Clinicopathological Analysis of miRNA Expression in Breast Cancer Tissues by Using miRNA In Situ Hybridization
Summary
Here we present a protocol to detect miRNA expression in breast cancer patient samples using miRNA in situ hybridization.
Abstract
In this article, we describe a detailed protocol for miRNA detection in breast cancer tissue using in situ hybridization with a digoxigenin-labelled LNA (Locked Nucleic Acid) probe. The probe was recognized by anti-DIG alkaline phosphatase antibodies and later developed using alkaline phosphatase substrate producing fluorescence signals. Here we utilized miRNA in situ hybridization (MISH) technique to analyze expression of miR-489 in tissues from breast cancer patients. This technique can detect the localization of miRNA of interest in individual tissue samples. This technique can be used to compare the expression of desired miRNA in tumor tissue with that in adjacent normal tissue and to identify the specific structures responsible for expressing this miRNA. This technique can be very useful in answering certain clinical questions, such as role of specific miRNA in the development of cancer. Our results indicate that mammary epithelial cells express significantly higher levels of miR-489 than adjacent tumor cells.
Introduction
Breast cancer is among the most common malignancies affecting the female population across the globe. Over 1.3 million cases of breast cancer are reported every year worldwide1,2. Although tumor cells have been traditionally regarded as biologically homogenous and highly proliferative, it has become evident that breast cancer is genetically and clinically heterogeneous. Resistance to prevalent anticancer drugs is a hallmark of advanced breast cancers, leading to the mortality in the majority of patients through its facilitation of cancer progression and distance metastasis3.
MicroRNAs (miRNAs) are a class of small RNA molecules approximately 22 nucleotides long that regulate gene expression by blocking mRNA translation or mediating mRNA degradation4. More than 2,000 different miRNAs have so far been identified in humans, which are linked to human diseases5. An increasing array of studies have demonstrated that miRNAs can function as oncogenes and tumor suppressors and are often dysregulated in tumors6. miRNAs strongly impact all of the primary tumorigenesis by controlling the major regulators of cell cycle progression, senescence, apoptosis and autophagy, along with those of tumor cell motility, invasion and metastasis7.
Aberrant miRNA expression has been also associated with clinical implications such as stage, differentiation, prognosis and response to adjuvant therapy8. In particular, increased miR-146 expression is correlated with poor prognosis in lung cancer patients9. Another study has demonstrated that lost expression of miRNA-let7b is linked to malignant phenotypes and, consequently, poor survival6. Understanding the types of cells and structures that express miRNAs is an important part of understanding the mechanics through which alterations in miRNA expression influence cell and tissue phenotypes10. Certain miRNAs found to be secreted in stromal cells are taken in by epithelial cells and specifically act upon their targets. In the same vein, miR-212 and miR-132 are secreted by stroma and regulate the epithelial-stromal interactions required for ductal outgrowth during mammary gland development11. The overall objective of this paper is to explain a detailed protocol to detect miRNA expression in breast cancer tissues through miRNA in situ hybridization. We have optimized all conditions to assay the levels of miRNA-489 expression in breast cancer patient samples. To this end, we used a DIG labelled locked nucleic acid (LNA) probe in our experiment. Some of its nucleotides had an LNA modification, that is, a methylene bridge connecting the 2' oxygen atom and 4' carbon atom of the ribose backbone which fixes the nucleotide such that it stays "locked in place" after hybridization. As a result, it is much more difficult for the hybridized complex to denature12,13.
Protocol
This study was approved by the Institutional Review Board of the Chonnam National University Hwasun Hospital and University of South Carolina. TMA samples composed of breast cancer and matched adjacent normal breast tissues were provided by the Biobank of Chonnam National University Hwasun Hospital, a member of the Korea Biobank Network.
1. Solution Preparation
- Prepare 1 L of DNase and RNase free water. Make all chemical solutions using diethylpyrocarbonate (DEPC) treated water. Treat 1 L of water with 500 µl of DEPC and incubate for 3 hr at room temperature. Autoclave the water to inactivate DEPC.
Caution: Do not inhale DEPC since it is known carcinogen. - Prepare 1x PBS from 10x PBS using DEPC treated water.
- Prepare 20x SSC (dissolve 175.3 g of NaCl and 88.2 g of Sodium Citrate in 800 ml water, adjust the pH to 7.0 with a few drop of 14 N solution of HCl and adjust the volume to 1 L with water), 4% paraformaldehyde, 95% ethanol, 80% ethanol using DEPC treated water.
- Add 8 ml of triethanolamine and 1.05 ml of HCl in 590 ml of DEPC-treated water and add 1.5 ml of acetic anhydride for acetylation.
- Prepare 50 µg/µl proteinase K in buffer (50 mM Tris-HCl, pH 8.0 in DEPC treated water).
- Prepare hybridization buffer composed of 500 µl of formamide, 250 µl of 20x SSC, 100 µl of 50x Denhardt's solution, 12.5 µl of t-RNA, 2.5 µl of herring sperm DNA, 30 µl of RVC, 0.02 g of blocking reagent and 50 µl of DEPC water. Prepare fresh.
2. Breast Tissue Sections
- Cut 5 µm formalin-fixed paraffin-embedded sections with microtome and apply the section to charge slide14.
Note: The breast tissue used here was prepared previously from the Korea Biobank Network.
3. Tissue Preparation
- Remove paraffin from the tissue by immersing slides into Coplin jar 2x in fresh xylene for 10 min each and rehydrate the tissue section by 5 min incubations in decreasing concentrations of ethanol (100%, 95% and 80%) in ddH2O. Perform this step in a fume hood.
- Immerse the tissue in DEPC treated water for 5 min. At this point make boundaries around tissue section with hydrophobic barrier pen.
- Fix the tissue with 4% PFA for 15 min followed by a wash with PBS.
- Immerse the tissue in 0.3% Triton X-100/PBS for 10 min followed by a wash with PBS.
- Incubate sample with 50 µg/µl Proteinase K solution for 15 min at 37 °C. Then incubate tissue sections with triethanolamine and acetic anhydride for 5 min at room temperature, followed by a wash with PBS. No PBS wash is required between Proteinase K treatment and triethanolamine/acetic anhydride treatment.
4. Hybridization
- Wash the tissue thoroughly and incubate the tissue section with hybridization buffer for 2 to 3 hr at room temperature. Typically 120 µl hybridization buffer is enough to cover the tissue section.
- Dilute 5'-DIG labelled LNA probe to final concentration of 3 pM in 120 µl hybridization solution. As the stock concentration usually is 40 ng/µl, add 3 µl of stock to 120 µl hybridization buffer and incubate mixture at 95 °C for 5 min.
- Incubate tissue section with probe overnight at 54 °C (Tm) in a humid chamber. Typically 120 µl is enough to cover one section. Create humidity by using two wet paper towels. The ideal size of chamber is 10 inches by 8 inches.
- Note: Tm may be different for different miRNA. 54 °C is mention for miR-489.
5. Stringency Wash
- Wash the tissue with 5x SSC for 7 min (after the hybridization step, RNase free conditions are no longer required).
- Wash the tissue section with 1x SSC for 7 min twice at 57 °C.
- Wash the tissue section with 0.2x SSC for 7 min at 57 °C.
- Wash the tissue section with 0.2x SSC for 7 min at room temperature.
- Incubate tissue section in PBS for 10 min.
6. Blocking
- Incubate the tissue section with blocking buffer at room temperature for 1 hr in a humid chamber (To make blocking buffer add 2 ml FBS, 10 µl of Tween-20 and 3 µl of BSA in 18 m of PBS. Store blocking buffer at 4 °C).
7. Primary Antibody Incubation and Development
- Dilute 1:300 anti-DIG-AP antibody in blocking buffer and incubate the tissue section with antibody at 4 °C overnight.
- Wash the sections with wash buffer 3 times, for 5 min each (buffer provided by development kit).
- Develop the tissue section with a commercial kit for alkaline phosphatase that produces CY-3 florescent light.
- Wash the section with wash buffer for 5 min. Stain section with 100 µl of 2.5 ng/µl Hoechst dye for 5 min followed by a 5 min wash with wash buffer.
- Mount the section with mounting buffer according to manufacturer's protocol. Use 15 µl per tissue section and wipe excessive mounting solution with tissue wipe. Seal the slide with clear nail polish to prevent leaking.
- Observe the section florescent microscope using with 10X lens so total magnification is 100X.
Representative Results
Human breast cancer tissue was used to determine miRNA-489 expression. Mammary gland duct and epithelial cells were found to express significantly higher miRNA-489 levels than adjacent tumor tissue in two patients (Figure 1A and 1B). This clearly demonstrates that miRNA-489 expression has been lost in tumor tissue, suggesting tumor suppressive activity of miRNA-489. To further confirm these results, tumor tissue and adjacent normal tissue from the same patient were compared. As observed in Figure 2A and 2B, normal tissue was found to express more miRNA-489 than its adjacent tumor tissue. 10 µm in the images indicates scale bar. Images were taken by using 10X lens.
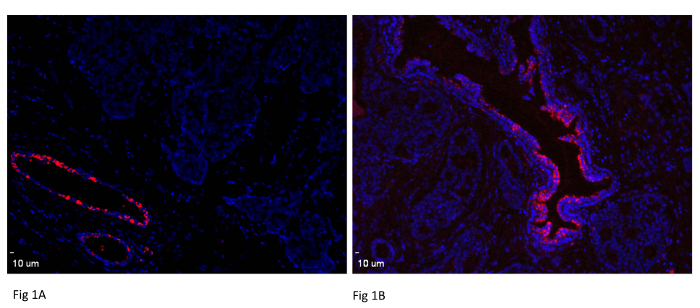
Figure 1: Expression of miR-489 in Normal Breast Tissue. Both normal duct structure and tumor cells are displayed on same section. Tissues are from different breast cancer patients. Normal mammary duct and mammary gland express miRNA-489 while absent in tumor tissue. Scale bar = 10 µm. Please click here to view a larger version of this figure.
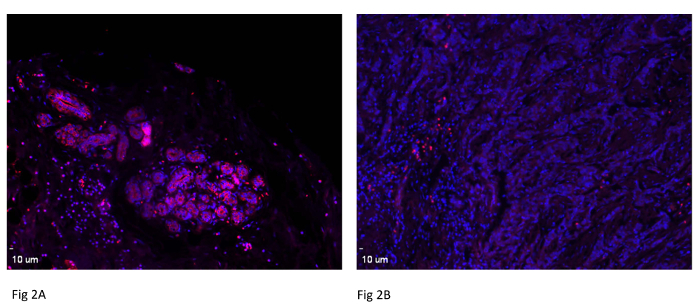
Figure 2: Expression of miR-489 in Breast Cancer Patient Pair. A tissue micro array was used to study miR-489 expression in a breast cancer patient. One representative patient pair is shown here. Scale bar = 10 µm. Please click here to view a larger version of this figure.
Discussion
The goal of this study was to determine the level of miRNA expression in breast cancer tissues using miRNA in situ hybridization. It must be noted that in this experiment, all conditions were optimized for breast cancer tissue. Further optimization might be required for other tissue types. All solutions must be made with DEPC water and all containers to be used should be rinsed with DEPC treated water and subsequently autoclaved. Even slight RNase contamination can interfere with the final outcome of the experiment. As miRNA is a very short sequence, it is necessary to use an LNA probe to facilitate better hybridization with the target. Furthermore, after tissue fixation, treatment with RNase-free Proteinase K is absolutely necessary otherwise proteins may interfere with hybridization of the probe with the endogenous miRNA. Additionally, before incubating the tissue section with the probe, the latter must be boiled to 95 °C to unfold any secondary structures.
After incubating overnight, it is important to wash with varying concentrations of SSC at the temperature mentioned above in protocol to reduce nonspecific binding. For the development of fluorescence, incubate tissue section with substrate for at least 1 hr. It is advisable to observe the section throughout incubation so that overdevelopment can be avoided. Be sure to mount the section with the mounting media provided with the kit.
There is one limiting factor that one should take in consideration. If target miRNA expression is lower in tissue it will be hard to detect its expression using this technique. It is always advisable to perform this protocol with a positive control such as U6 so the technique can be confirmed and all reagents are ensured to be working and they are RNase free.
It is possible to use biotin labelled probe instead of DIG labelled probe in case of weak signal with DIG labelled probe. Signal amplification is possible by using biotin-Avidin system. Avidin has strong affinity for biotin. By taking advantage of this affinity, it is possible to tag biotin probe with avidin-biotin duplex which is attach with alkaline phosphatase. By using this strategy, it is possible to amplify signal for low abundant miRNA as well.
The field of miRNA is rapidly emerging, since miRNAs play critical roles in key cellular processes and are strongly linked with cancer development. Moreover, these miRNAs can also be used as biomarker in cancer diagnosis. Several studies have indicated a strong correlation between miRNA expression and clinical parameters such as tumor stage suggesting possible application of miRNA expression in cancer prognosis. This technique also can be used for miRNA expression analysis in other cancer types. For instance, expression of miR-182 and miR-503 were analyzed in colon cancer patient tissues using in situ hybridization 15.
Traditional gene expression analysis can reveal miRNA expression level in tissue but it cannot reveal the location from which structure miRNA is being expressed. By using this technique, one can easily find the miRNA expression level as well as location.
Divulgazioni
The authors have nothing to disclose.
Acknowledgements
This work was supported by the NIH grant (5R01 CA178386-03) and the USC ASPIRE-1 grant to HC. We would like to thank Vrushab Gowda for his assistance with manuscript.
Materials
| ELF-97 | Life technology | E6604 | |
| 50X Denhard't | Life technology | 750018 | |
| t-RNA | Roche | 109541 | Reconstitute in DEPC water |
| Herring Sperm DNA | Promega | D1815 | |
| Roche Blocking | Roche | 11096176001 | |
| RVC | Fisher | 50-812-650 | Before use spin down it at 16.1 RCF and take supernatant |
| miR-489 probe | Exiqon | 38599-01 | |
| Nuclease free BSA | Roche | 711454 | |
| Primary antibody-anti DIG | Roche | 11093274910 | |
| Diethyl Pyrocarbonate | Sigma | 1609478 |
Riferimenti
- Shah, N. R., Chen, H. MicroRNAs in pathogenesis of breast cancer: Implications in diagnosis and treatment. World J Clin Oncol. 5, 48-60 (2014).
- Tang, H., et al. miR-185 suppresses tumor proliferation by directly targeting E2F6 and DNMT1 and indirectly upregulating BRCA1 in triple-negative breast cancer. Mol Cancer Ther. 13, 3185-3197 (2014).
- Ye, X. M., et al. Epigenetic silencing of miR-375 induces trastuzumab resistance in HER2-positive breast cancer by targeting IGF1R. BMC Cancer. 14, 134 (2014).
- Oneyama, C., et al. MicroRNA-mediated downregulation of mTOR/FGFR3 controls tumor growth induced by Src-related oncogenic pathways. Oncogene. 30, 3489-3501 (2011).
- McGuire, A., Brown, J. A., Kerin, M. J. Metastatic breast cancer: the potential of miRNA for diagnosis and treatment monitoring. Cancer Metastasis Rev. 34, 145-155 (2015).
- Quesne, J. L., et al. Biological and prognostic associations of miR-205 and let-7b in breast cancer revealed by in situ hybridization analysis of micro-RNA expression in arrays of archival tumour tissue. J Pathol. 227, 306-314 (2012).
- Fernandez, S., et al. miR-340 inhibits tumor cell proliferation and induces apoptosis by targeting multiple negative regulators of p27 in non-small cell lung cancer. Oncogene. 34, 3240-3250 (2015).
- Yu, L., Zhang, J., Guo, X., Li, Z., Zhang, P. MicroRNA-224 upregulation and AKT activation synergistically predict poor prognosis in patients with hepatocellular carcinoma. Cancer Epidemiol. 38, 408-413 (2014).
- Li, J., et al. microRNA-146 up-regulation predicts the prognosis of non-small cell lung cancer by miRNA in situ hybridization. Exp Mol Pathol. 96, 195-199 (2014).
- Kriegel, A. J., Liang, M. MicroRNA in situ hybridization for formalin fixed kidney tissues. J Vis Exp. , e50785 (2013).
- Ucar, A., et al. miR-212 and miR-132 are required for epithelial stromal interactions necessary for mouse mammary gland development. Nat Genet. 42, 1101-1108 (2010).
- Nuovo, G. J. In situ detection of microRNAs in paraffin embedded, formalin fixed tissues and the co-localization of their putative targets. Methods. 52, 307-315 (2010).
- Kierzek, E., et al. The influence of locked nucleic acid residues on the thermodynamic properties of 2′-O-methyl RNA/RNA heteroduplexes. Nucleic Acids Res. 33, 5082-5093 (2005).
- Fischer, A. H., Jacobson, K. A., Rose, J., Zeller, R. Paraffin embedding tissue samples for sectioning. CSH Protoc. 2008, (2008).
- Li, L., et al. Sequential expression of miR-182 and miR-503 cooperatively targets FBXW7, contributing to the malignant transformation of colon adenoma to adenocarcinoma. J Pathol. 234, 488-501 (2014).

