorbable 6-0 suture. Clean the area again with antiseptic solution.
- Place the animal in the recovery zone and administer an optimal dose of long-lasting analgesia.
4. In Vivo Fluorescent Imaging with an In Vivo Imaging System
- Turn on the in vivo imaging system device, initialize the program and allow the CCD camera to cool down for a few min.
- Prepare the inhalation anesthesia equipment. Open the isoflurane flow at 2% for a couple of min to fill the anesthetic chamber. Prepare the post-anesthesia recovery zone.
- Once the program has started and the CCD camera have cooled down, click the Imaging Wizard tool: A tutorial starts with a series of consecutive windows displayed, each one corresponding to a parameter of interest with several options available to be chosen by clicking in the corresponding box. Move forward through the tutorial by selecting the appropriate boxes in each window and click the OK button to move to the next set of parameters with the following sequence.
- Select the Epiluminescence box for the fluorescence parameter, select the mCherry box for the filter pairs parameter. Check the Photograph mode and Confirm Focus boxes and select the Automatic box for the exposure parameter.
- Once the instrument has been set up, move one animal inside the anesthesia chamber. When fully anesthetized, transfer the animal inside the in vivo imaging system cage and place it side up with its head inside a tubule connected to the anesthesia machine. Close the lid and click Acquire for monitoring.
- Acquire the images appearing (a total of five images, one image for each pair of filters selected) and save data by clicking the Save As button. Move the mouse to the area of post-anesthesia recovery. Repeat the process with the remaining animals.
- Repeat monitoring two or three times a week to follow-up the signal appropriately during the time course.
- Proceed to sacrifice the animal at the end of the time course by CO2 asphyxiation.
5. Quantification of In Vivo Fluorescence Images
- Segregation of actual fluorescence through image unmixing:
- Open the In Vivo Imaging Analysis Coupled Software Program.
- Choose the “Sequenceinfo” file to start the analysis. Two windows will appear: “Sequence view” and “Tool Palette”. Choose Tool Palette and once the menu has been displayed select the following options.
- Select Corrections and click on the box Adaptive FL Background Subtraction to remove the undesirable fluorescent signals from the luminescent image data. Choose the threshold of greatest interest and click Set.
- Click Spectral Unmixing, select the wavelengths of interest, the method chosen for unmixing (library, guided, automatic or manual) and then click Start Unmix.
- Select the Unmixed image corresponding to mCherry signal and double click. A new window will appear with the final image of the signal of interest.
- Repeat step 5.1.3.
- OPTIONAL: If a representative image (JPEG) of the unmix result is needed, choose the desired settings in Tool Palette | Image Adjust (color table, binning, contrast, etc.), and then click on Export Graphics in the unmix window to export the current image view as an image.
- Save the unmixed file: File | Save As | Choose Folder and Ok.
- Repeat the process with the rest of the monitoring days and with all animals.
- ROIs set up and signal quantification
- Click Browse and select the Unmixed file (see step 5.1.8) of interest to be analyzed. A new window will appear.
- Click Add To List to include all the unmixed files from each animal at different time points and then click on Load As A Group. All images must appear as a single sequence.
- Go to Tool Palette window: Click off the box Individual scale to obtain all images on the same scale.
- Double click in one image of the sequence and create a ROI on the zone of interest using the following sequence. Go to ROI Tools and select Countour and Auto 1 option, click on the circle shape appearing, place it on the center the fluorescent signal and then click Create on the displayed window.
Note: This automatically highlights pixels with an intensity of fluorescence above background values (i.e., lesion) and generates a shape whose area embraces the outlined pixels. - Copy the created ROI shape and paste on a background zone where there is no signal.
- Click on Measure ROIs | Select All to display values of fluorescence intensity. Proceed to select data values with the mouse, click right button, press copy and paste on a spreadsheet.
6. Data (fluorescent signal) Normalization
- Select the initial time point at which signal intensity is maximal. Proceed to normalize signal at each time point by using the formula:
Signal intensity at each time point / Maximal signal intensity observed during the time course) x 100.
Representative Results
Here, we describe the process for creating a heterologous model of endometriosis in which the architecture of lesions is preserved by implanting fluorescently labeled pieces of human endometrium into immunocompromised mice, thus allowing non-invasive monitoring of lesion progression. Labeling of endometrial fragments is achieved by infection with adenovirus engineered to express mCherry, a protein emitting fluorescence in the near infrared region. In Figure 1, we show representative images of human endometrial fragments infected with Ad-mCherry observed under the fluorescence microscope. For illustrative purposes, both labeled and non-labeled fragments are included so differences in fluorescence between infected and non-infected tissues (autofluorescence) can be noted. During monitoring, in addition to the reference wavelength for mCherry, fluorescent images are taken with different pairs of excitation/emission wavelengths filters (Figure 2) to define the characteristic fluorescent emission profile of tissues. The purpose of this action is to "unmix" actual fluorescence emitted by lesions from background and autofluorescence emitted by host tissues and scar originated during surgery respectively. An illustrative example of the unmix process is shown in Figure 3. Estimations of variation in lesion size is performed by quantifying and normalizing fluorescent signaling emitted by lesions during the time course. For this purpose, images of monitoring containing raw fluorescence emitted by animals during each time point are first brought together unnormalized (Figure 4) in a single file. Subsequently fluorescence is unmixed, normalized and represented as a false color image (Figure 5). Finally, ROIs corresponding to specific lesion and background signaling are automatically recognized by the program and quantified (Figure 6). Background ROI signaling is subtracted from lesion ROI signaling and results of intensity in each time point are normalized against the time point at which intensity is maximal (Figure 7). At the end of the monitoring process, several weeks after surgery mice are sacrificed and viable implant can be recovered attached to the mouse peritoneum (Figure 8).
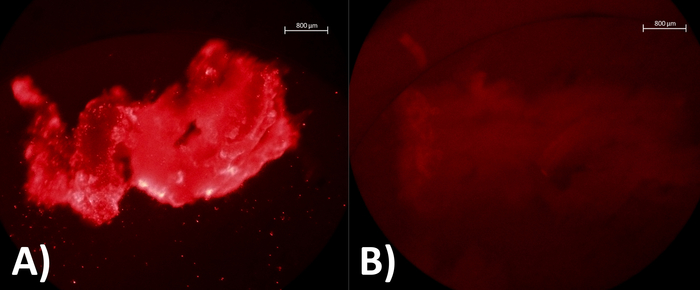
Figure 1: Visualization of endometrial fragments with fluorescence microscope after Ad-mCherry infection. (A) Human endometrial fragment incubated with Ad-mCherry at 37 °C and 5% CO2 during 24 h as a positive sample. (B) Human endometrial fragment incubated at 37 °C and 5% CO2 without Ad-mCherry as a negative control sample. Please click here to view a larger version of this figure.
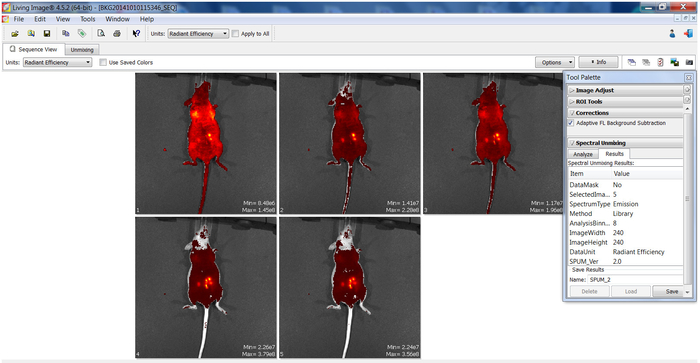
Figure 2: Raw imaging fluorescence emitted by labeled fragments implanted in mice. Picture shows representative imaging of raw fluorescence signal emitted by the same animal at a specific time point. Images correspond to screenshots obtained using software coupled to an in vivo imaging system device during a monitoring session. Each panel containing mice (numbered 1–5 in the left corner) corresponds to the fluorescence observed by using a different specific excitation/emission pair filter for acquiring images. Panel on the right (tool palette) show fluorescence parameters selected for acquisition of images Please click here to view a larger version of this figure.
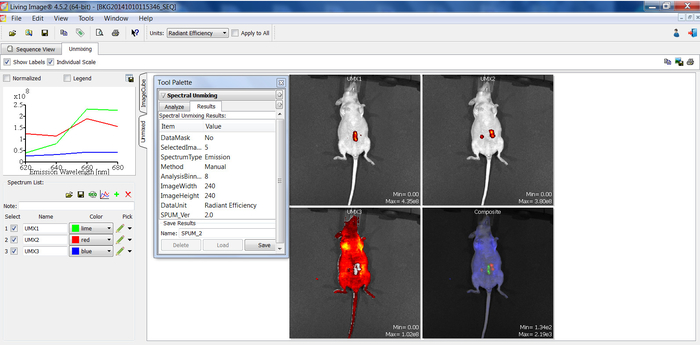
Figure 3: Unmixing of background vs specific fluorescence emitted by lesions. Picture shows representative images of the unmixing process performed to dissect actual fluorescence from lesions using the in vivo imaging system coupled software. Graph on the left panel denote normalized specific profiles of fluorescence emission by scar (green line, UMX1), lesions (red line, UMX2) and host tissue (blue line, UMX3 panel). Fluorescence intensity at different emission wavelengths (X-axis) is represented in units of radiant efficiency (Y-axis). Just note how each specific structure (i.e., scar, labeled lesions and host tissue) emits a different fluorescence profile which allows identifying and segregating them specifically form each other. On the right panel, fluorescence arising from launching specific emission profiles for scar (UMX1), lesions (UMX2) and host tissue (UMX3) are shown superposed on photograph images of mice. A composite image (Composite) is also included for illustrative purposes to denote the segmentation of fluorescence emitted by lesions from that emitted by scar or host tissues. Middle panel shows parameters selected for unmixing with the image software coupled to the in vivo imaging device Please click here to view a larger version of this figure.
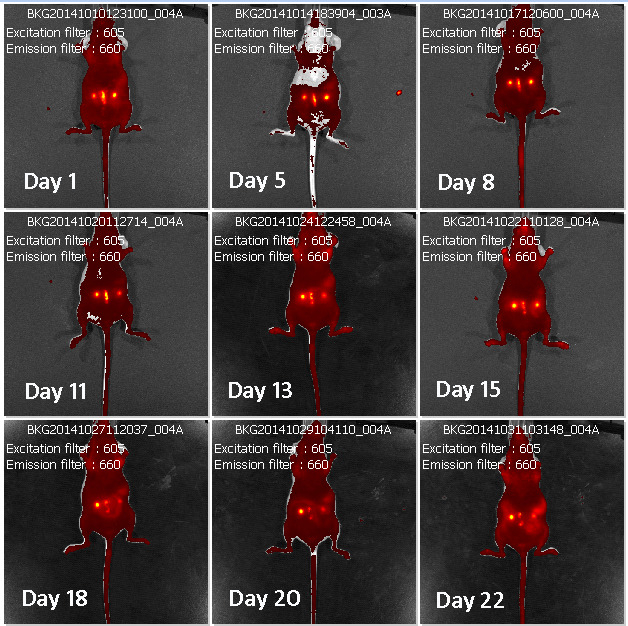
Figure 4: Time course monitoring of raw fluorescence emitted by lesions. Panel shows representative images of raw fluorescence emitted by a single mouse implanted with labeled human lesions (brilliant yellow spots) during the time course. Time points after surgery at which monitoring was performed are denoted as "Day (number)". Emission ad excitation pair filters used for monitoring are indicated in each panel/image. Each panel is identified by a specific code (BKG) in the upper part containing info related to the date at which fluorescence was acquired.
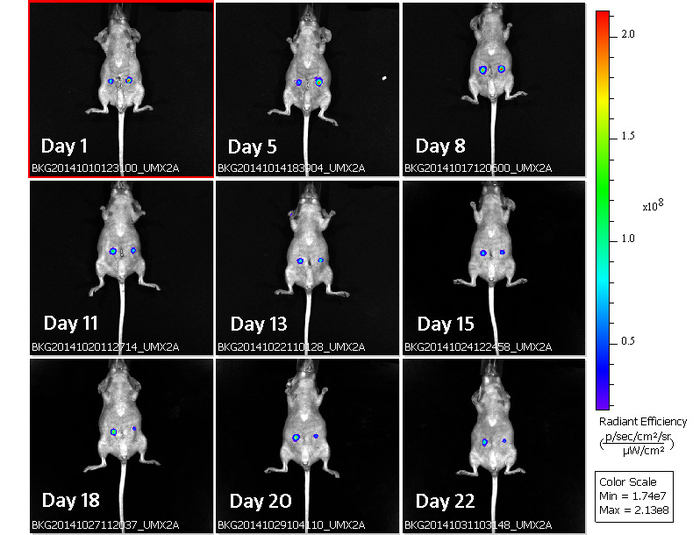
Figure 5: Time course monitoring of normalized fluorescence emitted by lesions. Representative images corresponding to unmixed, normalized fluorescent signaling emitted by human lesions (spots with rainbow color) superposed in a single mouse during the time course (days after surgery). Time points after surgery at which monitoring was performed are denoted as "Day (number)".Each panel is identified by a specific code (BKG) in the lower part containing info related to the date at which fluorescence was acquired. Rainbow palette color on the right side identifies fluorescence intensity (Radiant efficiency) emitted by lesions at each time point. Note how strong fluorescence intensity during the initial time points (i.e., red color in the center of lesions on days 1,5 and 8) declines during the time course (i.e., blue color in lesions on days 20 and 25). Please click here to view a larger version of this figure.
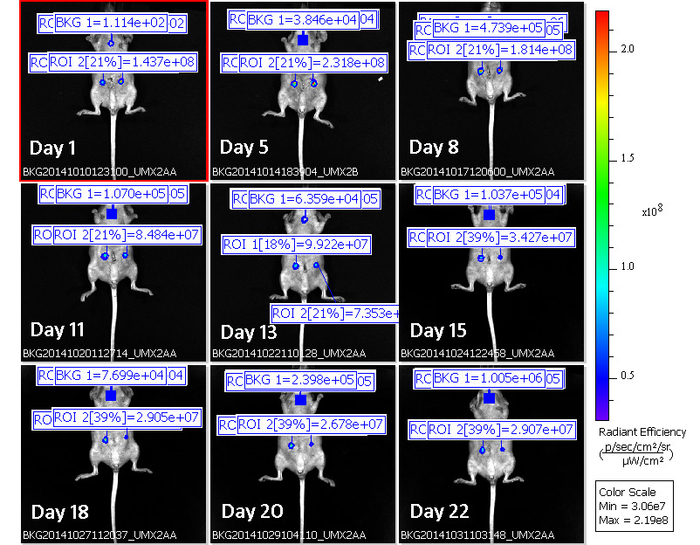
Figure 6: Use of ROIs for quantification of fluorescence intensity in lesions during the time course. Figure shows panel of images of normalized fluorescence emitted by lesions in a single mouse during the time course (i.e., Figure 5) with the addition of ROIs (delineating lesions and background) for quantification of fluorescence intensity. ROI 1 and ROI 2 identify the amount of fluorescence emitted by each of the two lesions during the time course. BKG identify the amount of fluorescence emitted by the host tissue (background fluorescence) during the time course. Background fluorescence is subtracted from ROIs for quantification purposes. Time points after surgery at which monitoring was performed are denoted as "Day (number)". Images acquired at each time point are labeled with a specific code (BKG) at the bottom of each one containing info related to the date at which fluorescence was acquired (first eight digits following BKG detail data for year(2014)-month (10) and -day(10 to 31)), an individual identification code (last 6 digits) and the specific profiles of fluorescence emission used for unmixing (UMX2) Rainbow palette color on the right side provides a visual scale of fluorescence intensity (Radiant efficiency) emitted by lesions at each time point. Note how fluorescence intensity values in the two lesions (ROI1 and ROI2) are higher at the initial time points (days 1 and 5) and decays during the time course to reach the lowest values at the end time points (days 20 and 25). Please click here to view a larger version of this figure.
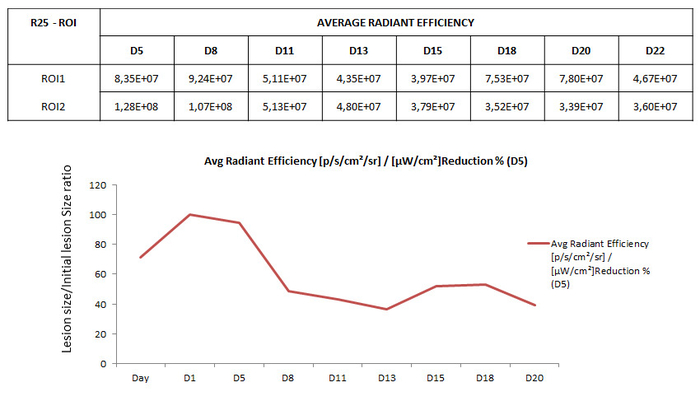
Figure 7: Normalization of fluorescence intensity during the time course. Table in the upper part shows illustrative example of the values of fluorescence intensity (radiant efficiency) emitted by two mCherry labeled lesions (ROI1 and ROI2) implanted in a mouse (R25). Monitoring of fluorescence was performed at different days after surgery (D5-D25) during the time course. D5- Graph at the bottom illustrates typical pattern of normalized fluorescence emitted by lesions infected with mCherry decaying during the time course. Y-axis shows values of fluorescence normalized to express the percentage of decay by using the formula (Signal intensity at each time point / Maximal signal intensity observed during the time course) x 100. Time points (Days (Dx) after implanting surgery) at which fluorescence was monitored are indicated in the X-axis. Note initial increase of signaling during the first 24 h after surgery, corresponding to stabilization of lesion, the peak in fluorescence intensity around D1-D5 and its subsequent decay due to episomal expression of mCherry during the time course Please click here to view a larger version of this figure.
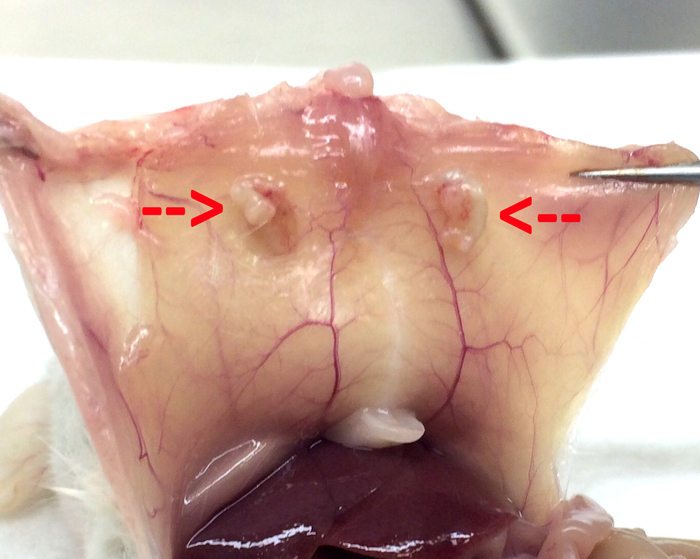
Figure 8: Macroscopic appearance of implanted endometriotic lesions. Representative images showing macroscopic appearance of endometriotic lesions implanted in mice at the end of the monitoring process upon sacrifice Please click here to view a larger version of this figure.
Discussion
The protocol herein detailed describes the implementation of an animal model of endometriosis in which the architecture of implanting lesions architecture is preserved whilst simultaneously allowing real time assessment of fluorescence emitted by mCherry labeled endometrial tissue. In this protocol, we describe the use of a specific in vivo imaging system and related software to non-invasively assess fluorescence emitted by the labeled lesion. Each user should adapt the protocol depending on the specific imaging device and related software available at their institution. Monitoring is performed in anesthetized animals through an isoflurane gas anesthesia machine coupled to an in vivo imaging system. To avoid interference with auto-fluorescence emitted by wounds, it is recommended to start monitoring lesions fluorescence at least three days after implantation surgery.
Heterologous mouse models of endometriosis similar to the one herein shown have been previously described consisting in the implantation endometrial fragments labeled with green fluorescent protein (GFP)18,19. The use of mCherry as a reporter for tagging the human tissue provides however an advantage over GFP because of the enhanced tissue penetration of the former. Due to its larger emission spectrum and higher photostability mCherry emits a brighter signal that is more appropriate for the visualization of intraperitoneal fragments.
Due to its small size adenoviruses are the vectors of choice for infection (i.e., labeling) of whole tissue pieces as those provide acceptable diffusion through 3D structures. Even with that the percentage of tissue infected cells is not higher than 30–35%. Thus, the limitation of this model relies on the inefficient labeling of tissue and additionally the transient expression achieved by adenoviruses. Indeed, fluorescence cannot be monitored beyond 4–6 week as it fades progressively due to the episomal transient expression of the Ad-virus. Efficiency of labeling and the period of monitoring might be increased by disrupting the donor tissue and infecting isolated single epithelial/stromal cells with ad-virus previous to being injected into recipient mice19,20. Such an approach, however, reduces the extent at which the animal model mimics the physiology of endometriosis provided that ectopic human lesions do not consist in a disorganized accumulation of single epithelial/stromal cells but rather in well-structured endometriotic tissue.
In this protocol, the most critical step is the labeling of the tissue and most specifically the determination of the appropriate concentration of ad-virus required for optimal infection. Indeed, the 1 x 1010 pfu/mL concentration pointed out in the protocol is mostly an orientative/consensus figure based on our experience. The optimal concentration might differ in each experiment depending on the type and quality of the biopsy and/or how quickly this is processed. We thus suggest testing at least three different (two-fold) titers in each experiment and choosing the one providing optimal labeling based on visualization under the fluorescence microscope.
Our protocol/model is useful to study the mechanisms implicated in the establishment and early development of the endometriotic lesions. In spite of the period of time of monitoring is constrained, the model is still useful to study and detect the effects of pharmacological compounds able to exert dramatic effects on lesion size in a short period of time such as antiangiogenic or antiestrogenic drugs. The development of more efficient and durable methods of labeling human tissue are expected to spread non-invasive monitoring as a consolidated technique to explore the potential therapeutic of a wider range of drugs in preclinical model endometriosis.
Disclosures
The authors have nothing to disclose.
Acknowledgements
This work was supported by Spanish Ministry of Economy and Competitiveness through the Miguel Servet Program [CP13/00077] cofounded by FEDER (European Regional Development Fund) and awarded to Dr R. Gómez as well as by Carlos III Institute of Health grants awarded to Dr R Gómez [PI14/00547 and PI17/02329] and to Prof A. Cano [PI12/02582].
Materials
| Endosampler™ | Medgyn | 22720 | Cannula for sampling the uterine endometrium |
| DMEM Medium | VWR | HYCLSH30285.FS | Medium |
| Ad-mCherry | Vector Biolabs | 1767 | Adenoviral vector expressing mCherry |
| PBS, 1X solution, sterile, pH 7,4 | VWR | E504-500ML | Buffer for washes |
| Pellets 17-B-Estradiol 18 mg/ 60 days | Innovative Research of America | SE-121 | Hormone pellets for rodents |
| Vetbond™ Tissue Adhesive | 3M | 780-680 | Tissue adhesive |
| Petri dishes in polystyrene crystal | Levantina | 367-P101VR20 | Petri dishes |
| Penicillin-Streptomicin | Sigma | P4333-100ML | Antibiotics |
| Syringes, medical 10 ml 0,5 ml | VWR | CODA626616 | Syringes |
| Nitrile gloves, powder-free | VWR | 112-2754 | Gloves |
| Soft swiss nude mice | Charles River | SNUSSFE05S | Mice for animal experiment |
| Ivis Spectrum In vivo Imaging system | Perkin Elmer | 124262 | In vivo Monitoring equipment |
| Living Image® (Ivis software) | Perkin Elmer | — | In vivo monitoring software |
| Fetal Bovine Serum | Gibco | 10082147 | Enrichment serum |
| 96-well cell culture treated plates | Life technologies | 167008 | Culture plates |
| Urine flasks | Summedical | 4004-248-001 | Flasks for washes |
| Sterile surgical blades | (Aesculap Division) Sanycare | 1609022-0008 | Surgical blades |
| Isovet 1000 mg/g | B-BRAUN | — | Isoflurane (Anesthetic) |
| Buprex® 0.3 mg | Schering Plough S.A. | — | Buprenorphine (Analgesic solution) |
| Injectable morphine solution 10 mg/mL | B BRAUN | — | Morphine (Analgesic solution) |
| Monofyl® Absorbable Sutures | COVIDIEN | — | Sutures |
| Desinclor chlorhexidine | Promedic SA | — | Antiseptic solution |
| Microscopy DMi8 | Leica Mycrosystems | — | fluorescence microscope |
| Hera Cell 150 Incubator | Thermo Scientific | 51026282 | Incubator |
References
- Nap, A. W., Groothuis, P. G., Demir, A. Y., Evers, J. L., Dunselman, G. A. Pathogenesis of endometriosis. Best Practice & Research: Clinical Obstetrics & Gynaecology. 18, 233-244 (2004).
- Holoch, K. J., Lessey, B. A. Endometriosis and infertility. Clinical Obstetrics and Gynecology. 53, 429-438 (2010).
- Eskenazi, B., Warner, M. L. Epidemiology of endometriosis. Obstetrics and Gynecology Clinics of North America. 24 (2), 235-258 (1997).
- Giudice, L. C., Kao, L. C. Endometriosis. Lancet. 364 (9447), 1789-1799 (2004).
- Donnez, J., et al. The efficacy of medical and surgical treatment of endometriosis-associated infertility and pelvic pain. Gynecologic and Obstetric Investigation. 54, 2-7 (2002).
- D’Hooghe, T. M., Bambra, C. S., Cornillie, F. J., Isahakia, M., Koninckx, P. R. Prevalence and laparoscopic appearance of spontaneous endometriosis in the baboon (Papio anubis, Papio cynocephalus). Biology of Reproduction. 45 (3), 411-416 (1991).
- Dick, E. J., Hubbard, G. B., Martin, L. J., Leland, M. M. Record review of baboons with histologically confirmed endometriosis in a large established colony. Journal of Medical Primatology. 32 (1), 39-47 (2003).
- Donnez, O., et al. Induction of endometriotic nodules in an experimental baboon modelmimicking human deep nodular lesions. Fertility & Sterility. 99 (3), 783-789 (2013).
- Grummer, R. Animal models in endometriosis research. Human Reproduction Update. 12 (5), 641-649 (2006).
- Rossi, G., et al. Dynamic aspects of endometriosis in a mouse model through analysis of implantation and progression. Archives of Gynecology and Obstetrics. 263 (3), 102-107 (2000).
- Grummer, R., et al. Peritoneal endometriosis: validation of an in-vivo model. Human Reproduction. 16 (8), 1736-1743 (2001).
- Becker, C. M., et al. A novel non-invasive model of endometriosis for monitoring the efficacy of antiangiogenic therapy. The American Journal of Pathology. 168 (6), 2074-2084 (2006).
- Laschke, M. W., Giebels, C., Nickels, R. M., Scheuer, C., Menger, M. D. Endothelial progenitor cells contribute to the vascularization of endometriotic lesions. The American Journal of Pathology. 178 (1), 442-450 (2011).
- Nap, A. W., et al. Antiangiogenesis therapy for endometriosis. The Journal of Clinical Endocrinology & Metabolism. 89 (3), 1089-1095 (2004).
- Wang, C. C., et al. Prodrug of green tea epigallocatechin-3-gallate (Pro-EGCG) as a potent anti-angiogenesis agent for endometriosis in mice. Angiogenesis. 16 (1), 59-69 (2013).
- Delgado-Rosas, F., et al. The effects of ergot and non-ergot-derived dopamine agonists in an experimental mouse model of endometriosis. Reproduction. 142 (5), 745-755 (2011).
- Al-Jefout, M., Andreadis, N., Tokushige, N., Markham, R., Fraser, I. A pilot study to evaluate the relative efficacy of endometrial biopsy and full curettage in making a diagnosis of endometriosis by the detection of endometrial nerve fibers. American Journal of Obstetrics & Gynecology. 197 (6), 578 (2007).
- Fortin, M., et al. Quantitative assessment of human endometriotic tissue maintenance and regression in a noninvasive mouse model of endometriosis. Molecular Therapy. 9 (4), 540-547 (2004).
- Liu, B., et al. Improved nude mouse models for green fluorescence human endometriosis. Journal of Obstetrics and Gynaecology Research. 36 (6), 1214-1221 (2010).
- Wang, N., et al. A red fluorescent nude mouse model of human endometriosis: advantages of a non-invasive imaging method. European Journal of Obstetric Gynecologic and Reproductive Biology. 176, 25-30 (2014).

