Light-Sheet Imaging to Reveal Cardiac Structure in Rodent Hearts
Summary
The protocol utilizes advanced light-sheet microscopy along with adapted tissue clearing methods to investigate intricate cardiac structures in rodent hearts, holding great potential for the understanding of cardiac morphogenesis and remodeling.
Abstract
Light-sheet microscopy (LSM) plays a pivotal role in comprehending the intricate three-dimensional (3D) structure of the heart, providing crucial insights into fundamental cardiac physiology and pathologic responses. We hereby delve into the development and implementation of the LSM technique to elucidate the micro-architecture of the heart in mouse models. The methodology integrates a customized LSM system with tissue clearing techniques, mitigating light scattering within cardiac tissues for volumetric imaging. The combination of conventional LSM with image stitching and multiview deconvolution approaches allows for the capture of the entire heart. To address the inherent trade-off between axial resolution and field of view (FOV), we further introduce an axially swept light-sheet microscopy (ASLM) method to minimize out-of-focus light and uniformly illuminate the heart across the propagation direction. In the meanwhile, tissue clearing methods such as iDISCO enhance light penetration, facilitating the visualization of deep structures and ensuring a comprehensive examination of the myocardium throughout the entire heart. The combination of the proposed LSM and tissue clearing methods presents a promising platform for researchers in resolving cardiac structures in rodent hearts, holding great potential for the understanding of cardiac morphogenesis and remodeling.
Introduction
Heart failure remains the leading cause of mortality worldwide, primarily due to the lack of regenerative capacity of mature cardiomyocytes1. The intricate architecture of the heart plays a crucial role in its function and provides insights into developmental processes. A profound understanding of cardiac structure is essential for elucidating the fundamental processes of cardiac morphogenesis and remodeling in response to myocardial infarction. Recent progress has demonstrated that neonatal mice can restore cardiac function following injury, while adult mice lack such regenerative capacity2. This establishes a foundation for investigating cues associated with structural and functional abnormalities in mouse models. Traditional imaging methods, such as confocal microscopy, have technical limitations, including restricted penetration depth, slow point-scanning scheme, and photo damage from prolonged exposure to laser light. These hinder comprehensive three-dimensional (3D) imaging of the intact heart. In this context, light-sheet microscopy (LSM) emerges as a powerful solution, offering the advantages of high-speed imaging, reduced photo damage, and exceptional optical sectioning capabilities3,4,5. The unique features of LSM position it as a promising method to overcome the limitations of conventional techniques, providing unprecedented insights into cardiac development and remodeling processes6,7,8.
In this protocol, we introduce an imaging strategy that combines advanced LSM with adapted tissue clearing approaches9, allowing for the imaging of entire mouse hearts without the need for specific labeling and mechanical sectioning. We further propose that conventional LSM imaging can be enhanced through multiview deconvolution10 or axially swept light-sheet microscopy (ASLM) techniques11,12,13,14,15 to improve axial resolution. Additionally, the integration of image stitching with either of these methods can effectively overcome the trade-off between spatial resolution and field of view (FOV), thereby advancing the imaging of adult mouse hearts. The incorporation of numerous tissue clearing approaches, including hydrophobic, hydrophilic, and hydrogel-based methods, enables deeper light penetration for capturing the morphology of the entire heart16,17,18,19.
While multiple clearing methods are compatible with current LSM systems, the goal is to minimize photon scattering and enhance light penetration in tissues, like the heart, by replacing lipids with a medium that closely matches its refractive index. iDISCO was chosen as the representative20,21 and adapted for autofluorescence imaging in this protocol due to its rapid processing and high transparency (Figure 1A). Collectively, the integration of the advanced LSM approach with tissue clearing techniques offers a promising framework to unravel intricate cardiac anatomy in rodent hearts, holding significant potential for advancing our understanding of cardiac morphogenesis and pathogenesis.
Protocol
Animal protocols and experiments have been approved and conducted under the oversight of the University of Texas at Dallas Institutional Animal Care and Use Committee (IACUC #21-03). C57BL6 mice, including neonates at postnatal day 1 (P1) and 8-week-old adults, were used in this study. No difference was observed between males and females. All data acquisition and image post-processing were carried out using open-source software or platforms with research or educational licenses. The resources are available from the authors upon reasonable request.
1. Sample preparation and tissue clearing (6 – 10 days)
- Prepare all chemical solutions in the Table of Materials required for the tissue clearing method before initiating the tissue clearing procedure.
NOTE: Perform all procedures in a fume hood while wearing proper personal protective equipment, especially when handling toxic chemicals. - Euthanize animals with CO2 by placing them in a CO2 chamber at the desired time points, such as P1 and 8 weeks, for whole heart collection and immerse the freshly isolated heart at room temperature (RT) in 0.2 M KCl to arrest in diastole16.
- Fix the heart by immersing in 4% paraformaldehyde (PFA) in 1x phosphate buffered saline (PBS) with gentle agitation on a rocker overnight to preserve the cardiac anatomy at 4 °C, see Table 1.
CAUTION: PFA is a toxic chemical and should be performed under a fume hood. - Wash the heart with 1x PBS at RT for 30 min, 3x.
NOTE: Optional: Embed the heart in agarose gel to create a heart assembly for the custom-built LSM system in step 2 as described below.- Prepare the 1% agarose solution by dissolving agarose in PBS and heat the mixture in the microwave until complete dissolution. Pour the solution into a mold to create a bottom layer until the solution cools down to 40 °C.
- Place the heart in the mold and fill the mold with agarose gel until it solidifies. Eliminate any bubbles in the agarose gel to minimize artifacts in the imaging.
NOTE: Embedding the sample reduces the risk of potential damage to the heart anatomy during mounting.
CAUTION: Utilize low-melting-point agarose and avoid placing the heart directly in heated agarose to mitigate the risk of exposing the sample to elevated temperatures.
- Dehydrate the heart using gradients of methanol mixed with deionized water (Figure 1B), processing through sequential concentrations of 20%, 40%, 60%, 80%, and 100% as outlined in Table 1. Repeat with fresh 100% methanol 1x to ensure complete dehydration. Dehydrate neonatal mouse hearts for 1 h in each methanol mixture with shaking. Adjust the incubation time to 2 h for 8-week-old mouse hearts and rat hearts.
CAUTION: Methanol is a volatile, irritating, and flammable chemical. - Replace and chill the heart with fresh 100% methanol for 10 min at 4 °C.
- Replace the solution and bleach the heart with 5% hydrogen peroxide (H2O2) in methanol using a volume ratio of 1:5 (30% H2O2 to 100% methanol) overnight at 4 °C to remove the pigments.
NOTE: The removal of pigments allows for minimizing photon absorption, leading to improved image quality with reduced artifacts. - Replace the solution and incubate the heart in 100% methanol for 1 h at RT.
- Delipidate the heart with the solution containing 66% dichloromethane (DCM) and 33% methanol for 3 h with shaking. Ensure the heart sinks to the bottom of the vial by the end of this step. If not, renew the solution (66% DCM / 33% methanol) and extend the incubation time (e.g., 1 more hour) to achieve full delipidation. Transfer the heart into a fresh solution quickly to prevent desiccation, as DCM is highly volatile.
CAUTION: Use polypropylene or glassware for the experiment, as DCM is not compatible with polystyrene. - Utilize dibenzyl ether (DBE) for refractive index matching (RI = 1.56). For neonatal mouse hearts, the process of refractive index matching takes 2 days, while for 8-week-old mouse hearts, it takes 5 days with mild shaking. Replace it with fresh DBE and extend the incubation time until the heart achieves complete transparency.
CAUTION: DBE is hazardous. Avoid any contact with the skin.
2. Sample mounting (1 day)
NOTE: In case a commercial LSM system is used, follow its specific protocol provided by the company to fix the heart and skip steps 2.1 – 2.9.
- Customize a chamber and a holder resistant to the refractive index-matching solution (e.g., DBE in the adapted iDISCO approach) using a 3D printer and Onyx materials (Figure 2A)16.
- Affix a magnet to the bottom of the chamber to facilitate quick connection and accurate localization under the LSM system (Figure 2B).
- Attach 25 mm x 50 mm x 0.17 mm cover glasses to the chamber using transparent silicone glue and verify the chamber's waterproof integrity by filling it with DBE and checking for leaks after 1 day.
- Customize a clamp holder using a 3D printer and affix a magnet for mounting the heart assembly inside the chamber (Figure 2C) or insert a needle into the agarose gel for mounting the heart assembly.
- Carry out the development of an in-house LSM system using a cylindrical lens and continuous-wave diode-pumped solid-state (DPSS) lasers as previously reported16,22. Use the excitation wavelength of 532 nm to capture autofluorescence from the myocardium (Figure 2D).
- Mount the chamber on a 6-dimensional stage and locate the optimal position where the chamber is perpendicular to the direction of laser propagation.
- Use a magnetic clamp holder to position the sample in the middle of the chamber and connect the holder to a 4-dimensional motorized stage (X, Y, Z, and Yaw; Figure 2D).
- Submerge the heart assembly into the chamber filled with DBE using the motorized stage and determine the scanning range along the detection axis (Z-direction).
- Determine the scanning velocity (Vz) of the motor along the Z-direction based on the step size (S) and acquisition time of an image (T) using the following equation:
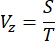
NOTE: The relationship between the step size and the axial resolution of the imaging system adheres to the Nyquist-Shannon sampling theorem. For instance, the step size was set at 1 µm, given that the axial resolution was 2.91 µm, as indicated in the previous report16. - To verify the spatial resolution and minimize potential opacity issues at different depths, measure the point spread function (PSF) of the system by imaging fluorescent beads with the diameter of 0.53 µm diluted in 1% low melting point agarose gel with a concentration of 1:1.5 x 105. Calculate the PSF across the entire sample by measuring the full width at half maximum (FWHM)16.
3. Image stitching (4-8 h)
- Calculate the number of tiles required to cover the entire mouse heart, considering the limited FOV. For each tile, the dimensions correspond to the FOV of the LSM system. For instance, a neonatal mouse heart with a cross-sectional measurement of 3 x 3.6 mm2 necessitates 2 x 2 tiles for coverage in the customized LSM (Figure 3A). In contrast, an 8-week-old mouse heart measuring 5 x 8 mm2 requires 3 x 4 tiles to cover the entire cardiac structure (Figure 3B).
NOTE: Image stitching is required if the conventional LSM has limited FOV for imaging the whole heart. - Capture images of the heart starting from tile 1, i.e., the top left corner tile of the heart (Figure 3A-B).
- Sequentially capture images of the remaining tiles with a 10% overlap between consecutive tiles until the whole heart is covered, as illustrated in Figure 3C.
- Download and install Fiji23 open-source software. Download and install the Fiji plugin, BigStitcher24.
- Navigate to the Help menu, select Update, choose Manage update site, and opt for BigStitcher. Subsequently, click Close, followed by Apply changes. Upon completing the plugin download, restart the Fiji software.
- Open the BigStitcher plugin and import all necessary tiles through the image file directory. Save the dataset as an HDF5 file.
- Organize tiles by choosing Move Tile By Regular Grid, select the pattern that is used for moving the heart, and select a 10% overlap between each tile. Stitch the tiles using the Stitching Wizard option.
- Export the stitched data using Image Fusion, to generate a tiff file.
4. Multiview deconvolution (5 days)
- Utilize fluorescent beads for image registration of multiview reconstruction along the heart (Figure 3D).
- Prepare resin25 by mixing Bisphenol-A diglycidyl ether (D.E.R. 332), Isophorone diamine, 5-amino-1,3,3-trimethylcyclohexanemethylamine (IPDA), and Polypropylene glycol diglycidyl ether (D.E.R 736) at a volume ratio of 4:1:1.
- Centrifuge 10 µL of fluorescence beads at 161 x g for 5 min and add 20 µL of methanol to replace the storage solution of the beads. Repeat it 3x.
- Prepare a 10 mL solution with 1:1 x 105 bead concentration by mixing the methanol-based bead solution into the resin prepared in step 4.1.1.
- Degas the solution in a vacuum chamber for 3 h to remove air pockets and bubbles.
- Pour the solution into a silicone mold or a plastic drinking straw and wait for 2 to 3 days until it solidifies. After 1 day before the resin becomes solid, put the glass tube inside the mold and wait until the tube attaches to the resin. Remove the resin from the mold with the glass tube (Figure 3D).
- Place the heart into the glass tube, fill it with DBE, and mount the glass tube in the chamber filled with DBE.
- Capture sequential images of the mouse heart along with the beads from the illuminated section by scanning the heart assembly across the detection axis (Z-axis; Figure 3E-F).
NOTE: Implement image stitching in step 3 to generate individual stacks of images if the heart size exceeds the FOV. - Rotate the mouse heart 60° along the Y-axis to capture subsequent stacks from multiple angles, i.e., 0°, 60°, 120°, 180°, 240°, and 300° (Figure 3E-F).
- Open the BigStitcher plugin and import all images through the image file directory. Save the dataset as an HDF5 file.
- Choose Detect Interest Points and manually select beads of interest for registration.
- Select Affine transformation model for registration. Choose Point Spread Function and assign PSF to all views.
- Click on Multiview Deconvolution and choose the type of iteration as Efficient Bayesian. Define Number of iterations as 10 and execute it26.
- Click on Image Fusion to export a tiff file.
NOTE: More detailed information about the multiview deconvolution can be found in other reports or protocols24,27,28.
5. Axially swept light-sheet system hardware (1 day)
- Install an electric tunable lens (ETL) on the Fourier plane of the cylindrical lens (CL)27 (Figure 2D) to scan the laser beam axially14. Ensure the liquid interface in ETL is horizontal to prevent wavefront distortion caused by gravity. Align ETL precisely to minimize beam decentering and higher-order optical aberrations30.
- Install a DAQ card in the workstation and connect a BNC cable to the DAQ card to synchronize camera exposure and ETL scanning.
- Open the housing of the ETL driver and remove the cover (Figure 4).
- Solder the signal wire of the BNC cable to the Analog in B pin, as labeled on the bottom of the PCB board (Figure 4).
- Solder the ground wire of the BNC cable to the GND pin; after that connect the other end of the BNC cable to AO (Analog Output) in the DAQ card (Figure 4).
- Plug the ETL Driver into the USB port of the workstation, connect the driver to the ETL using a hirose cable, and install Lens Driver Controller Software.
- Change Operation mode in Lens Driver Controller Software to Analog to control the ETL with a trigger generated from the DAQ card.
- In the software of the sCMOS camera, switch Capture mode to External (Light-sheet). Ensure that the active pixel scanning direction is perpendicular to the laser scanning direction to assist the synchronization.
- Connect the external trigger port on the back of the sCMOS camera to the AO of the DAQ card using a BNC cable (Figure 4).
6. Axially swept light-sheet system synchronization (7 days)
- Define exposure time based on the acceptable signal-to-background ratio (SBR) which for the proposed study, is at least 10. Increase the exposure time if SBR is lower than 10.
NOTE: Exposure time ranging from 10 to 50 ms provides an acceptable SBR in this project. - Define the acquisition time of an image (T) and the line interval (L) based on the exposure time (E) for the camera and the ETL frequency (f) using the following equations:
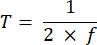
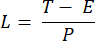
Where P indicates the number of pixel columns on the camera sensor. The representative parameters for the reference are f = 1 Hz, P = 2048, E = 10 ms, and L = 243 µs, respectively. - In LabVIEW program Trigger Generator.vi, generates both square and triangle triggers for synchronization (Figure 5).
NOTE: The customized LabVIEW program is available from the authors upon reasonable request. - Mount fluorescent beads diluted in 1% low melting point agarose gel16 with a concentration of 1:1 x 103 to locate the position of the beam waist in the image (Figure 6A).
- In the program, identify the starting and ending points of the laser beam under the FOV by changing the voltage along the scanning direction.
- To synchronize the sCMOS camera and ETL, scan the laser beam along the X-axis and active pixels along the Y-axis to generate a 2D image (Figure 6B). The brighter and sharper beads along the diagonal in the image indicate the precise synchronization between the sCMOS and ETL.
NOTE: The synchronization of the scanning speed of ETL and sCMOS pixel activation is critical for image quality. Various common asynchronous errors are summarized in Figure 6C-F. - After determining optimal synchronization parameters for synchronization, rotate the sCMOS camera 90° around the Z-axis to align the active pixel direction with the laser scanning direction.
- Repeat step 2.10, and measure the FWHM of the PSF. Use the same method as indicated in the previous report16, with the averaged lateral and axial resolutions of ASLM as 2.18 µm and 2.88 µm, respectively (Figure 7).
NOTE: Uniform illumination and resolution across the entire FOV are feasible under ideal conditions (Figure 7A-B). Asynchronous operation of the ETL with the sCMOS camera, as well as misalignment of the illumination and detection, can result in non-uniform spatial resolution (Figure 7C-D). - For cardiac imaging, after aligning the system and determining the optimal parameters for system synchronization, proceed with steps 2.6 to 2.9.
Representative Results
LSM has been demonstrated to foster cardiac studies31,32,33,34,35,36,37 due to the minimal risk of photo damage, high spatial resolution, and optical sectioning as opposed to other optical imaging methods such as brightfield and point-scanning techniques6,8,38,39,40. Thus, to better understand the 3D myocardial architecture, we have established the protocol based on primary approaches including adapted iDISCO clearing, image stitching, multiview deconvolution, and ASLM. While we presented results using mouse models, it is important to note that rats and other rodent models are also suitable for the proposed methods. The adapted tissue clearing protocol enables the rendering of the P1 mouse heart transparent within 4 days and the 8-week-old mouse heart transparent within 7 days (Figure 1B). This step serves as a starting point for minimizing photon absorption and scattering in the intact heart. Multiview deconvolution enables the improvement of axial resolution by fusing images from multiple perspectives into a single model. After sequentially recording images from 6 views of the same heart, the multiview deconvolution method achieves near-isotropic resolution with a lateral of 4.68 µm and axial of 5.06 µm, following 15 h of computation (Figure 8A). To obviate the computational complexity, we further customized an ETL-based ASLM to image from neonatal to adult mouse hearts. This method allows us to scan the myocardium with a tightly focused laser beam, minimizing the out-of-focus background and enhancing the image contrast in comparison to the conventional LSM methods (Figure 8B). With the current ASLM design, the averaged lateral and axial resolutions of the system achieve 2.18 µm and 2.88 µm, respectively, enabling in-depth exploration of myocardial trabeculation in ventricles. In addition, image stitching can be utilized independently, or in combination with multiview deconvolution or ASLM to expand the FOV for larger sample sizes. Integrating stitching with ASLM allows us to cover the entire 8-week-old mouse heart with uniform resolution (Figure 8C), providing an effective solution to investigate the cross-section of the entire heart.
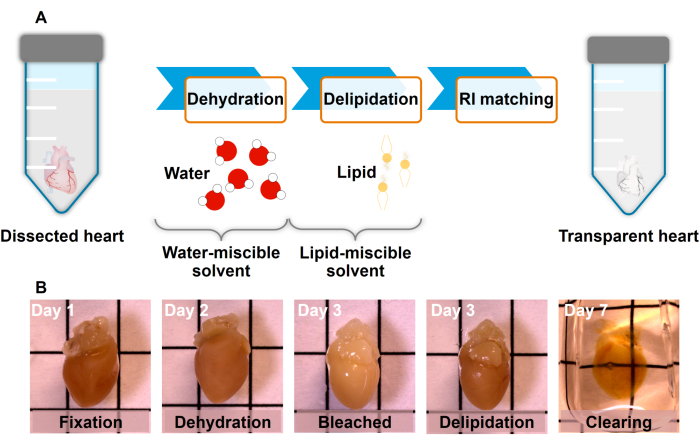
Figure 1: Tissue clearing process. (A) The hydrophobic method involves tissue dehydration, lipid extraction, and refractive index matching using dibenzyl ether (DBE) as the organic solvent. (B) Submerge the heart in a mixture of 4% formaldehyde (PFA) solution and dehydrate it with a gradient of methanol and deionized (DI) water mixtures. Bleach the heart using 5% H2O2 in methanol at 4 °C. Delipidate the heart with dichloromethane (DCM) and methanol solution until the sample sinks to the bottom of the tube. Lastly, incubate the heart in DBE in order to obtain the desired refractive index. Grid lines 5 mm. Please click here to view a larger version of this figure.
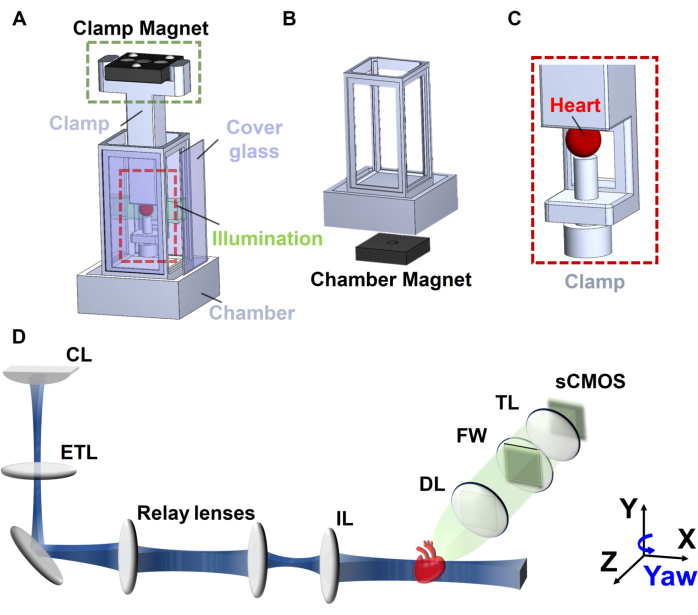
Figure 2: Schematic of axially swept light-sheet microscopy. (A) Customized 3D printed chamber, (B) chamber magnet, (C) clamp holder, and (D) ASLM system. Abbreviations: CL: cylindrical lens; ETL: electronic tunable lens; IL: illumination lens; DL: detection lens; FW: filter wheel; TL: tube lens. This figure has been modified from Ref16. Please click here to view a larger version of this figure.
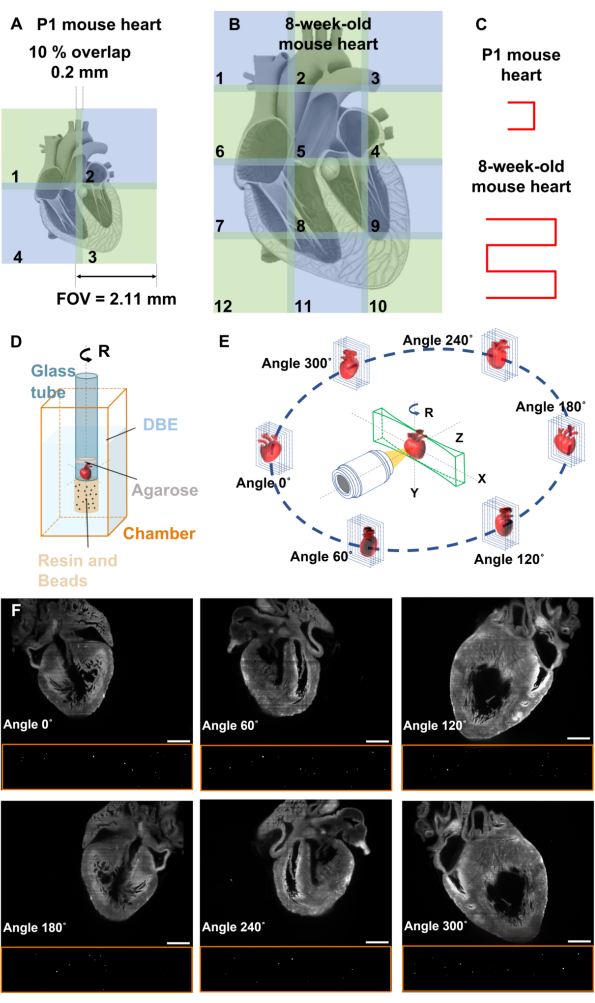
Figure 3: Image stitching and multiview deconvolution. (A-B) Image stitching is used to cover the entire mouse hearts, with each tile having a 10% FOV overlap with its adjacent tiles. (C) Movement pattern of the mouse heart for image stitching. (D) Fluorescent beads are affixed to the end of the glass tube to facilitate image registration, while the heart is positioned inside the glass tube containing DBE, enabling concurrent imaging of both the mouse heart and fluorescent beads. (E) Multiview deconvolution involves the acquisition of images from six distinct perspectives, each gathering images from unique directions. (F) Raw data of P1 mouse heart and beads at different angles of 0°, 60°, 120°, 180°, 240°, and 300°. Scale bars: 500 µm. This figure has been modified from16. Please click here to view a larger version of this figure.
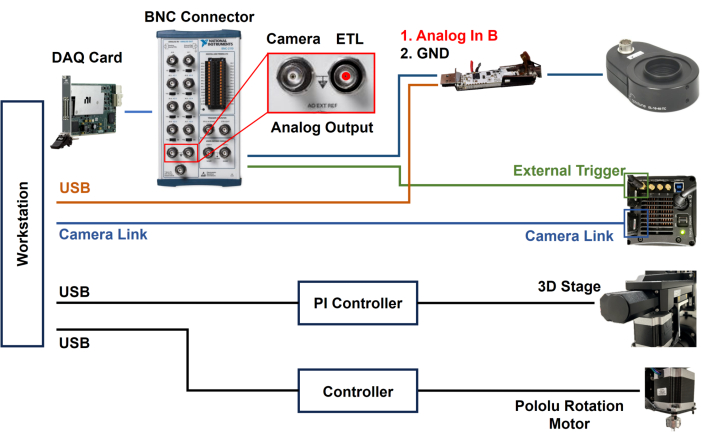
Figure 4: Block diagram of ASLM system. Utilizing DAQ card for synchronizing ETL and sCMOS camera. The housing of the ETL driver has been opened for soldering of signal and ground wires to the PCB. Please click here to view a larger version of this figure.
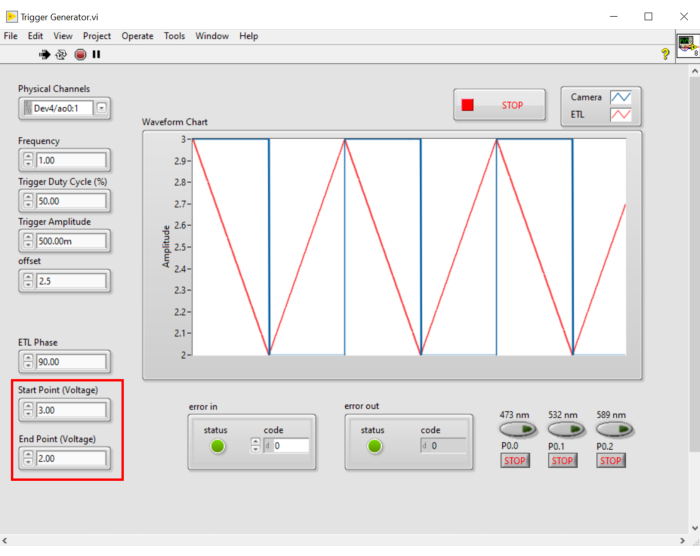
Figure 5: LabView control panel. LabVIEW control panel for generating triggers to synchronize ETL and activated pixels of sCMOS camera. Please click here to view a larger version of this figure.
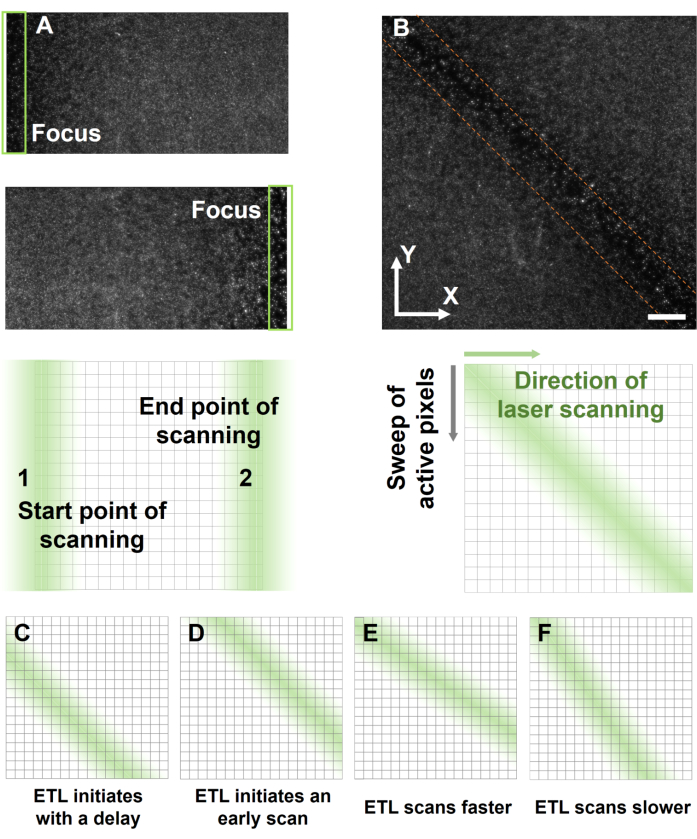
Figure 6: Synchronizing the focus of light-sheet with activated pixels. (A) The identification of the starting and ending points of the light-sheet scanning range is achieved by configuring the appropriate voltage for the ETL trigger. (B) The synchronous result of fluorescent beads is evident along the diagonal of the image. (C-F) The non-uniform spatial resolution across the entire FOV arises from the asynchronous operation of the ETL with the sCMOS camera. The ETL initiates scanning (C) later or (D) earlier than the activated pixel. (E-F) The focal area is not parallel to the diagonal of the image, indicating an incompatibility between scanning and activation speeds. ETL scans (E) faster or (F) slower than the sweeping of active pixels in sCMOS. Scale bar: 200 µm. Please click here to view a larger version of this figure.
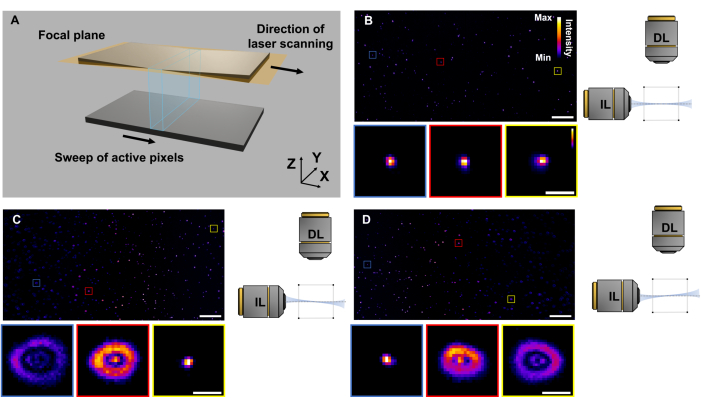
Figure 7: Troubleshooting of ASLM. (A) The light-sheet is placed exactly at the focal distance of the detection lens. The ETL scans the focus of the light-sheet along the propagation direction while synchronizing with the activated pixel of the sCMOS sensor. (B) XY view of fluorescent beads when the ETL is correctly aligned and synchronized. (C-D) Results of fluorescent beads on the focal plane and out-of-focus in cross-sectional images, indicating the misalignment between the ETL scanning and laser propagation. Scale bars: 200 µm and 10 µm in the inset. Please click here to view a larger version of this figure.
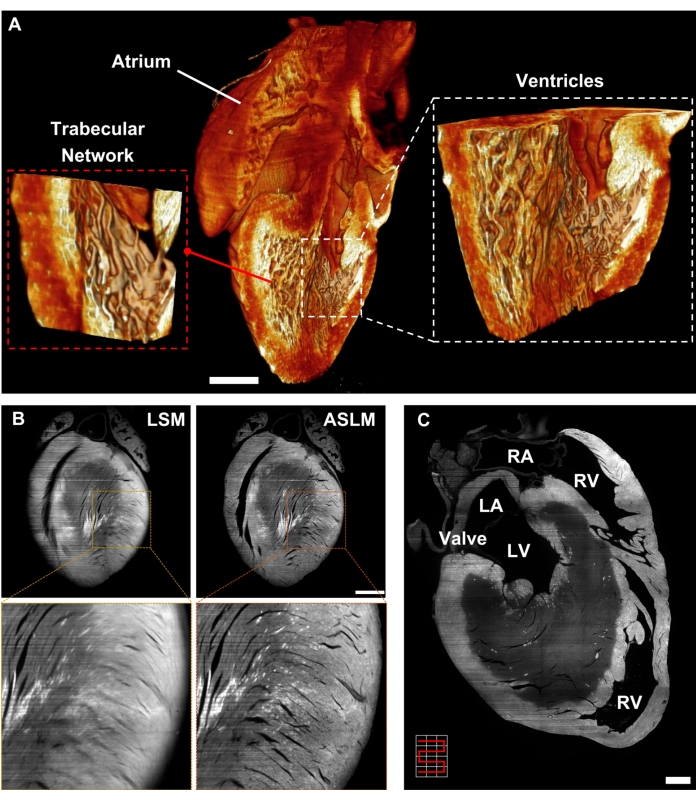
Figure 8: Light-sheet imaging of mouse hearts. (A) Volume-rendered image of the multiview deconvolution P1 mouse heart highlighting the trabeculation within the ventricular cavity. (B) Cross-sectional images of the 8-week-old mouse heart generated by conventional LSM (left) and ASLM (right). Images are presented as raw data. (C) Combination of Image stitching and ASLM for imaging an 8-week-old mouse heart, comprising 12 tiles arranged in 3 tiles horizontally and 4 tiles vertically. Scale bar: 500 µm. Abbreviations: LV: left ventricle, LA: left atrium, RV: right ventricle, and RA: right atrium. This figure has been modified from Ref16. Please click here to view a larger version of this figure.
| Procedure | Time (P1 mouse heart) | Time (8-week-old mouse heart) | Temperature | |||
| Fix in 4% PFA | Overnight | Overnight | 4°C | Sample preparation | ||
| Wash with 1x PBS | 30 min | 30 min | RT | |||
| Wash with 1x PBS | 30 min | 30 min | RT | |||
| Wash with 1x PBS | 30 min | 30 min | RT | |||
| Embed with agarose gel | 1 h | 2 h | RT | |||
| Incubate in 20% methanol/80% DI water | 1 h | 2 h | RT | Dehydration | ||
| Incubate in 40% methanol/60% DI water | 1 h | 2 h | RT | |||
| Incubate in 60% methanol/40% DI water | 1 h | 2 h | RT | |||
| Incubate in 80% methanol/20% DI water | 1 h | 2 h | RT | |||
| Incubate in 100% methanol | 1 h | 2 h | RT | |||
| Incubate in fresh 100% methanol | 1 h | 2 h | RT | |||
| Incubate in fresh 100% methanol | 10 min | 10 min | 4°C | Depigmentation | ||
| Bleach with 5% H2O2/methanol | Overnight | Overnight | 4°C | |||
| Incubate in 100% methanol | 1 h | 1 h | RT | |||
| Delipidate with 66% DCM/33% methanol on the shaker | 3 h | 3 h | RT | Delipidation | ||
| Incubate in DBE on the shaker | 2 days | 5 days | RT | RI matching | ||
Table 1: Customized tissue clearing steps for autofluorescence imaging.
Discussion
The advancement of imaging, computation, and tissue clearing methods has provided an unparalleled opportunity to extensively investigate cardiac structure and function. This holds great potential for deepening our understanding of cardiac morphogenesis and pathogenesis using an intact rodent heart model. In contrast to in vivo studies of zebrafish heart using a similar approach40,41,42,43, the integration of advanced LSM techniques and tissue clearing methods enables us to overcome challenges between spatial resolution and penetration depth when imaging the rodent model9,44. Tissue clearing techniques render hearts optically transparent by removing lipids and other light-scattering elements, enabling deeper imaging penetration45. When coupled with LSM, we are able to image the myocardium. This combination enhances the imaging depth and allows for the observation of intricate myocardial micro-structure. While numerous clearing methods have been established18, we adapted the iDISCO20 to rapidly render the intact heart transparent in comparison with other tissue clearing methods15, making it readily available for the imaging of autofluorescence from the myocardium. For specific fluorescence labeling, the original iDISCO and Adipo-Clear46 protocols can be combined with immunostaining in this customized imaging system, despite the longer incubation. Meanwhile, methods that preserve endogenous fluorescence and require shorter processing time are preferred for the proposed study. In contrast to other methods, this platform offers the following advantages. Firstly, we utilize an adapted iDISCO protocol for tissue clearing, enabling rapid rendering of the heart transparent in less than 1 week while retaining myocardium autofluorescence. Secondly, we incorporate computational methods such as image stitching and multiview deconvolution into this established light-sheet system for cardiac imaging. Lastly, we design and synchronize the ASLM system to enhance axial resolution across a large FOV, enabling multiscale imaging and analysis of myocardial compaction and trabeculation at a near isotropic resolution.
To capture cardiac images of the rodent model, we have provided four approaches based on LSM: i) the conventional method for screening purposes19, ii) image stitching, iii) multiview deconvolution, and iv) ASLM scanning. Considering the variation in heart size across different models, we sought to address the fundamental trade-off in spatial resolution, acquisition speed, and FOV. While image stitching enables the large FOV with compromised spatial resolution, combining it with multiview deconvolution or ASLM provides near-isotropic resolution across the extended FOV to cover the entire adult mouse heart. Furthermore, the integration of these methods enhances image quality and contrast in the deep tissue, allowing for a comprehensive digital reconstruction of 3D models of micro-structures such as myocardial trabeculation and compaction inside ventricles. However, the deluge of datasets, extensive computational processing cost, the need for precise image registration, and the potential for artifacts in multiview deconvolution and imaging stitching hinder their broad applications. While ASLM is an emerging alternative, the complexity of the system synchronization, laser scanning strategies, and reliance on tissue transparency pose challenges that may affect system robustness and increase the risk of photo damage to the heart.
Collectively, this methodology including LSM imaging and tissue clearing provides a starting point to explore cardiac structure from neonates to adults. It holds the potential to localize the distribution of specialized cardiac lineages and their functions during cardiac development47. With the development of computational analysis tailored for cardiac applications6,48,49,50,51,52,53 , this framework can be extended to investigate cardiac remodeling processes, particularly in response to myocardial infarction. Ultimately, this holistic strategy holds the potential to unravel the underlying mechanism of cardiac morphogenesis and advance the development of therapeutic interventions.
Disclosures
The authors have nothing to disclose.
Acknowledgements
We express our gratitude to Dr. Eric Olson's group at UT Southwestern Medical Center for generously sharing the animal models. We appreciate all the constructive comments provided by D-incubator members at UT Dallas. This work was supported by NIH R00HL148493 (Y.D.), R01HL162635 (Y.D.), and UT Dallas STARS program (Y.D.).
Materials
| 1% Agarose | |||
| Low melting point agarose | Thermo Fisher | 16520050 | |
| Deionized water | – | – | |
| Chemicals for tissue clearing | |||
| 5-Amino-1,3,3-trimethylcyclohexanemethylamine, mixture of cis and trans | Sigma-Aldrich | 118184 | |
| D.E.R.™ 332 | Sigma-Aldrich | 31185 | |
| D.E.R.™ 736 | Sigma-Aldrich | 31191 | |
| Dibenzyl ether (DBE) | Sigma-Aldrich | 33630 | |
| Dichloromethane (DCM) | Sigma-Aldrich | 270997 | |
| Fluorescent beads | Spherotech | FP-0556-2 | |
| Hydrogen peroxide (H2O2) | Sigma-Aldrich | 216736 | |
| Methanol | Sigma-Aldrich | 439193 | |
| Paraformaldehyde (PFA) | Thermo Fisher | 47392 | |
| Phosphate Buffered Saline (PBS) | Sigma-Aldrich | 79383 | |
| Potassium Chloride (KCl) | Sigma-Aldrich | P3911 | |
| Software and algorithms | |||
| Amira | Thermo Fisher Scientific | 2021.2 | |
| BigStitcher | Hörl et al.22 | ||
| Fiji-ImageJ | Schindelin et al.20 | 1.54f | |
| HCImage Live | Hamamatsu Photonics | 4.6.1.2 | |
| LabVIEW | National Instruments Corporation | 2017 SP1 | |
| Key components of the customized light-sheet system | |||
| 0.63 – 6.3X Zoom body | Olympus | MVX-ZB10 | |
| 10X Illumination objective | Nikon | MRH00105 | |
| 1X detection objective | Olympus | MV PLAPO 1X/0.25 | |
| 473nm DPSS Laser | Laserglow Technologies | LRS-0473-PFM-00100-05 | |
| 532nm DPSS laser | Laserglow Technologies | LRS-0532-PFM-00100-05 | |
| 589 nm DPSS laser | Laserglow Technologies | LRS-0589-GFF-00100-05 | |
| BNC connector | National Instrument | BNC-2110 | |
| Cylindrical lens | Thorlabs | ACY254-050-A | |
| DC-Motor Controller, 4 axes | Physik Instrumente | C-884.4DC | |
| ETL | Optotune | EL-16-40-TC-VIS-5D-1-C | |
| ETL Cable | Optotune | CAB-6-300 | |
| ETL Lens Driver | Optotune | EL-E-4i | |
| Filter | Chroma | ET525/30 | |
| Filter | Chroma | ET585-40 | |
| Filter | Chroma | ET645-75 | |
| Filter wheel | Shutter Instrument | LAMBDA 10-B | |
| Motorized translation stage | Physik Instrumente | L-406.20DG10 | |
| Motorized translation stage | Physik Instrumente | L-406.40DG10 | |
| Motorized translation stage | Physik Instrumente | M-403.4PD | |
| NI multifunction I/O | National Instrument | PCIe-6363 | |
| sCMOS camera | Hamamatsu | C13440-20CU | |
| Stepper motor | Pololu | 1474 | |
| Tube lens | Olympus | MVX-TLU |
References
- Sadek, H., Olson, E. N. Toward the goal of human heart regeneration. Cell Stem Cell. 26, 7-16 (2020).
- Porrello, E. R., et al. Transient regenerative potential of the neonatal mouse heart. Science. 331, 1078-1080 (2011).
- Stelzer, E. H. K. K., et al. Light sheet fluorescence microscopy. Nat Rev Methods Prim. 1, 73 (2021).
- Girkin, J. M., Carvalho, M. T. The light-sheet microscopy revolution. J Opt. 20, 053002 (2018).
- Power, R. M., Huisken, J. A guide to light-sheet fluorescence microscopy for multiscale imaging. Nat Methods. 14, 360-373 (2017).
- Ding, Y., et al. Multiscale light-sheet for rapid imaging of cardiopulmonary system. JCI Insight. 3, e121396 (2018).
- Ding, Y., et al. Light-sheet fluorescence imaging to localize cardiac lineage and protein distribution. Sci Rep. 7, 42209 (2017).
- Fei, P., et al. Cardiac light-sheet fluorescent microscopy for multi-scale and rapid imaging of architecture and function. Sci Rep. 6, 1-12 (2016).
- Richardson, D. S., Lichtman, J. W. Clarifying tissue clearing. Cell. 162, 246-257 (2015).
- Stelzer, E. H. K., Huisken, J., Swoger, J., Greger, K., Verveer, P. Multi-view image fusion improves resolution in three-dimensional microscopy. Opt Express. 15 (13), 8029-8042 (2007).
- Dean, K. M., Roudot, P., Welf, E. S., Danuser, G., Fiolka, R. Deconvolution-free subcellular imaging with axially swept light sheet microscopy. Biophys J. 108, 2807-2815 (2015).
- Dean, K. M., et al. Isotropic imaging across spatial scales with axially swept light-sheet microscopy. Nat Protoc. 17, 2025-2053 (2022).
- Hedde, P. N., Gratton, E. Selective plane illumination microscopy with a light sheet of uniform thickness formed by an electrically tunable lens. Microsc Res Tech. 81, 924 (2018).
- Voigt, F. F., et al. The mesoSPIM initiative: open-source light-sheet microscopes for imaging cleared tissue. Nat Meth. 16, 1105-1108 (2019).
- Giardini, F., et al. Mesoscopic optical imaging of whole mouse heart. J Vis Exp. (176), e62795 (2021).
- Sodimu, O., et al. Light sheet imaging and interactive analysis of the cardiac structure in neonatal mice. J Biophotonics. 16, e202200278 (2023).
- Ariel, P. A beginner’s guide to tissue clearing. Int J Biochem Cell Biol. 84, 35-39 (2017).
- Richardson, D. S., et al. Tissue clearing. Nat Rev Methods Prim. 1, 1-24 (2021).
- Ueda, H. R., et al. Tissue clearing and its applications in neuroscience. Nat Rev Neurosci. 21, 61-79 (2020).
- Renier, N., et al. iDISCO: a simple, rapid method to immunolabel large tissue samples for volume imaging. Cell. 159, 896-910 (2014).
- Kirchner, K. N., et al. A hydrophobic tissue clearing method for rat brain tissue. J Vis Exp. (166), e61821 (2020).
- Ding, Y., et al. Light-sheet fluorescence microscopy for the study of the murine heart. J Vis Exp. (139), e57769 (2018).
- Schindelin, J., Rueden, C. T., Hiner, M. C., Eliceiri, K. W. The ImageJ ecosystem: An open platform for biomedical image analysis. Mol Reprod Dev. 82, 518-529 (2015).
- Hörl, D., et al. BigStitcher: reconstructing high-resolution image datasets of cleared and expanded samples. Nat Methods. 16, 870-874 (2019).
- Becker, K., et al. Reduction of Photo Bleaching and Long Term Archiving of Chemically Cleared GFP-Expressing Mouse Brains. PLoS One. 9, e114149 (2014).
- Preibisch, S., et al. Efficient Bayesian-based multiview deconvolution. Nat. Methods. 11, 645-648 (2014).
- Guo, M., et al. Rapid image deconvolution and multiview fusion for optical microscopy. Nat. Biotechnol. 38, 1337-1346 (2020).
- Tomer, R., Khairy, K., Amat, F., Keller, P. J. Quantitative high-speed imaging of entire developing embryos with simultaneous multiview light-sheet microscopy. Nat Methods. 9, 755-763 (2012).
- Fahrbach, F. O., Voigt, F. F., Schmid, B., Helmchen, F., Huisken, J. Rapid 3D light-sheet microscopy with a tunable lens. Opt Express. 21, 21010-21026 (2013).
- Liu, Y., Rollins, A. M., Jenkins, M. W. CompassLSM: axially swept light-sheet microscopy made simple. Biomed Opt Express. 12, 6571-6589 (2021).
- Kolesová, H., Olejníčková, V., Kvasilová, A., Gregorovičová, M., Sedmera, D. Tissue clearing and imaging methods for cardiovascular development. iScience. 24 (4), 102387 (2021).
- Sands, G. B., et al. It’s clearly the heart! Optical transparency, cardiac tissue imaging, and computer modelling. Prog Biophys Mol Biol. 168, 18-32 (2022).
- Wilson, A. J., Sands, G. B., LeGrice, I. J., Young, A. A., Ennis, D. B. Muscle mechanics and ventricular function: Myocardial mesostructure and mesofunction. Am J Physiol – Hear Circ Physiol. 323, H257 (2022).
- Lee, S. E., et al. Three-dimensional cardiomyocytes structure revealed by diffusion tensor imaging and its validation using a tissue-clearing technique. Sci. Reports. 8, 1-11 (2018).
- Sereti, K. I., et al. Analysis of cardiomyocyte clonal expansion during mouse heart development and injury. Nat Commun. 9, 754 (2018).
- Olianti, C., et al. 3D imaging and morphometry of the heart capillary system in spontaneously hypertensive rats and normotensive controls. Sci. Reports. 10, 1-9 (2020).
- Olianti, C., et al. Optical clearing in cardiac imaging: A comparative study. Prog Biophys Mol Biol. 168, 10-17 (2022).
- Baek, K. I., et al. Advanced microscopy to elucidate cardiovascular injury and regeneration: 4D light-sheet imaging. Prog Biophys Mol Biol. 138, 105-115 (2018).
- Merz, S. F., et al. Contemporaneous 3D characterization of acute and chronic myocardial I/R injury and response. Nat Commun. 10, 1-14 (2019).
- Zhang, X., et al. 4D Light-sheet imaging and interactive analysis of cardiac contractility in zebrafish larvae. APL Bioeng. 7, 26112 (2023).
- Lee, J., et al. 4-Dimensional light-sheet microscopy to elucidate shear stress modulation of cardiac trabeculation. J Clin Invest. 126, 1679-1690 (2016).
- Zhang, X., Alexander, R. V., Yuan, J., Ding, Y. Computational Analysis of Cardiac Contractile Function. Curr Cardiol Rep. 24, 1983-1994 (2022).
- Zhang, X., et al. 4D Light-sheet Imaging of Zebrafish Cardiac Contraction. J Vis Exp. (203), e66263 (2024).
- Huisken, J., Swoger, J., Del Bene, F., Wittbrodt, J., Stelzer, E. H. K. K. Optical sectioning deep inside live embryos by selective plane illumination microscopy. Science. 305, 1007-1009 (2004).
- Molbay, M., Kolabas, Z. I., Todorov, M. I., Ohn, T., Ertürk, A. A guidebook for DISCO tissue clearing. Mol Syst Biol. 17, 9807 (2021).
- Chi, J., Crane, A., Wu, Z., Cohen, P. Adipo-clear: a tissue clearing method for three-dimensional imaging of adipose tissue. J Vis Exp. (137), e58271 (2018).
- Wan, Y., McDole, K., Keller, P. J. Light-sheet microscopy and its potential for understanding developmental processes. Ann Rev Cell Dev Biol. 35, 655-681 (2019).
- Yuan, J., et al. Extended reality for biomedicine. Nat Rev Methods Prim. 3, 1-1 (2023).
- Ding, Y., et al. Saak transform-based machine learning for light-sheet imaging of cardiac trabeculation. IEEE Trans Biomed Eng. 68, 225-235 (2020).
- Buffinton, C. M., Benjamin, A. K., Firment, A. N., Moon, A. M. Myocardial wall stiffening in a mouse model of persistent truncus arteriosus. PLoS One. 12 (9), e0184678 (2017).
- Trincot, C. E., et al. Adrenomedullin induces cardiac lymphangiogenesis after myocardial infarction and regulates cardiac edema via Cx43. Circ Res. 124, 101 (2019).
- Yokoyama, T., et al. Quantification of sympathetic hyperinnervation and denervation after myocardial infarction by three-dimensional assessment of the cardiac sympathetic network in cleared transparent murine hearts. PLoS One. 12, e0182072 (2017).
- Coram, R. J., et al. Muscleblind-like 1 is required for normal heart valve development in vivo. BMC Dev Biol. 15, 36 (2015).

