Agrobacterium tumefaciens-Mediated Genetic Engineering of Green Microalgae, Chlorella vulgaris
Summary
This protocol outlines the utilization of Agrobacterium tumefaciens-mediated transformation (AMT) for integrating gene(s) of interest into the nuclear genome of the green microalgae Chlorella vulgaris, leading to the production of stable transformants.
Abstract
Agrobacterium tumefaciens-mediated transformation (AMT) serves as a widely employed tool for manipulating plant genomes. However, A. tumefaciens exhibit the capacity for gene transfer to a diverse array of species. Numerous microalgae species lack well-established methods for reliably integrating genes of interest into their nuclear genome. To harness the potential benefits of microalgal biotechnology, simple and efficient genome manipulation tools are crucial. Herein, an optimized AMT protocol is presented for the industrial microalgae species Chlorella vulgaris, utilizing the reporter green fluorescent protein (mGFP5) and the antibiotic resistance marker for Hygromycin B. Mutants are selected through plating on Tris-Acetate-Phosphate (TAP) media containing Hygromycin B and cefotaxime. Expression of mGFP5 is quantified via fluorescence after over ten generations of subculturing, indicating the stable transformation of the T-DNA cassette. This protocol allows for the reliable generation of multiple transgenic C. vulgaris colonies in under two weeks, employing the commercially available pCAMBIA1302 plant expression vector.
Introduction
Agrobacterium tumefaciens, a gram-negative soil-borne bacterium, possesses a unique interkingdom gene transfer ability, earning it the title "natural genetic engineer"1. This bacterium can transfer DNA (T-DNA) from a tumor-inducing plasmid (Ti-Plasmid) into host cells through a Type IV secretion system, resulting in the integration and expression of the T-DNA within the host genome1,2,3,4. In the natural setting, this process leads to tumor formation in plants, commonly known as crown gall disease. However, Agrobacterium can also transfer T-DNA into various other organisms, including yeast, fungi, algae, sea urchin embryos, and even human cells under laboratory conditions5,6,7,8.
Exploiting this natural system, Agrobacterium tumefaciens-mediated transformation (AMT) enables the random integration of gene(s) of interest into a host cell's nuclear genome by modifying the T-DNA region of the Ti-plasmid. For this purpose, a widely used AMT plant expression vector is pCAMBIA13029. Researchers can employ simple cloning workflows in E. coli before transferring the desired vector into A. tumefaciens for subsequent transfer to the host of interest.
Green microalgae are eukaryotes that share many similarities with land plants but are highly recalcitrant to genetic modification. However, genetic transformation plays a crucial role in both fundamental and biotechnological research of microalgae. In several microalgae species, particularly Chlamydomonas reinhardtii, genetic transformation via AMT has successfully introduced transgenes such as human interleukin-2 (hIL-2), the severe acute respiratory syndrome coronavirus 2 receptor-binding domain (SARS-CoV-2 RBD), and two antimicrobial peptides (AMPs)10,11,12,13. Among these, Chlorella vulgaris, a less fastidious and fast-growing green algae species, holds significant potential for the sustainable production of carbohydrates, proteins, nutraceuticals, pigments, and other high-value compounds14. However, the lack of reliable tools for creating transgenic strains of C. vulgaris hampers its commercial progress. Since there have been only a limited number of published works utilizing AMT in C. vulgaris15, and given the considerable differences between plant and microalgae cultivation, optimizing the AMT protocol becomes essential.
In this study, researchers inserted green fluorescent protein (mGFP5) downstream of the Cauliflower Mosaic Virus (CamV) 35S promoter and added a histidine tag to use it as a reporter gene for protein expression. Transformants were selected using Hygromycin B, and after subculturing for over twenty generations, the transformation remained stable. The pCAMBIA1302 plasmid employed in this work can be readily adapted to contain any gene of interest. Furthermore, the method and materials presented can be adjusted for other green algae species with an active CamV35S promoter, as this promoter is used for Hygromycin selection.
Protocol
All media and solutions must be autoclaved prior to use unless otherwise stated. All centrifuge tubes, pipette tips, etc., should be sterile or autoclaved before use. For easy reference, the media recipes used in this protocol are listed in Table 1.
1. Preparation of A. tumefaciens electrocompetent cells
- Inoculate Agrobacterium (AGL-1) into a 25 mL sterile shaker flask of LB media (supplemented with rifampicin, 20 mg/L-1) from a frozen glycerol stock and grow overnight at 28-30 °C and 150 rpm.
NOTE: The Agrobacterium strain AGL-1 is (see Table of Materials) resistant to rifampicin, ampicillin, chloramphenicol, and streptomycin and can be obtained from several suppliers. - The following day, dilute the culture 1,00,000-fold by adding 0.5 µL of the overnight culture to 50 mL autoclaved ultra-pure water and spread 10 µL of this diluted culture on an LB plate, and then incubate at 28-30 °C for 1-3 days.
- Pick a colony and start a 20 mL overnight culture in LB with rifampicin at 28-30 °C.
- The next day, inoculate 500 mL LB media (no antibiotic) in a shaker flask with 9 mL of the overnight culture and grow at 28-30 °C until the OD600 reaches 0.5.
- Divide the culture evenly into ten 50 mL centrifuge tubes and chill on ice for 30 min. Pre-cool the centrifuge to 4 °C.
- Centrifuge the tubes at 4 °C and 4000 x g for 15 min. Subsequently, remove as much supernatant as possible using a pipette and resuspend the pellet in each tube in 50 mL of ice-cold water.
- Centrifuge the cells at 4 °C and 4000 x g for 15 min. Remove the supernatant and resuspend the pellet in 25 mL of ice-cold water in each tube.
- Centrifuge the cells at 4 °C and 4000 x g for 15 min. Remove the supernatant and resuspend the pellet in 1 mL of ice-cold 10% (w/v) glycerol in each tube.
- Combine all tubes into one 50 mL tube and centrifuge the cells at 4 °C and 4000 x g for 15 min. Remove the supernatant and resuspend the pellet in 400 µL ice-cold 10% (w/v) glycerol. The cell concentration should be about 1-3 x 1010 cells/mL.
- Distribute the cells as 50 µL aliquots in sterile prechilled microtubes. Freeze in liquid nitrogen and store in a -80 °C freezer.
2. Electroporation of A. tumefaciens
- Add 50 µL of AGL-1 A. tumefaciens electrocompetent cells into a prechilled 0.1 mm cuvette and add 2 µL of pCAMBIA1302 or similar plasmid (conc 15 ng/µL) (see Table of Materials).
- Electroporate at 2400 V using an electroporator (200 Ω, capacitance extender 250 µFD, capacitance 25 µFD, exponential decay, see Table of Materials). The time constant should be >4.5 ms for successful electroporation with exponential decay.
- Immediately add 1 mL of LB media to the cuvette and gently pipette up and down to mix. Transfer the 1 mL of resuspended cells into a 1.5 mL microtube and incubate it at 28-30 °C for at least 1 h to recover.
- Plate the cells on LB agar containing the appropriate antibiotic (i.e., 50 mg/L of kanamycin for pCAMBIA1302 for prokaryotic selection).
- Incubate at 28-30 °C for 2-3 days.
- Select one colony, grow overnight in LB with the appropriate selection (50 mg/L of kanamycin for pCAMBIA1302), and cryopreserve the plasmid-containing strain using 50% glycerol at -80 °C for future use.
3. AMT of C. vulgaris
NOTE: Prepare C. vulgaris (UTEX 395, see Table of Materials) and A. tumefaciens cultures in parallel for co-cultivation. C. vulgaris cultures should be started 3 days before preparing A. tumefaciens cultures. Protocol was modified based on that published by Kumar et al.7.
- Prepare C. vulgaris culture
- C. vulgaris is traditionally stored on slants or maintained in log-phase cultures in illuminated conditions. From a log phase culture at OD600 = 1.0, spread 0.5 mL onto a Tris-Acetate-Phosphate (TAP, see Table 1) agar plates (1.2% w/v agar A) and grow for 5 days at 25 °C with illumination (50 µE/m2s). This will yield approximately 5 x 106 cells.
- Prepare A. tumefaciens culture
- Grow A. tumefaciens AGL-1 pCAMBIA1302 strain in a shaker flask by inoculating 10 mL of LB media supplemented with 15 mM glucose, 20 mg/L rifampicin, and 50 mg/L kanamycin (see Table of Materials) using the glycerol stock. Incubate at 28-30 °C and 250 rpm overnight.
- Inoculate 1 mL of overnight culture into 50 mL of LB with 15 mM glucose, 20 mg/L rifampicin, and 50 mg/L kanamycin and incubate at 28-30 °C and 250 rpm until the OD600 reaches 1.0.
- Cocultivate C. vulgaris and A. tumefaciens
- To prepare the A. tumefaciens culture for co-cultivation, transfer the grown culture to a 50 mL tube and centrifuge at 4000 x g for 30 min at room temperature. Remove the supernatant using a pipette and wash the cells twice with the induction medium.
NOTE: The induction media used is TAP media adjusted to pH 5.5 and supplemented with F/2 medium trace metals and vitamins with 200 µM acetosyringone (AS, see Table of Materials). Resuspend the cells in induction media such that OD600 = 0.5. - To prepare the C. vulgaris culture for co-cultivation, add 25 mL of induction media to the C. vulgaris plate containing ~ 5 x 106 cells. Transfer to a 50 mL tube. Centrifuge the cells at 4000 x g for 15 min at room temperature and discard the supernatant.
- Mix the algal cell pellet with 200 µL of the bacterial suspension (OD600: 0.5) and incubate them in a rotary shaker at 21-25 °C at 150 rpm for 1 h.
- Spread the mixed culture (200 µL) onto induction media plates supplemented with 15 mM glucose and incubate at 21-25 °C in the dark for 3 days.
- After 3 days, use 10 mL of TAP media supplemented with 400 mg/L cefotaxime or 20 mg/L of tetracycline (see Table of Materials) to collect the microalgae and incubate in the dark at 21-25 °C for 2 days to eliminate A. tumefaciens from the culture.
- Plate 500 µL of the culture onto selective media, e.g., TAP media supplemented with 20-70 mg/L of Hygromycin B (when using pCAMBIA1302) and 400 mg/L-1 cefotaxime or 20 mg/L of tetracycline. Incubate 21-25 °C in the dark for 2 days before placing them in an illuminated chamber.
NOTE: Resistant colonies will appear 5-7 days after light exposure. - Pick single colonies from the transformation plate and re-streak them on TAP agar plates supplemented with 400 mg/L of cefotaxime or 20 mg/L of tetracycline and 20-70 mg/L of Hygromycin B.
- To prepare the A. tumefaciens culture for co-cultivation, transfer the grown culture to a 50 mL tube and centrifuge at 4000 x g for 30 min at room temperature. Remove the supernatant using a pipette and wash the cells twice with the induction medium.
4. Colony PCR (cPCR) to confirm gene integration in C. vulgaris transformants
- Extract genomic DNA from a C. vulgaris transformant after subculturing at least twice from a single colony using a plant genomic DNA extraction kit (see Table of Materials). Using this as a template for PCR, design primers for the mgfp5 region of the T-DNA (mgfp5-Fwd: 5' CCCATCTCATAAATAACGTC 3', and M13-Rev: 5' CAGGAAACAGCTATGAC 3').
- Using a Q5 high-fidelity DNA polymerase master mix kit (see Table of Materials), perform polymerase chain reaction (PCR) to validate transgene integration.
NOTE: The following conditions were used for the present study: 98 °C for 3 min; 35 cycles (98 °C for 10 s, 57 °C for 30 s, 72 °C for 1 min); 72 °C for 5min. Use pCAMBIA1302 as a positive control and wild-type C. vulgaris genomic DNA as a negative control.
- Using a Q5 high-fidelity DNA polymerase master mix kit (see Table of Materials), perform polymerase chain reaction (PCR) to validate transgene integration.
- To confirm that there is no pCAMBIA1302 present in the sample due to residual contamination by A. tumefaciens, use colony PCR. Add a small amount of transformant cells to 10 µL of sterile water and boil at 98 °C for 15 min. Use this as a template for PCR.
- Design primers for a region outside of the T-DNA on pCAMBIA1302 (B1-Fwd: 5'AGTAAAGGAGAAGAACTTTTC 3' and B1-Rev: 5'CCTGATGCGGTATTTTCTC 3').
NOTE: The following conditions were used for the present study: 94 °C for 3 min; 30 cycles (94 °C for 30 s, 48 °C for 30 s, 68 °C for 5 min); 68 °C for 7 min. Use a boiled colony of A. tumefaciens AGL-1 (pCAMBIA1302) as a positive control and a boiled colony of wild-type C. vulgaris as a negative control.
- Design primers for a region outside of the T-DNA on pCAMBIA1302 (B1-Fwd: 5'AGTAAAGGAGAAGAACTTTTC 3' and B1-Rev: 5'CCTGATGCGGTATTTTCTC 3').
- To confirm that there is no A. tumerfaciens contamination present in the sample, use colony PCR. Add a small amount of transformant cells to 10 µL of sterile water and boil at 98 °C for 15 min. Use this as a template for PCR.
- Design primers for the virE2 gene contained on the AGL-1 virulence plasmid (virE2-Fwd: 5' AGGGAGCCCTACCCG 3' and virE2-Rev: 5' GAACCAGCCTGGAGTTCG 3').
NOTE: The following conditions were used for the present study: 94 °C for 3 min; 30 cycles (94 °C for 30 s, 53 °C for 30 s, 68 °C for 3 min); 68 °C for 7 min. Use a boiled colony of A. tumefaciens AGL-1 (pCAMBIA1302) as a positive control and a boiled colony of wild-type C. vulgaris as a negative control.
- Design primers for the virE2 gene contained on the AGL-1 virulence plasmid (virE2-Fwd: 5' AGGGAGCCCTACCCG 3' and virE2-Rev: 5' GAACCAGCCTGGAGTTCG 3').
- Run PCR samples on a DNA agarose gel along with a ladder (1 Kbp DNA ladder, see Table of Materials) to confirm the size of the resulting fragments16.
5. Measuring the fluorescence of transformants
- Inoculate each transformant from a log-phase maintenance culture or TAP agar slant into 50 mL of TAP supplemented with 200 mg/L-1 cefotaxime and 25 mg/L-1 Hygromycin B such that the starting OD600 = 0.1. Inoculate one culture with wild-type C. vulgaris and only 200 mg/L-1 cefotaxime as the negative control. Incubate at 25 °C at 150 rpm with 150 µmol/m2s of photosynthetically active light (see Table of Materials).
- Every 24 h, take a 300 µL of sample and measure the absorbance and fluorescence (excitation 488 nm, emission 526 nm) in a 96-well plate reader or UV/Vis spectrometer and fluorometer (see Table of Materials). Use a black plate with a transparent bottom for simultaneous measurements.
6. Crude protein extraction, protein purification, and SDS-PAGE electrophoresis
- Inoculate transformant (#50) from a log-phase maintenance culture into 300 mL of TAP supplemented with 200 mg/L-1 cefotaxime and 25 mg/L-1 Hygromycin B to reach the OD680 = 0.1. Inoculate one culture with wild-type C. vulgaris and only 200 mg/L-1 cefotaxime as the negative control. Incubate at 25 °C at 150 rpm with 150 µmol/m2s of photosynthetically active light.
- Extract the crude protein sample from the transformant (#50) and wild-type strains using 5 mL lysis buffer (see Table 1), followed by sonication for 4 min.
- Purify crude protein sample with a Ni-NTA resin through 5 mL polypropylene columns (see Table of Materials).
- Lyophilize the 15 mL purified protein samples and resuspend them in 1 mL lysis buffer for SDS-PAGE analysis17.
Representative Results
To show successful transformation using the method above, C. vulgaris was cocultured with either AGL-1 containing the pCAMBIA1302 plasmid or without the plasmid (wild-type and plated on TAP agar supplemented with Hygromycin B and cefotaxime (Figure 1A). The leftmost plate shows the transformed colonies capable of growth on Hygromycin B/cefotaxime plates, and the middle plate shows that wild-type AGL-1 cannot grow on the Hygromycin B/cefotaxime plates. The rightmost plate shows that when no selection is used (cefotaxime only), C. vulgaris transformants and wild-type can grow. Single colonies were streaked onto TAP agar with Hygromycin B and cefotaxime (Figure 1B). Escaped colonies from the rightmost plate (Figure 1B) were inoculated onto TAP liquid with Hygromycin B and cefotaxime (Figure 1C). Due to variable expression levels after random integration of T-DNA, transformants were originally recovered on plates with increasing concentrations of Hygromycin B ranging from 25-70 mg/L. One colony from each plate was grown in liquid TAP media containing the same concentration of Hygromycin B along with the wild-type C. vulgaris at the lowest concentration (Figure 1D). The wild-type C. vulgaris cannot grow in the presence of Hygromycin, while colonies resistant up to 70 mg/L were obtained.
To confirm that the T-DNA cassette was stably integrated into the genome of C. vulgaris, one colony selected from each of the 20, 25, 30, 50, and 70 mg/L Hygromycin B plates were each subcultured by repeated streaking on TAP agar plates with selection two times and then routinely subcultured in TAP media over ten times, named transformant #20, #25, #30, #50, #50, and #70 respectively. Using these cultures, both colony PCR was performed to rule out contamination by pCAMBIA1302 in AGL1, and genomic DNA was extracted to confirm the mGFP5 gene was present in the C. vulgaris genome. In Figure 2A, colony PCR was performed using primers targeting a 5 kbp region outside of the T-DNA left and right borders were able to amplify this region from pCAMBIA1302 plasmid as a positive control, but this region was not present in any of the C. vulgaris samples indicating that pCAMBIA1302 DNA was not present in any of the cultures. The wild-type C. vulgaris sample was used as a negative control, and no amplification was seen.
Genomic DNA was extracted from transformants #25, #50, and #70, and PCR was performed, amplifying a 1763 bp region containing the mgfp5 gene of the T-DNA to confirm integration in the transformants (Figure 2B). pCAMBIA1302 was used as a positive control, and amplification was detected in all of the transformants, indicating that the mgfp5 gene was present in all of the transformants' genomic DNA. The wild-type C. vulgaris was used as a negative control, and no amplification was seen. Figure 2C shows the amplification of a 600 bp fragment of the virulence protein E2 (VirE2) which is found in the tumor-inducing plasmid (Ti plasmid) of the AGl1 strain. The positive control, consisting of the AGl1 strain containing pCAMIA1302, was successfully amplified with primers, indicating the presence A. tumefaciens in the sample. However, all of the C. vulgaris samples tested negative for this region, indicating that repeated subculturing on cefotaxime successfully eliminated contamination of AGL1 from the culture. The wild-type C. vulgaris sample was employed as a negative control, and no amplification was observed. Taken together, these three tests confirm that the T-DNA containing mgpf5 was inserted in the genome of C. vulgaris, and these results were not due to the contamination of AGL1 in the samples nor due to the presence of pCAMBIA1302 in the cells. These results were repeated after 20 and approximately 100 subcultures of the transformants confirming the long-term integration of the T-DNA into the genome.
Finally, the phenotype was confirmed by growing the transformants in TAP media with selection and measuring the fluorescence. This was compared to the wild-type C. vulgaris. Due to the background absorbance/autofluorescence of chlorophyll, the fluorescence was compared by normalizing the data to the wild-type strain. In Figure 3A, it can be observed that there is a noticeable difference in growth between the transformants and the wild-type C. vulgaris. This could potentially be attributed to the multiple random integrations of the T-DNA into the genome, which may be impacting other cellular processes and leading to slower growth. Nonetheless, Figure 3B highlights that despite the lower growth, all of the selected strains exhibit higher fluorescence levels when normalized for cell density. Notably, strain #70 demonstrated significantly higher fluorescence levels than the others, with a p-value of <0.01. This emphasizes the importance of screening numerous colonies to identify transformants that express the desired protein without negatively impacting overall growth behavior.
To confirm the expression of mGFP5-6xHis, SDS-PAGE was used to analyze the crude protein extract and his-tag purified proteins from both C. vulgaris (WT) and C. vulgaris transformant (#50). Subsequently, his-tag protein purification was done using a Ni-NTA resin gravity flow kit following the manufacturer's protocol18. Figure 4 shows the presence of mGFP5-6xHis protein (~30 kDa) in the transformant sample #50 and no protein in the negative control (WT C. vulgaris).
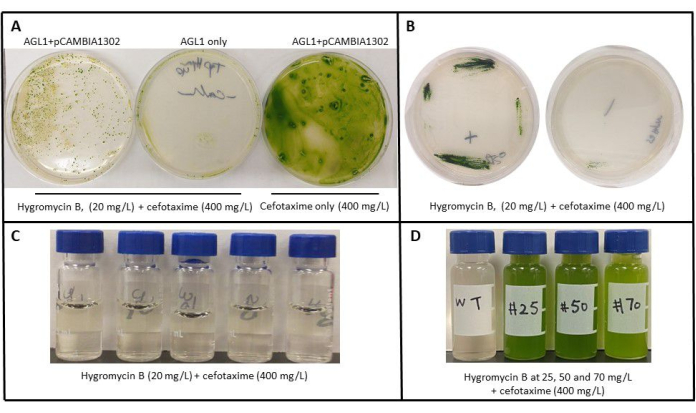
Figure 1: Recovery of C. vulgaris transformants after AMT. (A) After co-cultivation of C. vulgaris, transformants are AGL-1 containing the pCAMBIA plasmid or without the plasmid (AGL-1 only) on selective plates. (B) Re-streaked single colonies selected from Hygromycin B (20 mg/L) plates on selective TAP agar. (C) Escaped colonies lack growth from AGL1 only plate on Hygromycin B (20 mg/L) liquid media. (D) Growth of single colonies #25, #50, and #50 from Hygromycin selection plates grown in liquid TAP media containing Hygromycin and wild-type C. vulgaris that cannot grow in Hygromycin media. Please click here to view a larger version of this figure.
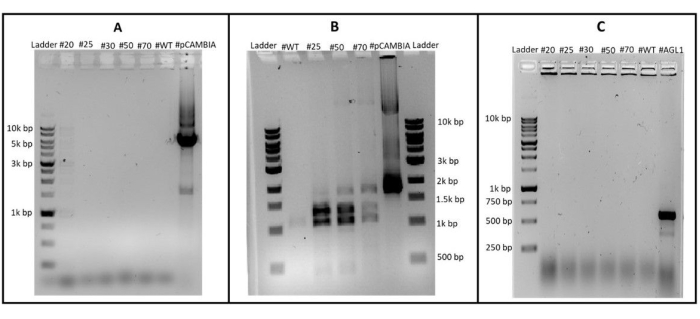
Figure 2: Detection of pCAMBIA1302 contamination. (A) Integration of T-DNA in transformants (B) and detection of A. tumefaciens contamination (C). WT indicates wild-type C. vulgaris, PC is the positive control (pCAMBIA1302) and AGL1 indicates A. tumefaciens AGL1 strain. Please click here to view a larger version of this figure.
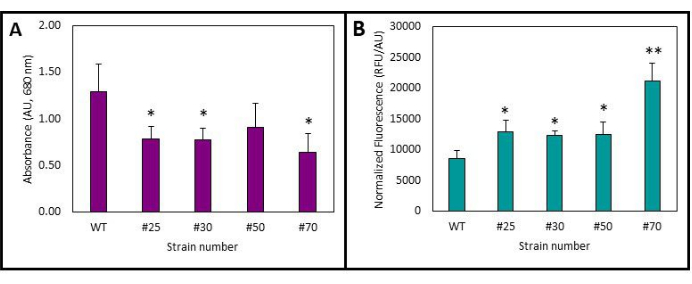
Figure 3: Growth and fluorescence of the transformants at 3 days post-inoculation. (A) Growth as measured by absorbance at 680 nm. (B) Fluorescence normalized to the wild-type strain. Average of 3-4 biological replicates, error bars represent 1 σ. *p < 0.05, **p < 0.01 when compared to the wild-type (WT). Please click here to view a larger version of this figure.
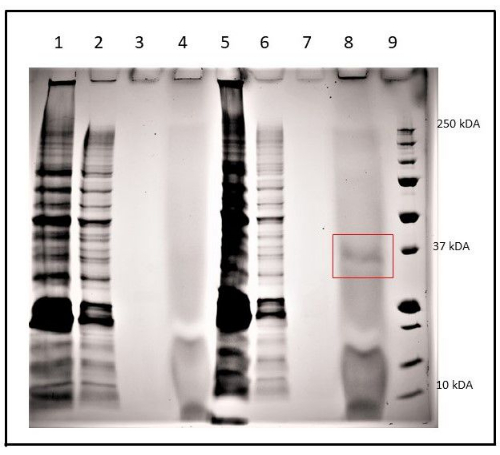
Figure 4: SDS-PAGE analysis of mGFP5-6xHis expression in C. vulgaris (WT) and transformed C. vulgaris sample #50. Extracts were prepared from crude, purified (first wash), purified (diluted), and purified (concentrated) from C. vulgaris (WT), lanes 1-4, respectively. Crude, purified (first wash), purified (eluted), and purified (lyophilized) from C. vulgaris (WT), lanes 5-8, respectively. The red box indicates purified GFP-6xHis protein from transformant #70. Proteins were separated on a polyacrylamide 12% gel. The molecular weights (MWs) of the protein standards (Std.) are shown on the right. Please click here to view a larger version of this figure.
| Name of media/buffer | Composition | ||
| LB | 5 g/L yeast extract, 10 g/L tryptone, 5 g/L NaCl | ||
| TAP | 20 mM Tris base, 1.58 mM K2HPO4, 2.4 mM KH2PO4, 7.0 mM NH4Cl, 0.83 mM MgSO4, 0.34 mM CaCl2, 1 mL/L glacial acetic acid, and 1 mL/L of each F/2 medium trace metals and F/2 vitamins | ||
| F/2 medium trace metals | 22 mg/L ZnSO4·7H2O, 180 mg/L MnCl2·4H2O, 6.3 mg/L Na2MoO4·2H2O, 14 mg/L CoCl2·6H2O, 9.8 mg/L CuSO4·5H2O, 3.15 g/L FeCl3·6H2O, 4.36 g/L Na2EDTA·2H2O | ||
| F/2 medium vitamins | 0.1 mM Vitamin B12 (cyanocobalamin), 25 mg/L Biotin, 335 mg/L Thiamine, 50 mM HEPES Buffer pH 7.8 | ||
| Lysis Buffer | 20 mM Na2HPO4, 300 mM NaCl, pH 7.4 | ||
Table 1: Media and buffer recipes.
Discussion
The efficiency of transformation is associated with several different parameters. The choice of A. tumefaciens strains used for AMT is crucial. AGL-1 is one of the most invasive strains discovered and, for this reason, has been routinely used in plant AMT. Supplementing the induction media with glucose (15-20 mM) is also important for AMT efficiency. Considering C. vulgaris can grow in both phototrophic and heterotrophic conditions, glucose or other carbon sources are often omitted from microalgae media to prevent contamination. For example, Bold's Basal Medium (BBM) is the most commonly used media for the phototrophic cultivation of C. vulgaris19. However, BBM does not support A. tumefaciens growth due to the lack of an organic carbon source and ammonium as a nitrogen source it prefers (BBM uses nitrate)20. Therefore, TAP media was used in this work because it contains acetate as a carbon source for C. vulgaris, glucose for A. tumefaciens, and ammonium as a nitrogen source both species can use. TAP agar plates also allow for much faster growth rates for C. vulgaris, minimizing the time to recover transformants after AMT. To induce Ti-plasmid virulence proteins, 200 µM of acetosyringone must also be added to the induction media.
To eliminate A. tumefaciens from the culture after AMT, cefotaxime, along with serial passaging, was used. AGL-1 is resistant to ampicillin, chloramphenicol, rifamycin, and streptomycin, and it is sensitive to cefotaxime, tetracycline, spectinomycin, and kanamycin. In plants, cefotaxime is often used to eliminate A. tumefaciens because it has a lower phytotoxic effect for shoot regeneration in many plant species than other antibiotics21. However, we also confirmed that tetracycline (which costs about 100 times less) could be used instead of cefotaxime in this protocol when combined with serial passaging from a single colony (data not shown). This is likely because antibiotics like tetracycline would inhibit protein synthesis in the chloroplast in plants which are dependent on photosynthesis22. However, C. vulgaris can grow in the dark when supplemented with acetate, meaning chloroplast dysfunction is not necessarily toxic to this species under carefully chosen conditions.
Although 20 mg/L of Hygromycin B was sufficient to prevent the growth of wild-type C. vulgaris, a range of Hygromycin B concentrations (20-70 mg/L) were tested. It is possible that colonies isolated from higher concentration plates may have multiple integrated copies of the T-DNA or the T-DNA is located in a more transcriptionally active region of the genome resulting in higher expression of the Hygromycin B phosphotransferase resulting in faster inactivation of the herbicide and less lag time in culture growth23. Multiple copies might be the reason for the differences in the intensity of the PCR products when amplifying mgfp5 from the genome (Figure 2B). The presence of GFP-6xHis protein in the purified sample during SDS-PAGE analysis (Figure 4, Lane 8) and its absence in the wild-type strain (Figure 4, lane 4) confirms that AMT and pCAMBIA1302 can be used for successful expression and stable integration of transgene in C. vulgaris.
In this work, we developed and optimized a protocol for reliable C. vulgaris UTEX 395 transformation using A. tumefaciens AGL-1 and a pCAMBIA system plasmid. As all of the strains and plasmids needed for this work are available from several different sources, this method will be easy to replicate in any laboratory. As one of the most important industrial strains of microalgae, a standard reference method for reliable AMT will be crucial to engineering C. vulgaris for biotechnological applications.
Disclosures
The authors have nothing to disclose.
Acknowledgements
The authors would like to thank Prof. Paul Hooykaas for kindly providing the pCAMBIA1302 vector and Agrobacterium tumefaciens AGL1 from the Institute of Biology Leiden, Leiden University, the Netherlands. The authors would also like to thank Eva Colic for her help in growing the fluorescent transformants. This work was funded by the Natural Sciences and Engineering Research Council of Canada and the Mitacs Accelerate program.
Materials
| 1 Kb Plus DNA ladder | FroggaBio | DM015 | |
| Acetosyringone | Fisher Scientific | D26665G | |
| Agrobacterium tumefaciens | Gold Biotechnologies | Strain: AGL-1; Gift from Prof. Paul Hooykaas | Genotype: C58 RecA (RifR/CarbR) pTiBo542DT-DNA |
| Biotin | Enzo Life Sciences | 89151-400 | |
| CaCl2·2H2O | VWR | BDH9224-1KG | |
| Cefotaxime | AK Scientific | J90010 | |
| Chlorella vulgaris | University of Texas at Austin Culture Collection of Algae | Strain: UTEX 395 | Wildtype strain |
| CoCl2·6H2O | Sigma Aldrich | C8661-25G | |
| CuSO4·5H2O | EMD Millipore | CX2185-1 | |
| FeCl3·6H2O | VWR | BDH9234-500G | |
| Gene Pulser Xcell Electroporator | Bio-Rad | 1652662 | Main unit equipped with PC module. |
| GeneJET Plant Genome Purification Kit | Thermo Scientific | K0791 | |
| Glacial acetic acid | VWR | CABDH3093-2.2P | |
| Glycerol | BioBasic | GB0232 | |
| HEPES Buffer | Sigma Aldrich | H-3375 | |
| Hygromycin B | Fisher Scientific | AAJ6068103 | |
| K2HPO4 | VWR | BDH9266-500G | |
| Kanamycin | Gold Biotechnologies | K-250-25 | |
| KH2PO4 | VWR | BDH9268-500G | |
| MgSO4·7H2O | VWR | 97062-134 | |
| MnCl2·4H2O | JT Baker | BAKR2540-01 | |
| Na2CO3 | VWR | BDH7971-1 | |
| Na2EDTA·2H2O | JT Baker | 8993-01 | |
| Na2MoO4·2H2O | JT Baker | BAKR3764-01 | |
| NaCl | VWR | BDH7257-7 | |
| NaH2PO4 H2O | Millipore Sigma | CA80058-650 | |
| NaNO3 | VWR | BDH4574-500G | |
| NEBExpress Ni Resin | NewEngland BioLabs | NEB #S1427 | |
| NH4Cl | VWR | BDH9208-500G | |
| pCAMBIA1302 | Leiden University | Gift from Prof. Paul Hooykaas | pBR322, KanR, pVS1, T-DNA(CaMV 35S/HygR/CaMV polyA, CaMV 35S promoter/mgpf5-6xhis/NOS terminator) |
| Polypropylene Columns (5 mL) | QIAGEN | 34964 | |
| Precision Plus Protein Unstained Protein Standards, Strep-tagged recombinant, 1 mL | Bio-Rad | 1610363 | |
| Rifampicin | Millipore Sigma | R3501-1G | |
| SunBlaster LED Strip Light 48 Inch | SunBlaster | 210000000906 | |
| Synergy 4 Microplate UV/Vis spectrometer | BioTEK | S4MLFPTA | |
| Tetracycline | Thermo Scientific Chemicals | CAAAJ61714-14 | |
| TGX Stain-Free FastCast Acrylamide Kit, 12% | Bio-Rad | 1610185 | |
| Thiamine | TCI America | T0181-100G | |
| Tris Base | Fisher Scientific | BP152-500 | |
| Tryptone | BioBasic | TG217(G211) | |
| Vitamin B12 (cyanocobalamin) | Enzo Life Sciences | 89151-436 | |
| Yeast Extract | BioBasic | G0961 | |
| ZnSO4·7H2O | JT Baker | 4382-01 |
References
- Smith, E. F., Townsend, C. O. A plant tumor of bacterial origin. Science. 25 (643), 671-673 (1907).
- Chilton, M. D., et al. Stable incorporation of plasmid DNA into higher plant cells: The molecular basis of tumorigenesis. Cell. 11 (2), 263-271 (1977).
- De Cleene, M., De Ley, J. The host range of crown gall. The Botanical Review. 42, 389-466 (1976).
- Hooykaas, P. J., Schilperoort, R. A. Agrobacterium and plant genetic engineering. Plant Molecular Biology. 19, 15-38 (1992).
- Bundock, P., den Dulk-Ras, A., Beijersbergen, A., Hooykaas, P. J. J. Transkingdom T-DNA transfer from Agrobacterium tumefaciens to Saccharomyces cerevisiae. The European Molecular Biology Organization. 14 (13), 3206-3214 (1995).
- Piers, K. L., Heath, J. D., Liang, X., Stephens, K. M., Nester, E. W. Agrobacteriumtumefaciens-mediated transformation of yeast. Proceedings of the National Academy of Sciences of the United States of America. 93 (4), 1613-1618 (1996).
- Kumar, S. V., Misquitta, R. W., Reddy, V. S., Rao, B. J., Rajam, M. V. Genetic transformation of the green alga-Chlamydomonas reinhardtii by Agrobacteriumtumefaciens. Plant Science. 166 (3), 731-738 (2004).
- de Groot, M. J., Bundock, P., Hooykaas, P. J., Beijersbergen, A. G. Agrobacteriumtumefaciens-mediated transformation of filamentous fungi. Nature Biotechnology. 16 (9), 839-842 (1998).
- Hajdukiewicz, P., Svab, Z., Maliga, P. The small, versatile pPZP family of Agrobacterium binary vectors for plant transformation. Plant Molecular Biology. 25 (6), 989-994 (1994).
- Dehghani, J., Adibkia, K., Movafeghi, A., Pourseif, M. M., Omidi, Y. Designing a new generation of expression toolkits for engineering of green microalgae; robust production of human interleukin-2. BioImpacts. 10 (4), 259-268 (2020).
- Berndt, A. J., Smalley, T. N., Ren, B., Simkovsky, R., Badary, A., Sproles, A. E., Fields, F. J., Torres-Tiji, Y., Heredia, V., Mayfield, S. P. Recombinant production of a functional SARS-CoV-2 spike receptor binding domain in the green algae Chlamydomonas reinhardtii. PLoS One. 16, 0257089 (2021).
- Li, A., Huang, R., Wang, C., Hu, Q., Li, H., Li, X. Expression of anti-lipopolysaccharide factor isoform 3 in Chlamydomonas reinhardtii showing high antimicrobial activity. Marine Drugs. 19 (5), 239 (2021).
- Xue, B., Dong, C. M., Hu, H. H., Dong, B., Fan, Z. C. Chlamydomonas reinhardtii-expressed multimer of ToAMP4 inhibits the growth of bacteria of both Gram-positive and Gram-negative. Process Biochemistry. 91, 311-318 (2020).
- Khan, M. I., Shin, J. H., Kim, J. D. The promising future of microalgae: current status, challenges, and optimization of a sustainable and renewable industry for biofuels, feed, and other products. Microbial Cell Factories. 17, 36 (2018).
- Cha, T. S., Yee, W., Aziz, A. Assessment of factors affecting Agrobacterium-mediated genetic transformation of the unicellular green alga, Chlorella vulgaris. World Journal of Microbiology and Biotechnology. 28, 1771-1779 (2012).
- Lee, P. Y., Costumbrado, J., Hsu, C. Y., Kim, Y. H. Agarose gel electrophoresis for the separation of DNA fragments. Journal of Visualized Experiments. (62), e3923 (2012).
- Bio-Rad Laboratories Inc. A Guide to Polyacrylamide Gel Electrophoresis and Detection. Bulletin 6040, Rev C. Bio-Rad Laboratories Inc. Accessed. , (2023).
- NEBExpress Ni Resin Gravity Flow Typical Protocol. New England Biolabs Inc Available from: https://international.neb.com/protocols/2019/09/10/nebexpress-ni-resin-gravity-flow-typical-protocol (2023)
- Ward, V. C. A., Rehmann, L. Fast media optimization for mixotrophic cultivation of Chlorella vulgaris. Scientific Reports. 9, 19262 (2019).
- Morton, E. R., Fuqua, C. Laboratory maintenance of Agrobacterium. Current Protocols in Microbiology. , (2012).
- Haddadi, F., Abd Aziz, M., Abdullah, S. N., Tan, S. G., Kamaladini, H. An efficient Agrobacterium-mediated transformation of strawberry cv. Camarosa by a dual plasmid system. Molecules. 20 (3), 3647-3666 (2015).
- Wang, X., Ryu, D., Houtkooper, R. H., Auwerx, J. Antibiotic use and abuse: a threat to mitochondria and chloroplasts with impact on research, health, and environment. Bioessays. 37 (10), 1045-1053 (2015).
- Gelvin, S. B. Plant DNA repair and Agrobacterium T-DNA integration. International Journal of Molecular Sciences. 22 (16), 8458 (2021).

